Abstract
Astroviruses (AstVs) are important pathogens associated with enteric diseases in humans and other animals. However, most animal AstVs, including feline astrovirus (FAstV), are poorly understood. The aim of the present study was to investigate the prevalence and association of FAstV with enteric diseases in cats, and to conduct a molecular analysis of FAstVs, in Korea. Eleven faecal samples from 62 hospitalised cats at animal hospitals in the Moran market in South Korea tested positive for FAstV. The prevalence of FAstV was higher in cats <2 months old (25%) than in cats >2 months old (14.3%) (P = 0.31). Diarrhoea and normal faeces were observed in 19% (8/42) and 15% (3/20) of cats with FAstV, respectively (P = 1.00). Amino acid sequences alignment and phylogenetic tree analysis showed that FAstVs, including Korean strains, formed a single clade within the mamastroviruses.
Short Communication
Astroviruses (AstVs; family Astroviridae) are small, non-enveloped, spherical viruses, which have a star-shaped structure that is visible by electron microscopy. 1 The genome is a positive single-stranded RNA genome of approximately 7 kb in length that comprises a 5′ untranslated region, three open reading frames (ORFs) and a poly-A tail. 2
AstVs have been classified into two genera, Mamastrovirus and Avastrovirus, infecting mammalian and avian species, respectively. Mamastroviruses have a broad host range, including humans, 3 sheep, 4 deer, 5 dogs, 6 cats, 7 mice, 8 cows, 9 mink, 10 bats, 11 cheetahs, 12 pigs, 13 sea lions 14 and rabbits. 15 AstVs include duck astrovirus 1, turkey astrovirus 1 and 2, and avian nephritis virus. 2
Although AstV infections are a common cause of gastroenteritis in humans, particularly in young children, the association of AstVs with enteric diseases in other animals is not well understood, with the exception of mink and turkeys.10,16
Feline astrovirus (FAstV) was first identified by electron microscopy in faecal samples of cats with diarrhoea in the USA in 1981, 7 and has since been reported in Australia, 17 England, 18 Germany, 19 Italy 20 and New Zealand. 21 Also, an earlier study reported that there is little association between age and excretion of FAstV in cats. 17
FAstV sequence data available in the GenBank database at the National Center for Biotechnology Information are limited and, to date, the genotypes of FAstV have not been analysed. The aim of this study was to investigate the association of FAstV with enteric disease in cats, and to delineate Korean FAstV strains by genotype.
To investigate FAstV in cats, a total of 62 faecal samples were collected from hospitalised cats, including 20 cats <2 months old and 42 cats >2 months old (range, 3–132 months; interquartile range, 10–60 months) in three animal hospitals in the Moran live animal market between January and March 2011. The Moran live animal market, located in Gyenggi-do, is one of the biggest animal markets in Korea, where a variety of animals, including dogs, chickens, ducks, rabbits and cats, are gathered and sold. Cats were chosen randomly from all hospitalised cats. Clinical signs observed in the selected cats included coughing, fever, abdominal pain and diarrhoea (Supplementary material). Of the 62 cat faecal samples collected, 42 were classified as diarrhoea, while the others were classified as normal faeces. Feline coronavirus (FCoV) was diagnosed in 12 of the hospitalised cats using polymerase chain reaction (PCR) with the primer sets (p205 and 211) as previously described. 22
Viral RNA was extracted using the Trizol method according to manufacturers’ instructions (Invitrogen). Reverse transcription PCR (RT-PCR) was performed in a reaction volume of 25 μl using a commercially available RT-PCR kit (One-Step RT-PCR kit; Qiagen) with three sets of primers. Two primer sets from previous studies, primers panAstVfor1 and panAstVrev, and primers Mon269 and Mon270, were used to amplify the ORF1b and ORF2 regions, respectively.11,23 To amplify the region between the 3′ end of ORF1b and the 5′ end of ORF2, a third primer set (forw-ard: 5′-TCTGGAAGAGCTATGACACC-3′; reverse: 5′-GTTGAGGAGTATGCACGCT-3′) was designed based on nucleotide sequences obtained from Korean FAstV strains in the current study. PCR was performed under the following conditions: 94°C for 2 mins; 35 cycles of 94°C for 1 min, 50°C for 1 min and 68°C for 1 min; and a final extension step of 68°C for 10 mins. PCR products were analysed on a 1% agarose gel and cloned using pGEM-T Easy Vector System I (Promega). The cloned fragments were sequenced using the T7 and SP6 universal primers and an ABI Prism 3730xi DNA Sequencer (Applied Biosystems) at Macrogen (Seoul, Korea). The ORF1b and ORF2 amino acid sequences of AstV from different host species were compared using the BioEdit program. 24
Of the 62 cat faecal samples tested, 11 tested positive for FAstV (Table 1). Several risk factors (faecal status, age and gender) associated with FAstV infections were analysed (Table 2). FAstV occurred in cats with diarrhoea (8/42; 19%) and in cats with normal faeces (3/20; 15%). In a previous study of cats in animal shelters in the USA, 2% of cats with normal faeces and 8% of cats with diarrhoea were FAstV positive. 25 In Australia, AstV-like particles were observed by electron microscopy in 3% of cats with normal faeces and 10% of cats with diarrhoea. 17 In the present study, the prevalence of FAstV in cats >2 months old (14.3%) was lower than in cats <2 months old (25%) (Table 2), suggesting that susceptibility to FAstV infection may be related to age. The prevalence of FAstV was not associated with gender or the clinical signs of the cats.
Summary of details for the 11 cats that tested positive for feline astrovirus (FAstV) in this study*

FAstV strains were identified according to sequence comparisons as follows: FAstV-K6 with FAstV-K2, FAstV-K27 with FAstV-K26, FAstV-K40 with FAstV-K37, FAstV-K38 and FAstV-K43, and FAstV-K46 with FAstV-K53 and FAstV-K56
Risk factors associated with astrovirus infection in the present study*

FAstV = feline astrovirus; OR = odds ratio; CI = confidence interval
Risk factors (faecal status, age and gender) were evaluated using Fisher’s exact test. P <0.05 was considered significant
In the current study, 12 cats with diarrhoea tested positive for FCoV; however, no co-infections of FAstV and FCoV were observed (Supplementary material). FCoV is an important pathogen of cats that can cause mild or subclinical enteric diseases. 26 Kittens are particularly susceptible to FCoV, which can also cause feline infectious peritonitis. 27
Analysis of partial ORF1b (759 nt) and ORF2 (656 nt) sequences of the 11 Korean FAstV-positive samples identified four individual strains of FAstV: FAstV-K6, FAstV-K27, FAstV-K40 and FAstV-46. The nucleotide sequences of these strains were identical to those of FAstV in the remaining seven samples: FAstV-K2, FAstV-K26, FAstV-K37, FAstV-K38, FAstV-43, FAstV-K53 and FAstV-K56.
The ORF1b amino acid sequences of AstVs isolated from four host species (feline astrovirus FAstV 443/07A, porcine astrovirus PoAstV12-3, human astrovirus Ihar/2011/kor and canine astrovirus 3/05) shared 92.2%, 83.3%, 74.8% and 73.2% identity, respectively, with the Korean FAstV-K6 strain identified in the present study (Table 3). The ORF2 amino acid sequences of the Korean FAstV-K6 strain shared 95.3% identity with that of the FAstV 443/07A strain isolated in Italy in 2007 (Table 3). This similarity was higher than with a FAstV strain (87.8%) isolated from Bristol, UK, in 1987 (data not shown).
Comparison of the ORF1b and ORF2 amino acid sequences of astroviruses (AstVs) found in four species*
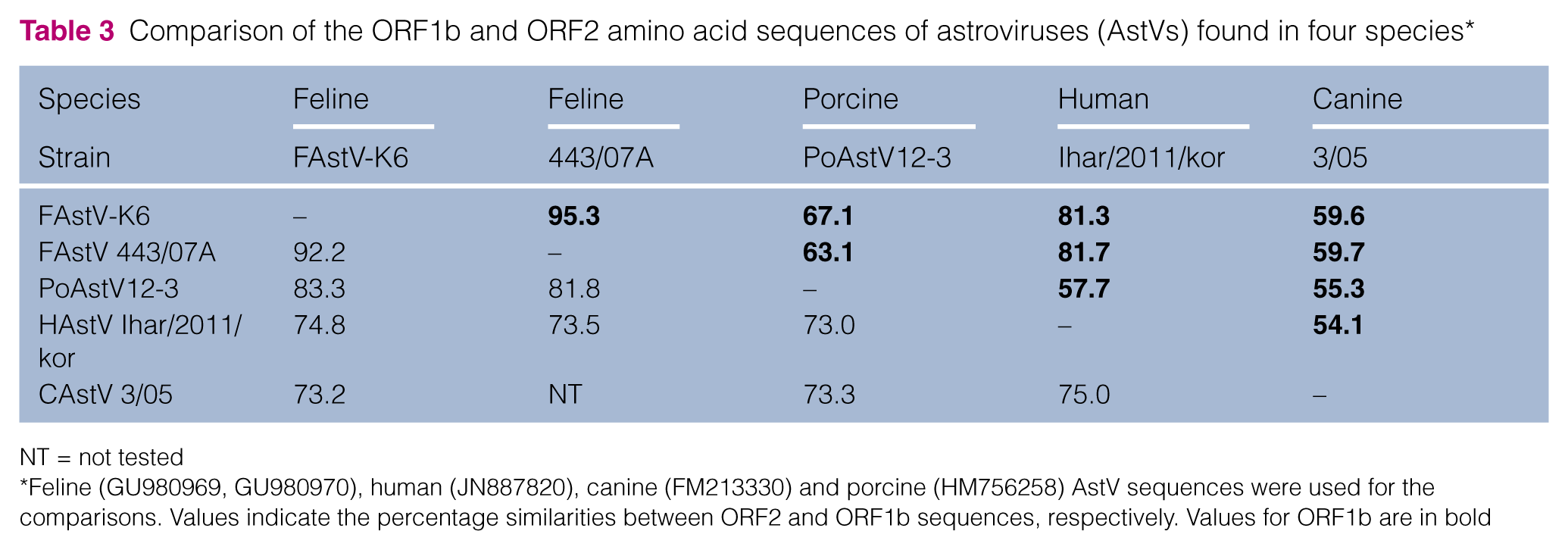
NT = not tested
Feline (GU980969, GU980970), human (JN887820), canine (FM213330) and porcine (HM756258) AstV sequences were used for the comparisons. Values indicate the percentage similarities between ORF2 and ORF1b sequences, respectively. Values for ORF1b are in bold
The neighbour-joining (NJ) phylogenetic trees of both ORF1b (Figure 1a) and ORF2 (Figure 1b) show that the Korean FAstVs formed a single clade with the FAstV 443/07A strain. This clade also included the Bristol FAstV strain in the ORF2 NJ tree (Figure 1b). The Korean FAstVs were more closely related to the 443/07A/Italy/2007 strain than to the Bristol/UK/1986 strain, and the FAstVs also had a close relationship with human AstVs on the ORF2 phylogenetic tree (Figure 1b).
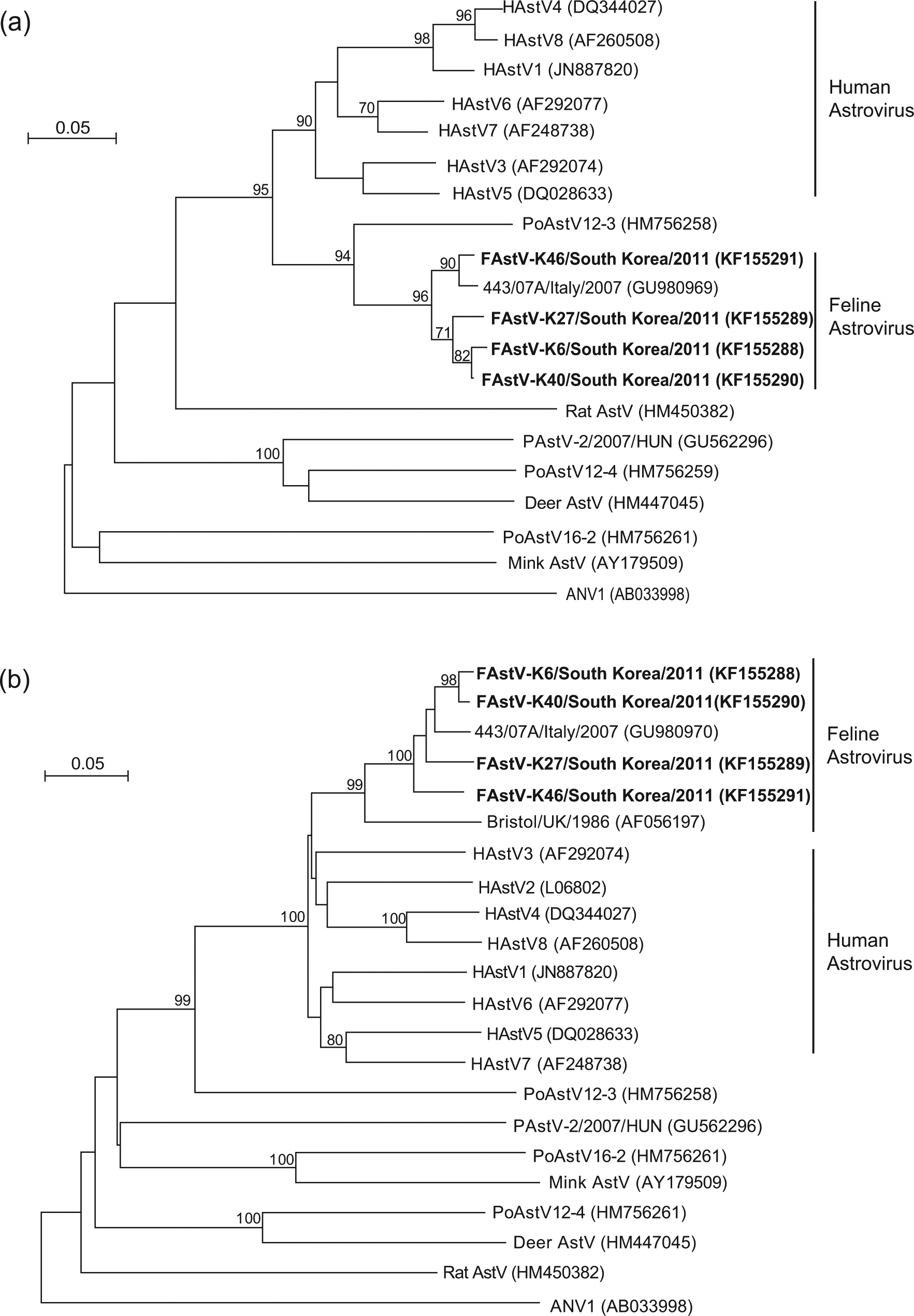
Neighbour-joining (NJ) tree showing the phylogenetic relationships of feline astrovirus (FAstV) to other astrovirus (AstV) species based on partial ORF1b (a) and ORF2 (b) amino acids sequence analysis. The trees were constructed using the NJ method in Mega 4.1 software with 1000 bootstrap replicates. Bootstrap values >70% were observed on the branches. Korean FAstVs are highlighted in bold. The scale bar indicates the number of amino acid changes per site. HAstV = human AstV; PAstV/PoAstV = porcine AstV; ANV = avian nephritis virus
Conclusions
We did not observe a clear association between FAstV infection and diarrhoea, but we did observe that FAstV infection occurred more frequently in younger than in older cats. The FAstVs examined all belonged to one clade.
Supplemental Material
Click here for Supplementary Table
Properties of cats sampled
Footnotes
Acknowledgements
We are grateful to Mr Seong-Hee Lee and Ms Eun-Hye Park for technical assistance.
Supplementary material
Properties of cats sampled.
Funding
This study was supported by a grant from the Animal and Plant Quarantine Agency (QIA), Ministry of Food, Agriculture, Forestry and Fisheries, Republic of Korea in 2012 (Project Code No. N-AD20-2010-19-01).
Conflicts of interest
The authors do not have any potential conflicts of interest to declare.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
