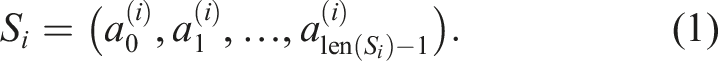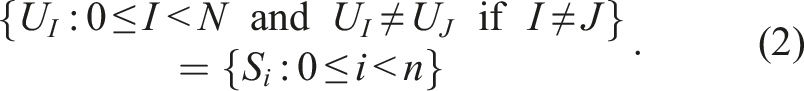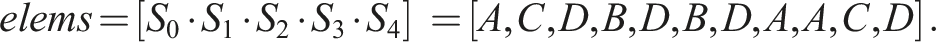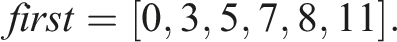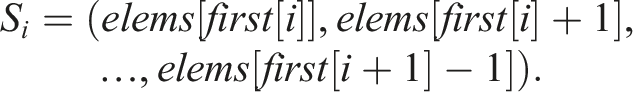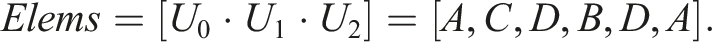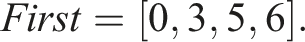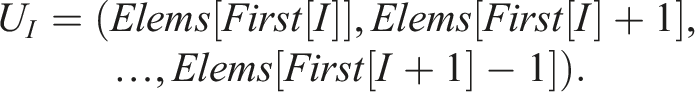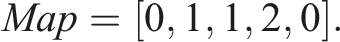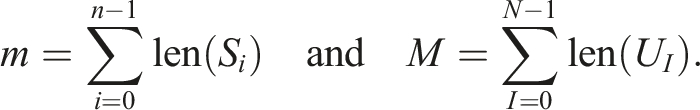Abstract
High performance computing (HPC) applications that work with redundant sequences of data can benefit from their deduplication. We study this problem on the symmetry-adapted no-core shell model (SA-NCSM), where redundant sequences of different kinds naturally emerge in the data of the basis of the Hilbert space physically relevant to a modeled nucleus. For a fast solution of this problem on multicore architectures, we propose and present three multithreaded algorithms, which employ either concurrent hash tables or parallel sorting methods. Furthermore, we present evaluation and comparison of these algorithms based on experiments performed with real-world SA-NCSM calculations. The results indicate that the fastest option is to use a concurrent hash table, provided that it supports sequences of data as a type of table keys. If such a hash table is not available, the algorithm based on parallel sorting is a viable alternative.
Keywords
1. Introduction
This work deals with the problem of deduplication of sequences of computationally processed data, which can be of any nature. Let len(X) denote the length of a sequence X. We say that two sequences X and Y are equal, and write X = Y, if they are both of the same length and all their corresponding elements (elements with the same index) are equal.
For the sake of consistency with algorithm pseudocode and computer code, we stick to the zero-based indexing even in the mathematical notation throughout this text. Let us consider a programming problem of mapping indices 0 ≤ i < n − 1 to variable-length sequences S
i
, where
Let n = 5, S0 = (A, C, D), S1 = (B, D), S2 = (B, D), S3 = (A), and S4 = (A, C, D). Then, S0 = S4 and S1 = S2. The number of unique sequences is thus N = 3 and one possible way to enumerate them is U0 = (A, C, D), U1 = (B, D), and U2 = (A). Let X ⋅ Y denote the operation of concatenating elements of two sequences X and Y. In Example 1, storing all n = 5 sequences in a computer memory would require an array of length 11 The mapping of indices to sequences can then be implemented by another array Here 1. first[i] is the index of the first element of S
i
in elems, 2. first[n] equals the length of elems, and 3. the length of S
i
can be derived as len(S
i
) = first[i + 1] − first[i]. Consequently, Storing N = 3 unique sequences would require an array of length 6 Similarly to first, we can construct the corresponding array First for unique sequences as follows This array is now related to unique sequences Therefore, we additionally need to map the indices i of S
i
to the indices I of U
I
, which can be implemented with another array Let m and M denote the total number of elements of the sequences S
i
and U
I
, respectively The lengths of the arrays elems and first, which represent the sequences S
i
, are m and n + 1, respectively. The lengths of the arrays Elems, First, and Map, which represent the unique sequences U
I
, are M, N + 1, and n, respectively. The memory requirements for storing the sequences S
i
and the unique sequences U
I
are thus O(m + n) and O(M + N + n), respectively. In our example, m + n = 16 and M + N + n = 15, so it does not seem that introducing an extra level of indirection with unique sequences (Map) would be of much benefit. Our work addresses applications where n and m are very large numbers, and where N ≪ n and M ≪ m. In such applications, the memory savings can be significant. However, reducing all the sequences S
i
into a set of unique sequences U
I
, that is, their deduplication, represents some application overhead. To reduce this overhead as much as possible, we need an efficient sequences deduplication algorithm. This work focuses on HPC applications and multicore runtime environments (such as supercomputer nodes), and proposes and evaluates three parallel multithreaded algorithms for deduplication of data sequences. This article is an extended version of our conference paper Langr and Dytrych (2023), where we introduced a parallel multithreaded algorithm for identification of unique sequences based on parallel sorting. Here, we additionally • present two alternative parallel algorithms based on concurrent hash tables, • experimentally evaluate and compare all three proposed algorithms, and • provide more thorough analysis and discussion of all the algorithms in relation to the characteristics of input data.
2. Related work
The defined problem may be considered as a specific instance of the generic problem domain referred to as data deduplication. Data deduplication describes methods that are mostly used to reduce the amount of data stored in a storage system or transferred over a network by eliminating redundant parts; see, for example, Meister (2013); Xia et al. (2016); Zhang and Deng (2017). Our problem represents data deduplication for particular types of data (sequences). Problems of this type are relatively rarely addressed in the literature. We have found some relevant papers applying deduplication methods in order to compress the memory footprints of sparse matrices; see Karakasis et al. (2013); Kourtis et al. (2008); Willcock and Lumsdaine (2006). However, they focus on the data reduction results and acceleration of subsequent matrix operations. In contrast to our work, they do not propose any efficient parallel deduplication algorithms.
3. Application
Our need for fast parallel multithreaded deduplication of data sequences comes from our nuclear structure calculations that leverage the symmetry-adapted no-core shell model (SA-NCSM); see, for example, Barrett et al. (2013); Launey et al. (2016). With SA-NCSM calculations, we try to model atomic nuclei, that is, to obtain their nuclear wave functions. The SA-NCSM first forms a basis for the Hilbert space physically relevant to a given nucleus. Since each basis is infinite-dimensional, only its finite subset of basis states/functions is taken into consideration. This subset is selected by a single even number—so-called basis cutoff parameter Nmax (it represents the maximum allowed number of harmonic oscillator quanta above the minimum for a given nucleus). The higher Nmax, the more basis functions are selected. The subspace of the Hilbert space that is spanned by the selected basis functions is called a model space. For each SA-NCSM calculation, the model space is defined by three main input parameters: the number of protons, the number of neutrons, and Nmax.
Once the basis for a model space is formed, a Hamiltonian matrix is constructed such that its rows and columns are spanned by the basis functions. Finally, the resulting wave functions are obtained in the form of linear combinations of basis functions. The coefficients of these linear combinations are elements of the eigenvectors of the Hamiltonian matrix. The entire SA-NCSM workflow for a given nucleus consists of a series of calculations with increasing Nmax. This process is stopped when the eigenvalues converge for all the energy states in which one is interested.
Our implementation of the SA-NCSM is the software framework called LSU3shell; see Dytrych et al. (2016). It is written in the C++ programming language, and its parallelization is based on the hybrid MPI + OpenMP programming model, which makes it applicable to practically any contemporary large-scale supercomputer; see Message Passing Interface Forum (2021). The SA-NCSM employs a sophisticated mathematical background provided by group theory and the theory of representation, and the group of particular interest is the special unitary group SU(3); see Heine (1960).
Within LSU3shell, a basis is organized into so-called IpIn blocks. Each IpIn block is related to a particular distribution of protons and neutrons in the harmonic oscillator shells. A combination of two IpIn blocks forms a submatrix of a Hamiltonian matrix. Internally, an IpIn block consists of multiple SU(3) proton-neutron irreducible representations (PN irreps). Each PN irrep is mapped to a contiguous chunk of matrix rows/columns. Their number is given by the PN irrep multiplicity factor in the IpIn block. A particular combination of PN irreps thus forms a matrix block in the submatrix given by two IpIn blocks. For more details on the construction of the basis and its organization; see Langr et al. (2018, 2019).
The total number of PN irreps for all IpIn blocks is typically large, but the total number of distinct PN irreps is much smaller. LSU3shell, therefore, stores the information about PN irreps in a standalone database. Then, for each IpIn block, its PN irreps are represented by a sequence of PN irreps database indices (which are integer numbers).
To process submatrix blocks independently—for instance, to enable finer level parallelization—the following information is required for each PN irrep of given IpIn blocks: 1. Its index in the PN irreps database. 2. Its multiplicity factor. 3. First matrix row/column of the corresponding matrix block.
For each IpIn block, its PN irrep indices, their multiplicity factors, and block rows/columns represent three sequences of data (all of them are integer numbers). Fast and independent processing of submatrix blocks would require having these sequences stored in memory for all IpIn blocks. However, this is infeasible in practice, since the total number of elements of these sequences is typically too large. For illustration, the SA-NCSM basis for the 12C, Nmax = 12 model space has n = 73,676,583 IpIn blocks, and the total amount of their PN irreps is m = 1,214,960,841. Storing the sequences of PN irrep indices, multiplicity factors, and block rows/columns for all IpIn blocks would thus require tens of gigabytes of memory.
Counts of all and unique sequences and their elements for the12C nucleus and Nmax = 12 SA-NCSM basis.

The same analysis for multiple different model spaces produced similar results. This means that, in general, if we deduplicate the sequences, then we can store them in memory in all 3 cases (II, MF, and RCO) with affordable memory footprints. This conclusion brought us to the problem of how to quickly identify and remove duplicated sequences and reduce them into sets of sequences that are unique (i.e., deduplicate them). Generally, in SA-NCSM calculations, this problem is solved by each MPI process independently (for its mapped parts of the basis). Therefore, we focused on developing algorithms for fast and scalable parallel multithreaded deduplication of data sequences.
4. Algorithms
Let us start with a single-threaded (ST) solution of the sequences deduplication problem. We can find such a solution relatively easily provided that there is a hash table that can use variable-length sequences as table keys available. In this table, we perform lookup for each input sequence. If the lookup fails, it implies that a new unique sequence has been identified. Then, we need to: 1. Insert the sequence into the hash table and map it to the next available unique sequence index. 2. Append the position of the elements of the identified unique sequence into the array First. 3. Append the elements of the identified unique sequence into the array Elems.
Additionally, the index of the unique sequence is mapped to the index of the input sequence. The pseudocode of the described method is presented as Algorithm 1. In this pseudocode as well as in the following ones, we assume the output arrays are implicitly defined and are initially empty.
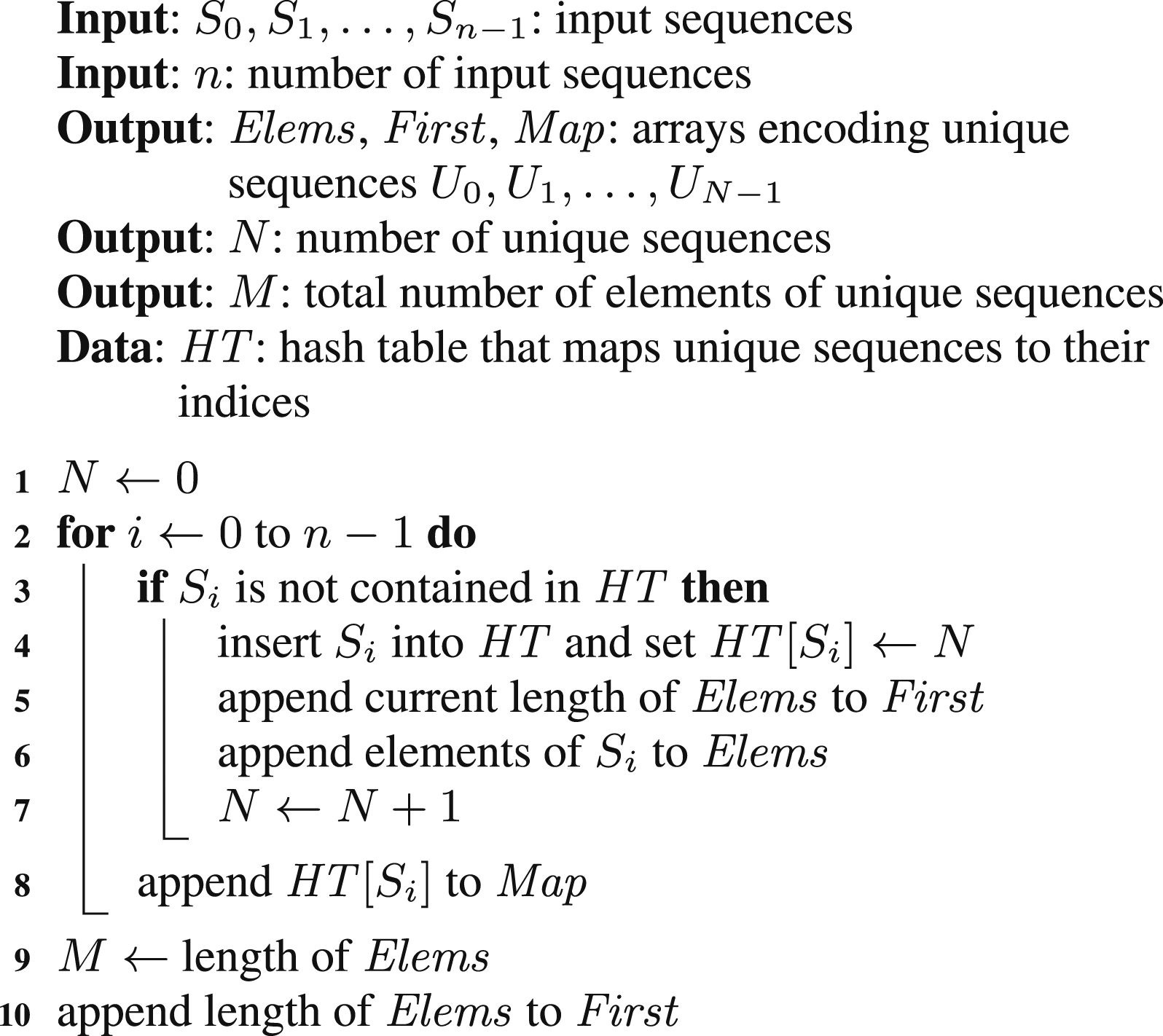

This algorithm cannot be directly parallelized by splitting the loop iterations between multiple threads. First, this loop is inherently sequential due to appending data mainly to the Elems and First arrays with initially unknown lengths. Moreover, implementations of hash tables provided by performance-aware programming languages (such as
Below, we present three parallel multithreaded algorithms for deduplication of data sequences. The first two are based on parallelization of Algorithm 1, where a concurrent hash table is employed. The third algorithm is an alternative that does not require a concurrent hash table but is based on parallel sorting.
4.1. Parallel algorithm MTSK
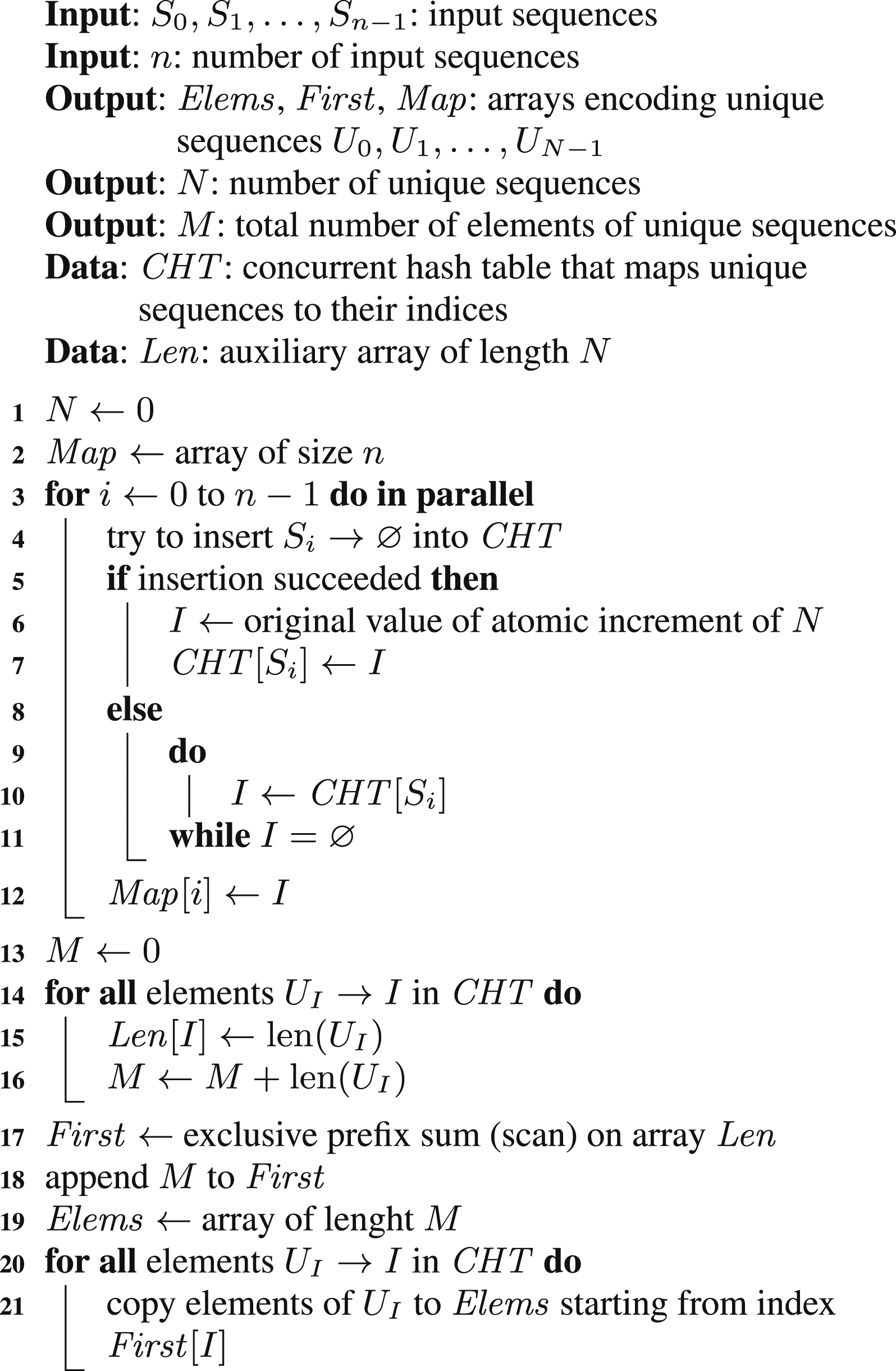
Our first parallel algorithm is based on a concurrent hash table that, similarly to the single-threaded solution, can use variable-length sequences as table keys; we call this algorithm MTSK (multithreaded; sequence keys). The pseudocode of this algorithm is presented as Algorithm 2.
The main loop is parallelized such that its iterations are split between all the threads. In contrast to the single-threaded algorithm, we now cannot perform lookup and insertion as two subsequent operations since another thread could do the insertion in between. Instead, we try to insert each input sequence and check if this operation was successful. If it was, then a unique sequence has been identified and now, we can evaluate its index by atomically incrementing a shared counter. This index is then set to the inserted table record. If the insertion failed, then we need to check whether the unique sequence index has already been set, and, eventually, wait for its update.
In the parallelized loop, we cannot set the content of the Elems array, since its length is originally unknown. This also implies that we cannot set the content of the First array. Instead, we construct both these arrays after the main loop is finished. To find the values of the elements of the First array, an auxiliary array with the lengths of unique sequences is used and the prefix sum operation is applied to it.
4.2. Parallel algorithm MTIK
Our second parallel algorithm is based on a concurrent hash table as well, but now, we do not use sequences as table keys. Instead, we use the indices of input sequences as keys and consider two keys i and j being equal when S i = S j ; we call this algorithm MTIK (multithreaded; index keys). The advantage of this algorithm is that it can be implemented even if there is no concurrent hash table that supports variable-length sequences as table keys available. However, instead, the hash table needs to support a user-provided operation for evaluation of the equality of two keys.

The pseudocode of this algorithm is not presented since it is almost the same as Algorithm 2. The main differences are as follows: 1. The concurrent hash table CHT is “indexed” by i instead of by S
i
. 2. The records of CHT are i → I instead of U
I
→ I, and U
I
is represented by S
i
.
The downside of this algorithm is that it generally requires multiple accesses to the sequences S i . This may be less efficient, especially in cases where these sequences are not stored in memory, but generated on demand instead. In our SA-NCSM calculations, this case applies for the MF and RCO sequences.
4.3. Parallel algorithm MTPS
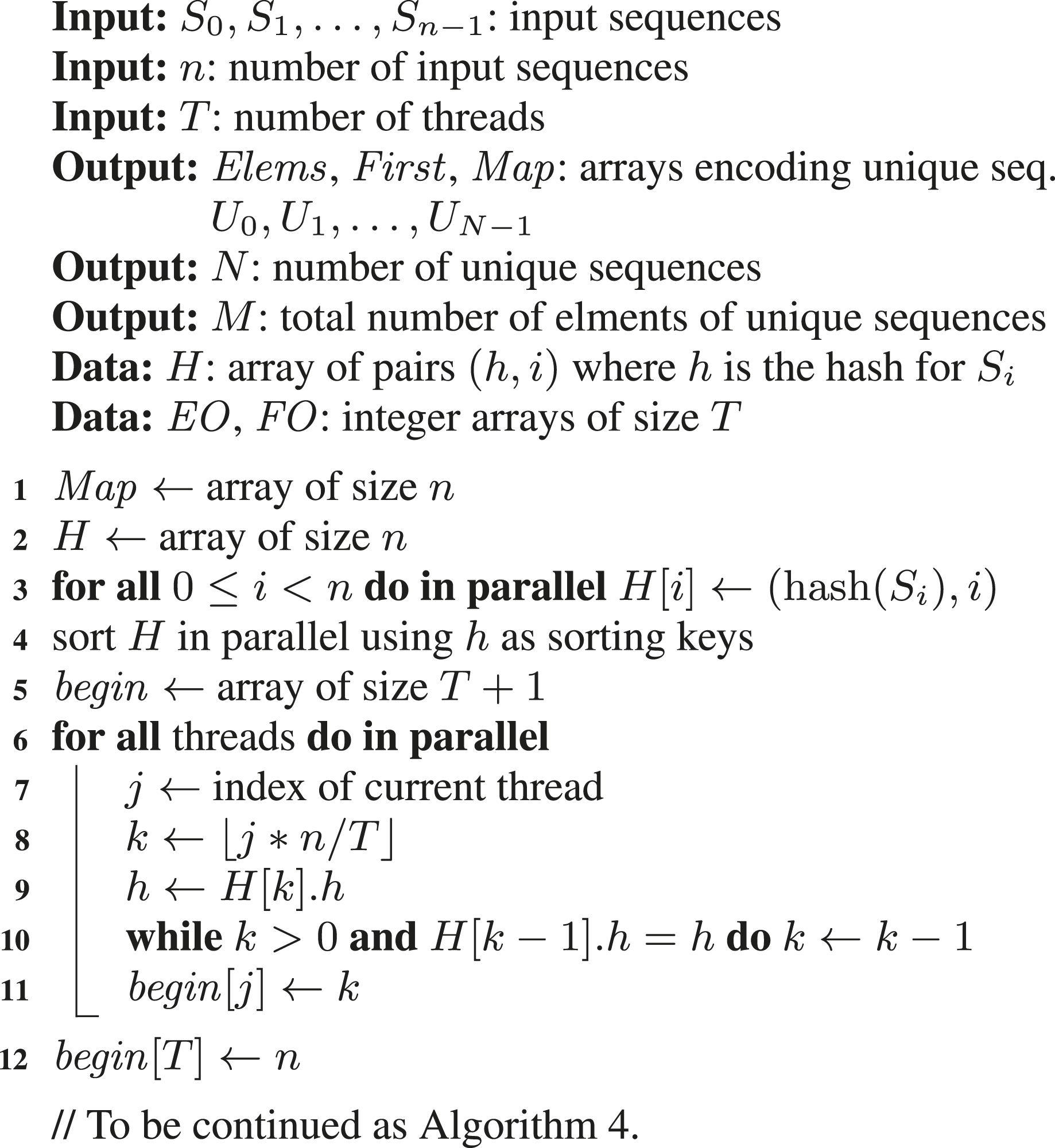
In contrast to the first two parallel algorithms, our third parallel algorithm does not require a concurrent hash table. This algorithm was originally proposed in our conference paper; see Langr and Dytrych (2023), and we call it here MTPS (multithreaded; parallel sorting). It is based on calculating the hashes for all the input sequences S i and their subsequent logical grouping with respect to the same hashes. This then enables independent processing of each group of sequences by a single thread, since equal sequences (having the same elements) belong to a single group. Moreover, if a sufficient hash function is used, then all the sequences in the same group are likely to be equal.
The pseudocode of the proposed algorithm is presented as Algorithm 3, Algorithm 4, and Algorithm 5. First, an auxiliary array H is constructed, where each element contains information about a sequence index and its hash. Then, this array is sorted considering hashes as sorting keys, which effectively places the information about sequences with the same hashes next to each other. Each part of H with the same hash defines a single group of sequences. In the next step, these groups are divided into T segments, where T denotes the number of threads. Now, each thread can operate independently on its segment. Since each unique sequence corresponds to a single group, each thread can process its local groups to identify its unique sequences. The information about them is encoded in thread-local arrays Elems j and First j . When thread-local unique sequences are identified, sequences within a group are compared for equality. This is necessary since two sequences with different elements may have the same hash. Information about index mapping is stored directly in the Map array, but this mapping now applies only to thread-local unique sequences. Finally, thread-local unique sequences are merged into global unique sequences, that is, into the resulting arrays Elems and First. This requires the indexes of the elements and the indexes in Map to be incremented accordingly. The increments are computed as prefix sums (scans) on the lengths of the thread-local arrays Elems j and First j .
Almost all the steps of the algorithms are executed in parallel. There are a few single-threaded operations, which are computationally trivial (prefix sums are performed on T values only). The advantage of this algorithm is that it does not require a concurrent hash table. However, the price for this benefit is that the input sequences need to be accessed twice—once when their hashes are calculated and second time where unique sequences in groups are identified.
5. Implementation issues
To evaluate all the proposed algorithms, we implemented them in C++ with OpenMP threading. This allowed us to

use these implementations on the data generated in SA-NCSM calculations by the LSU3shell program. For the single-threaded algorithm (ST), we used the hash table provided by the C++ standard library in the form of the
For the implementation of the parallel sort-based algorithm (MTPS), we chose the parallel multi-way mergesort, in particular, its OpenMP implementation provided by the libstdc++ parallel mode; see Singler and Konsik (2008). Initially, we also tried to use a few variants of parallel quicksort. However, it turned out that all of them were significantly slower than the mentioned mergesort. We attribute this behavior to the fact that quicksort is generally much more data-sensitive and does not efficiently deal with cases where there are only relatively few distinct sorting keys in the sorted data.


For calculation of the hashes of the sequences S
i
, we adopted the method provided by the C++ Boost library. This method combines hashes of individual sequence elements in the following way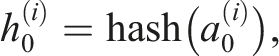
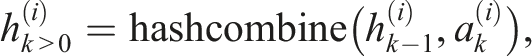
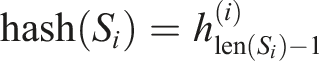
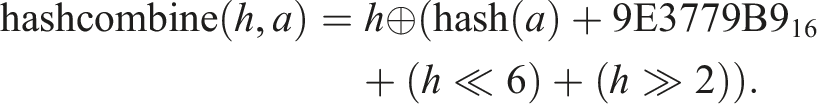
The ⊕, ≪, and ≫ operations denote the exclusive bit OR (XOR), the left bit shift, and the right bit shift, respectively. The hashes of individual input sequence elements were calculated in our case by the C++ standard library
6. Experiments and results
Relevant characteristics of the model spaces and their sequences used for experiments.
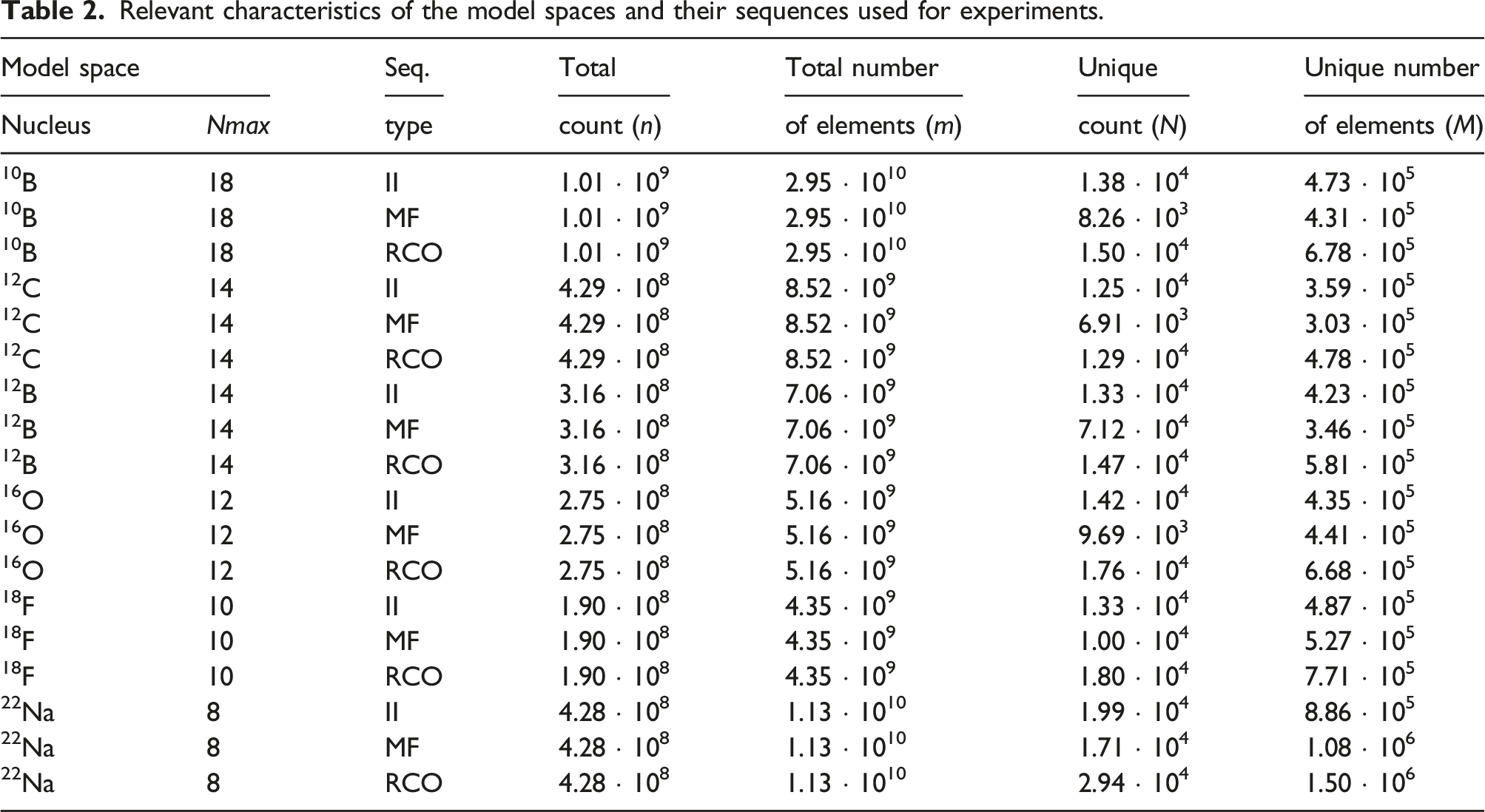
As a testbed for performance evaluation, we used nodes of the Karolina supercomputer operated by IT4Innovations in Ostrava, Czech Republic. A single Karolina node consisted of two AMD EPYC 7H12 CPUs each having 64 computational cores. Each CPU was additionally configured to contain four NUMA nodes, each with 16 CPU cores. 1
On this system, we measured the sequences deduplication runtime for all the combinations of the following experiment parameters: 1. Different SA-NCSM model spaces: 10B, 12C,12 B,16O, 18F, and 22Na. 2. Different types of sequences: II, MF, and RCO. 3. Different deduplication algorithms: ST, MTSK, MTIK, and MTPS. 4. Different number of threads mapped to specific parts of the computational node: 1 thread (T = 1) mapped to a single NUMA node, 16 threads (T = 16) mapped to a single NUMA node, 64 threads (T = 64) mapped to a single CPU (single socket), 128 threads (T = 128) mapped to the entire computational node (both its CPUs/sockets).
Deduplication runtime for different SA-NCSM model spaces, different types of sequences, and different algorithms measured for the single NUMA node setup.
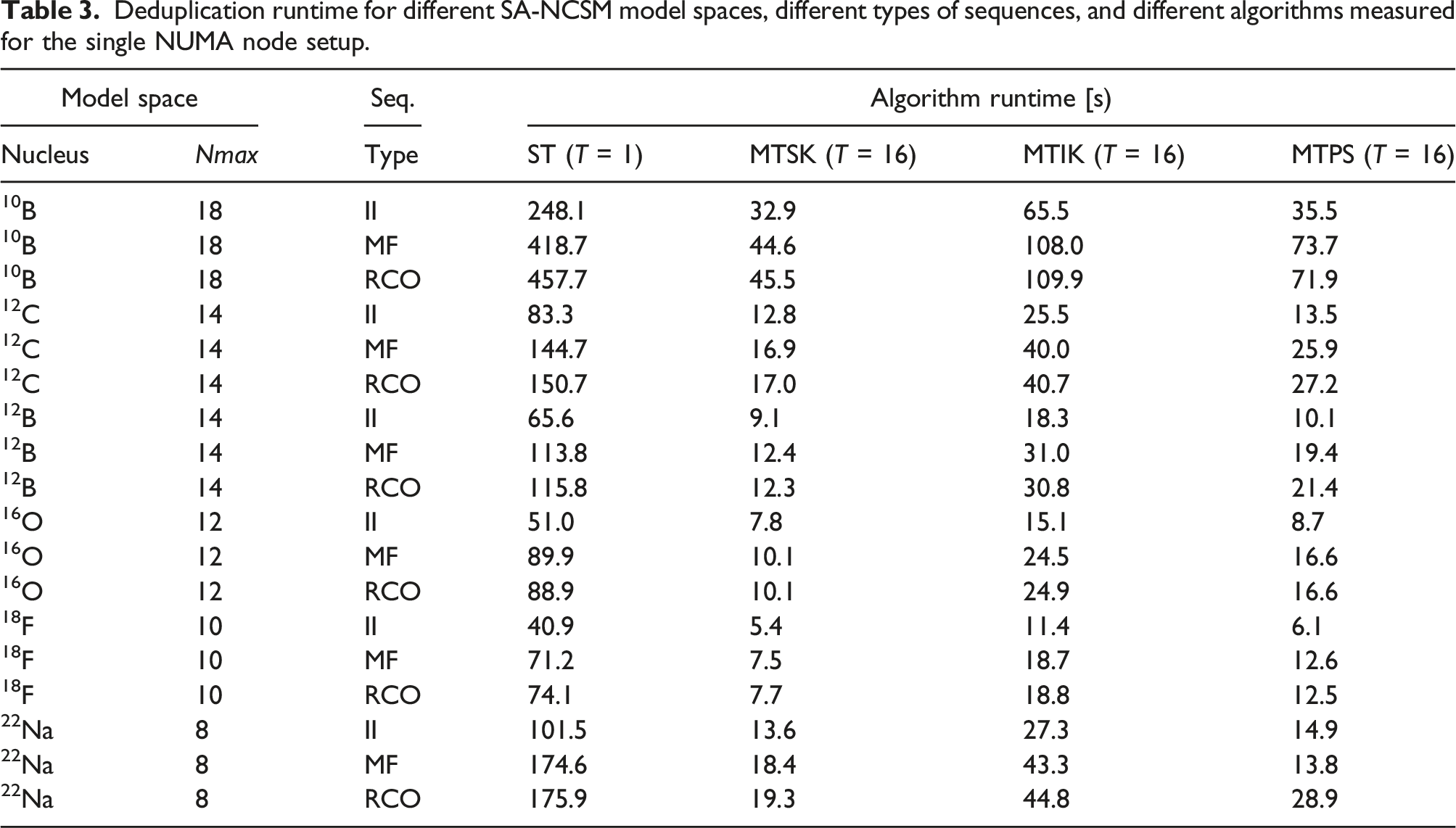
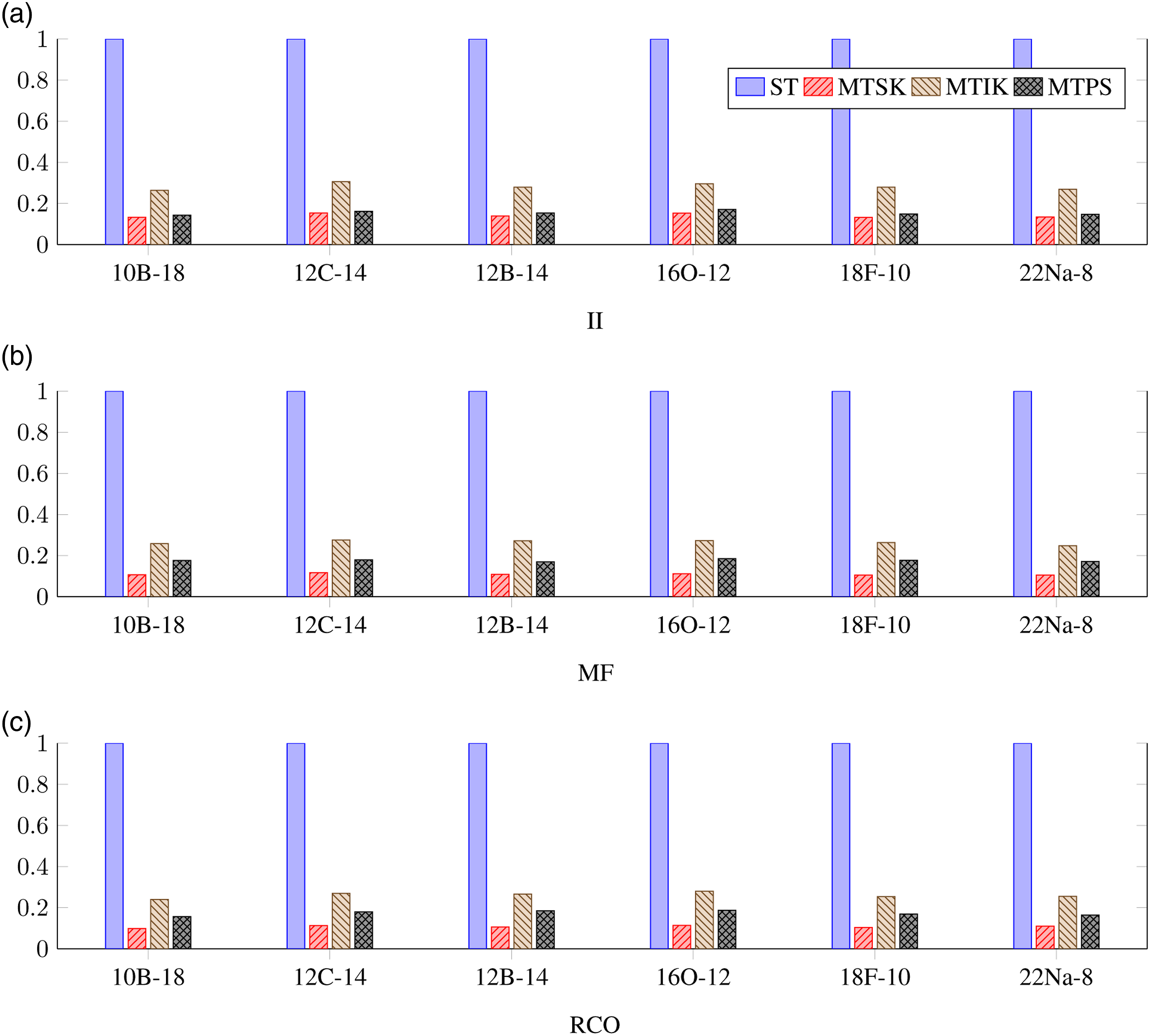
Normalized deduplication runtime for different SA-NCSM model spaces, different types of sequences, and different algorithms measured for the single NUMA node setup. (a) II, (b) MF, and (c) RCO.
Next, in Figure 2, we show the results of the same measurements performed for a single-threaded algorithms runs (T = 1). These results allow us to evaluate the overhead of each parallel algorithm. Obviously, the MTSK algorithm provides runtime practically equivalent to the ST algorithm; sometimes, it is even faster. This indicates that the implementation of the concurrent hash table provided by the oneTBB library is very efficient. The single-threaded overhead of the MTIK and MTPS algorithms is much higher. Specifically, for the on-the-fly generated sequences (MF and RCO), the measured times are around twice in comparison with the ST algorithm. Normalized deduplication runtime for different SA-NCSM model spaces, different types of sequences, and different algorithms measured for the single-threaded algorithms runs. (a) II, (b) MF, and (c) RCO.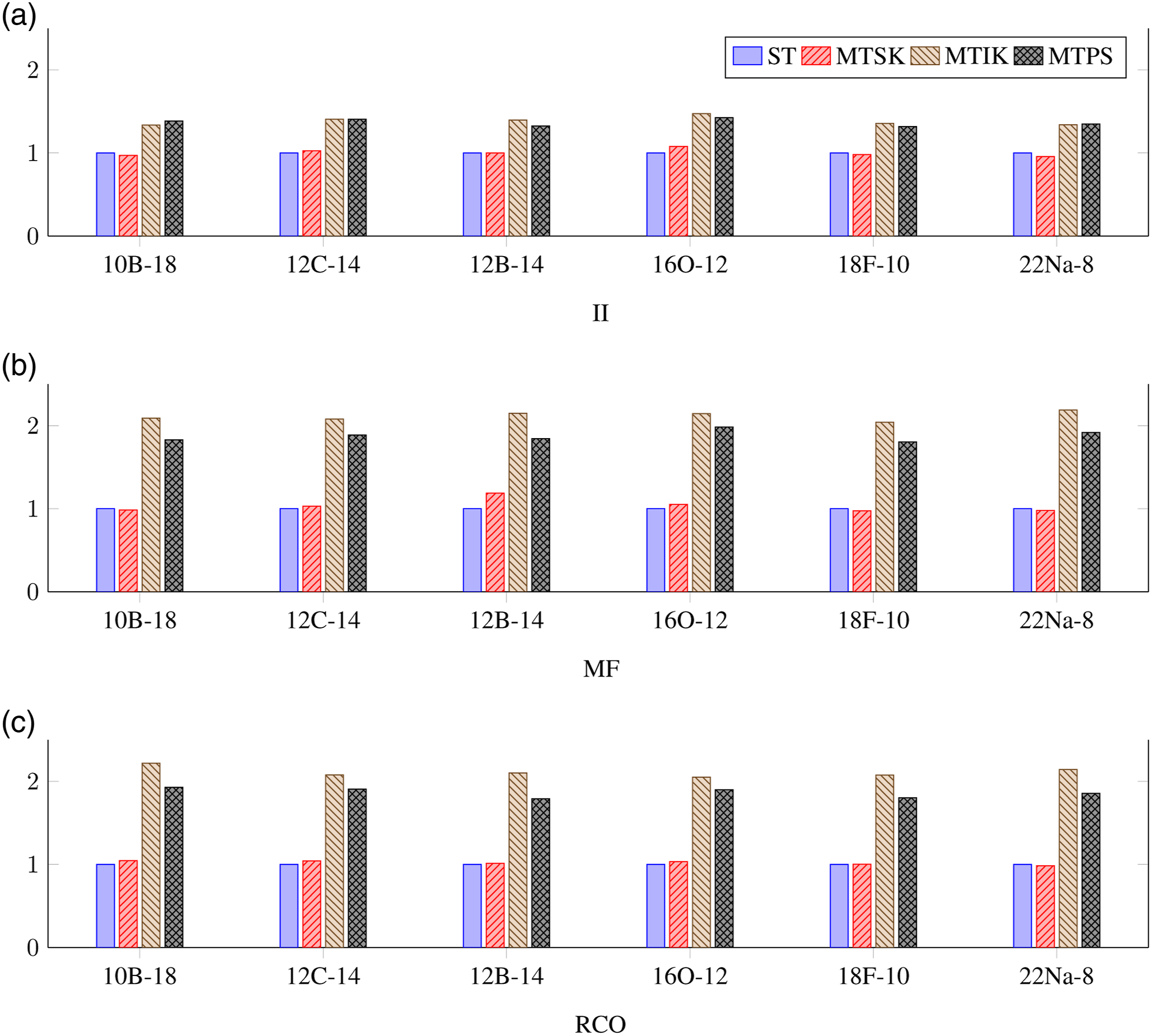
Next, we evaluated the strong scalability (fixed problem size) for all the parallel algorithms with respect to the different number of threads mapped to a single NUMA node, single CPU (socket), and the entire computational node (both CPUs). The results obtained for the 10B, Nmax = 10 model space are shown in a logarithmic scale in Figure 3. These results indicate that the MTSK and MTIK algorithms scale well with the increasing number of threads, but the scalability of the MTPS algorithm is worse, which is significant specifically for the entire-node setup. It seems that this algorithm is more memory-bound, therefore, it suffers from the configurations where the number of cores highly exceeds the number of memory controllers. Normalized deduplication runtime for different number of threads, different types of sequences, and different algorithms measured for the 10B, Nmax = 10 model space. (a) II, (b) MF, and (c) RCO.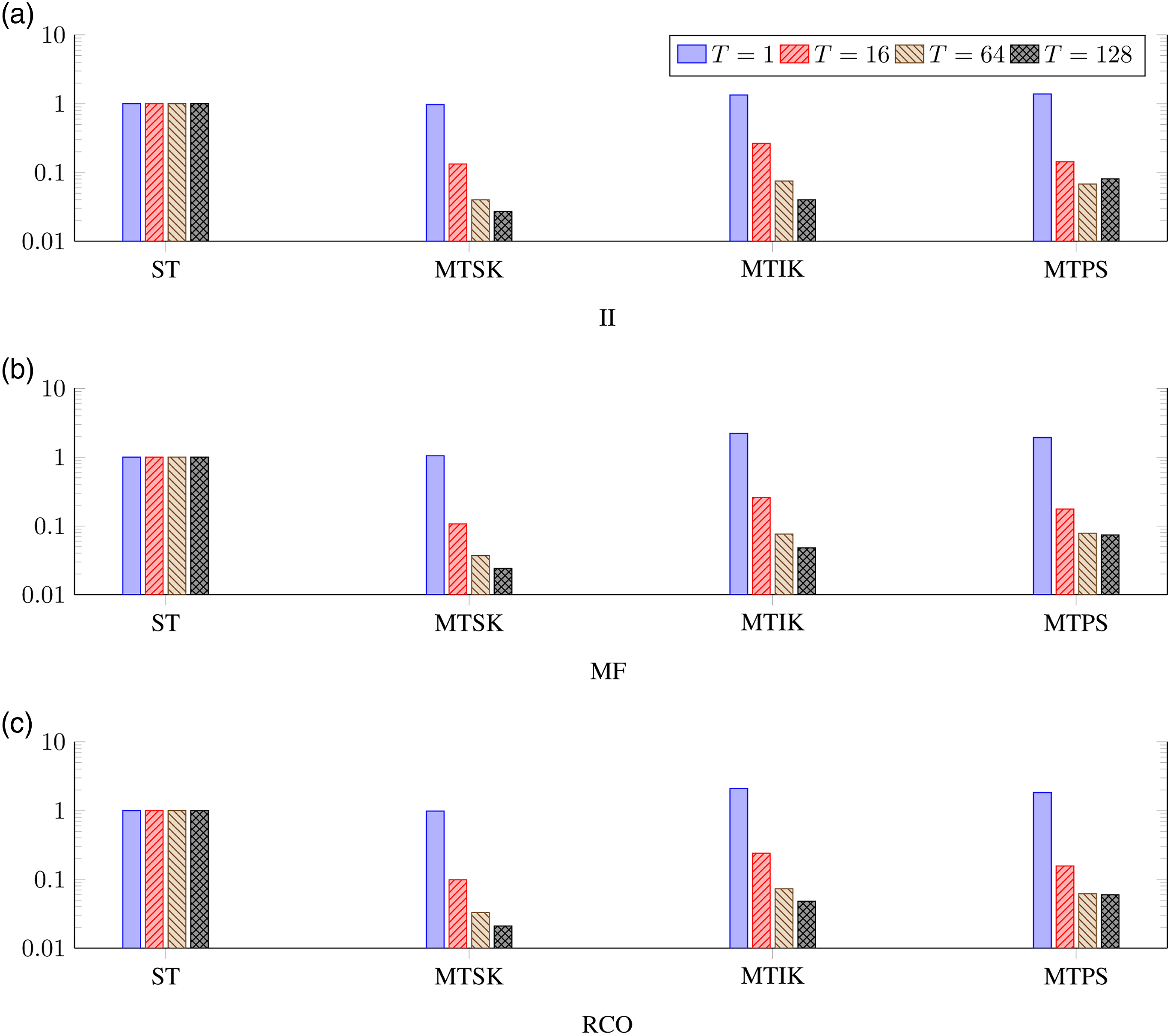
Finally, we compared the proposed parallel algorithms among themselves for different numbers of used threads. The results obtained for the 10B, Nmax = 10 model space are shown in a logarithmic scale in Figure 4. As already noted, the MTSK algorithm was the fastest one in all cases. For the single NUMA node runs, the MTPS algorithm gave similar performance for the II sequences, and it was only slightly slower for the MF and RCO sequences. In the single-CPU cases, the MTIK and MTPS algorithms provided comparable runtimes, but for the entire-node cases, the MTPS algorithm was mostly the slowest one. Normalized deduplication runtime for different algorithms, different types of sequences, and different numbers of threads measured for the 10B, Nmax = 10 model space.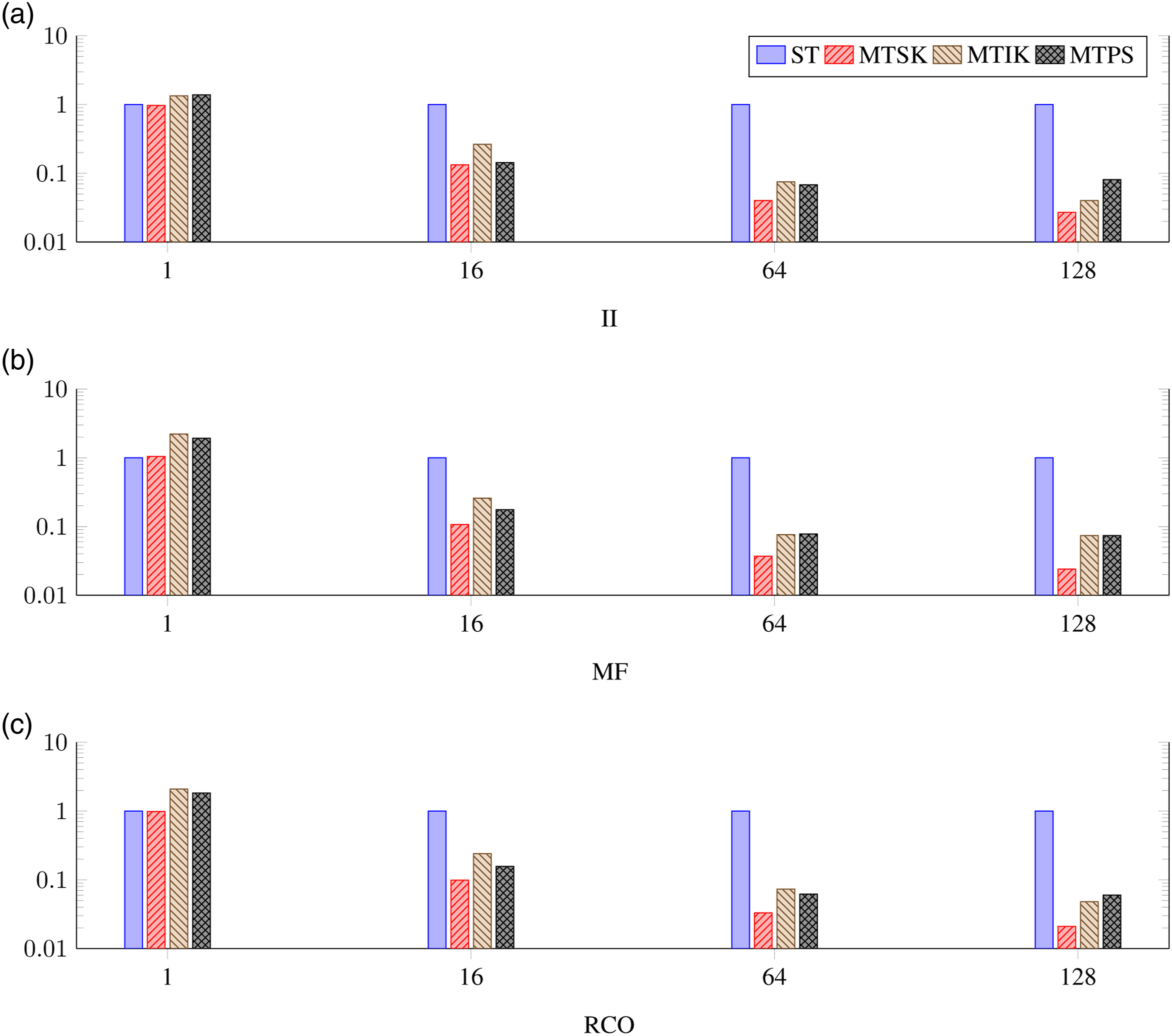
7. Discussion
Our measurements indicate that all the proposed parallel multithreaded algorithms considerably reduce the time needed for the solution of the sequences deduplication problem in SA-NCSM applications. The highest speedup was achieved by the MTSK algorithm, which appears to be generally superior. The only drawback of this algorithm is that it requires a concurrent hash table that can use variable-length sequences as sorting keys. In case of the C++ programming language, such a hash table is not part of the standard library; therefore, the implementation of this algorithm needs some third-party solution.
On the contrary, the MTPS algorithm is hash table-free. Instead, it requires a parallel sorting method, which, since the C++17 standard, is a part of the standard library itself; see ISO/IEC (2017). For some applications, this may be a significant advantage, since it may provide better portability of application code. The drawback of this algorithm is the dual access of each input sequence. First, the sequences are accessed in order of their growing indices (when their hashes are calculated). However, in the second case (when unique sequences in groups are identified), the sequences are accessed in an irregular order. This second access is generally slower due to less efficient prefetching and related lower utilization of caches. These effects are significant in setups where the number of used cores exceeds the number of memory controllers. The MTPS algorithm is thus best suitable for single NUMA node runs, where its performance and efficiency is similar to the MTSK algorithm.
Out of all the multithreaded algorithms, the MTIK algorithm was in our measurements mostly the least efficient one. Its only advantage in comparison with MTSK is that the unique sequences do not need to be stored in memory twice (once in the hash table and second in the resulting arrays). In our application, where the number of unique sequences was very low, these effects were practically negligible. However, in applications where the number of unique sequences (N) would not be that low compared to the number of input sequences (n), the difference might be noticeable.
8. Conclusions
The contribution of this article is the proposal of three parallel multithreaded algorithms for solution of the problem of deduplication of sequences of data. In addition, we presented an experimental evaluation and comparison of these algorithms using data from a real-world HPC application—nuclear structure calculations that adopted the SA-NCSM. In these calculations, the redundant sequences of various kinds naturally emerge in the computer representation of model space bases. We demonstrated that, for multiple different model spaces, the proposed algorithms could significantly reduce the runtime required to deduplicate the sequences of basis data, which, in practice, resulted in significant memory savings for each MPI process. Finally, we compared the proposed algorithms and discussed their advantages and drawbacks, as well as their target suitable application use cases.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Czech Science Foundation under Grant No. 22-14497S, by the Ministry of Education, Youth and Sports of the Czech Republic under Grant No. CZ.02.1.01/0.0/0.0/16_019/0000765, and by the Ministry of Education, Youth and Sports of the Czech Republic through the e-INFRA CZ (ID:90140).
Note
Author biographies
Daniel Langr is a research assistant professor at the Department of Computer Systems, Faculty of Information Technology, Czech Technical University in Prague. His research interests include sparse matrix computations, parallel sorting algorithms, efficient implementations of parallel algorithms, and tuning codes for large supercomputers.
Tomáš Dytrych is a researcher at the Nuclear Physics Institute of the Czech Academy of Sciences. His research interests include the physics of atomic nuclei, computational group theory, and the application of modern programming techniques for high performance computing of nuclear structure.