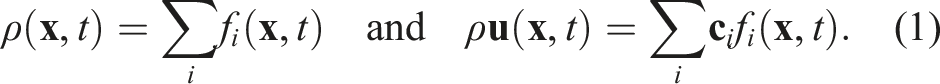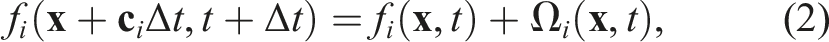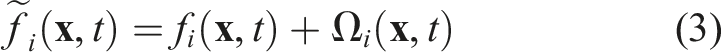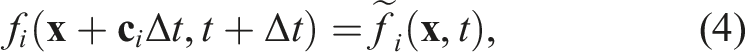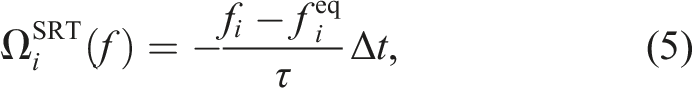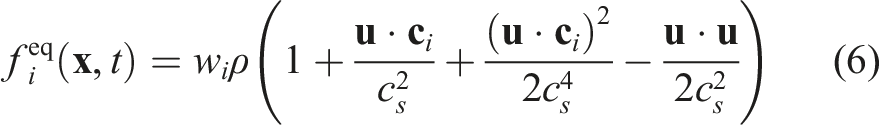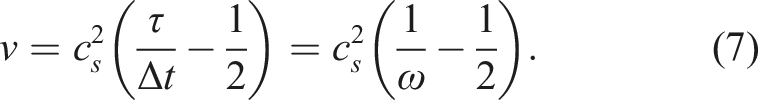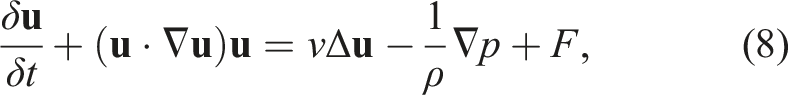Abstract
We implement and analyse a sparse/indirect-addressing data structure for the Lattice Boltzmann Method to support efficient compute kernels for fluid dynamics problems with a high number of non-fluid nodes in the domain, as encountered in porous media flows. The data structure is integrated into a code generation pipeline to enable sparse Lattice Boltzmann Methods with a variety of stencils and collision operators and to generate efficient code for kernels on CPU as well as on AMD and NVIDIA accelerator cards. To further enhance performance, we optimize these sparse kernels with an in-place streaming pattern to save memory accesses and memory consumption and we implement a communication hiding technique to demonstrate strong scalability. We provide a comprehensive, systematic performance analysis comparing the sparse and the traditional dense data structure. We present single GPU performance results for the sparse approach with up to 99% of maximal bandwidth utilization. We integrate the optimized generated kernels in the high performance framework
Keywords
1. Introduction
The increasing power of high-performance computing (HPC) systems enables computational fluid dynamic (CFD) simulations, which were still out of scope some years ago. The leading HPC systems in the Top500 list 1 reach a peak performance of ExaFLOPs, so they can perform 1018 floating point operations per second. This trend is also caused by the utilization of accelerators, such as NVIDIA, AMD, or INTEL Graphics Processing Units (GPUs). With these new computing capabilities, computational fluid dynamic (CFD) problems, which were out of scope before, can now be tackled, such as fully resolved porous media simulations at relevant scales, as presented in Mattila et al. (2016) or entire body arterial flows as in Randles et al. (2015). However, it is not trivial to fully utilize the maximum performance of these HPC systems, especially for systems with accelerators (Brodtkorb et al. (2013); Hijma et al. (2023); Lai et al. (2020); Rak (2024)).
The Lattice Boltzmann method (LBM) (Chen and Doolen (1998); Qian et al. (1992)) is an efficient inherently parallel method to solve CFD problems for complex geometries. On many HPC systems, the LBM shows excellent performance as presented in Liu et al. (2023), Spinelli et al. (2023), Watanabe and Hu (2022) or Godenschwager et al. (2013).
There are two common ways to store data for LBM simulations. The direct-addressing LBM stores and computes all cells of the domain. We call this technique “dense” or ”direct-addressing” data structure in the following.
It is shown to be very fast and efficient for most simulation setups and therefore used for example in the LBM frameworks Palabos (Latt et al. (2021)), ProjectPhysX (Lehmann et al. (2022)), Sailfish (Januszewski and Kostur (2014)) and OpenLB (Krause et al. (2021)). However, the direct-addressing data structure struggles in performance and memory consumption for simulation domains with a high number of non-fluid nodes, such as in porous media flows. In the following, we call such domains “sparse domains”. In contrast, simulations with a high percentage of fluid nodes, such as a free channel flow, we will refer to as “dense domain”.
The second way to store data for LBM simulations is the indirect-addressing storage format. We call it “sparse” data structure in the following. This approach stores only fluid cells and can therefore save a significant amount of memory. Additionally, the sparse approach reaches superior performance for sparse complex geometries as compared with the dense approach. Therefore, it is used in multiple LBM frameworks such as HARVEY (Randles et al. (2013)), HemeLB (Mazzeo and Coveney (2008)), Musubi (Hasert et al. (2014)), ILBDC (Zeiser et al. (2009)) and MUPHY (Bernaschi et al. (2008)), just to mention a few. In particular, the sparse data structure is employed successfully to simulate porous media flows as in Pan et al. (2004), Zeiser et al. (2009), Wang et al. (2005) and Vidal et al. (2010).
In this work we study the code generation for highly efficient LBMs and their performance on large HPC systems. The sparse data structure is realised with the code generation framework of lbmpy (Bauer et al. (2021)), allowing it to run on a variety of architectures, such as all common CPUs as well as NVIDIA and AMD accelerators. The generated sparse compute kernels are integrated in the multiphysics HPC framework
We compare the performance of the generated sparse kernels with the dense approach and present the scaling performance of the sparse data structure on modern HPC systems such as JUWELS Booster (Alvarez (2021)) and LUMI 2 . Further, we show performance results for realistic model problems such as a flow in a porous media, a flow over a packed bed, and a coronary artery flow on a high number of accelerator cards.
2. Lattice Boltzmann method
The lattice Boltzmann method is a mesoscopic method established as an alternative to classical Navier-Stokes solvers (Krüger et al. (2017)). The simulation domain is usually discretized by a lattice of square cells. A cell at position
A standard set of discrete three-dimensional velocity directions would be the D3Q19 stencil, which results in Q = |{
The Boltzmann equation discretized in time, space, and velocity space reads
The simplest collision operator is the single relaxation time (SRT) operator
The incompressible Navier-Stokes equation including the kinematic viscosity reads
For a detailed discussion of the theoretical foundations of the LBM and its derivation of the Navier–Stokes equations as the macroscopic limit, see Chen and Doolen (1998) and Krüger et al. (2017).
3. Data structures in
waLBerla
In the Exemplary setup of a sparse simulation domain in 2D with a low percentage of fluid covering the domain (light blue), and a high number of obstacle cells. Visualisation of the block partitioning with extraction of blocks without fluid. Illustration of a dense and a sparse data structure for an exemplary setup of 5 × 5 cells per block and a D2Q5 stencil. While the dense data structure stores PDFs and operates on all cells, the sparse data structure only stores and operates on fluid cells.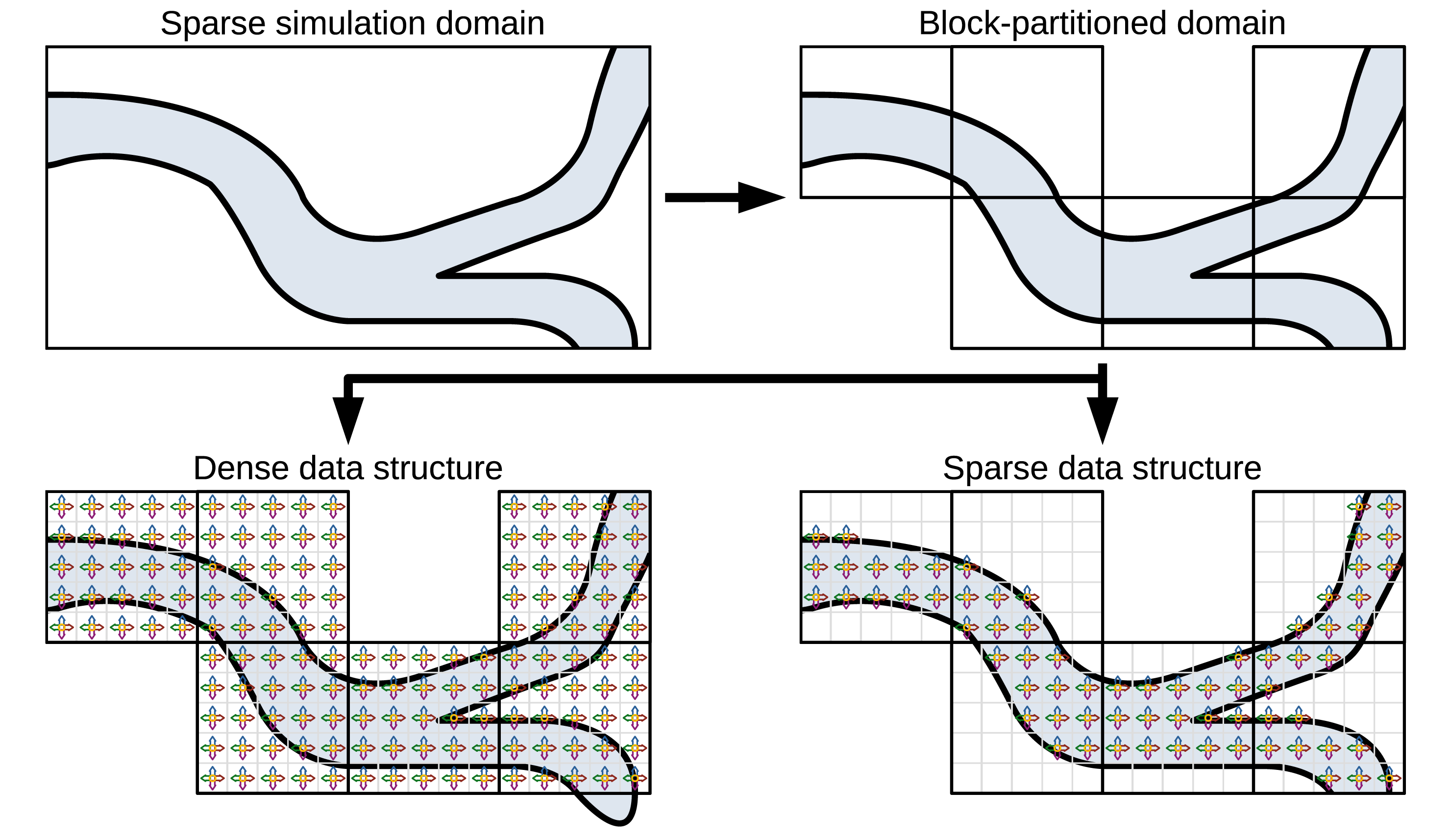
As indicated in Figure 1, the dense/direct-addressing data structure implemented in
3.1. Sparse data structure
To avoid the disadvantages of the dense data structure, we have developed a sparse data structure in
The idea of the indirect-addressing data structure is to only store fluid cells in a one-dimensional array, we call
A second data structure, the
To still have the possibility to access the actual position of the cell
While the
The exact structure of the Structure of the 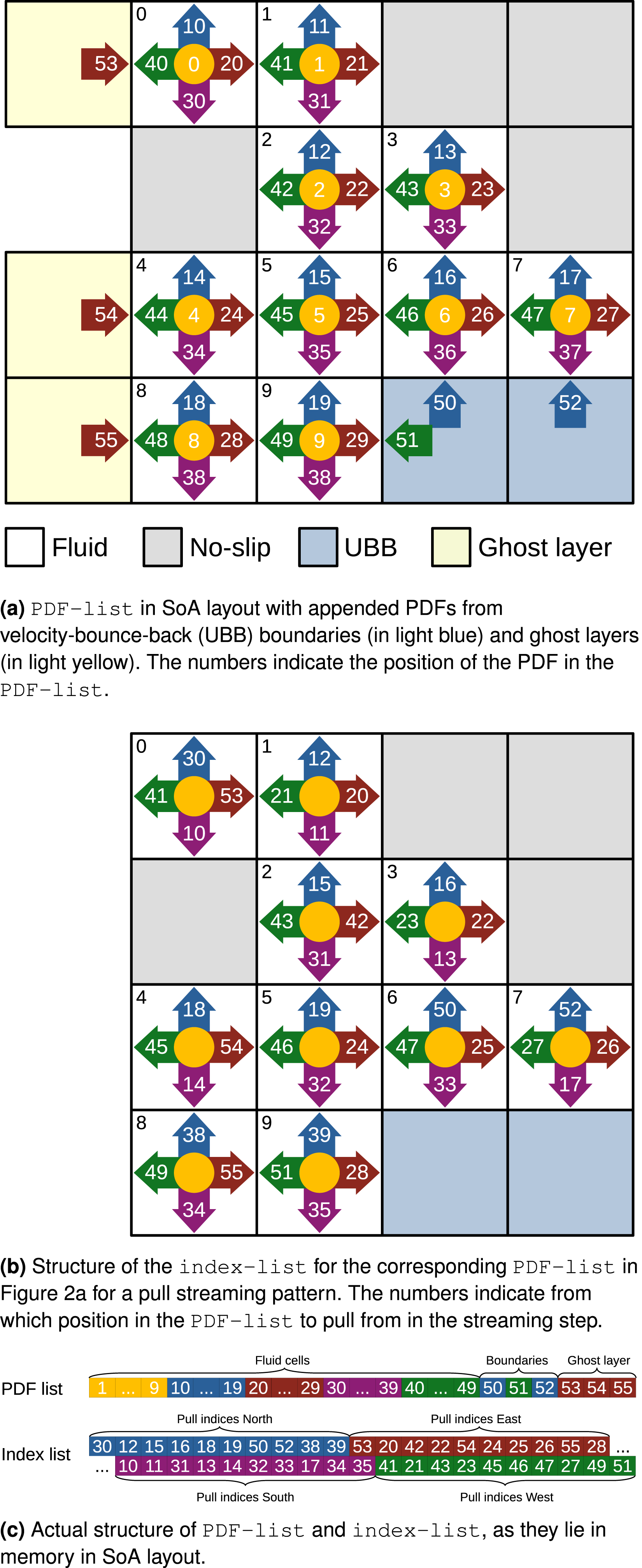
In Figure 2(a) the PDF of cell 0 in direction west
As presented in Figure 1, a computational domain is usually split into multiple
3.2. Sparse boundary conditions
Some modifications to the list data structures are made to support the implementation of boundary conditions. No-slip boundary conditions can easily be realised by setting the pull indices of the PDF, which would pull from a no-slip boundary to the inverse direction of the PDF. This is also illustrated in Figure 2(b). For illustration we focus on
For boundary conditions other than no-slip or periodic, PDFs, which correspond to a boundary cell but point to a fluid cell, must be appended to the
3.3. Sparse communication
For the dense data structure, every block has a ghost layer of at least one cell all around in which it stores the PDFs traveling in the corresponding direction. This ghost layer is used to communicate PDF information between neighboring blocks. For the communication between sparse blocks, we also have to append these ghost layer PDFs to the
To enable periodic boundary conditions on a domain decomposed into multiple blocks and distributed over multiple MPI processes, the block on the periodic boundary is treated as a neighbor of the block on the opposite side of the domain. Thereby, these blocks communicate PDF information into the ghost layer cells of each other. The cell next to the periodic boundary is then able to pull PDFs from the ghost layer cells of the same block, which correspond to the PDFs of the cell on the opposite side of the periodic domain.
As we only append boundary and ghost-layer PDFs pointing to fluid, the memory overhead of the additionally stored PDFs for handling the boundary conditions and the communication is relatively low. In LBM kernels, we still only need to iterate over fluid cells.
3.4. Code generation for sparse kernels
Many variants of the LBM have been developed over the last decades, which vary in complexity, accuracy, and computational cost. The code generation framework lbmpy is capable to generate kernels for most of these LBMs. To get an overview of lbmpy and the provided functionalities and LBM variants, see Bauer et al. (2021) and Hennig et al. (2022).
For example, the classical collision models such as single-relaxation time (SRT, Qian et al. (1992)), two-relaxation time (TRT, Ginzburg (2005)), and multi-relaxation time (MRT, Dhumieres et al. (2002)) operators are available. However, more advanced collision models are also supported, such as the central moment operator or the cumulant operator. The cumulant LBM, e.g., provides superior accuracy and stability for high Reynolds number flows (see Geier et al. (2015)). The complexity of the collision models increases from the SRT model to the cumulant model in terms of complexity and the number of moment transfers, so e.g. the transfer from moment space to central moment space or to cumulant space. This can increase the number of floating point operations in the collision step significantly. However, due to optimizations such as common sub-expression elimination (CSE), the number of operations per cell lies between only 200 and 400 FLOPS for a D3Q19 stencil irrespective of the collision model (Hennig et al. (2022)). Therefore, the performance of the compute kernels remains memory-bound, as the number of memory accesses stays constant for all collision operators. Consequently, we can report a similar performance for all collision operators in the following in Figure 5.
To profit from the functionalities of the code generation pipeline lbmpy, we integrated the generation of new sparse LBM kernels, boundary handling kernels, and communication kernels. The main difference of the code generation for sparse kernels is the iteration loop and the indexing. In dense kernels, the loop iterates over all cells in a three dimensional way. The sparse kernels, on the other side, only iterate over the one dimensional
To save memory accesses, the code generation usually generates a fused stream-collide step, so every PDF has to be loaded and stored only once per time step.
All together, lbmpy is now able to generate efficient sparse kernels for various velocity sets and collision operators, which can run on all common CPUs and NVIDIA and AMD GPUs.
3.4.1. Single node performance
In Figure 3, we present the performance of the generated sparse LBM kernel in comparison to the generated direct-addressing kernel. The diagram shows the mega fluid lattice updates per second (MFLUPs) depending on the porosity ϕ as defined in equation (9). The LB method uses a D3Q19 velocity set and the SRT collision operator. The benchmark measures the LBM kernel performance without boundary handling or communication, and it is performed on a single NVIDIA A100 GPU. Single GPU benchmark for sparse, dense and hybrid data structure with varying porosity on a NVIDIA A100 with 2563 cells, D3Q19 velocity set and SRT collision operator. The theoretical performance is calculated from the bandwidth of a streaming benchmark (1361 GB/s) and the theoretical number of memory accesses of the kernels, as LBM code is usually memory bound.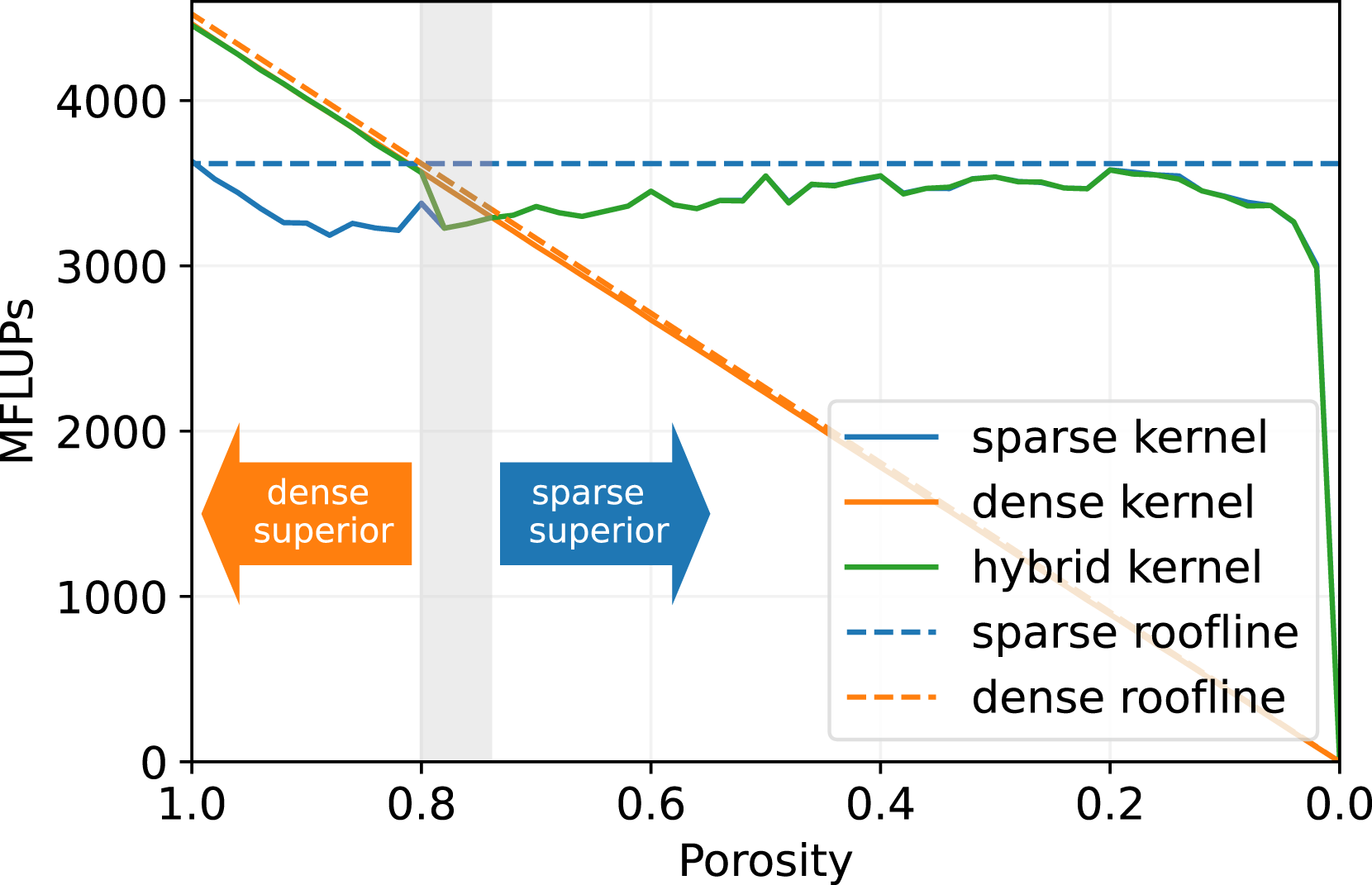
We observe that the single GPU performance of the sparse and the dense kernel is quite close to the theoretical peak performance. As the pure LBM stream-collide step is commonly limited by the memory bandwidth of the architecture, the performance of an efficient LBM kernel should be close to the theoretical peak performance, which is calculated from the memory bandwidth divided by the number of memory accesses needed for updating one LBM cell. This means that the presented kernels sufficiently saturate the memory bandwidth of the NVIDIA A100 GPU. Efficient LBM implementations with a performance close to bandwidth peak are also reported in Zacharoudiou et al. (2023-01), Lehmann et al. (2022) and Wittmann et al. (2013).
Furthermore, Figure 3 shows, that the MFLUPs performance for the sparse kernel remains essentially constant for decreasing porosity. However, we also observe, that the sparse kernel performs worse for ϕ ≥ 0.75. This is caused by the extra memory accesses of the sparse data structure. A dense kernel has to read and write every PDF of a cell per time step. This results in a memory access volume of 2Q ⋅ BPDF bytes per cell on GPUs, with Q as stencil size, here 19, and BPDF as the bytes per stored PDF, here 8 bytes for double precision. The sparse kernel, on the other hand, accesses 2Q ⋅ BPDF + (Q − 1) ⋅ Bidx bytes per cell, because it needs to read neighboring information from the
Nevertheless, the performance for the dense kernel decreases linearly when porosity decreases. The reason for this behavior is that dense kernels in
As indicated in Figure 3, the theoretical break-even point for sparse and dense kernels is approximately ϕ ∼ 0.75.
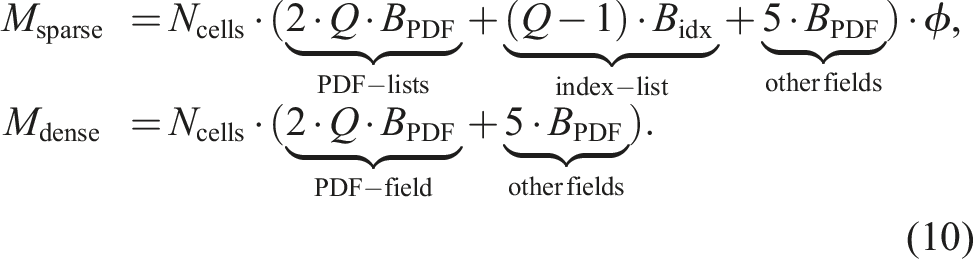
On the same NVIDIA A100 GPU we present a comparison of the memory consumption for a lattice of 2563 cells in Figure 4. The memory usage is measured with the NVIDIA monitoring tool nvidia-smi, showing that the sparse data structure consumes linearly less memory with decreasing porosity. On the other hand, the dense data structure exhibits a constant memory footprint because it stores all cells in the domain, regardless of whether a cell is fluid or boundary. For a porosity of 1.0, where all cells in the domain are fluid, the sparse data structure consumes more memory, since it also has to store the Memory consumption benchmark for 2563 cells on a single NVIDIA A100 GPU for D3Q19 stencil and pull streaming pattern. The theoretical memory consumption is calculated in equation (10).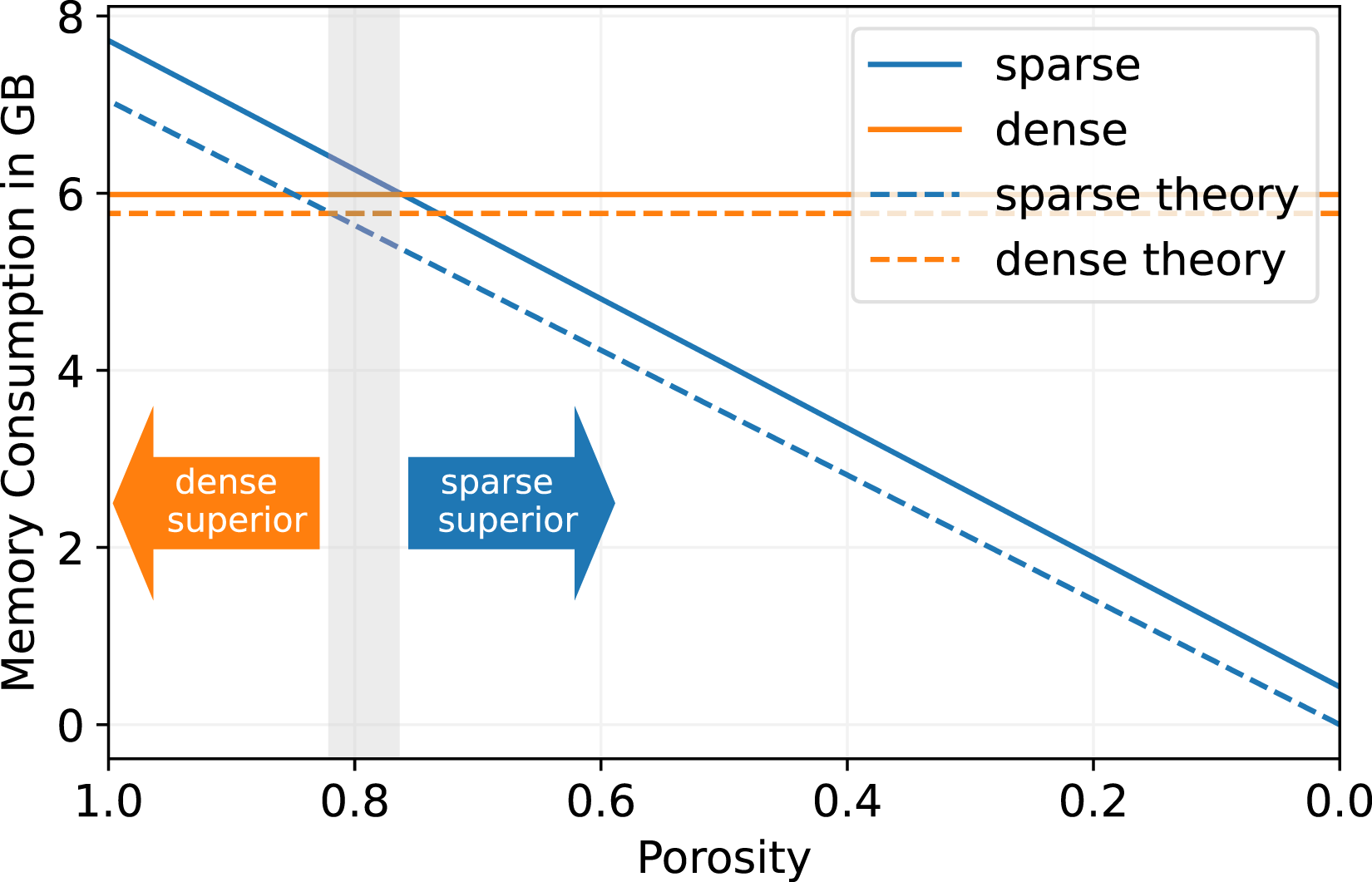
The additional fields stored are a velocity field (3D), a density field (1D), and a flag field to indicate boundary cells (1D).
We see that for the theoretical as well as for the measured memory footprint, the break-even point of the sparse and dense data structure is at a porosity of around ϕ ∼ 0.8, which is a similar result as for the performance comparison. For a higher porosity, the dense data structure is more suitable in terms of memory consumption, and for a lower porosity, the sparse structure becomes superior.
The measured memory consumption for the sparse as well as for the dense LBM in Figure 4 is close to the theoretical memory consumption, so that there is only a small overhead of less than 10% coming from other data structures than the necessary pure PDF data.
3.5. Hybrid data structure
In certain application scenarios, a hybrid data structure may be advantageous. As an example, consider a free flow over a particle bed as depicted in Figure 11. After the domain partitioning, some blocks contain only fluid cells, while other blocks consist primarily of non-fluid cells. In this case, neither the sparse nor the dense data structure seems to fit the given scenario perfectly.
Therefore, we implement the hybrid simulations in
In Figure 3, we display the performance for the hybrid data structure in green with ϕ
S
= 0.8. As expected, the performance reflects that of the dense kernel for ϕ ≥ ϕ
S
and, that of the sparse kernel for ϕ < ϕ
S
. Consequently, the hybrid approach can always reach the maximum possible
To our knowledge, the presented hybrid approach is a novelty for LBM frameworks, and can be very beneficial for increasing performance and reducing the memory consumption of an application. These benefits depend heavily on the sparsity of the domain.
4. Optimizations to sparse LBM
The high-performance framework
4.1. In-place streaming: AA pattern
The most common streaming patterns for LBM are the two-grid algorithms, where either PDFs of a cell are pushed into the neighbor cells (push scheme), or PDFs are pulled from the neighbor cells (pull scheme) (Wellein et al. (2006)). These algorithms have in common that a temporary PDF field is needed. This is because PDFs are stored in a different position than where they are read from. These two-grid streaming patterns read from PDF field A, then they propagate (push/pull) the PDFs, and, lastly, they store the propagated results at a different location in the temporary PDF field B. Therefore, these streaming methods are also called AB patterns. After the propagation, a field swap of fields A and B is needed.
The in-place streaming AA pattern on the other hand, enables writing and storing PDF values in the same positions of the PDF field, so that PDFs can be read from field A and also be written to field A without creating data dependencies. This saves the memory of the temporary PDF field B. Additionally, it also saves memory accesses. We integrated the AA streaming pattern in the code generation pipeline of lbmpy. For details about the functionality of the AA streaming pattern see Bailey et al. (2009).
Memory accesses per cell for Pull and AA pattern on CPU with size of PDFs Bpdf = 8 Byte, size of indices in the
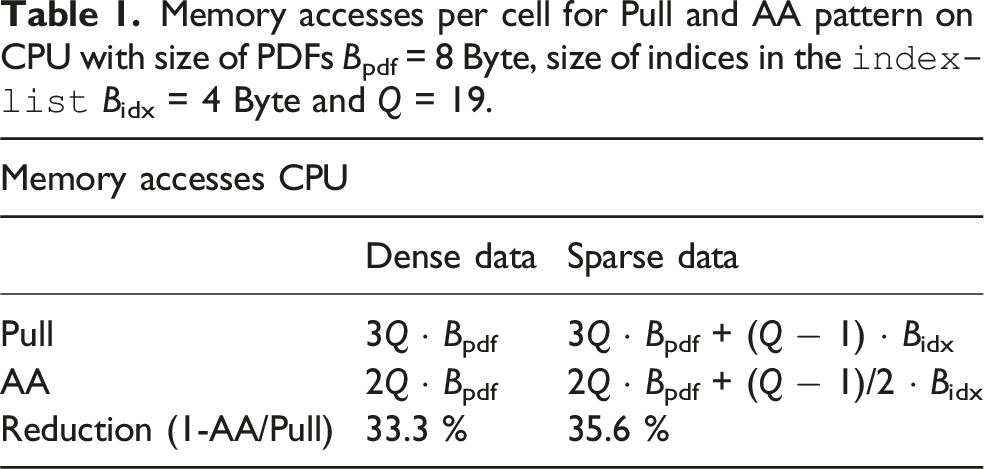
For sparse kernels, utilizing the AA pattern has even more advantages in terms of memory access. For the pull pattern, in addition to the
Therefore, this results in 2Q + (Q − 1)/2 memory accesses for sparse LBM kernels with the AA streaming pattern. We see in Table 1 that the AA pattern on a CPU reduces the memory accesses compared to the pull pattern by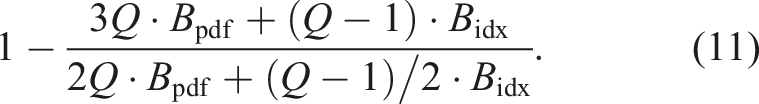
Because well-optimized LBM codes are usually memory-bound, an increase in the performance of the LBM by the same ratio can be expected.
Memory accesses per cell for Pull and AA pattern on GPU with size of PDFs Bpdf = 8 Byte, size of indices in the
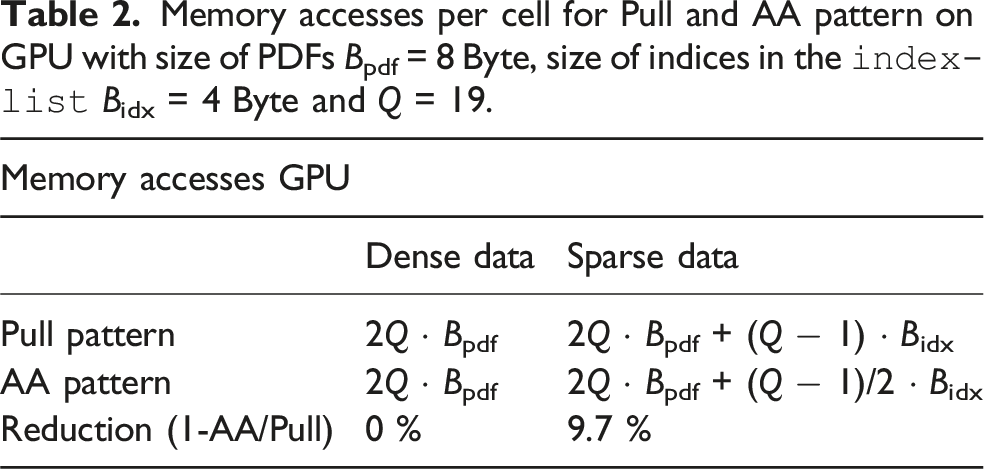
On the other hand, the condition to store the temporary PDF field can be avoided on CPUs and accelerators. So the memory consumption for the sparse LBM in equation (10) shrinks to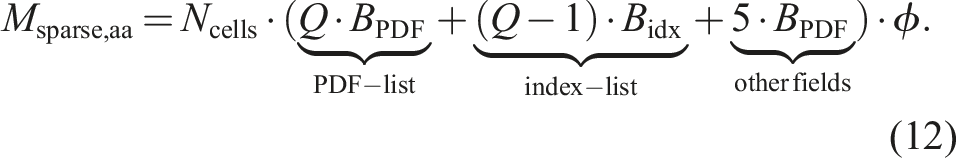
So for a D3Q19 stencil, double precision PDFs, and an integer
4.1.1. Benchmarking results
In Figure 5 the single GPU benchmarking results for a sparse LBM kernel on a NVIDIA A100 with a D3Q19 stencil and 2563 lattice cells are presented. We compare the pull streaming pattern with the AA pattern for various collision operators. The theoretical peak performance is calculated by the bandwidth, which is 1367 GB/s found by streaming benchmarks (Siefert et al. (2023)), and the number of theoretical memory accesses from Table 2. Single GPU Benchmark for pull versus AA streaming pattern for single relaxation time (SRT), two relaxation time (TRT), multi relaxation time (MRT), central moment (CM) and cumulant collision model on a D3Q19 stencil with 2563 cells on a single NVIDIA A100.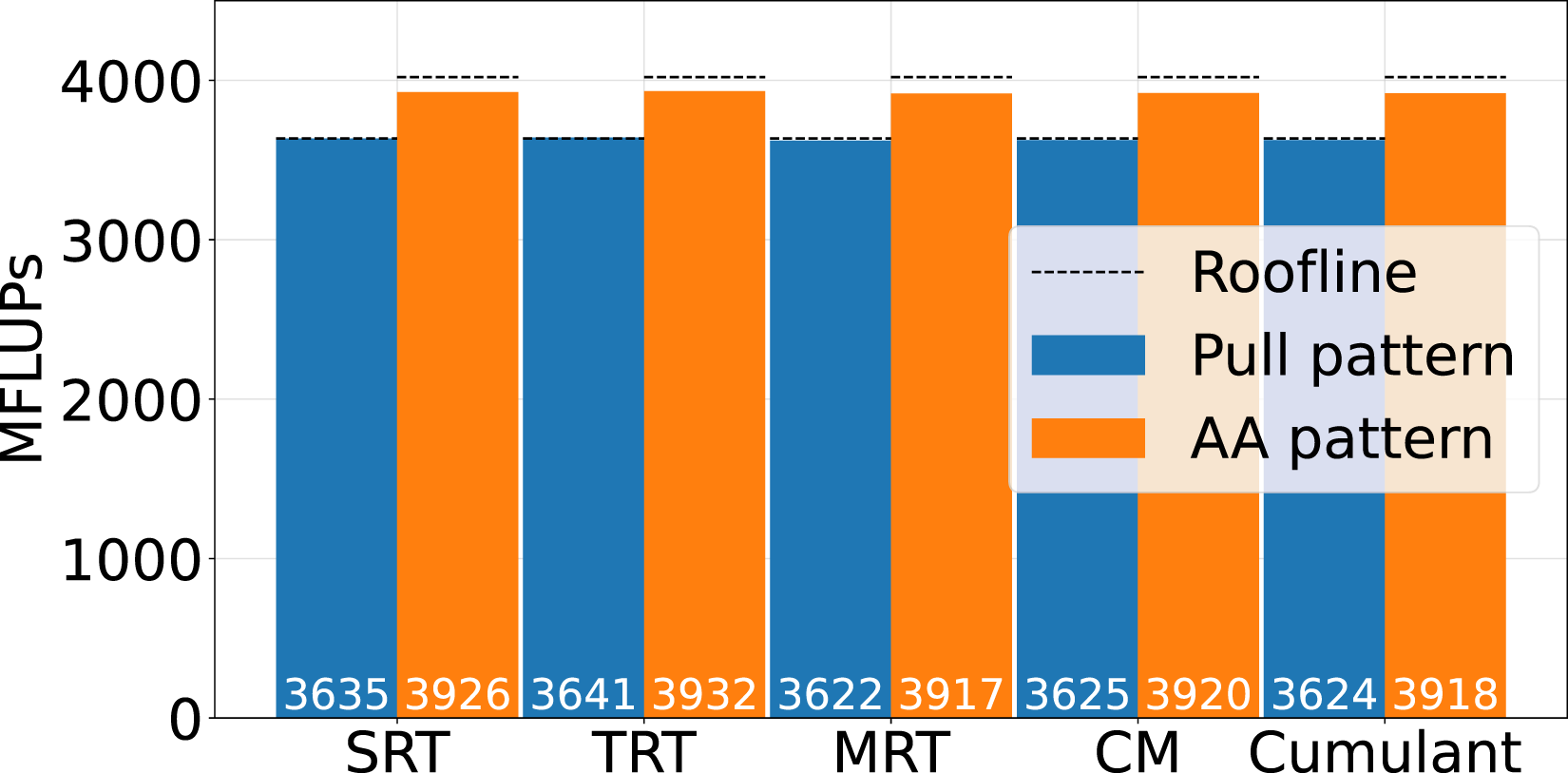
We observe, that the performance of both streaming patterns is close to the theoretical performance for all of the presented collision operators. The average performance increase of the AA pattern compared to the pull pattern on accelerators is
4.2. Communication hiding
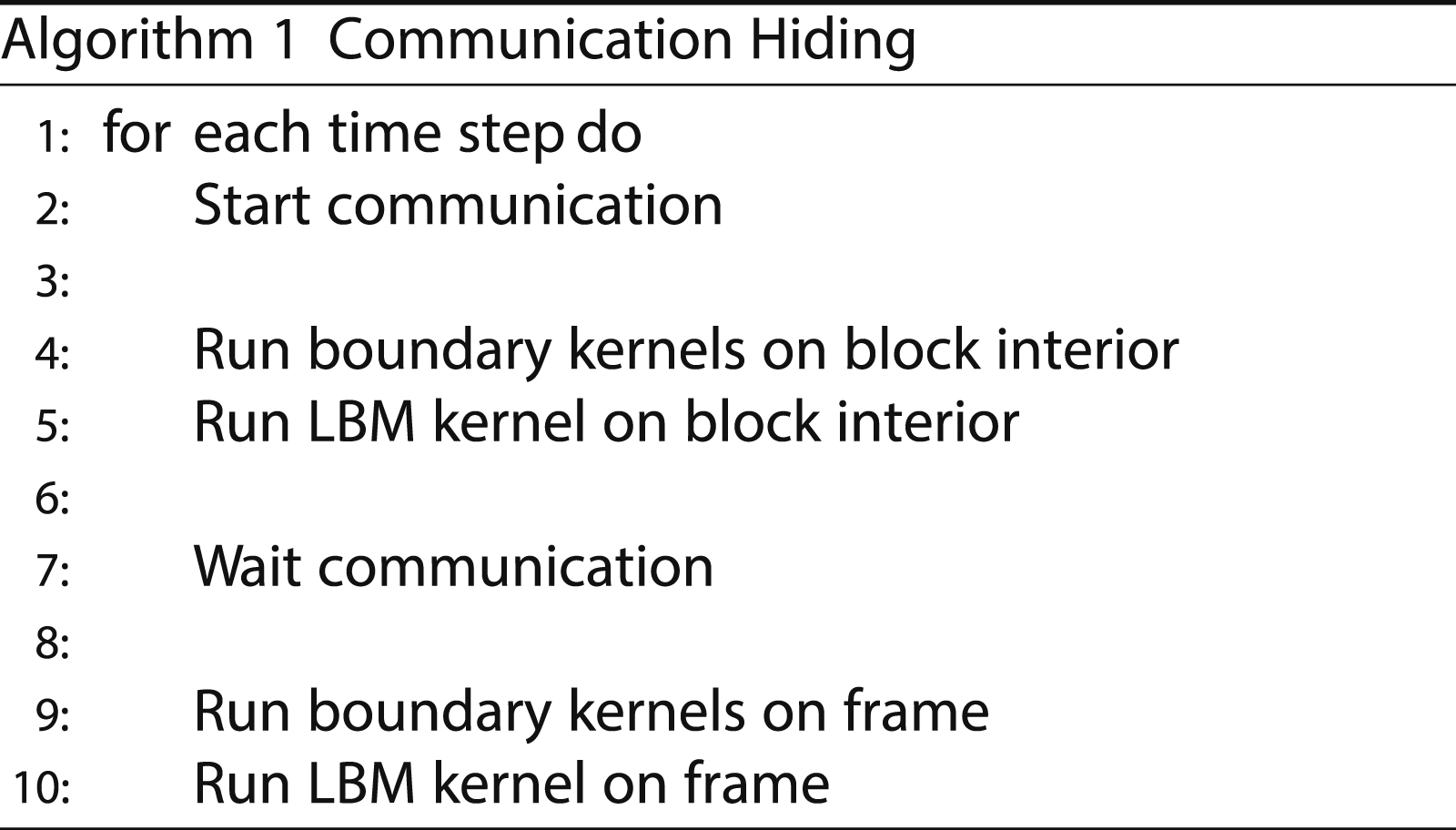
Communication hiding is used to overlap communication with computation. For this, the domain on every block has to be divided into a ”block interior” and a ”frame”, as illustrated in Figure 6. The frame only consists of the outermost cells, while the block interior consists of all other cells. An exemplary code to achieve communication hiding is shown in Algorithm subsection 1. At first, the communication is started. Every block packs its outermost PDFs in an MPI buffer and performs a non-blocking MPI-Send to its neighbors. Now, the block interior cells can be updated because the information from neighbour blocks is not needed for these cells. After this step, the algorithm must wait for the communication to complete and write the information of the MPI buffers to the ghost layers. Lastly, with the updated information in the ghost layers, the LBM and boundary kernels can now be executed on the cells of the frame. Subdivision of the PDF field in a frame and the block interior to enable communication hiding. In this example, the frame width is 5 in x, and 3 in y-direction.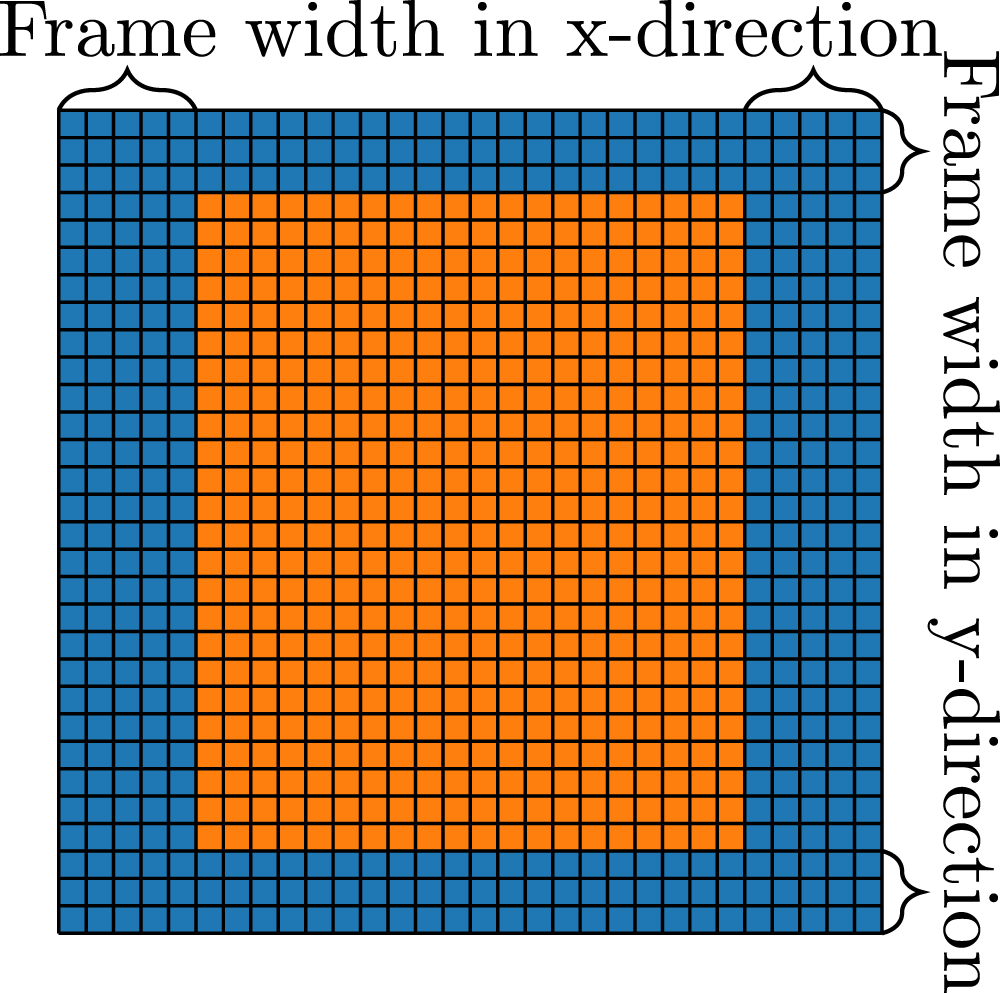
With this algorithm, the communication of the simulation can be overlapped with the kernel on the interior, which leads to higher performance because of better scalability on an increasing number of MPI processes. The width of the frame in all three dimensions has to be chosen suitably to achieve best possible performance. A thinner frame width would increase the number of cells in the interior, providing more time to overlap the communication. On the other hand, a small frame width results in small kernels. Especially on GPUs, small kernels can not fully utilize the GPU, which can lead to performance drops. Additionally, consecutive memory access in one dimension is not possible for a thin frame, which can also reduce the simulation’s performance.
4.2.1. Communication hiding for sparse data structures
Implementing communication hiding for a sparse data structure is not straightforward because there is no topological information for the cells. This means that a cell has no direct information about whether it is inside the block interior or part of the frame. To compensate this, we store two additional index lists, one for the interior and one for the PDFs on the frame. These index lists are initialized by the flag field at the start of the simulation, where spacial information of cells is still present. Furthermore, the pull index for the center PDF of every fluid cell must be stored. This index is used to get the correct write access for kernels on the interior and frame cells. These modifications on the list structure are integrated into the code generation so that they can be turned on and off and allow generated code to run on different architectures.
In the LB framework HemeLB, a comparable communication hiding approach is implemented (Carver et al. (2012); Shealy et al. (2021)).
4.2.2. Scaling results
In Figure 7, the weak scaling of the sparse data structure on the JUWELS Booster HPC cluster is presented. JUWELS Booster is currently place 21 of the Top500 HPC systems (June 2024) and consists of 936 compute nodes, each equipped with 4 NVIDIA A100 GPUs (see Alvarez (2021)). Weak scaling benchmark on NVIDIA A100 GPU cluster JUWELS Booster with different configurations for the communication hiding. The roofline is obtained by a stream benchmark (Siefert et al. (2023)). The runs are executed with 3203 cells per GPU, with a D3Q19 stencil and SRT collision model on an empty channel setup.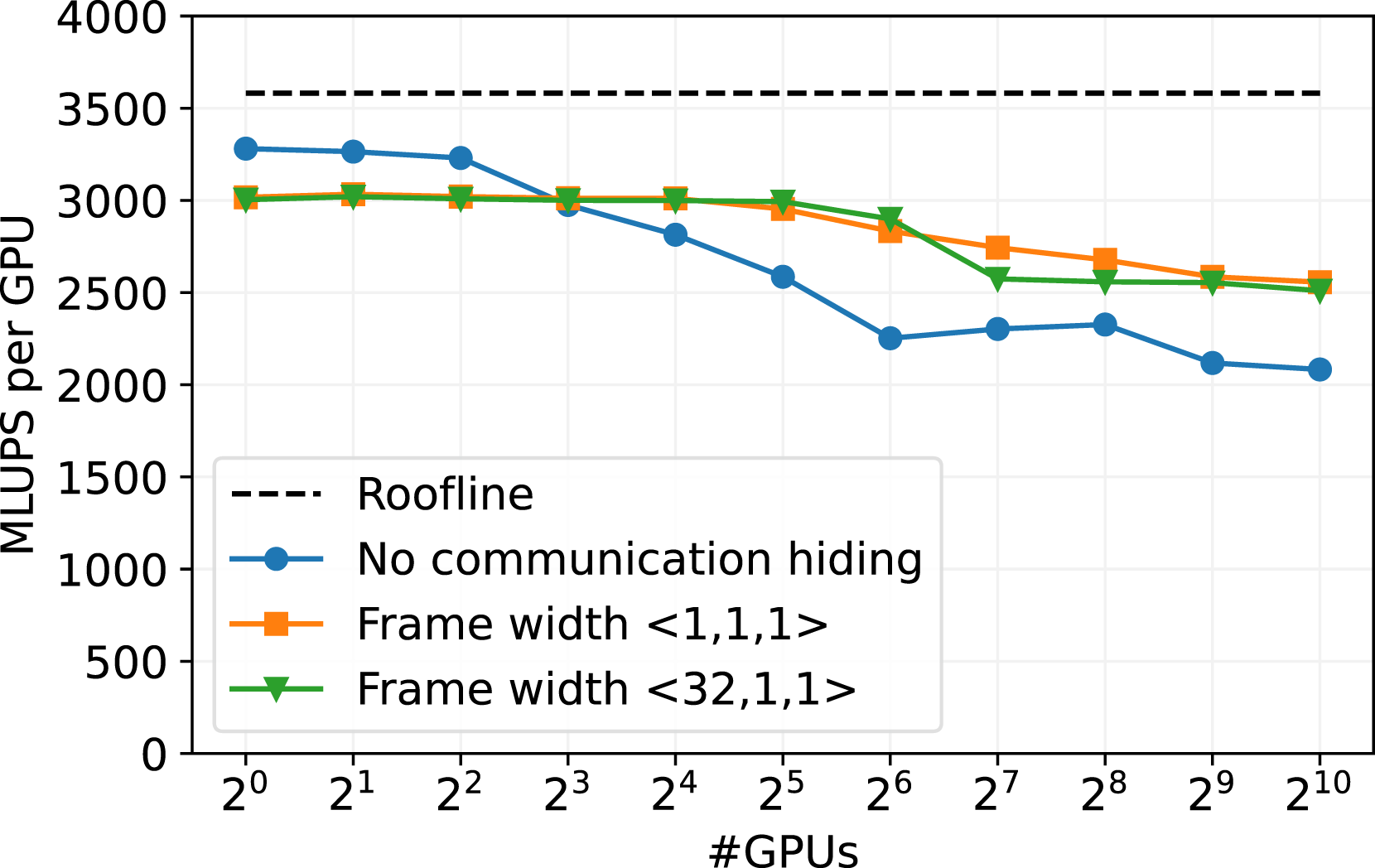
We tested three versions of the communication. One is without communication hiding, one with the minimum frame size of one cell in every direction, and one with a frame thickness of 32 cells in x direction and one cell in y and z direction. This option is promising, since consecutive memory accesses are still enabled in x-direction, while the frame size is still small enough to allow a good communication overlap. In general, a smaller frame size increases the work of the kernels on the interior cells and, therefore, should increase the effectiveness of the communication hiding. On the other hand, the GPU utilization of the kernels on the frame is quite low for a small frame size, and consecutive memory accesses are not secured.
The version without communication hiding performs best up to one node (4 GPUs), see Figure 7. There the intra-node communication speed is quite high since we can exploit the high bandwidth of NVIDIA GPU-to-GPU connections. However, the performance without communication deteriorates for more than 4 GPUs when the inter-node communication speed becomes relevent.
The benchmark runs with communication hiding start with worse performance on single node, as the overhead of the kernel call on the frame cells limits the performance. Nevertheless, these versions exhibit excellent scalability for up to 32 GPUs. Beyond 32 GPUs, the performance drops to 83% scaling efficiency on 1024 GPUs. This can possibly be explained by the InfiniBand network architecture of JUWELS Booster, which is implemented as a DragonFly + network. The drop in performance could be caused by the need for communication between different switch islands of the system when more than 32 GPUs are employed.
In these cases, the size of the frame only has a negligible impact. The two scenarios for communication hiding behave similarly. The smaller frame size of
Additionally, we tested the scaling efficiency on the GPU partition of LUMI, which is in the top 5 on the current Top500 list (June 2024). The HPC cluster comprises 2978 nodes with 4 AMDMI250X GPUs per node. Further, every AMD MI250X GPU consists of two Graphical Compute Dices (GCDs) (Pearson (2023)), so we create one MPI process per GCD and show the scaling over the GDCs. As already studied in Holzer et al. (2024), Lehmann (2022) and Martin et al. (2023), it seems not to be possible to achieve significantly better performance than equivalent to approximately 50% of memory bandwidth for LBM codes on a single AMDMI250X. We observe the same behavior in Figure 8. Weak scaling benchmark on GPU cluster LUMI-G with different configurations for the communication hiding. The roofline is obtained by a stream benchmark (Siefert et al. (2023)). The AMD MI250X GPUs have two compute chips per GPU (GCDs). The runs are executed with 2563 cells per GCD, with a D3Q19 stencil and SRT collision model on an empty channel setup.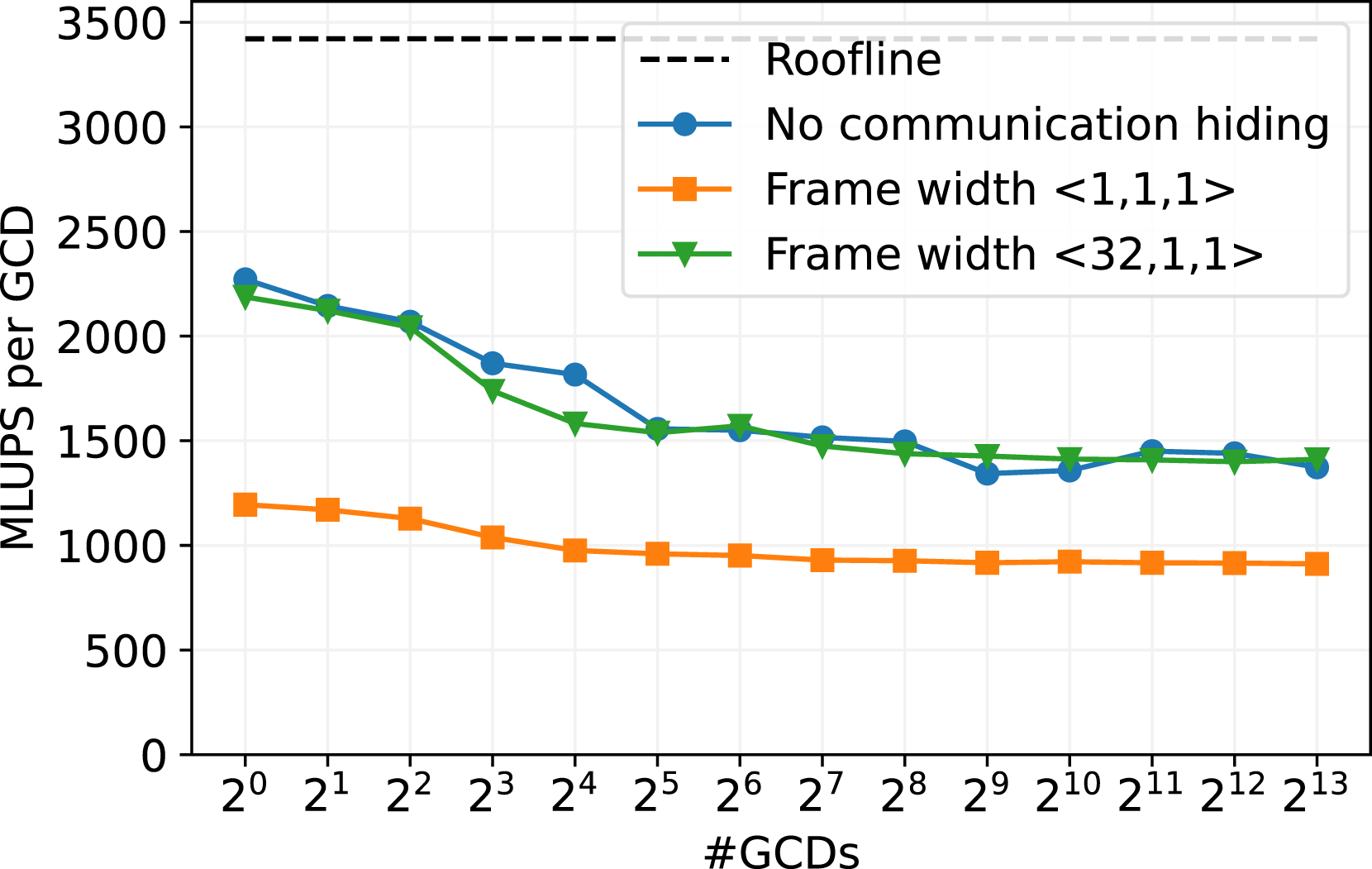
We tested the three communication routines similar to the benchmarks on JUWELS Booster. For the runs without communication hiding, the scaling behavior is similar as for the larger frame size of
5. Applications
To evaluate the performance of the sparse data structure in a more realistic scenario than the artificial porosity benchmark as in Figure 3 or the weak scaling of an empty channel on JUWELS Booster (Figure 7) and on LUMI (Figure 8), we set up three different applications. The first is a flow through a porous medium consisting of a stationary particle bed. The second one is an extended version of the first application, where the bottom part of the domain consists of the same particle bed, while the upper domain is a free flow, such that we simulate the interaction of a free flow with a porous sediment bed. The last application is the flow through a geometry of coronary arteries, which also results in a complex and sparse domain.
5.1. Flow through porous media
The efficient simulation of fluid flow through porous media is an ongoing research topic, for example in Pan et al. (2004), Yang et al. (2023), Han and Cundall (2013) or Ambekar et al. (2023), to mention a few. For this porous application, we generated a particle bed with the Flow through a particle bed consisting of 21580 particles of 0.0041 m diameter, which corresponds to an average porosity of 0.356663. The size of the domain is 0.1 m in every dimension.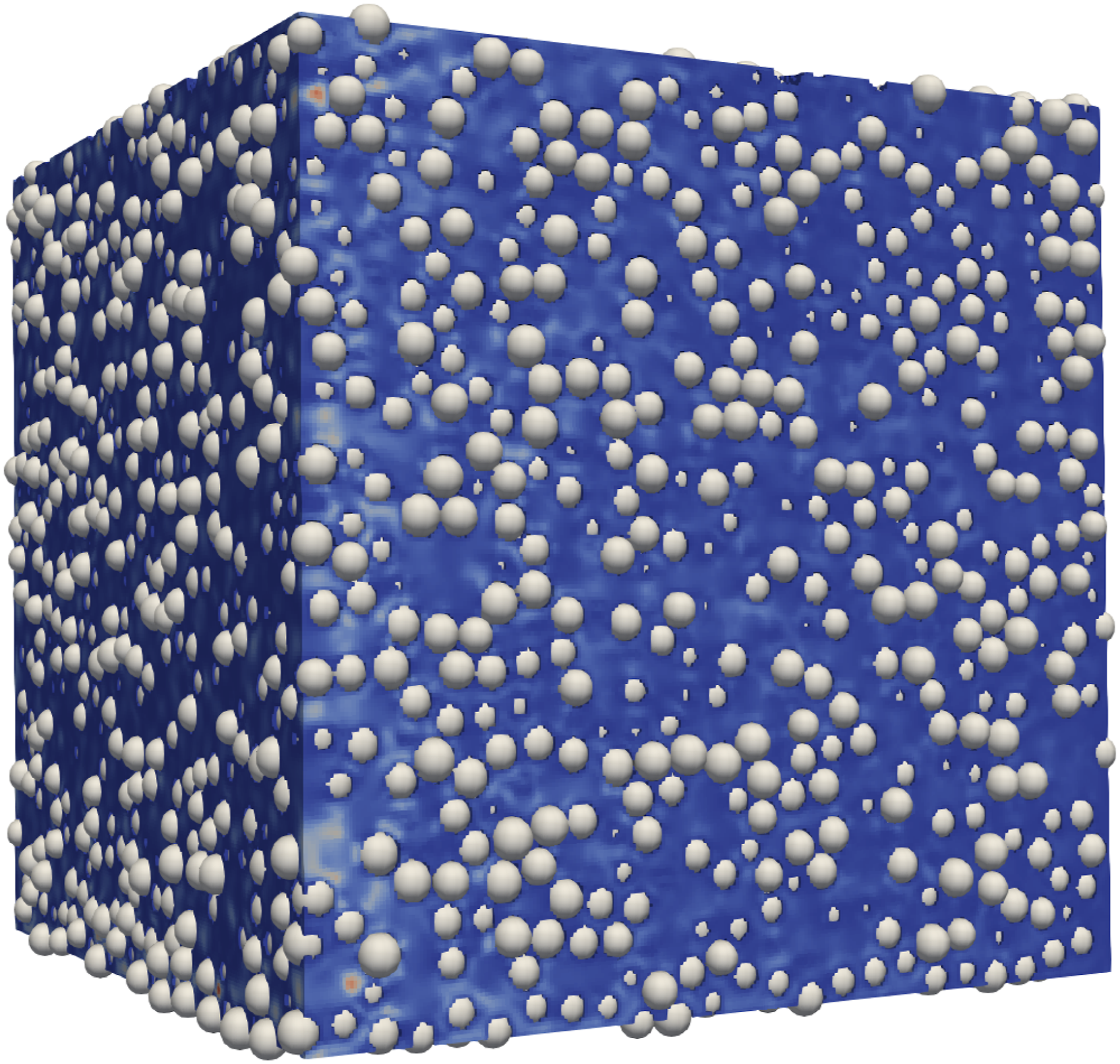
We decomposed the domain with 64 blocks in a 4 × 4 × 4 arrangement to run on 64 NVIDIA A100 GPUs on JUWELS Booster. While we fixed the number of blocks to 64, we increase the cells per block and therefore also the resolution of the domain and the number of total cells, as shown in Figure 10. We performed this benchmark for the sparse data structure, and compare it to the dense data structure. Comparison of the sparse and the dense data structure for the flow through the particle bed in Figure 9. The number of 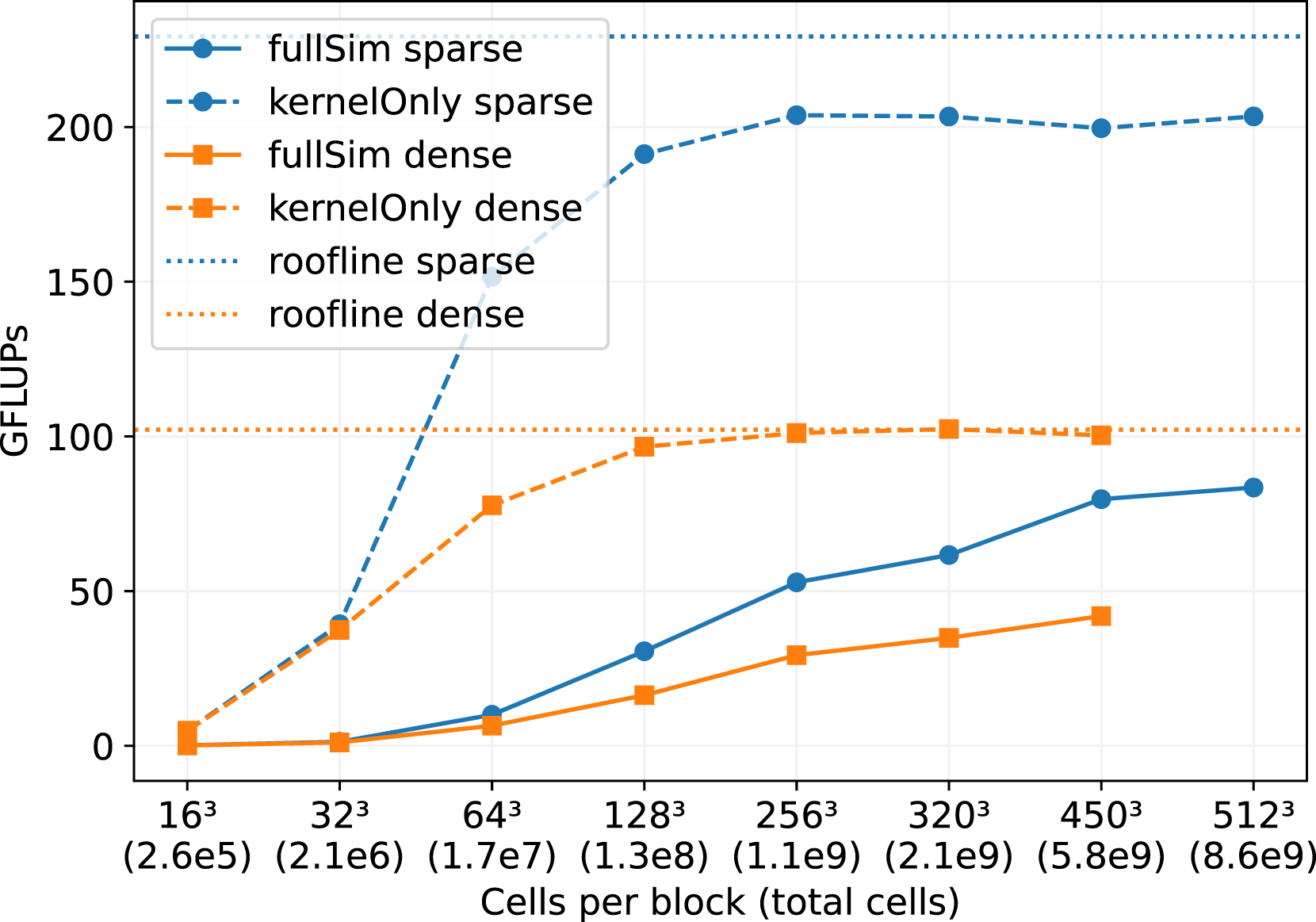
5.1.1. Kernel-only performance
We first focus on at the ”kernel-only” results in Figure 10, which only show the LBM kernel performance without handling boundaries or performing communication. These results give good insights in the raw performance improvements that we achieved by implementing the sparse data structure, because the performance of the full simulation may be dominated by communication or boundary handling routines. We observe, that for block sizes below 643, the GPU utilization is too low to achieve good performance. Both, the sparse and the dense kernel saturate at a block size of 2563. Further, we note that the performance of the sparse data structure is approximately two times higher than for the dense structure. This also fits quite well to the results in Figure 3 for a porosity of ϕ ∼ 0.35. The sparse kernel-only performance does not quite reach the theoretical bandwidth limit. This could be caused by some in-balances, since the porosity varies between the blocks, from a minimum of 0.337816 to a maximum of 0.392441. This means, that some blocks, and therefore processes, have more workload in terms of cells. We measure the performance at the end of the simulation run, when all processes finished their work, so the performance is determined by the slowest processor. The performance of the dense kernel on the other side is not affected by the porosity differences, and therefore performs exactly the same work on every MPI process and thus does not suffer from load in-balance issues.
5.1.2. Full-simulation performance
When running the full simulation including boundary handling and communication routines, the overall performance shrinks by at least 50% for both data structures compared to the kernel-only performance. For sparse simulations, the boundary handling has only a small effect on the performance, as the boundary conditions for the particles are implicitly handled by the streaming step (see section about sparse boundary handling). So the main factor is the communication of ghost layer cells between the MPI processes. This communication overhead can be optimized, for example by communication hiding as discussed before, but it can not be avoided completely.
For the full-simulation results in Figure 10, we again observe low performance for small block sizes, which can be explained by the low utilization of GPUs. However, this time, there is no saturation for a block size of 2563 because a more extensive block size results in better communication hiding when the ratio between computational work and communication improves in favor of computational work. For the dense structure, it was not possible to perform simulations with a block size of 5123, as the memory consumption exceeded the 40 GB of the GPU RAM of a single A100. For the sparse structure, this is not a problem, as for a porosity of 0.356,663 we save around 50% of memory compared to the dense structure.
For this application, we significantly benefit from the sparse data structure, as it achieves a performance increase of
5.3. Free flow over river bed
The second application is the simulation of a free flow over a river bed similar to Kemmler et al. (2023) or Fattahi et al. (2016). In Figure 11, the bottom part of the domain consists of the same porous medium/particle bed as the first application in Figure 9, with the same average porosity of about Free Flow over a particle bed. The porosity of the blocks in the particle bed on the bottom (blue blocks) have a porosity of about 0.35, while the upper blocks (red blocks) consists of fluid cells only.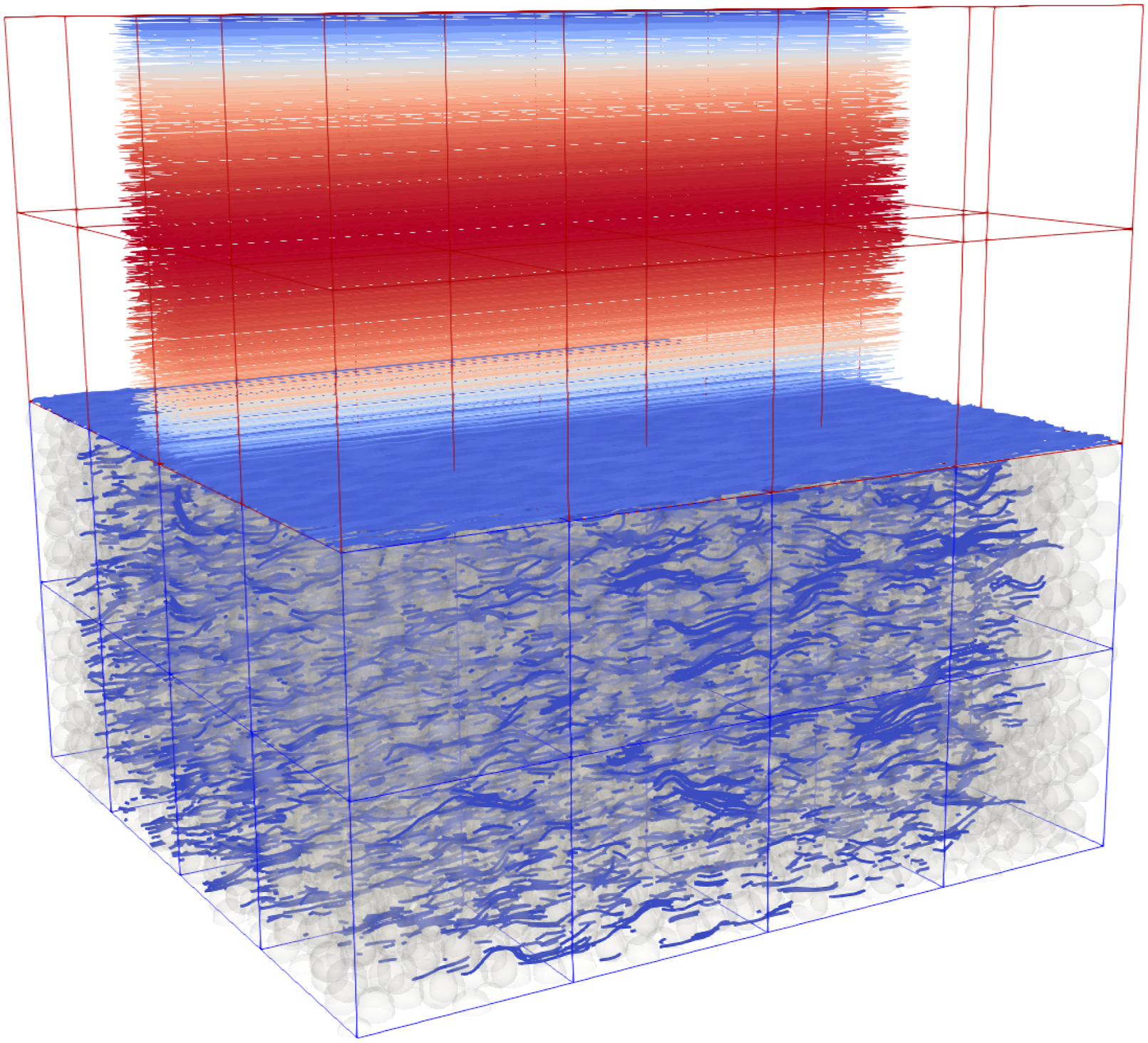
This application is suitable for utilizing the hybrid data structure. As already indicated in Figure 11, the blocks in the upper part of the domain should hold their data in a dense structure (red blocks). In contrast, the blocks in the porous part of the domain should be stored with a sparse data structure (blue blocks). The framework includes the functionality to select the appropriate data structure for each block, the user only has to specify an appropriate porosity threshold.
In Figure 12, a comparison of the sparse, the dense and the hybrid data structure is shown. The simulations are again executed on the JUWELS Booster GPU cluster, with a fixed number of NVIDIA A100 GPUs and one MPI process per GPU. The performance of the raw LBM kernel (kernel-only) is plotted, as well as the entire simulation run, including communication between the MPI processes. We fixed the number of GPUs to 64 as well as the problem size to 109 cells and only vary the cells per block, resulting in a high number of blocks for small cells per block and one block per GPU for the largest block size of 2563 cells. Comparison of the sparse, dense and hybrid data structure for the free flow over a particle bed in Figure 11 on 64 NVIDIA A100 GPUs on Juwels Booster. The problem size is fixed to 1.07∗109 cells, while the number of cells per block and therefore also the number of blocks vary. The LBM kernel-only performance is shown as well as the performance of the whole simulation including boundary handling and communication between MPI processes.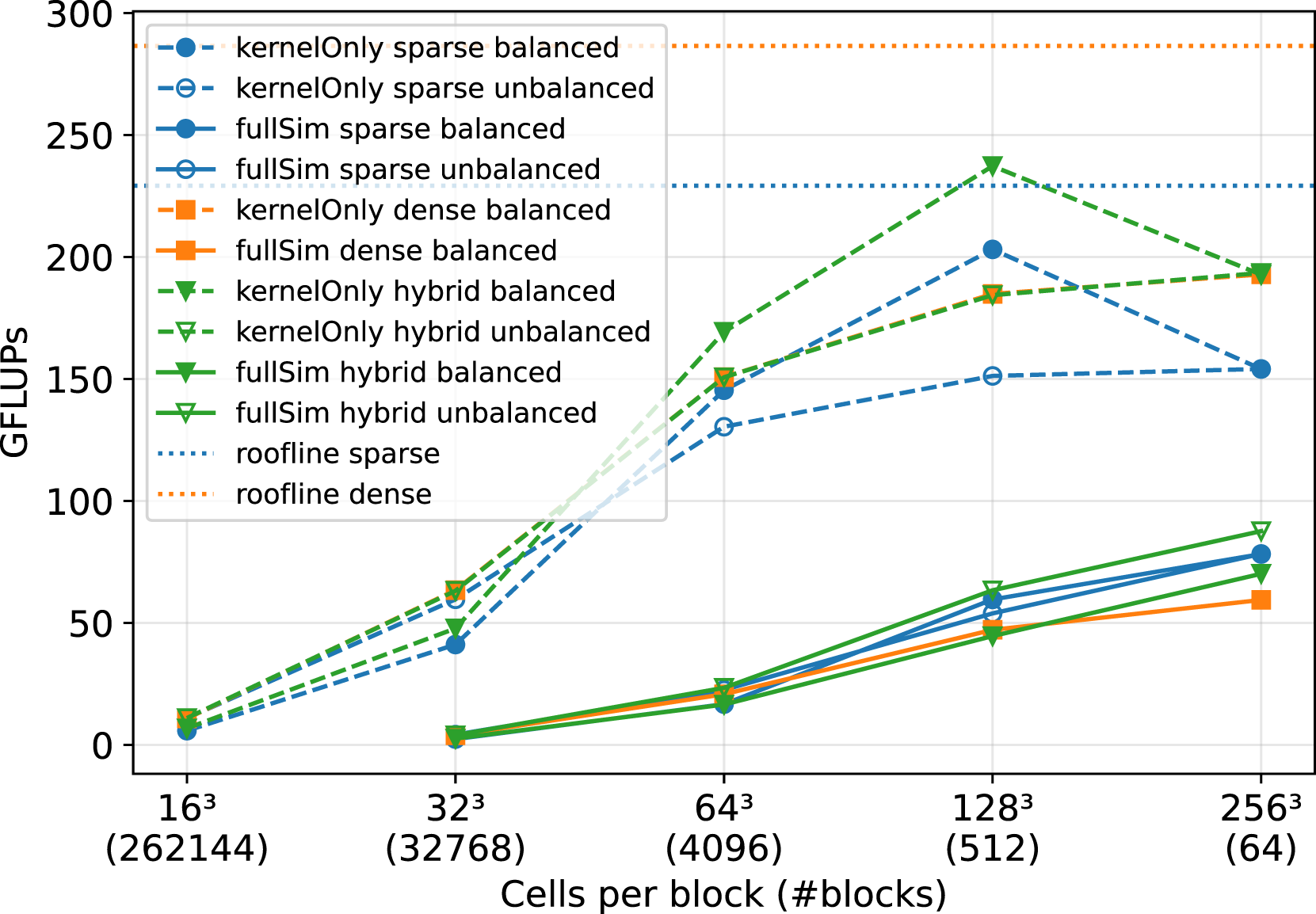
Evaluating the raw kernel performance first, we again observe that a more significant number of cells per block leads to a better utilization of the GPU.
Load-balancing issues can emerge when using a sparse or a hybrid data structure with multiple blocks per GPU. This is because the block partitioning of a domain can lead to a wide span of porosity values on the blocks. When decomposing the river bed simulation in Figure 11 with sparse blocks only, this results in half of the blocks yielding a porosity of 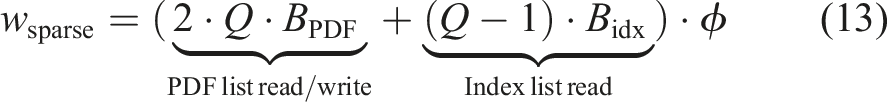
The workload for the dense block on a GPU is
For the simulation run with sparse data blocks only, the workload of the blocks differs significantly depending on their porosity. Therefore, we employ a load-balancing algorithm to balance the blocks over the MPI processes to reach a better workload distribution. We used a space-filling-Hilbert-curve approach, as described in Schornbaum and Rüde (2018).
5.3.1. Kernel-only performance
In Figure 12, we observe that the load-balancing works well for the sparse kernel-only runs. Especially for the block sizes of 643 and 1283, the load-balancing seems to reduce the workload unbalance significantly and, therefore, increases the performance. For 2563 cells per block, there is only one block per GPU, so no load-balancing is possible in this case.
No load balancing is necessary for the dense data structure, as every block has the same workload. We see that the kernel-only runs of the dense data structure outperform the unbalanced sparse runs. This is because the sparse data structure is slower than the dense structure on the blocks with a porosity of 1.0 (see again Figure 3), and all MPI processes must wait for the processes holding these blocks. However, when balancing the workload for the sparse kernels, at least for a block size of 1283, we can achieve a higher performance than the dense kernels.
The kernel-only runs for the hybrid data structure without load balancing closely follow the results of the dense kernels in Figure 12. Without load balancing, the low performance of the dense blocks in the porous region heavily dominates the runtime. However, here we can balance the workload to achieve better results. So for the kernel-only runs, using the workload-balanced hybrid data outperforms the other kernels for all block sizes. The only exception is for 2563 cells per block. There no load-balancing is possible, because there is only one block per GPU.
5.3.2. Full-simulation performance
Comparable to the particle bed, the performance of the full-simulation results in Figure 12 is again heavily trimmed by the communication routines, which is not avoidable. For all data structures, the performance increases with increasing block sizes, as the number of blocks and therefore the necessary communication decreases.
Workload per MPI process/GPU before and after load balancing for the hybrid data structure run with 1283 cells per block in Figure 12.
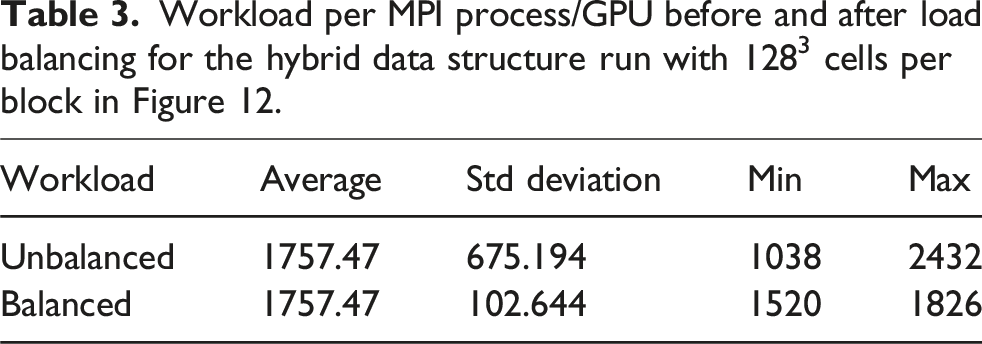
Nevertheless, for a complex domain setup such as a free flow over a riverbed, we also manage to improve the application’s performance by utilizing the sparse or the hybrid data structure. Additionally, we note the reduced memory consumption as presented in Figure 4. Overall, when half of the domain consists of a porous medium, such as in Figure 11, we save about 25% of memory with the hybrid data structure.
5.4. Coronary artery
Finally, we present performance results for the flow in a coronary artery. This is a topic of high interest in medical engineering and has been studied, for example, in Axner et al. (2009), Afrouzi et al. (2020), Bernaschi et al. (2010) and Godenschwager et al. (2013). The flow in a coronary artery results in a complex and highly sparse domain. An example setup for this application is shown in Figure 13. While the whole domain would consist of a very high number of blocks, we can discard all blocks that do not contain fluid cells. Still, the remaining blocks have a very low porosity. Of course, when lowering the size of the block, the block structure would converge better to the geometry and result in higher block porosities. However, small block sizes also lead to under-utilization of GPUs, as already shown in Figure 10. Domain partitioning of a coronary artery with 1.2 ⋅ 108 fluid cells, 531 blocks and 1283 cells per block.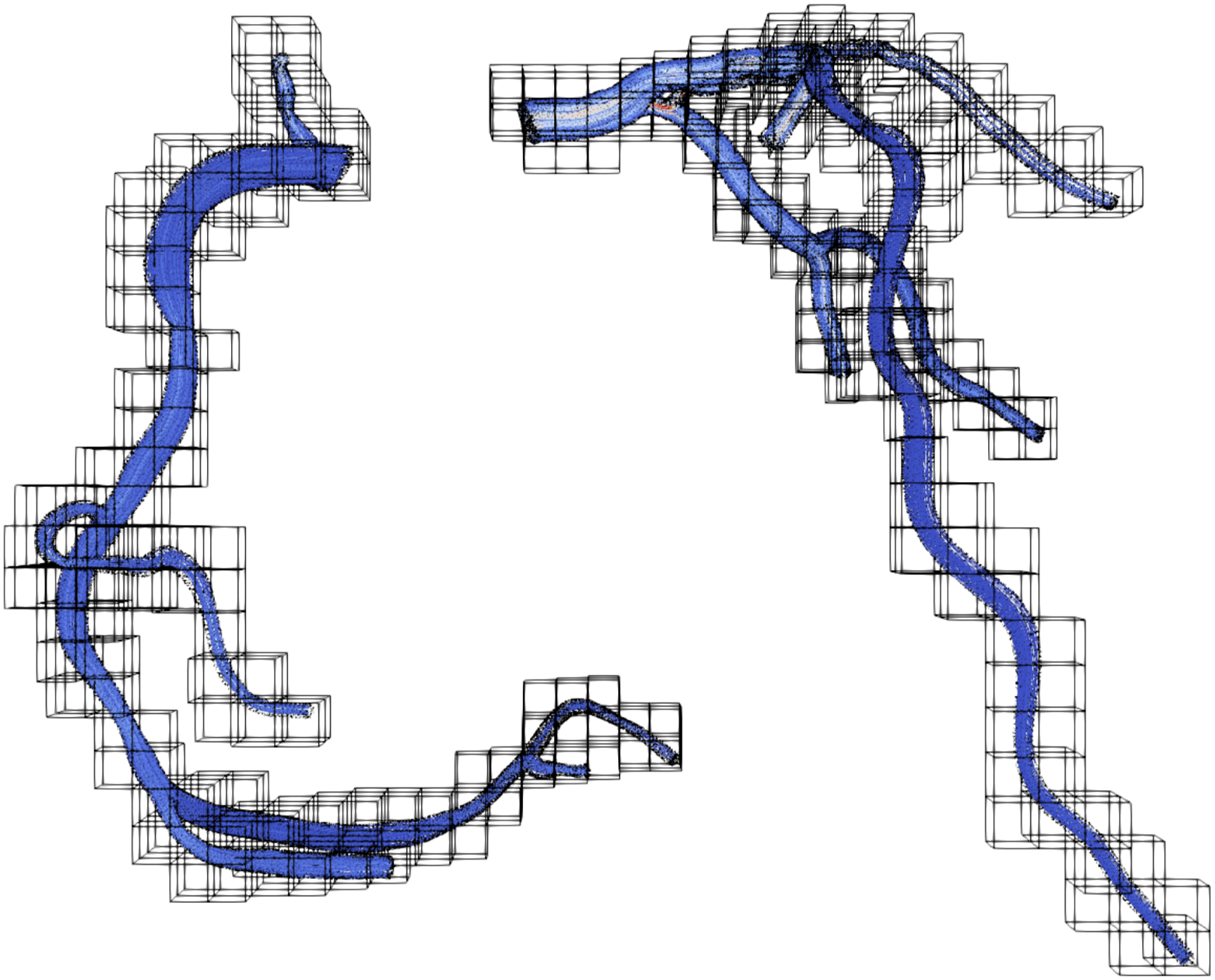
In Figure 14, we compare the performance of the sparse and the dense data structure on JUWELS Booster. We fixed the number of NVIDIA A100 GPUs to 60 and the problem size to 1.2 ⋅ 108 fluid cells while varying the block size and, therefore, also the number of blocks. Comparison of the sparse and the dense data structure for the artery flow in Figure 13 on 60 NVIDIA A100 GPUs on JUWELS Booster. The problem size is fixed to a number of 1.2 ⋅ 108 fluid cells, while the block size and therefore also the number of blocks varies.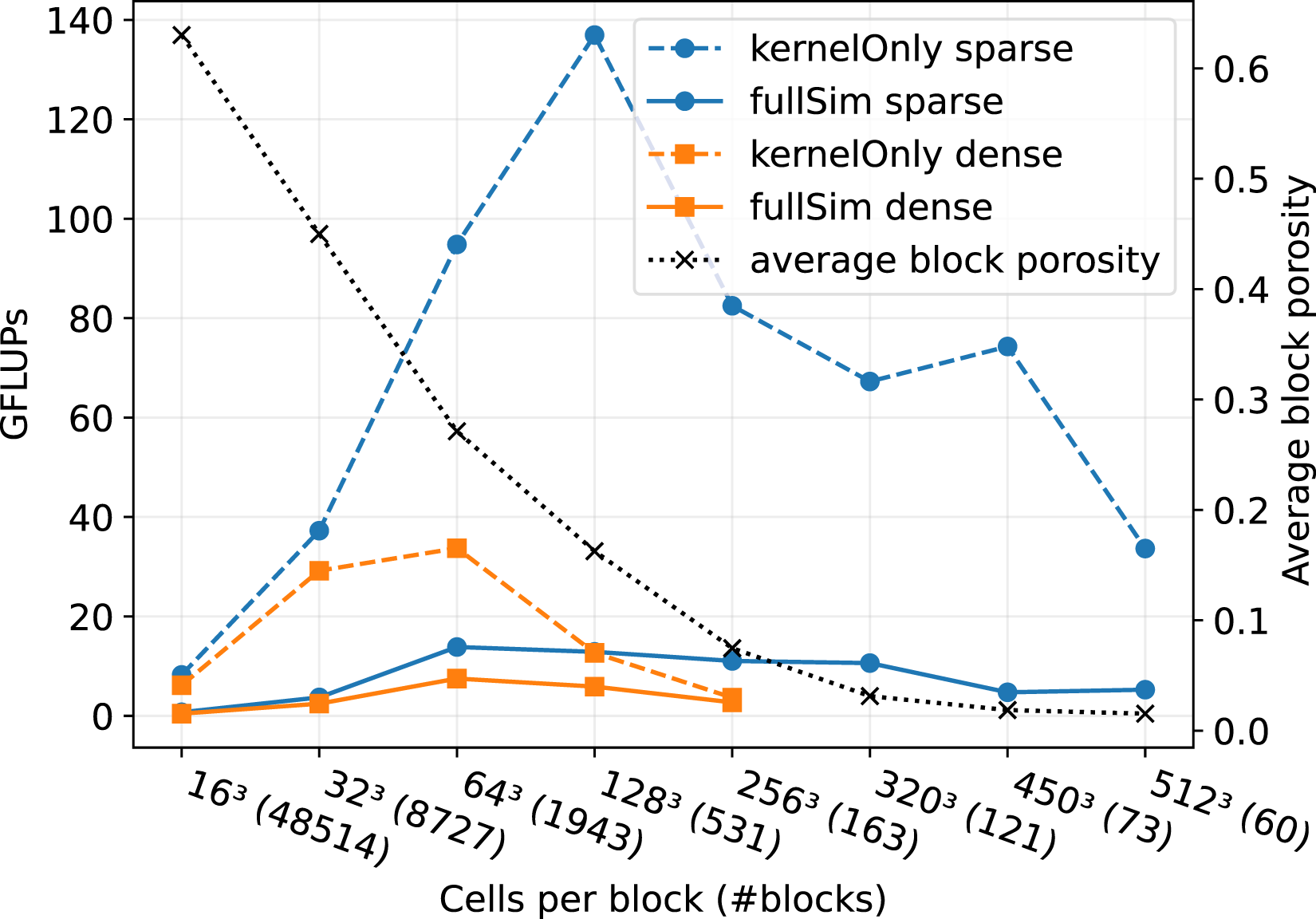
5.5. Kernel-only performance
Again, we observe the behavior of a small block size, which results in low performance for the kernel-only runs of the sparse data structure. We experience that increasing the block size up to 1283 cells per block leads to higher performance of the raw sparse LBM kernel. For block sizes larger than 1283, the porosity drops below ϕ < 0.1. We observe in Figure 3 that the sparse LBM performance starts to deteriorate for a porosity smaller than 0.1. Therefore, for block sizes larger 1283, the performance of the sparse kernel in Figure 14 shrinks because of the low block porosities. The sweet spot between large block sizes for good GPU utilization and a porosity greater than 0.1 for good kernel performance is a block size of 1283 cells per block for this setup.
5.5.1. Full-simulation performance
The performance for the sparse full simulations is relatively stable for moderate block sizes. This is because the performance is heavily dominated by communication, which is especially high in this application because of the high number of blocks per MPI process.
Nevertheless, the sparse data structure shows significantly higher performance than the dense data structure for all block sizes. This is true for the kernel-only runs and the full simulation runs. For a block size of 1283 cells, we achieve a speed-up of about 11 for the kernel-only run and still a speed up of 2 for the full simulation.
5.5.2. Memory consumption
Also remarkable is the amount of memory we can save for the artery setup. So when choosing a block size of 1283 cells per block to reach maximum raw performance; then, we end up with an average block porosity of ϕ ∼ 0.16. This means that according to Figure 4, we save about 75% of memory when using the sparse instead of the dense data structure. We were not able to acquire results for the dense structure for block sizes greater than 2563 because the NVIDIA A100 RAM ran out of memory.
6. Discussion
Parts of this work are comparable to Martin et al. (2023). In this article, the framework HARVEY is ported to different programming models such as CUDA, HIP, SYCL and Kokkos. The authors compare the performance on different hardware, among others also on NVIDIA A100 and AMD MI250X GPUs. HARVEY is a LBM based CFD software with a focus on simulations of blood flow in patient-derived aortas, also utilizing the indirect addressing approach.
Martin et al. (2023) measure the performance of HARVEY and a proxy app for NVIDIA A100 GPUs on the HPC System Polaris
3
. Their proxy app is used to show the maximum achievable performance for simple test cases. Their results of the proxy app for an empty channel flow on one node (4 NVIDIA A100 GPUs) reaches around 12,000 MLUPS, equivalent to 3000 MLUPS per A100 GPU. This is close to what we achieve, as shown in Figure 7 for the communication-hiding cases. The actual HARVEY framework performs slightly worse for the same empty channel with around 2250 MLUPS per GPU. On the maximum scaling size of 1024 NVIDIA A100 GPUs, the proxy-app reaches around 104 MLUPS, equivalent to 976 MLUPS per GPU. The HARVEY framework achieves around 6∗104 MLUPS, equivalent to 585 MLUPS per A100 GPU. In Figure 7
When comparing the results for the artery geometry with a resolution of 55 μm, HARVEY achieves around 4∗104 MLUPS for 64 A100 GPUs, so 625 MLUPS per GPU. In Figure 14
Further, Martin et al. (2023) show comparable results on the AMD MI250X GPUs, which were tested on the Frontier
4
HPC system. We compare the results for 4 GCDs. The proxy app of shows around 2250 MLUPS per GCD, while
Also comparable are the results from Zacharoudiou et al. (2023-01). The authors conducted blood flow simulations with their LBM framework HemeLB on various HPC systems including JUWELS Booster. They were able to achieve a performance of around 200 MLUPS per GPU on the NVIDIA A100 GPUs of JUWELS Booster with a comparable number of cells per GPU
7. Conclusion
In this article, we presented the benefit of sparse LBM kernels, especially when using accelerator cards. We presented the integration of the sparse data structure into lbmpy, to be capable of generating efficient compute kernels for various architectures. We compared the sparse and the dense data structure on a single GPU and found that for a domain porosity of
The sparse kernels show excellent performance for the pull streaming pattern for various collision operators such as single-/two-/multi-relaxation times, central moments, or cumulants on a single GPU. We managed to further increase the performance by
We proposed a novel hybrid data structure, that is able to flexibly switch between the sparse and the dense approach, depending on which subdomains are handled more efficiently with the corresponding data structure.
We set up a porous media flow simulation and achieved a speed-up of 1.9 and reduced the memory consumption by 50%. For an artery blood flow simulation, we gained a speed-up of 2 dependent on the block sizes and achieved a decrease of memory consumption of about 75%. We experienced imbalances in the distribution of the work over the MPI processes when using the sparse data structures. To maximize the efficiency of the sparse LBM, we employed load balancing, but more research is needed to fully optimize the workload distribution while maintaining the spacial locality of the neighboring blocks on neighboring processes.
To further increase the flexibility of the code generation with lbmpy, future work is planned to support the emerging INTEL GPUs by having a SYCL 5 back-end. As SYCL is available on most currently available systems, the code generation could used to generate optimized sparse LB kernels for all existing and upcoming hardware such as CPU, accelerator cards, or even exotic hardware such as Accelerated Processing Units (APUs) or Field Programmable Gate Arrays (FPGAs).
Footnotes
Author contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the SCALABLE project (https://www.scalable-hpc.eu/). This project has received funding from the European High-Performance Computing Joint Undertaking (JU) under grant agreement No 956000. The JU receives support from the European Union’s Horizon 2020 research and innovation program and France, Germany, and the Czech Republic. The authors gratefully acknowledge the Gauss Centre for Supercomputing e.V. (![]() ) for funding this project by providing computing time through the John von Neumann Institute for Computing (NIC) on the GCS Supercomputer JUWELS at Jülich Supercomputing Centre (JSC). We acknowledge the EuroHPC Joint Undertaking for awarding this project access to the EuroHPC supercomputer LUMI, hosted by CSC (Finland) and the LUMI consortium through a EuroHPC Regular Access call. The authors gratefully acknowledge the scientific support and HPC resources provided by the Erlangen National High Performance Computing Center (NHR@FAU) of the Friedrich-AlexanderUniversität Erlangen-Nürnberg (FAU).
) for funding this project by providing computing time through the John von Neumann Institute for Computing (NIC) on the GCS Supercomputer JUWELS at Jülich Supercomputing Centre (JSC). We acknowledge the EuroHPC Joint Undertaking for awarding this project access to the EuroHPC supercomputer LUMI, hosted by CSC (Finland) and the LUMI consortium through a EuroHPC Regular Access call. The authors gratefully acknowledge the scientific support and HPC resources provided by the Erlangen National High Performance Computing Center (NHR@FAU) of the Friedrich-AlexanderUniversität Erlangen-Nürnberg (FAU).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.