Abstract
CityCOVID is a detailed agent-based model that represents the behaviors and social interactions of 2.7 million residents of Chicago as they move between and colocate in 1.2 million distinct places, including households, schools, workplaces, and hospitals, as determined by individual hourly activity schedules and dynamic behaviors such as isolating because of symptom onset. Disease progression dynamics incorporated within each agent track transitions between possible COVID-19 disease states, based on heterogeneous agent attributes, exposure through colocation, and effects of protective behaviors of individuals on viral transmissibility. Throughout the COVID-19 epidemic, CityCOVID model outputs have been provided to city, county, and state stakeholders in response to evolving decision-making priorities, while incorporating emerging information on SARS-CoV-2 epidemiology. Here we demonstrate our efforts in integrating our high-performance epidemiological simulation model with large-scale machine learning to develop a generalizable, flexible, and performant analytical platform for planning and crisis response.
Justification for ACM gordon bell special prize for HPC-based COVID-19 research
We implement a distributed COVID-19 ABM in Chicago, modeling 2.7 million individuals moving between 1.2 million places with hourly activity schedules; run mixed Bayesian calibration and ABM workflows at full machine scale and 84% utilization on ALCF Theta; provide forecasts and counterfactual analyses to city and state public health departments.
Performance attributes
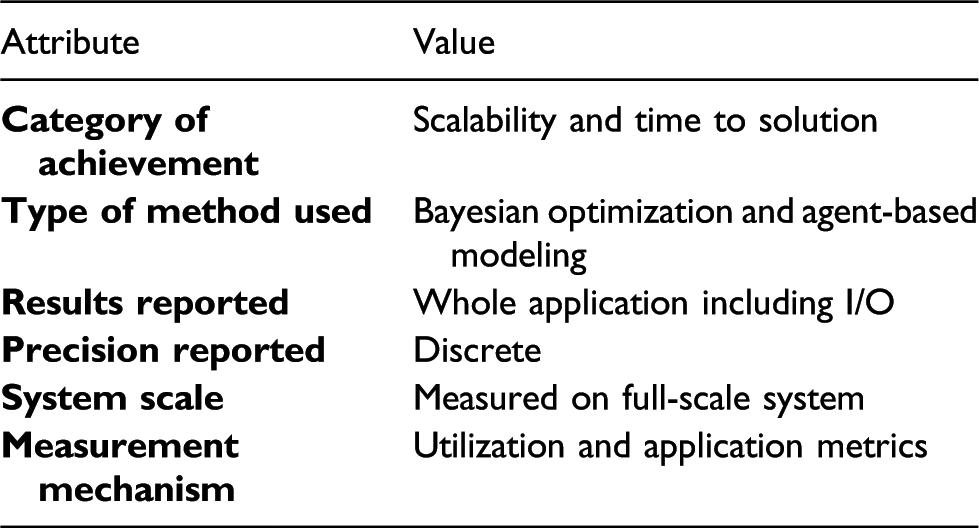
1. Overview
The COVID-19 epidemic has brought epidemiological modeling to the forefront of discussions on how evidence-based decision-making can be supported in times of crisis and uncertainty. Since the beginning of March 2020 we have applied CityCOVID, a detailed agent-based model (ABM) that represents the 2.7 million residents of Chicago in terms of people (behaviors and social interactions), places (1.2 million unique geolocations including households, schools, workplaces, and hospitals), and hourly activity schedules, with the aim of understanding COVID-19 transmission dynamics and potential effects of nonpharmaceutical interventions (NPIs). CityCOVID is based on the ChiSIM framework (Macal et al., 2018) and Repast HPC (Collier and North 2013), which enable the creation of efficient, urban-scale MPI-distributed agent-based models (ABMs). Each individual, or agent, in the model includes its own individualized disease progression dynamic that determines transitions between possible COVID-19 disease states. Transitions have functional dependence on heterogeneous agent attributes, exposure through colocation over time with infected individuals, and other factors such as the effects of protective behaviors on viral transmissibility.
Our detailed modeling approach is substantially more computationally demanding than other types of modeling methods being applied to COVID-19, such as statistical models and compartmental models. However, in contrast to statistical models, which tend to be retrospective, we are able to investigate novel interventions that have not been implemented yet. When compared with compartmental models, we can simulate more specific and realistic NPIs, such as imposing restrictions on specific types of businesses, or detailed school attendance models, closely resembling those being considered by public health officials.
A central element in the application of epidemiological models is running in silico experiments for model calibration, scenario analyses, and uncertainty quantification, what we refer to as model exploration (ME) (Ozik et al., 2018). These processes often require sophisticated machine learning (ML) algorithms to automate iterative, or sequential, strategic explorations of possible model behaviors, since brute-force approaches are not generally feasible even with the largest computing resources. Substantial advancements are being made in sequential ML algorithms that can be applied to ME. However, implementing dynamic ME processes at the necessary computational scales poses serious challenges, not the least of which is the need for efficiently dispatching and coordinating mixed workloads related to ML algorithms and complex models such as CityCOVID.
We developed the EMEWS framework (Ozik et al., 2016), built on Swift/T (Wozniak et al., 2013a), expressly to enable large-scale ME as ML-driven HPC workflows, which can be run on leadership-scale resources. Throughout the epidemic, members of our group have been regularly meeting with city, county, and state stakeholders and providing them with CityCOVID model outputs in response to their evolving priorities. Given the dynamic nature of information on the epidemiology of SARS-CoV-2 and the shifting foci of NPIs, we have engaged in a concerted effort to address computational efficiency across multiple dimensions and to reduce the “time to analysis.”
The work we present here covers the approaches we took, including the following: Parallelization, load balancing, and caching within CityCOVID Applying multiple large-scale ML algorithms, such as multiobjective approximate Bayesian computation and Bayesian optimization, for model calibration and uncertainty quantification Coordinating single-node ML tasks and ordered multinode MPI tasks in a scalable HPC workflow Connecting the system to the real world via observed data and database access
We demonstrate the scaling performance of our integrated approach through whole-machine runs on the Argonne Leadership Computing Facility (ALCF) Theta supercomputer. Furthermore, we discuss the potential impact of this work in developing novel use cases for HPC resources beyond COVID-19 to a generally applicable decision-support platform.
2. State of the art
2.1. Epidemiological modeling
Current epidemiological modeling for disease spread is based on three intrinsically different types of modeling approaches: (1) so-called SIR/SEIR compartmental models, which are based on differential equations that specify mathematically rates of change between compartments, for example, susceptible (S), exposed but not infected (E), infected (I), and recovered (R) states; (2) agent-based models, in which the interactions of individual agents (people) and their explicit behaviors and contacts are directly simulated; and (3) statistical forecasting, which makes no assumptions about the details of the disease, such as the Institute for Health Metrics and Evaluation (IHME) model. Each of these approaches is capable of, for example, predicting the expected daily number of deaths, hospitalizations, and infections for COVID-19 in a given geographical region, as well as providing uncertainty bounds for these estimates, if stochastic elements are included in the model formulations.
The basic idea of “mechanistic” epidemiological models is to explicitly model the mechanisms of disease spread between individuals by modeling two processes: (1) the frequency of co-located contacts between individuals and (2) the nature of contacts between individuals that, probabilistically, facilitate or mitigate disease transmission. Agent-based models allow for these processes and all relevant variables to be included at the finest grain level of detail required to answer the questions posed by decision-makers. The CityCOVID model can simulate entire cities with millions of individuals, necessitating the use of HPC resources. The detailed agent-based modeling approach facilitates testing hypotheses concerning transmission, understanding optimal policies for reducing rates of transmission, and providing estimates for the impact of loss of staffing for critical services.
2.2. Agent-based modeling
Agent-based models and the toolkits with which they are developed, such as Repast Simphony (North et al., 2013), MASON (Luke et al., 2005), and NetLogo (Wilensky 1999), have been confined to a single CPU for most of their development, with occasional forays into multithreaded parallelism. Within the past decade, however, we have seen increasing support for multiprocess distributed toolkits. Here, performance is achieved primarily by dividing the global population of agents among processes. A typical simulation can loop through all the agents during each iteration of a simulation. The fewer agents to iterate over, the faster the simulation. Toolkits such as D-MASON (Cordasco et al., 2014) initially attempted to avoid the complexity of MPI (at least in the Java environment) in distributing and coordinating agents across processes in favor of sockets and the Java Message Service before turning to MPI as a more performant solution for interprocess communication. Other ABM toolkits such as Pandora (Rubio-Campillo 2014), Care HPS (Borges et al., 2017), and Repast HPC (Collier and North 2013) also use MPI as the message-passing layer, while Flame GPU (Richmond and Chimeh 2017) distributes agents across a GPU such that their behavior can be executed in parallel. Other areas of emphasis are strategies for load balancing and the optimal partitioning of agents using different topologies and ghosting techniques (Borges et al., 2017; Collier et al., 2015).
2.3. Model exploration
The rise of artificial intelligence has provided broad and renewed interest in the development of statistical and ML algorithms that can address the evolving scale of ME tasks, resulting in vibrant ecosystems of free and open source libraries that are continually added to and updated as research frontiers are expanded. The relevant algorithms applicable to ME include various active learning (Settles 2012) approaches, including Bayesian optimization (BO) approaches using Gaussian process (GP) surrogate models (Binois et al. 2018, 2019) and ML models such as random forest (Breiman 2001; Ozik et al., 2018); approximate Bayesian computation (ABC) approaches (Hartig et al., 2011; Beaumont 2019), including sequential Monte Carlo (Beaumont et al., 2009) and Incremental Mixture ABC (Rutter et al., 2019); evolutionary approaches, including single (Holland 1992) and multiobjective (Deb et al., 2002) genetic algorithms; and data assimilation approaches, including ensemble Kalman filtering (Evensen 2009; Pei et al., 2018) and particle filters (Arulampalam et al., 2002; Gordon et al., 1993). Libraries that provide cutting-edge implementations of these algorithms or the components from which to implement them include scikit-learn (Pedregosa et al., 2011), scikit-opt (Head et al., 2018), BOTorch (Balandat et al., 2020), ABCpy (Dutta et al., 2017), ELFI (Lintusaari et al., 2018), and DEAP (Fortin et al., 2012) in Python; caret (Kuhn 2008), mlrMBO (Bischl et al., 2017), randomForest (Liaw and Wiener 2002), EasyABC (Jabot et al., 2013), IMABC (Maerzluft et al., 2021), DiceKriging (Roustant et al., 2012), hetGP (Binois and Gramacy 2019), laGP (Gramacy 2016), and GPareto (Binois and Picheny 2019) in R. However, a vast majority of these libraries and algorithms have not been developed specifically with HPC scales or batch sampling in mind. That is, batch sizes often stay lower than a dozen (e.g., Chevalier and Ginsbourger (2013)), especially for multiobjective Bayesian optimization Daulton et al. (2020). Exceptions include Wang et al. (2018), which uses an ad hoc filtering to select batches of a few hundred candidates a posteriori, without considering repeating experiments and thus decreasing the noise at evaluated configurations. In addition, approaches combining ABC and GP methodologies have also been considered (see, e.g., Wilkinson (2014), Meeds and Welling (2014)), although the approach we take here differs in that ABC is used to provide initial evaluations and the GP optimization process provides refining. Moreover, toolkits specifically aimed at model exploration on HPC resources exist (see, e.g., Adams et al. (2014b), Groen et al. (2021)); however, they are not geared toward integrating external, multilanguage libraries or large-scale mixed ML and simulation workflows.
2.4. Workflow integration and MPI task management
Coordination among the ML and ABM modules in this workflow requires deep integration of components written in Python, R, and C++/MPI. The overall workflow shares hardware resources for these systems and rapidly switches among them. The workflow generates parameters in R-based optimizers, distributes these parameters to C++/MPI simulations, and registers progress via database operations called via Python—all on the same resources. Thus, building this complex application requires a workflow system with the capability to manage in-memory user libraries and third-party packages such as scripting language interpreters.
Other workflow solutions are generally constructed to orchestrate Unix-like program invocations (Zhao et al., 2007) or whole-job scheduler submissions (Thain et al., 2005; Deelman et al., 2005); some recent systems support in-memory Python snippets (Babuji et al., 2017), while others have been extended to handle more complex ensembles and heterogeneous compute resources (Di Natale et al., 2019). To our knowledge, Swift/T is the only system that offers innate support for calling functions in Python and R, as well as tasks that are libraries that use MPI. In addition to coordinating ML and simulation tasks, our approach further allows for external ME algorithms to control the overall progression of complex workflows (see § 3.3).
Launching large numbers of MPI jobs is difficult on today’s HPC systems. Typical solutions treat this as a shell programming problem and use a shell-scripting-like technique (or Python, for example) to manage calls to vendor-supplied job launchers (mpiexec, srun, aprun, jsrun, etc.), each with different command line interfaces. In contrast, the Swift/T approach seeks to use a pure MPI model for managing workflow tasks that use MPI. More generally, an MPI-based solution is needed (Wozniak et al., 2019).
As a quantitative comparison of these approaches, consider running a minimal 64-process C/MPI application that simply does
3. Innovations realized
The innovations within this work are in two areas: distributed simulation with CityCOVID and application and workflow integration of large-scale machine learning.
A schematic for the CityCOVID Hypervolume Refinement (HVR) workflow, our most complex workflow for which we provide full machine performance results (§ 5), is shown in Figure 1. Empirical data from hospital systems and public health departments are used to parameterize CityCOVID and to provide targets for calibration. Each iteration of the HVR algorithm (described in full in § 3.2.2) consists of three ML phases, each using an ensemble of single-node, many-core R methods distributed across the machine. The first phase is for fitting Gaussian process, or kriging, metamodels to the evaluated simulation runs, where evaluations are errors in fitting both hospitalization and death time series model outputs to empirical data. The second phase calculates the Expected Hypervolume Improvement of new candidate parameter points, and the third phase optimizes the best new candidate points. The calculations from the three ML phases produce simulation allocations, which are assigned across the existing Pareto front and new candidate points for evaluation. These simulations are instantiated and distributed across the system as multinode C++/MPI programs. The HVR algorithm proceeds through multiple iterations as it refines the Pareto front (i.e., jointly optimizes multiple objectives). Simulation parameters are written to a Postgres database as the algorithm progresses for real-time feedback and final reporting. Conceptual progress model in the HVR workflow. The workflow loops until convergence.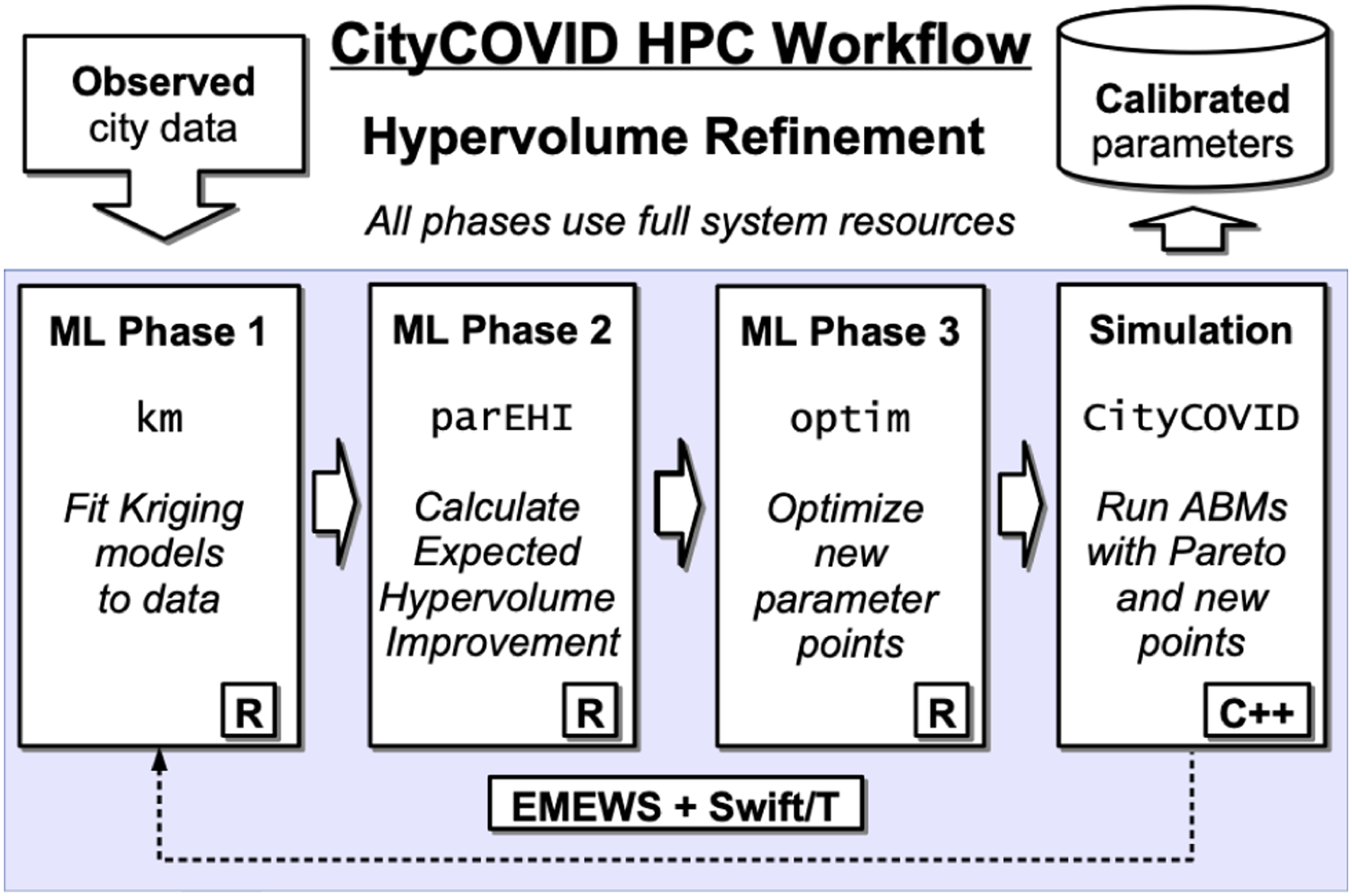
3.1. CityCOVID
3.1.1. Model overview
Models built with the ChiSIM framework (Macal et al., 2018) take three main inputs: (1) synthetic people, (2) places, and (3) activity schedules. In the case of CityCOVID, we model the 2.7 million residents of Chicago as they move between 1.2 million places based on their hourly activity schedules. The synthetic population of agents extends an existing general-purpose synthetic population (Cajka et al., 2010) and statistically matches Chicago’s demographic composition. Agents can colocate at the geolocated places, which include households, schools, workplaces, hospitals, and group quarters (e.g., nursing homes, dormitories, and jails). The agent hourly activity schedules are derived from the American Time Use Survey and the Panel Study of Income Dynamics and assigned based on agent demographic characteristics. CityCOVID includes COVID-19 disease progression within each agent, including differing symptom severities, hospitalizations, and age-dependent probabilities of transitions between disease stages.
CityCOVID calibration parameters

3.1.2. Model implementation
CityCOVID implements detailed agent behaviors and individualized disease progression and differentiates the movement of agents to places with how they behave when colocating with others in those places. For example, agents may engage in protective behaviors such as wearing a mask or keeping a six-foot distance, thereby reducing viral transmissibility while participating in activities outside of their homes. CityCOVID contains explicit representations of people and places, implemented as C++ classes. Places are assigned to a process rank, and persons move between places, and thus between processes, according to their hourly activity schedules. When a person is moved between processes, it needs enough of its state available on the target process to continue to follow its activity schedule and to transition between disease states.
The potential performance costs of this model are threefold: (1) serializing of a person’s state on the source process and deserializing of that state on the destination process; (2) MPI related overhead in transferring the person’s state; and (3) uneven distribution of persons among processes that causes uneven work loads. For the last factor, processes with less load (i.e., those with fewer persons to transition between disease states) would be idle waiting for those with greater loads to finish, resulting in longer runtimes. For (1), CityCOVID (via ChiSIM (Collier et al., 2015; Macal et al., 2018)) mitigates the cost by caching person objects and their static attributes on every process that the person has visited. This minimizes the amount of data transferred and avoids any overhead associated with the frequent creation of new C++ objects and the concomitant memory allocation. For (2) and (3), CityCOVID minimizes the amount of cross-process movement and load balances the number of persons on each process through graph partitioning, where each place is a vertex and each edge between two vertices represents the volume of person movement between those two places. The edge weight represents the number of persons that travel between the two places, and a vertex weight, the total number of persons to visit that place. By partitioning this graph into groups of vertices (places), such that the edge weights connecting these groups are minimized and the vertex weights are roughly equal among all the partitions, highly connected places are placed on the same process, and we avoid any issues caused by uneven computational load. For the full Chicago scenario used here, the places graph consists of approximately 1.2 million vertices (places) and approximately 1.9 million edges (trips between places). To partition the graph, we used the METIS toolkit (Karypis and Kumar, 1999), assigning each place to the process rank corresponding to its graph partition. This assignment was performed during the development of the synthetic population (the collection of agents, places, and activity schedules), external from any simulation execution. While the agent behavior is itself stochastic, the number of places that an agent can select from as part of this behavior is tightly bounded, and thus effective load balancing using graph partitioning can be performed as a preprocessing step rather than during the simulation itself.
We also eliminated the time spent reading the input data (approximately 2.5 min on Theta) by caching it after the first read. The input data reflect the load balancing and assign persons and places to particular ranks. Each rank caches the data assigned to it. Consequently, we ensure that when Swift/T allocates a group of 256 processes for each simulation run that the rank assignment within each group is repeatable.
3.2. Model Exploration algorithms
3.2.1. Sequential ABC
The initial workflow we constructed was aimed at parameter estimation of so-called deep model parameters, those that are not readily observable. For example, the base hourly probability of transmission between one infectious and one susceptible person occupying the same location (θ2 in Table 1) is difficult to measure directly. Similarly, the reduction in transmission from individual protective behaviors, such as wearing a mask and social distancing (θ9 in Table 1), not only is difficult to directly measure but also can vary over time as behaviors change. These parameters, along with their joint distributions and uncertainties, can nonetheless be estimated through simulation-based inverse modeling approaches.
For simulators with intractable likelihoods, approximate Bayesian computation (ABC) methods, while computationally intensive, provide the ability to generate simulated posterior distributions of the model input parameters and, consequently, posterior model outputs that propagate data and model uncertainties to model results (Beaumont, 2019). There are a number of ABC methods and libraries implementing them (Jabot et al., 2013; Maerzluft et al., 2021; Dutta et al., 2017; Lintusaari et al., 2018). Sequential ABC methods (Beaumont et al., 2009; Moral et al., 2011; Rutter et al., 2019) improve the yield of parameter space sampling compared with one-shot rejection sampling approaches. In addition, batch sequential ABC methods can exploit the concurrency available on HPC resources, providing the ability to tune the batch sampling size to balance sample yield and time to solution.
We utilized a batch sequential ABC algorithm Lenormand et al. (2013) implemented in the EasyABC library Jabot et al. (2013), and we directly integrated it into an EMEWS workflow for our initial model calibration. The calibration was preceded by parameter prioritization analyses using an efficient global sensitivity method Morris (1991), identifying the nine parameters (a) Joint posterior distributions of CityCOVID input parameters (Table 1) from sequential ABC, (b) successive Pareto fronts of errors in deaths (x-axis) and hospitalizations (y-axis) from HVR workflow, (c) COVID-19 attributed hospitalization outputs from CityCOVID (red dots: empirical Chicago data, dark line: median simulation output, dark band: 50% simulation intervals, and light band: 95% simulation intervals), (d) CityCOVID zip code level snapshot of weekly infection outputs at 47 days after June 3, 2020, initial easing of restrictions in Chicago for two scenarios (strict: population wide adherence to protective behaviors, that is, θ9 is maintained as reopening occurs, relaxed: gradual increase of θ9 to a value corresponding to 80% viral transmission reduction).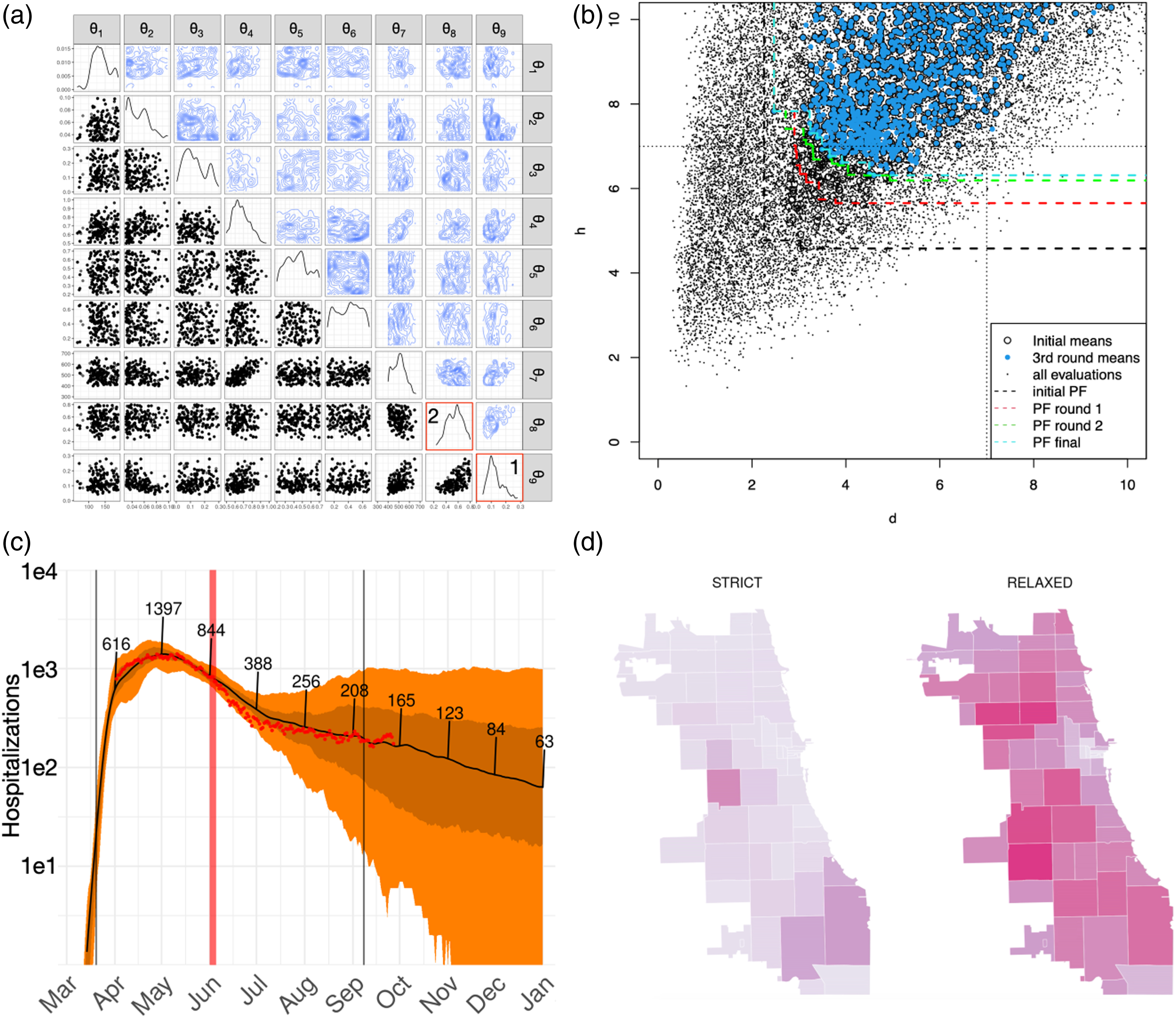
The diagonal panels in Figure 2(a) are the marginal distributions for each input parameter θ i , and the off-diagonal elements show contour (upper) and scatter plots (lower) of each two-dimensional parameter subspace θi,j. The calibration resulted in two main takeaways, both associated with agent behaviors during the stay-at-home period: (1) the reduction in transmission due to individual protective behaviors (θ9) exhibited a pronounced peak at about 90% transmission reduction (Figure 2(a) inset (1)) and (2) the average proportion of agent pre-COVID-19 out of household activities where agents instead opted to stay at home during the shutdown (θ8) showed a broader range of values between 40% and 60% (Figure 2(a) inset (2)). While the second result can be compared with empirical data on mobility based on cellphone data, providing constraints on the parameter estimation result, the first result encapsulates generally unobserved dynamics of person-to-person interactions across the population in response to public health messaging and perceptions of risk.
As the pandemic progressed, our team sought methods for improving the time to solution for generating parameter estimates. One such approach was utilizing another sequential ABC algorithm, IMABC (Rutter et al., 2019), which members of our team implemented as an R package Maerzluft et al. (2021). IMABC provided two main improvements over the original ABC algorithm. Because of a more directed sampling approach, IMABC increased the efficiency of sampling from the CityCOVID parameter space, which sped up algorithm convergence. The IMABC algorithm and package also included the ability to continue sampling from a checkpointed algorithm state, that is, to further refine a previously generated simulated posterior distribution. This allowed for improved robustness, for example, for restarting after HPC resource failure, and the possibility for optionally longer-running calibrations when convergence in the standard timeframe proved difficult.
Another approach we used for improving parameter estimation time to solution was to employ metamodels, also known as surrogate models, for multiobjective optimization, which we discuss next.
3.2.2. Hypervolume refinement
To complement the uncertainty estimation in the parameter space provided by the ABC approach, we also consider finding a set of best parameters matching simultaneously both COVID-19 attributable deaths and hospitalizations. Denote L
D
(resp. L
H
): 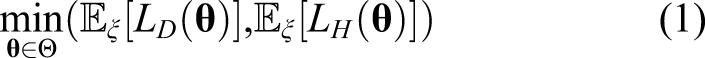
To handle the challenges associated with running hundreds of noisy simulations at once, we developed the multiobjective Bayesian optimization loop shown in Figure 1. To both accurately learn the input-dependent variance and reduce the computational complexity of GP modeling, we complement the exploitation side of BO by leveraging replication of runs at the same location. Every design
3.3. Machine learning in HPC
Running machine learning workloads in high-performance computing systems is at the core of the technological contributions of this effort. Our approach is to use Swift/T to coordinate and distribute in-memory library calls to ML and simulation codes in an HPC-oriented manner. While this technique had been prototyped earlier (Ozik et al., 2015), the CityCOVID workload effort involved the development and integration of the following innovations.
3.3.1 Pluggable algorithm workflow control
A key innovation of EMEWS is the ability to directly incorporate R- or Python-based model exploration algorithms. This is done by defining resident, or stateful, tasks to encapsulate the logic within iterating, state-preserving algorithms (Ozik et al., 2015). These in-memory calls are critical for ML-based workflows in which numerically oriented modules maintain state over long periods but are treated as discrete tasks by the workflow system. While certain workflow systems are designed from the ground up to perform numerical procedures such as optimization (Abramson et al., 2001; Adams et al., 2014a), these are generally difficult to extend and reprogram for arbitrary workflows. We have exploited this capability to implement a variety of workflows specifically geared toward the concurrency afforded by HPC resources, both in terms of the number of simultaneous simulation tasks and through the mixed use of worker pools to handle machine learning tasks as well. We used sequential ABC approaches (Lenormand et al., 2013; Rutter et al., 2019) and HVR (Binois et al., 2015) for uncertainty quantification (Figure 2(a)) and calibration (Figure 2(b)). The ML workloads within the HVR workflow (Figure 1) were implemented by using R-based parallelization via a new Application architecture for HVR workflow.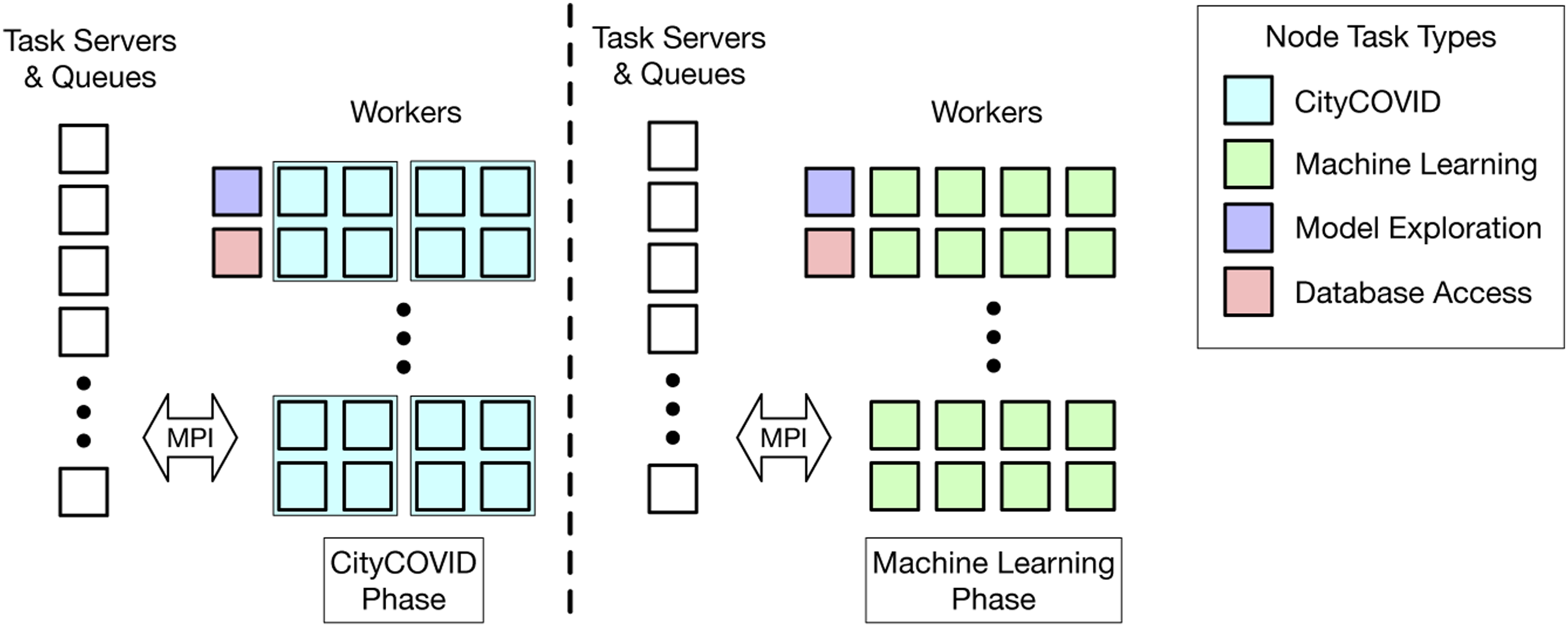
3.3.2 Load balancing and work distribution
From a computer systems perspective, a challenge for ML-based workflows is managing the differing resource requests from simulation and learning-based work items. In the CityCOVID/HVR workflow, simulation tasks require 256 ranks, and learning tasks require 64. The Swift/T model allows workflows to request tasks on any number of ranks from 1 to the number of workers allocated to the compute job. These tasks may optionally request that the ranks for a task occupy contiguous ranks starting at regular intervals or other constraints. For example, all CityCOVID workflows run the simulations on contiguous ranks starting on ranks such that rank mod 256 ≡ 0. The ML tasks run independent R-interpreters on each node of the system, each of which uses threads internally to utilize the 64 cores on each node. The database access tasks call into Python libraries using a single core at a time. Thus, Swift/T faces a significant work distribution challenge.
Swift/T translates workflows into ADLB (Lusk et al., 2010) programs. In the implementation used by Swift/T, ADLB divides the available ranks into workers and servers. Any number of servers and workers may be used, down to a singleton of each. The workers simply perform work requested from their server. Servers exchange work using the work stealing approach from Scioto Dinan et al. (2008), in which empty servers steal half of the work queue from another server selected at random. This explains the exponential ramp-up commonly seen in Swift/T performance plots. For the CityCOVID workload, we used increasing numbers of servers for increasing run sizes. Servers spent the bulk of their time looking for contiguous blocks of ranks for MPI tasks, and this work was easily shared by the work stealing approach.
3.4. Supporting public health
We have been using the CityCOVID model to forecast the spread of COVID-19 in Chicago and have been providing model results to support city and state public health officials and decision-makers. From the beginning, the decision-makers we work with have had the philosophy that their “decisions would be guided by data, science, and public health experts.” To meet these requirements, decision-makers recruited data analytics support for understanding the current COVID-19 situation and epidemiological modeling support for forecasting what might happen in the future based on possible NPIs, and they paired these groups with public health departments.
We have supported the Chicago Department of Public Health (CDPH) since the early stages of the pandemic, March 2020 (Thomas (2021)). As described in (§ 3.2.1), we have been able to estimate the extent to which individual protective behaviors, such as mask wearing and social distancing, have affected community SARS-CoV-2 transmission. These model-based insights were used by CDPH in their briefings with Chicago aldermen about the importance of maintaining COVID-19 mitigation measures in the lead up to the partial reopening on June 3, 2020.
We have also supported the Illinois Department of Public Health through the Illinois Governor’s COVID-19 Modeling Task Force (IGCMTF) (Mahr (2021)). The IGCMTF includes four modeling groups, including Argonne, that are applying various epidemiological modeling approaches to understand COVID-19 spread and explore what can be done to reduce the spread.
For interacting with public health officials, we established a weekly/biweekly routine that consists of (1) calibrating the model based on the most recent data; (2) designing and running experiments with the model to forecast effects on the spread by varying scenario parameters; (3) compiling and analyzing the model outputs from thousands of simulations; (4) applying quality control/assurance checks, which include independent subject matters experts such as epidemiologists, to the assumptions and results and identifying possible anomalies; (5) applying causal analysis to explain why the results came out as they did; and (6) reporting the results in a structured format that public health officials can easily discern and understand the implications. A standardized set of charts and graphs is provided in as succinct a form as possible and presented in live forums to public health officials by the modelers.
Since the fall of 2020 and continuing through today, questions have focused on the effects of vaccination programs and, more recently, the effects of variants of concern, in other words, mutations that, for example, result in increased disease severity, transmissibility, and vaccine escape. The fine-grained nature of the CityCOVID model allows us to track which variant an individual agent has been infected with and follow the disease course and subsequent transmission of that individual, where parameters unique to that variant can be adjusted. Our group continues to calibrate and run the model and report results to public health officials regularly.
4. How performance was measured
4.1. What applications were used to measure performance
All software was built with GCC 7.3.0 with the Cray MPI wrappers for Theta as appropriate. The CityCOVID model is based on the RepastHPC 2.3.1 agent-based modeling system and the ChiSIM Chicago model 0.4.2. The workflow system was Swift/T 1.4.3, which was linked to interpreters in Python 3.8.2.1, R 3.6.0, Tcl 8.6.6, and Cray MPI. All Python- and R-libraries used were the latest compatible with these interpreters as of October 1, 2020. The R-library size to perform these optimizations contains 112 packages totaling 321 MB. In order to improve the performance associated with numerical operations in the machine learning tasks, an external R with Cray LibSci v.20.03.1 providing vendor-optimized BLAS/LAPACK libraries was also used.
4.2. System and environment where performance was measured
All experiments were run on the Theta system at the ALCF. Theta is a Cray XC40 with 4392 compute nodes, each with an Intel KNL 7230 (Xeon Phi), aggregating 11.7 petaflops in total. Each node has 64 compute cores with access to 16 GB of high-bandwidth in-package memory, 192 GB of DDR4 RAM, and 128 GB of SSD. The system interconnect is a dragonfly network. The system has Lustre and GPFS filesystems; we used Lustre for software installations and GPFS for application data. We used the vendor-provided Cray MPICH 7.7.17.
4.3. How performance was measured
The technical details of our workflow solution are as follows. The workflow runs as a single job allocation in the Cobalt scheduler on Theta. The Swift/T runtime runs as a single MPI program across all nodes and evaluates the logic of the Swift workflow. Tasks in the workflow such as the invocation of R-code snippets (the ML algorithms) or MPI-enabled libraries (CityCOVID/C++) are placed in a distributed task queue in the ADLB library underlying Swift/T. Even database accesses are represented as Python code snippets (via Psycopg2); thus, they do not block the progress of the overall workflow. MPI tasks are launched by Swift/T using MPI 3.0 features developed previously (Wozniak et al., 2013b).
Performance was simply measured by using the clock libraries available in the programming models used and reported in log files. The precision of our measurements need be accurate only to the nearest second to describe the performance. The overall performance metric of our workflow is delivering high utilization to the compute-intensive ML and ABM modules.
5. Performance results
In this section, we present five types of performance results: an illustration of a full-scale run on Theta (§ 5.1), strong-scaling results for full-system scale (§ 5.2), time to solution results for the learning tasks (§ 5.3), communication and messaging rates (§ 5.4), and ABM specific performance results (§ 5.5).
5.1. System utilization at full scale
Figure 4 shows a full-scale run of the HVR workflow on 4098 nodes of Theta. Each colored phase corresponds to a phase shown in Figure 1. Two optimization iterations are shown from the beginning of a run. Utilization timeline on 4098 nodes of Theta consisting of machine learning and simulation phases.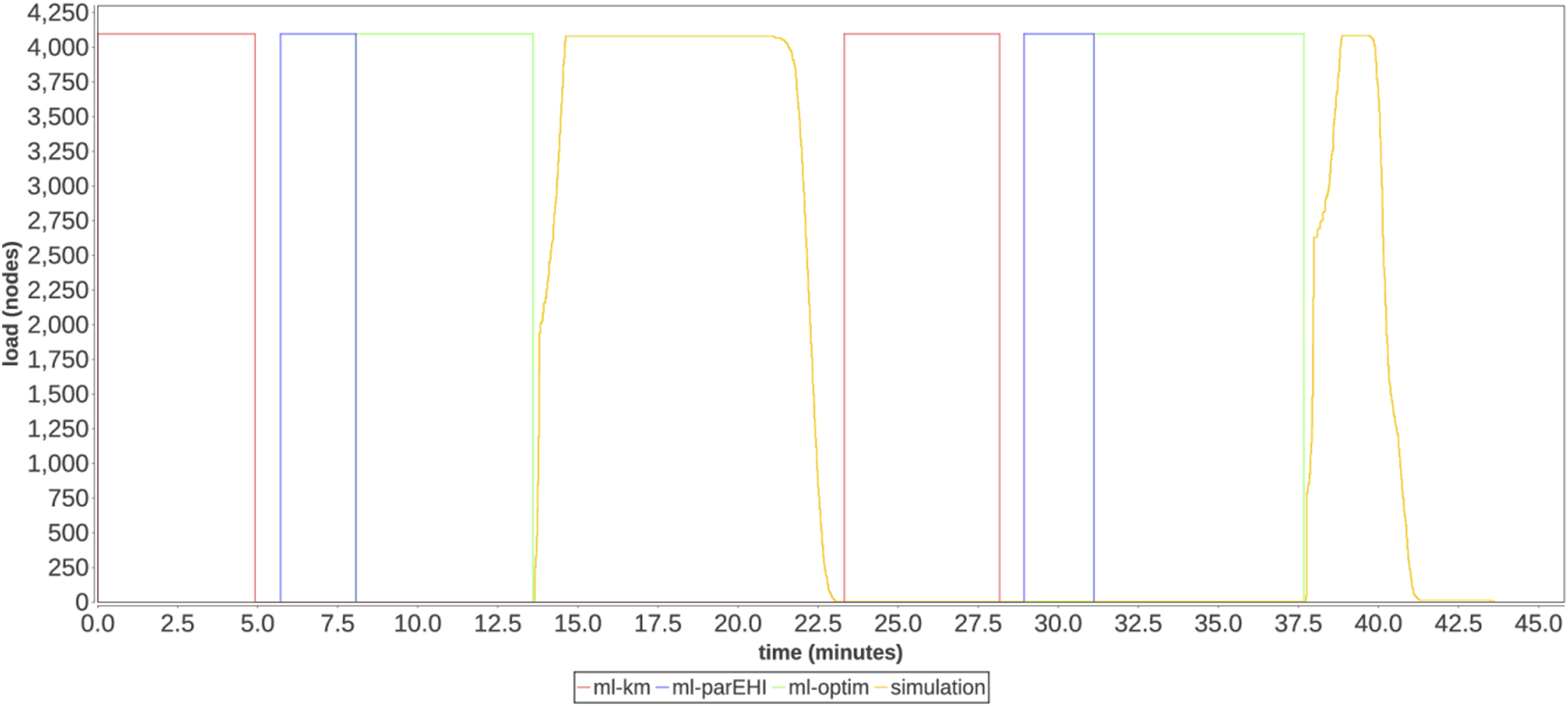
A key performance challenge of this workflow is juggling the 2040 MPI simulation tasks generated by the workflow, each a 256-core execution. As parameters for these tasks emerge from the HVR algorithm, this work must be rapidly distributed. Additionally, there are constraints on the task placement, since the MPI task should occupy contiguous cores across whole nodes to support fast communication and a data caching strategy used by CityCOVID. Nevertheless, the worker utilization across the two optimization iterations, including the learning and simulation tasks, is 84%. This is a good result considering that the Swift/T workflow scheduler operates at the rank granularity, and that 128 server ranks were distributing short (sub-minute), multiprocess tasks to 262,144 worker ranks.
The plot shows fine-grained load levels for the MPI-based simulation phase. As shown, the workflow rapidly ramps up as the simulation phase begins and ramps down more slowly as tasks exit. The first phase takes 9.4 min, whereas the second phase takes 5.9 min, demonstrating the benefit of the caching strategy. The ML task utilization is not measured directly; rather, we measure the overall time to solution in the following.
5.2. Strong scaling
The next key metric of our workflow is demonstrating strong scaling for a given problem as the computing allocation size increases. We ran the same workflow parameters on Theta in job node counts 257, 513, 1,025, 2,049, and 4098. We then measured the overall time to solution and calculated the corresponding task rates. This corresponds to 2 nodes or 128 ranks of ADLB servers (§ 3.3.2) for the largest case, and 1 node or 64 ranks of ADLB servers for the other cases. The remaining nodes were allocated as ADLB workers.
As shown in Figure 5, the time to solution and simulation rate both accelerate to full scale on Theta, and the likelihood of further scaling is encouraging. In our smallest case, the workflow takes 5 h and 54 min on 257 nodes, but at 4098 nodes it completes in under 44 min! Overall time to solution and strong-scaling plot for HVR workflow.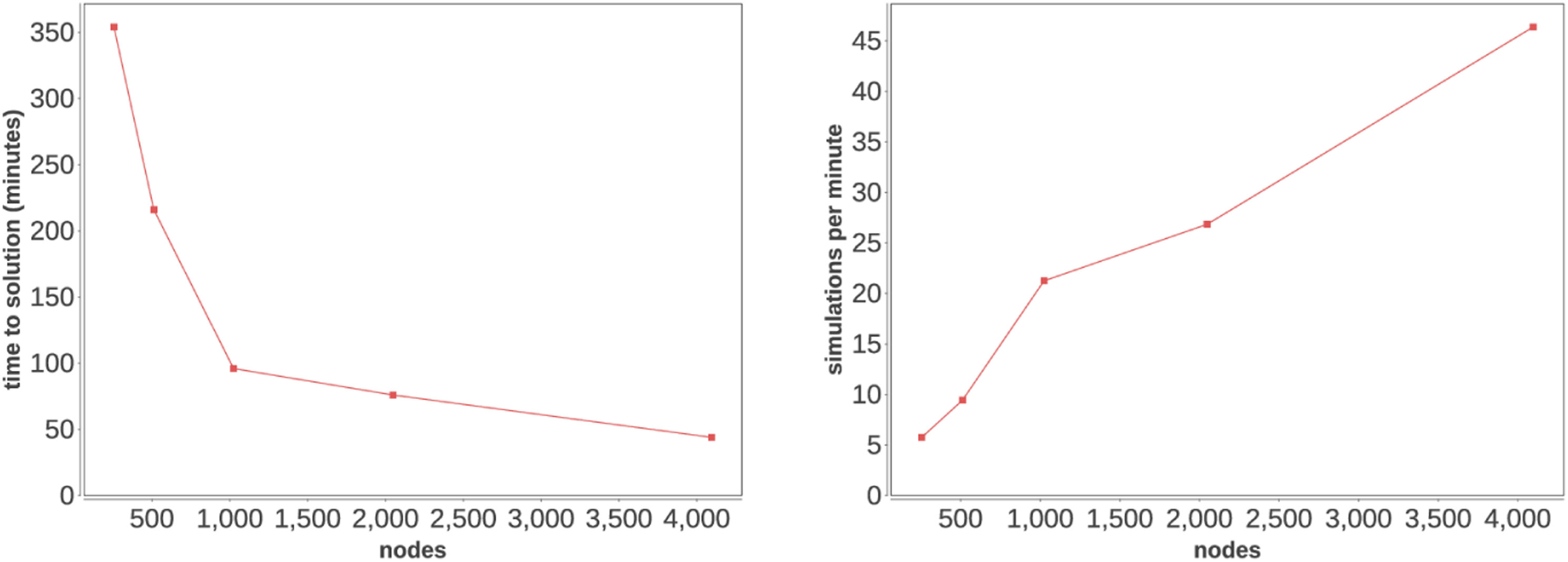
Further utilization improvements could be possible with asynchronous HVR algorithms, enabling overlap between workflow phases. The Swift/T programming model supports arbitrary data dependencies, enabling such patterns; and the flexibility of being able to directly integrating Python and R-code will facilitate such algorithmic investigations.
5.3. ML phase scaling
Here, we report the scaling results for the ML phases. The ML phase uses the novel
In Figure 6, we report the average time per phase for the ML workload. The most time-consuming phase is the Kriging model (ml-km) phase, which takes 177 s on 256 worker nodes and only 19 s on 4096 worker nodes. The least time-consuming phase is the Pareto front improvement phase (ml-parEHI), which takes 147 s on 256 worker nodes and 19 s on 4096 nodes. As shown in the plot, scaling efficiency drops off at the larger scales because the system runs low on tasks to execute, allowing the task distribution and task runtime imbalance overheads to negatively impact utilization. Time to solution for machine learning phases by node count.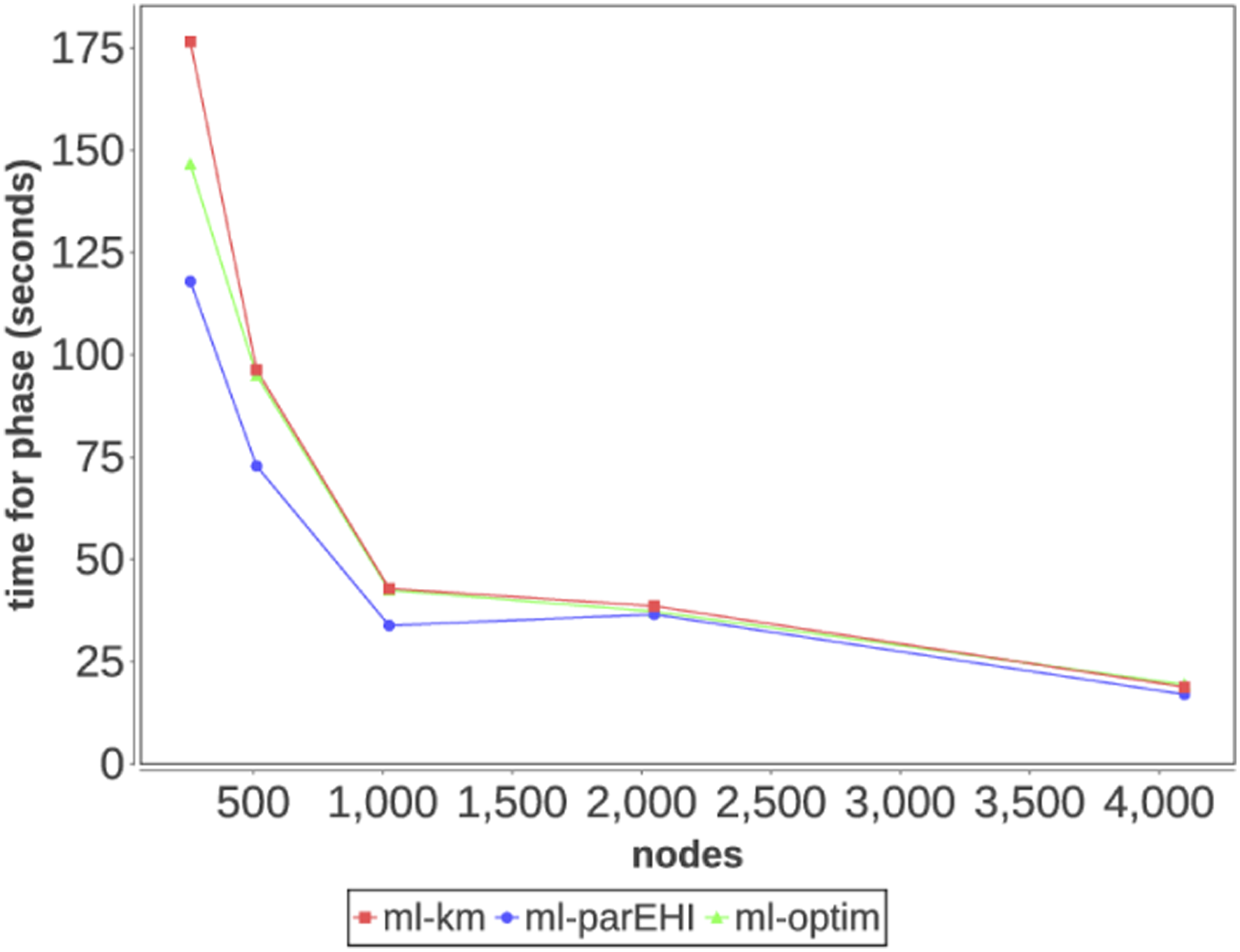
5.4. Messaging rates
Workflow variables in Swift/T are stored on the task servers and accessed as in a distributed hash table over MPI. Over the whole workflow, in the 4098-node run the system performed 722,485 retrieves in 2619 s, an aggregate rate of 276 data reads per second. During the peak period of the first simulation ramp-up, shown in Figure 4 near minute 13, the system performed 148,854 read accesses in 12.5 s, for a rate of 11,908 reads/second. This workload was spread over 128 task servers, for a rate of 93.0 accesses per second.
5.5. ABM specific performance results
Within the HVR workflow, each CityCOVID model ran for 69 simulated days (1656 h), corresponding to approximately 5 × 109 agent-hours. The significance of this metric is that it tracks how many hours agents had the potential to colocate with other agents and spread or be exposed to COVID-19. At the 4098-node scale and after the input data caching had occurred, 1020 concurrent simulations were launched and completed in 5.9 min. This corresponds to the simulation of 1.3 × 1010 potential transmissions/second. The runtime of the CityCOVID model could potentially be improved by parallelizing disease transmission and disease state transitions, either by exploiting intraprocess parallelism (threading) or reimplementing disease states and disease state transitions to be more GPU amenable. Both of these potential improvements are complicated by the necessity of logging disease states and transitions and thus could require a parallel logging capability. We have begun exploring how to leverage GPUs in an agent-based modeling toolkit with our work on Repast4Py (Collier et al., 2020), our next-generation distributed ABM toolkit in Python.
6. Implications
The implications of the success of this workflow applications are broad. Here, we describe the implications for COVID-19 modeling and public health, as well as for data science, learning, and simulation in the HPC space.
6.1. COVID-19 modeling and public health
Public health departments need guidance for prioritizing interventions in the context of limited resources and empirical data uncertainties. Through our regular meetings with city, county, and state stakeholders during the COVID-19 pandemic, we have first-hand experience in the dynamic nature of evolving mitigation priorities and the need for the detailed modeling that CityCOVID provides to rapidly coevolve. CityCOVID can simulate specific and realistic NPIs closely resembling those being considered by public health officials and can thereby guide local policy and intervention development. By integrating high-performance epidemiological simulation with large-scale machine learning, we have developed a generalizable, flexible, and performant analytical platform that can meaningfully support evidence-based decision-making during a public health crisis.
Furthermore, the COVID-19 pandemic has highlighted a number of issues, including drastic health inequities. Reducing COVID-19 morbidity and mortality will likely require an increased focus on the social determinants of health, given their disproportionate impact on populations most heavily affected by COVID-19. The detailed data-driven modeling approach that CityCOVID represents, coupled with the HPC solution developed here, has the potential to provide the robust exploration of underlying factors and reveal promising targeted interventions to reduce these observed inequities. The melding of advanced simulation, machine learning, and high-performance computing enables an in silico laboratory for identifying gaps where additional empirical data are needed and facilitates hypothesis generation.
6.2. Convergence of data science, learning, and simulation for HPC
This application demonstrates the capability and need to bring HPC closer to diverse scientific communities, including the public health sector. By using detailed local data as a starting point to parameterize and calibrate simulations and produce actionable analyses (i.e., from observational data to scenario outputs documented in an integrated Postgres database), we applied Theta as the numerical engine at the center of a data pipeline. This made the overall workflow more relevant to our public health collaborators and demonstrated the capability of the system to be an integral part of the decision-making process.
From a computer systems perspective, the complex integrations and high performance achieved here demonstrate that applications written in high-level workflow languages and mathematical systems can be executed at large scale. Additionally, they can be closely integrated with previously developed MPI-based scientific applications that are scalable in themselves. We also demonstrated that very large (thousands) ensembles of MPI program executions can be run on the system without negatively impacting the scheduler or other users by tapping into the capabilities of MPI, via the high-level model offered by Swift/T.
We expect that data+learning+simulation workflows will become more tightly integrated in the near future. As shown in Figure 4, learning is already half of the job. The capabilities shown here are initial steps to more general-purpose HPC-enabled decision-support platforms that can rapidly assess and forecast the course of disease outbreaks and other crises. We expect that incorporating real-time data streams and weather-forecast-like automated steering will drive further enhancements and increase the performance and robustness of the results obtained.
In the broad ME context of Gaussian process-based Bayesian optimization, parallel infill criteria have been developed to benefit from parallelism of modern architectures. Yet, most of these works cannot be directly transposed to HPC scale. More work is needed to leverage existing methods to scale GPs with millions of fixed observations to the sequential framework. Thus, developing algorithms that are specifically geared toward and can take advantage of HPC resources has the potential to advance computational science and open up previously unexplored areas of statistical research. There are trade-offs to consider, such as between accurately selecting the optimal points for a given task and the time it takes to make this selection at the specified accuracy. Besides the ability to tackle more complex problems with an increasing number of variables, higher concurrency raises challenges in coping with consideration of synchronous versus asynchronous algorithms. Support for asynchronous algorithms (see, e.g., Janusevskis et al. (2012)) is a promising perspective for future work.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the DOE Office of Science through the National Virtual Biotechnology Laboratory, a consortium of DOE national laboratories focused on response to COVID-19, with funding provided by the Coronavirus CARES Act. This work was also funded by an award from the c3.ai Digital Transformation Institute and the National Institute of Allergy and Infectious Diseases grant R01AI136056. This research used resources of the Argonne Leadership Computing Facility, which is a DOE Office of Science User Facility supported under Contract DE-AC02-06CH11357. We would like to thank the Argonne Leadership Computing Facility staff for their timely and critical support. This research was completed with resources provided by the Laboratory Computing Resource Center at Argonne National Laboratory. This material is based upon work supported by the U.S. Department of Energy, Office of Science, under contract number DE-AC02-06CH11357. This research was also supported by the Exascale Computing Project (17-SC-20-SC), a collaborative effort of the U.S. Department of Energy Office of Science and the National Nuclear Security Administration.
Author biographies
Jonathan Ozik is a Computational Scientist at Argonne National Laboratory and Senior Scientist with Public Health Sciences affiliation in the Consortium for Advanced Science and Engineering at the University of Chicago. He develops applications of large-scale agent-based models, including models of infectious diseases, healthcare interventions, biological systems, water use and management, critical materials supply chains, and critical infrastructure. He also applies large-scale model exploration across modeling methods, including agent-based modeling, microsimulation and machine/deep learning. He leads the Repast project for agent-based modeling toolkits and the Extreme-scale Model Exploration with Swift (EMEWS) framework for large-scale model exploration capabilities on high-performance computing resources. He has been awarded an R&D 100 award in 2018 for contributions to the Swift/T workflow software.
Justin M Wozniak is a Computer Scientist at Argonne National Laboratory and a Scientist-At-Large at the University of Chicago. He participates in the Exascale Computing Project in cancer applications, deep learning, workflow systems, and streaming data analysis, and is part of the Argonne COVID-19 project. He won an R&D 100 award in 2018 for the Swift/T workflow software. He is the co-founder and co-chair of the Workshop on Experiment-in-the-Loop Computing (XLOOP@SC) (2019-present) and co-organizer of the DOE Workflows Workshop (ongoing). His research interests are in high-performance computing, programming systems, and storage systems.
Nicholson Collier is a Software Engineer at Argonne National Laboratory, and Staff Software Engineer in the Consortium for Advanced Science and Engineering at the University of Chicago. He develops, architects, and implements large-scale agent-based models and frameworks in a variety of application areas, including the transmission of infectious diseases, biological systems, and critical materials supply chains. He also develops large-scale model exploration workflows across various domains, including agent-based modeling, microsimulation and machine/deep learning. He is the lead developer for the Repast project for agent-based modeling toolkits and co-developer of the Extreme-scale Model Exploration with Swift (EMEWS) framework for large-scale model exploration capabilities on high-performance computing resources.
Charles M Macal is a Distinguished Fellow, Senior Systems Engineer, and Group Leader for Social and Behavioral Systems at Argonne National Laboratory. He applies computational modeling and simulation tools to complex systems to solve problems in a variety of fields, including energy, infectious diseases, and emergency preparedness. He is the chief scientist for the Decision and Infrastructure Sciences Division, and is a principal investigator for the development of the widely used Repast agent-based modeling toolkit. He has appointments at the University of Chicago Consortium for Advanced Science and Engineering and the Northwestern-Argonne Institute for Science and Engineering. He is a registered professional engineer in the State of Illinois. He is a Senior Member of IEEE, ACM, and the Society for Computer Simulation, and serves on the editorial boards of several professional journals.
Mickaël Binois is a junior researcher at Inria Sophia Antipolis in the Acumes project-team. Prior to joining Inria, Dr Binois was a postdoctoral researcher in the MCS division of Argonne National Laboratory and with Robert Gramacy at the University of Chicago Booth School of Business. His research interests include surrogate models (especially Gaussian processes), computer experiments, Bayesian optimization (constrained, single or multi-objective), and uncertainty quantification. He has led and contributed to multiple open source R-packages for optimization including GPareto, hetGP, DiceOptim, GPGame, and activegp.
