Abstract
Bisphenol A (BPA) is a prevalent environmental endocrine disruptor with potential impacts to the neurological system in humans. This study used an integrated method combining network toxicology, molecular docking, and molecular dynamics simulations to explore the molecular mechanisms underlying BPA-induced neurotoxicity. We identified 255 potential neurotoxicity-related targets through the integration and comprehensive analysis of multiple data sources, including the Comparative Toxicogenomics Database (CTD), ChEMBL, STITCH, GeneCards, and the Online Mendelian Inheritance in Man (OMIM) database. Analysis of the protein-protein interaction (PPI) network unveiled 52 core targets, among which TNF, TP53, INS, ESR1, and PTGS2 emerged as pivotal hubs in the toxicity network. Functional enrichment analysis indicated that the core targets of BPA’s influence on neurotoxicity are predominantly enriched in vital signaling cascades, including inflammatory responses, pathways of neurodegeneration, MAPK signaling pathway, serotonergic synapse pathway, and pathways in cancer. Molecular docking results demonstrated that BPA exhibited stable binding interactions with core targets. Furthermore, molecular dynamics simulations provided insights into the interactions between BPA and key targets (ESR1, TNF, and TP53), supporting the potential conformational stability of these complexes. Collectively, these computational findings contribute to understanding the potential molecular mechanisms of BPA-induced neurotoxicity and are informative for generating hypotheses related to its pathogenesis.
Introduction
Bisphenol A (BPA), an environmental endocrine-disrupting chemical, poses significant health risks owing to its extensive application in producing various consumer goods such as heat-sensitive paper, polycarbonate plastics, and epoxy resins. 1 However, the demand for BPA-containing products has continued to rise over the past few decades, leading to widespread environmental contamination and near-continuous exposure through dietary intake, air inhalation, and dermal contact. 2 Due to its bioaccumulation in living tissues, prolonged exposure to BPA can exacerbate adverse health effects.3,4
There is growing evidence that BPA exposure affects obesity, 5 insulin, 6 glucose homeostasis, 7 metabolism, 8 and brain function. 9 Due to its lipophilic properties, BPA can cross the blood-brain barrier and accumulate in neural tissues, thereby exerting adverse effects on the nervous system. 10 Extensive human studies have demonstrated that BPA exposure during prenatal development, infancy, childhood, and adulthood can disrupt brain development and neurological function, leading to a wide range of neurological disorders. 11 Mechanistically, BPA has been shown to elevate oxidative stress and amyloid-beta accumulation in the brains of patients with Alzheimer’s disease (AD), while simultaneously promoting neuroinflammation and tau hyperphosphorylation. 12 Additionally, experimental evidence indicates that BPA exposure may induce neurotoxicity through the degeneration of dopaminergic neurons, thereby increasing susceptibility to Parkinson’s disease (PD). 13 Notably, epidemiological data also links BPA exposure to an elevated risk of neurovascular conditions such as stroke. 14 These findings collectively highlight that BPA-induced neurotoxicity is not an abstract phenomenon but rather operates through specific pathological pathways to contribute to distinct central nervous system disorders, including AD, PD, and stroke,15,16 providing a disease-level reference for understanding its toxic mechanisms.
Neurotoxicity refers to the harmful effects of external factors (such as chemical substances, physical factors, and biological factors) on the structure or function of the nervous system, which can damage neurons, synapses, or nerve conduction pathways and thereby induce neurological dysfunction. Recent studies have demonstrated that exposure of HT-22 cells to BPA significantly increases lactate dehydrogenase leakage rates, impair cell membrane integrity, and activate apoptotic pathways, ultimately leading to cell death. 17 BPA disrupts hippocampal neurogenesis by impairing the regulatory function of the ubiquitin-proteasome system. 18 Additionally, BPA exposure inhibits the Nrf2/HO-1 and AKT/mTOR signaling pathways, exacerbating oxidative stress, apoptosis, and excessive autophagy, thereby accelerating neuronal degeneration. 19 BPA also prematurely accelerates neuronal differentiation and migration, which may perturb normal neocortical development. 20 Neurotoxicity assessments confirm BPA’s harmful effects across species. Detailed observations of zebrafish embryos reveal that even a low dose of BPA can interfere with the normal upregulation of key neurodevelopmental genes (e.g., Ngn1, Elavl3, and GFAP), disrupting central nervous system development and causing behavioral abnormalities at key developmental stages. 21 In mammals, BPA exposure elevates NRG1-positive neuron populations in the colon, potentially linked to neurotoxic responses and the activation of compensatory protective mechanisms. 22
Network toxicology integrates multi-source biological data to investigate the toxic mechanisms of chemicals from a systems biology perspective. Molecular docking and molecular dynamics simulations are computational tools used to predict interactions between compounds and their potential targets. 23 These approaches provide a novel framework for in-depth exploration of complex biological mechanisms. Previous network toxicology studies on BPA have identified similar multi-target regulatory patterns in diverse conditions, including multiple sclerosis (MS), systemic lupus erythematosus (SLE), rheumatoid arthritis (RA), and AD.7,24 While other research groups have previously applied this approach to explore BPA-induced neurotoxicity, 25 our study extends this framework by integrating a broader range of target databases and incorporating molecular dynamics simulations to validate the binding stability of core targets. This enhancement provides a more comprehensive perspective on BPA’s multi-target regulatory network.
In this study, we employed network toxicology, molecular docking, and molecular dynamics simulations to analyze the potential mechanisms of BPA-induced neurotoxicity from systemic to molecular levels, aiming to provide theoretical support for the prevention and treatment of related neuropathological conditions. A detailed breakdown of the study’s research content is outlined in the flow chart (Figure 1). The flow chart of this study is to explore the potential molecular mechanism of BPA-induced neurotoxicity.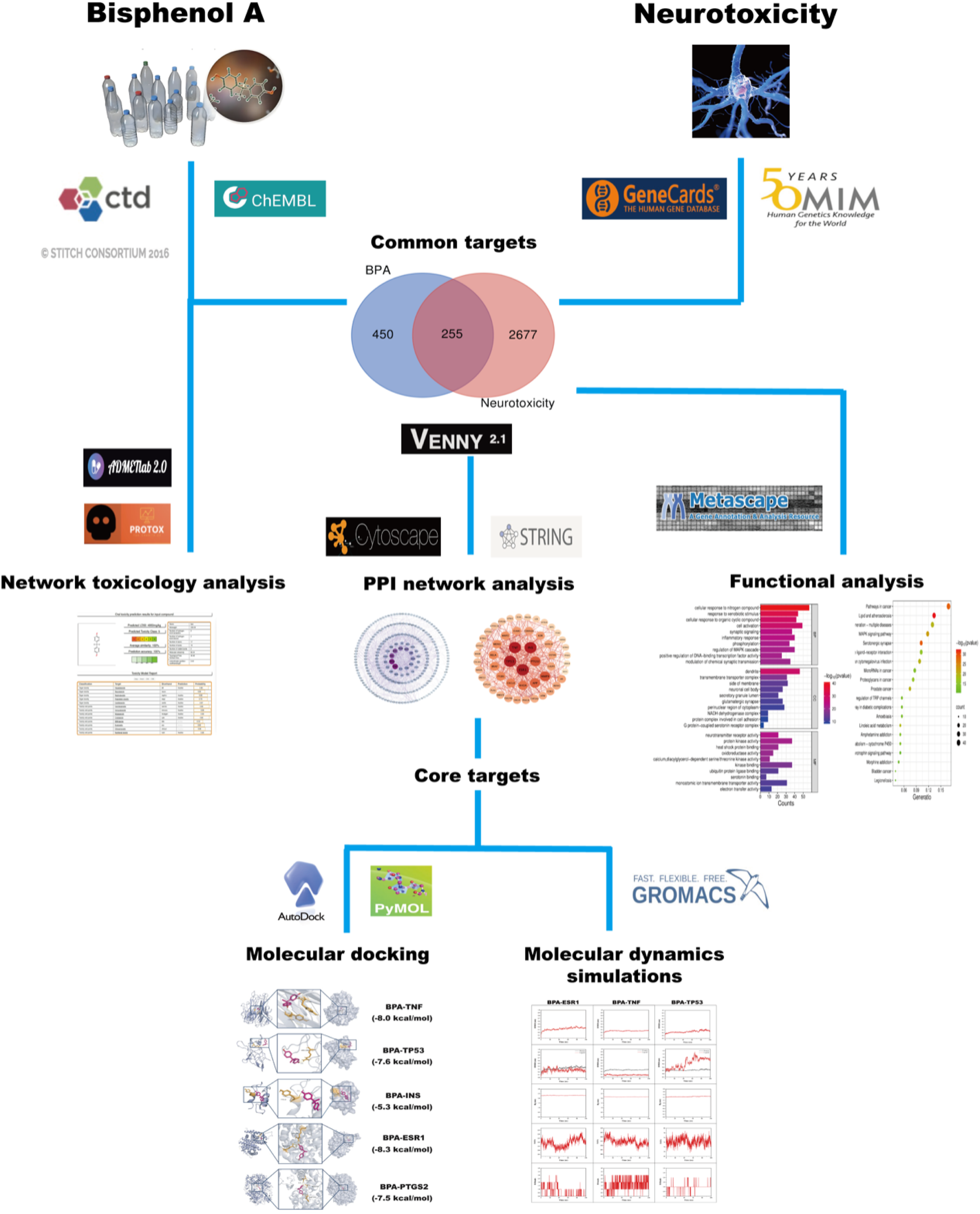
Methods
Network Toxicology Analysis of BPA
Based on the chemical structure of BPA, we used 2 established computational platforms, ADMETlab 2.0 26 and ProTox 3.0, 27 to comprehensively predict the toxicological properties of BPA. Through these 2 platforms, we obtained toxicological predictions for BPA, which were used in subsequent network analysis.
Collection of BPA-Related Targets
The structural formula and SMILES notation of BPA were retrieved from the PubChem database using the query term “bisphenol A.” Using this structural data, we extracted potential BPA targets from the ChEMBL and CTD databases by searching for the keyword “bisphenol A” and refining the results to “Homo sapiens.” In order to increase the biological significance of target selection, we applied a screening criterion of “interaction count” greater than the median in the CTD database. Additionally, BPA’s SMILES code was input into the STITCH database with the minimum interaction score set to “high confidence” to identify potential targets that might have been overlooked in initial searches. After merging targets and removing duplicates, the acquired target names were normalized against the UniProt database. The accuracy and consistency of the merged target list were verified through cross-referencing multiple databases, redundancy removal, and the application of biologically relevant consistency thresholds.
Selection of Target Networks Related to Neurotoxicity
Using GeneCards and OMIM databases, we conducted a keyword search to identify relevant targets related to “neurotoxicity.” To ensure the highest possible relevance to neurotoxicity, we applied strict screening procedures. This involved setting the “score” cutoff at the level of the median, eliminating genes with low “score” values, and creating a specialized neurotoxicity target gene library. Additionally, we used Venn diagram analysis to identify targets that overlap both BPA-related and neurotoxicity-associated targets, thereby highlighting potential targets mediating BPA-induced neurotoxicity.
Protein Interaction Network Construction and Core Targets Screening
Potential targets of BPA-induced neurotoxicity were input into the STRING database with the species selection restricted to “Homo sapiens.” For the purpose of analysis, the “minimum required interaction score” was set to “medium confidence >0.4.” The protein-protein interaction (PPI) network was extracted from the STRING database and imported into Cytoscape software (version 3.8.2) for visualization. Network topology parameters were computed utilizing the NetworkAnalyzer plugin within the Cytoscape software platform. We sifted through the genes based on the computed primary score file from NetworkAnalyzer, selecting those with a higher-than-median score for Betweenness Centrality (BC), Closeness Centrality (CC), and Degree Centrality (DC). A foundational subnetwork was created with sifted genes for further analysis.
Gene Function and Pathway Enrichment Analysis of Target Protein
To investigate the potential biological roles of BPA in neurotoxicity, we systematically retrieved data from the Metascape database to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses. A thorough GO analysis was performed, encompassing evaluations of biological processes (BPs), cellular components (CCs), and molecular functions (MFs). Additionally, KEGG pathway enrichment analysis was employed to identify critical pathways associated with BPA-induced neurotoxicity. This strategy focuses on both therapeutic and potential toxic pathways, enabling more precise target identification and a deeper comprehension of BPA’s mechanistic impact. To ensure reliability, strict filtering criteria were applied: the cutoff
Molecular Docking of BPA With Core Targets
We conducted molecular docking simulations to study the intermolecular forces and binding patterns between BPA and the identified key target proteins. This structure-based theoretical method allows for conformations and binding affinity of receptors and ligands to be predicted. The crystal structures of the core target proteins were obtained from the Protein Data Bank (PDB) at the RCSB. The crystal structures of the key target proteins selected from the RCSB PDB are to be obtained using these PDB identifiers: TNF (PDB ID: 1A8M), TP53 (PDB ID: 2G3R), INS (PDB ID: 1A7F), ESR1 (PDB ID: 2BJ4), and PTGS2 (PDB ID: 5F19). Using PyMOL, we processed the protein structures by eliminating water molecules and the original ligands. The processed proteins were then loaded into AutoDockTools 1.5.7 for hydrogenation, charge computation, and optimization of nonpolar hydrogen bonds. Employing precisely configured grid box dimensions and genetic algorithms, molecular docking was executed via AutoDock Vina using CMD commands. The docking outcomes were visualized with PyMOL offering detailed insights into the intricate interactions between BPA and the core target proteins.
Molecular Dynamics Simulations
Molecular dynamics (MD) simulations were executed with GROMACS (version 2022) utilizing the Amber99sb-ildn force field for proteins and GAFF for the small molecules. The starting structure was based on the most stable BPA-protein complex configuration uncovered during molecular docking. The system was encased in a cubic box, filled with the TIP3P water model and subjected to periodic boundary conditions, with chloride ions added for charge balance. Electrostatics were handled through the Particle-mesh Ewald technique, and energy was minimized using the steepest descent approach. Both electrostatic and van der Waals forces were capped at a distance of 1 nm. After equilibration in the NPT ensemble, a 100 ns molecular dynamics run was conducted at standard temperature and pressure. The non-bonded interactions were truncated at 10 Å. During the simulation, the temperature was kept at 300 Kelvin using the Langevin thermostat, and the pressure was controlled at 1 atmosphere using the Berendsen barostat. The system was evaluated using GROMACS built-in tools: gmx rms for RMSD calculations with the protein backbone as a reference, gmx gyrate for Rg analysis, gmx sasa for SASA calculations with a probe radius of 1.4 Å, and gmx hbond for hydrogen bond analysis, applying a default distance cutoff of 3.5 Å and an angle cutoff of 30°. Finally, the binding free energy between the protein and the small molecule ligand was analyzed using the generalized Born and surface area solvation (MM/GBSA) method.
Results
Fundamental Computational Toxicity Assessment of BPA
ProTox 3.0 database predictions reveal that BPA exhibits biologically active ecotoxicity, blood-brain barrier penetration capabilities, and a potent binding affinity with the estrogen receptor ESR1 (Supplemental Material S1). ADMETlab 2.0 evaluations identify BPA as having potential ocular and dermal toxicity, with predicted interactions with estrogen receptor α (ER) and its ligand-binding domain (ER-LBD) (Supplemental Material S2). Peer-reviewed studies corroborate the well-documented relationship between estrogen signaling and neurotoxic outcomes. 28 These analyses establish a critical foundation for subsequent systematic investigations into BPA-triggered neurotoxicity.
Identification of Targets of BPA Affecting Neurotoxicity
This study began with an initial screening process that yielded 705 BPA-related targets from the CTD, ChEMBL, and STITCH databases. Next, we used detailed information from the GeneCards and OMIM databases to identify 2,932 targets closely related to neurotoxicity. By carefully integrating and eliminating redundancy among these targets, we refined our focus to a core group of 255 cross-targets that emerged as potential candidates for BPA’s effects on neurotoxicity. The Venn diagram depicts BPA’s target genes associated with neurotoxicity (Figure 2A). The network diagram visualizes the interactions between these elements, where green squares indicate genes, blue circles symbolize BPA, and red squares represent neurotoxicity (Figure 2B). Venn diagram and network relationship between potential BPA targets and neurotoxicity. (A) Venn diagram of BPA and neurotoxicity targets; (B) BPA-target-neurotoxicity network diagram, with green boxes representing genes, blue circles representing BPA, and red boxes representing neurotoxicity.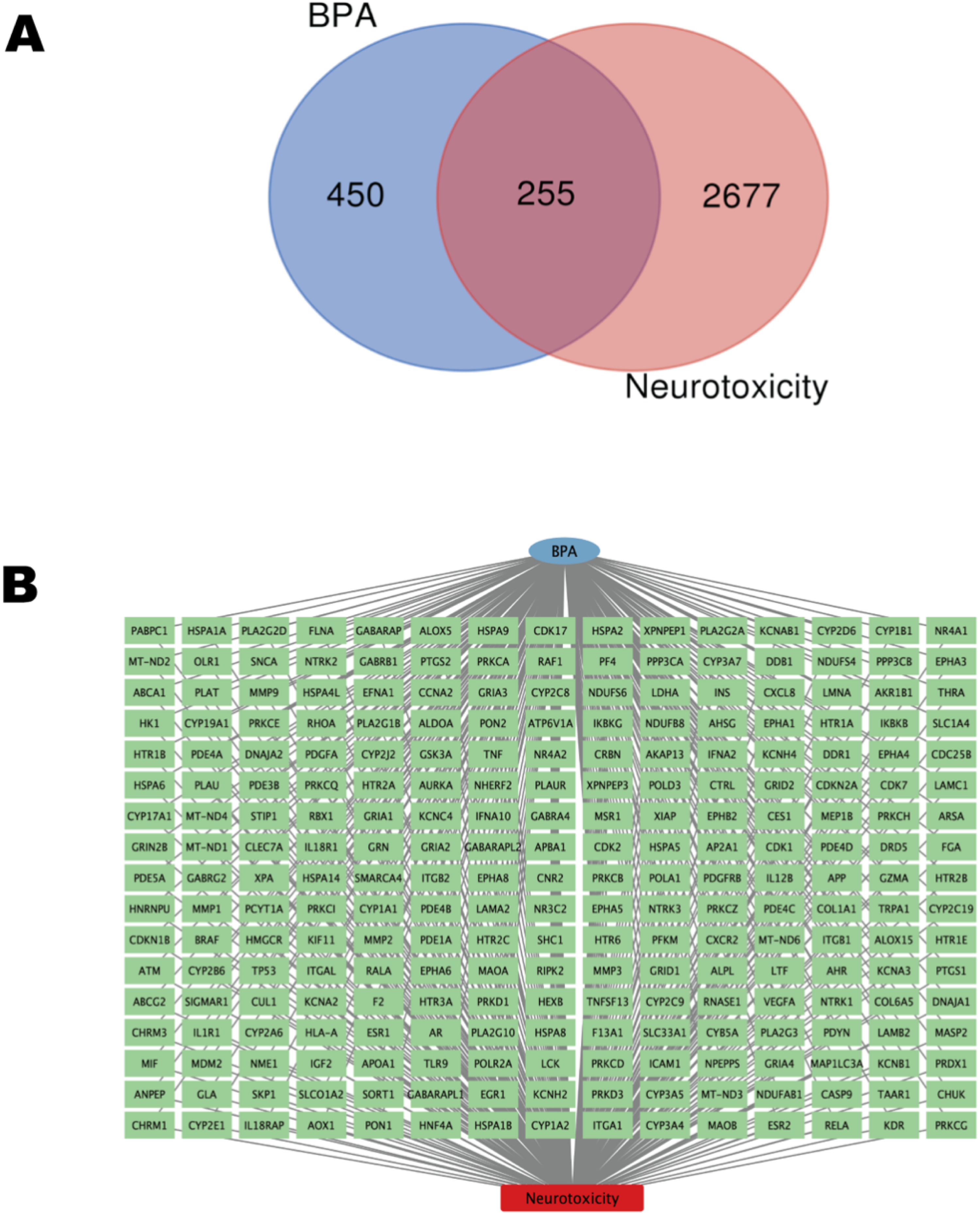
PPI Network Construction and Core Target Screening
Using the STRING database, we mapped potential targets linked to BPA-induced neurotoxicity onto the PPI network. This resulted in a densely connected network comprising 251 nodes and 2,437 edges. Subsequent analysis using Cytoscape software enabled us to examine the topological features of these nodes in detail, producing a refined, visually clear PPI network diagram (Figure 3A), which provides a foundation for subsequent functional investigations and mechanistic studies. This graphical representation illustrates the complex relationships between the potential targets identified in the study. It also visually encodes their relative importance: the size of the nodes and the intensity of their coloration are proportional to their degree of connectivity, with the largest and darkest elements corresponding to the most central and representative targets. Network analysis revealed 52 key targets linked to neurotoxicity caused by BPA exposure. We prioritized the top 5 nodes based on their degree centrality: TNF, TP53, INS, ESR1, and PTGS2. The network diagram showed the functional relationships within the core targets (Figure 3B). PPI network of potential targets of neurotoxicity induced by BPA. (A) PPI network of protein targets. (B) PPI network of core protein targets. Circles represent target proteins. The larger the diameter of the circle, the brighter the color, and the higher the degree value. Straight lines represent interactions between target proteins.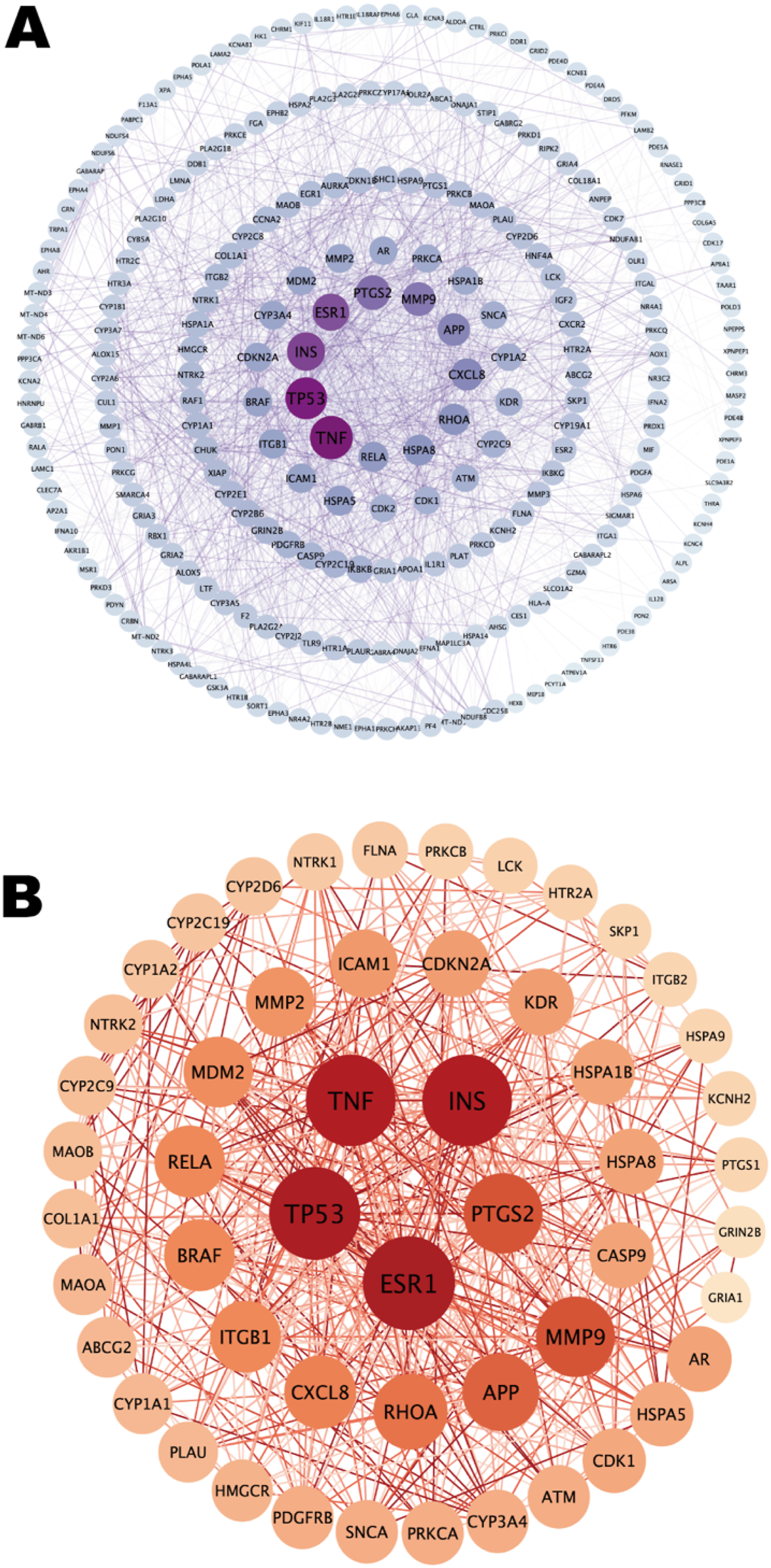
GO and KEGG Analyses of Potential Targets
Using the Metascape database with Homo sapiens as the restricted species, we performed GO analysis on 255 potential targets, which yielded 1926 statistically significant GO entries, including 1569 biological processes (BPs), 138 cellular components (CCs), and 219 molecular functions (MFs). The GO entries were arranged based on FDR values, and we show the top 10 BP, CC, and MF with the lowest FDR values (Figure 4A). Additionally, Metascape was used for KEGG analysis of the 255 targets to identify their roles in specific signaling pathways. Among the 224 enriched signaling pathways, the top 20 (by lowest FDR values) were visualized via statistical bubble plots (Figure 4B). GO and KEGG enrichment analyses of potential targets of BPA neurotoxicity. (A) The bar graph of the top 10 enriched entries for each GO category (BP, CC, and MF) in reverse order of FDR values. (B) The bubble diagram of the top 20 enriched KEGG signaling pathways in reverse order of FDR values.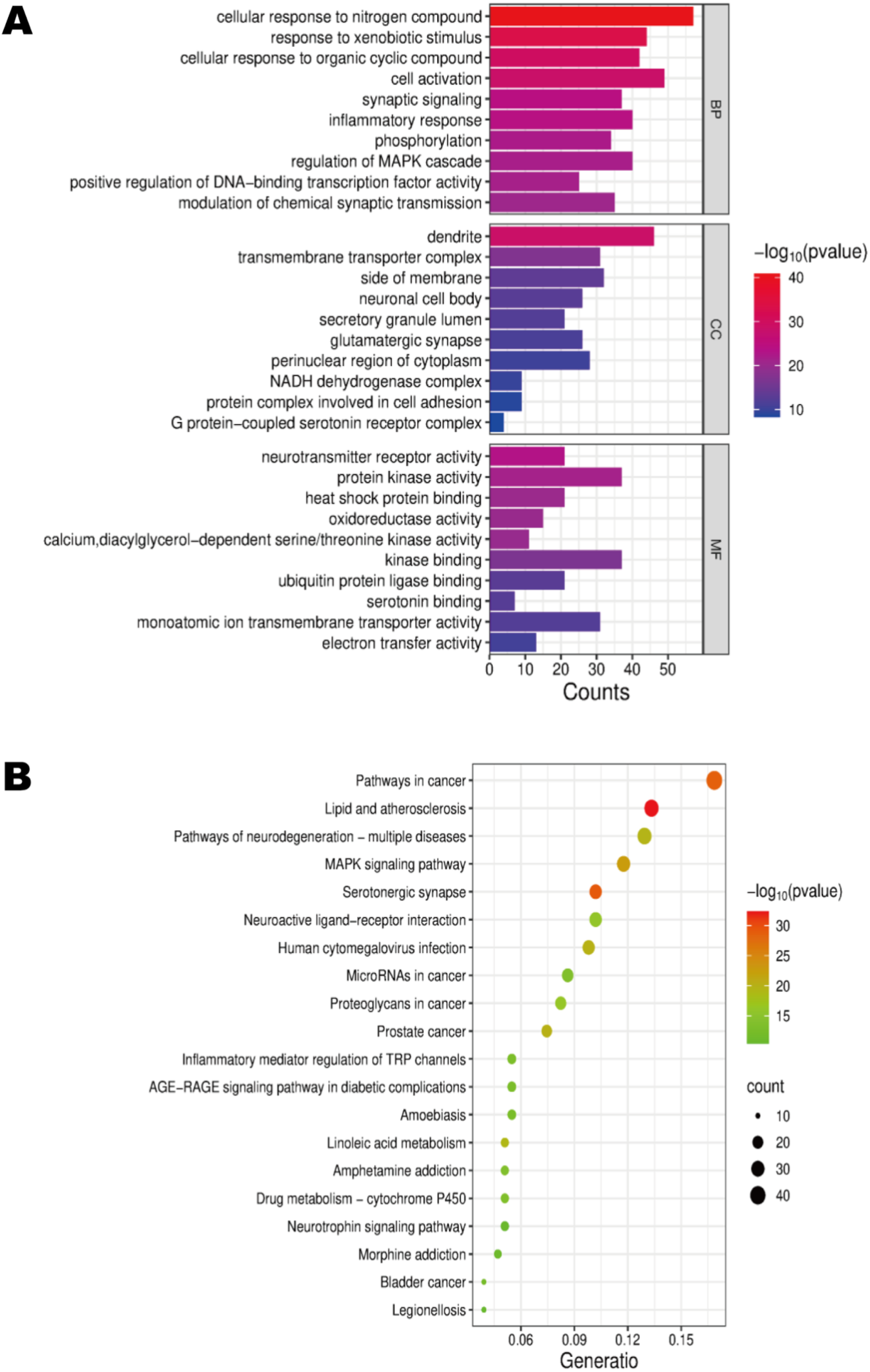
Based on the GO and KEGG analyses of potential targets, these analyses indicated that the BP terms primarily involved cellular response to nitrogen compound, response to xenobiotic stimulus, and cell activation. CCs encompassed dendrite, receptor complex, and postsynapse. The MFs are related to neurotransmitter receptor activity, protein kinase activity, and phosphotransferase activity. The KEGG signaling pathways were concentrated, showing significance across an array of pathways including lipid and atherosclerosis, serotonergic synapse, pathways in cancer, MAPK signaling pathway, PI3K-Akt signaling pathway, Ras signaling pathway, and pathways of neurodegeneration—multiple diseases.
Molecular Docking Validation Results
Molecular docking analysis was used in the study to explain the interaction among BPA and 5 core targets, TNF, TP53, INS, ESR1, and PTGS2. Based on the AutoDock Vina scoring functions, the computational assessment revealed numerical variations in the predicted binding affinity of BPA for these target proteins. The ESR1 exhibited the lowest docking score of −8.3 kcal/mol. This was followed by the TNF at −8.0 kcal/mol and the TP53 at −7.6 kcal/mol. Meanwhile, the PTGS2 and INS showed docking scores of −7.5 and −5.3 kcal/mol, respectively. These docking scores indicate differences in the predicted binding affinity of BPA for these core targets. To further elucidate the complex binding conformation, we performed visualization using PyMOL to demonstrate the lower energy binding conformation (Figure 5). Docking simulations identified specific interactions such as hydrogen bonds and van der Waals forces, clarifying the binding pattern and stability of BPA with its core protein targets. Visualization not only vividly demonstrated the close interaction between BPA and key targets but also provided important evidence for analyzing the structural basis of their interactions, underscoring the potential role of BPA in regulating neurotoxicity at the molecular level. Molecular docking results of BPA with the lowest binding energy in each target. (A) BPA and TNF; (B) BPA and TP53; (C) BPA and INS; (D) BPA and ESR1; (E) BPA and PTGS2.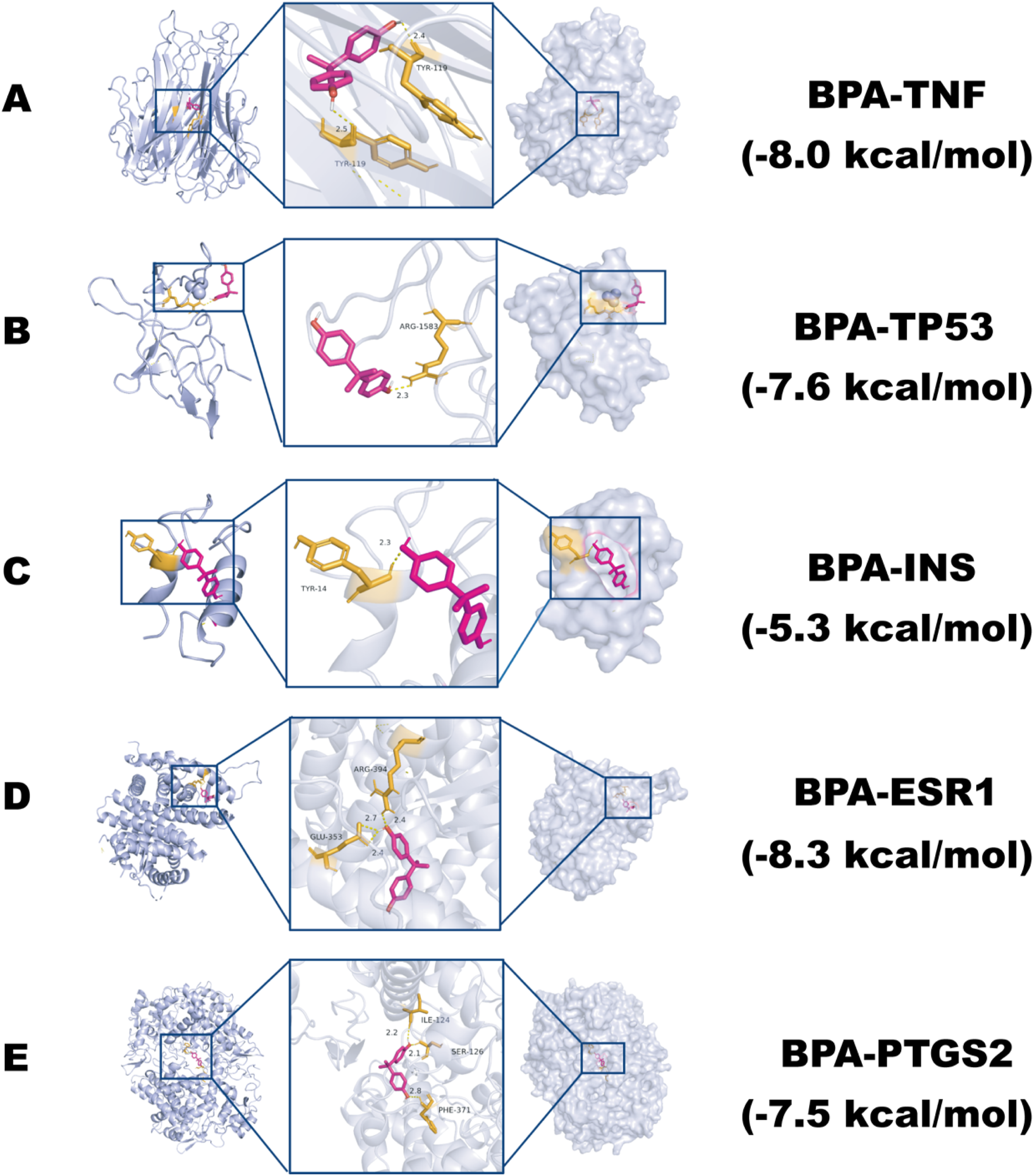
Molecular Dynamics Simulations and Binding Free Energy Assessment Results
Building on the earlier predictions of BPA toxicity targets and molecular docking binding energy analysis, ESR1, TNF, and TP53, which exhibit lower predicted binding energies, were chosen for molecular dynamics simulations with BPA. The root-mean-square deviation (RMSD) is a key indicator of system stability, and its fluctuations are directly related to atomic displacement. The BPA-TP53, BPA-TNF, and BPA-ESR1 complexes showed slight RMSD fluctuations in the initial 20 ns, but these fluctuations gradually stabilized over time and eventually reached equilibrium (Figure 6A). The complexes maintained relatively stable RMSD values in the equilibrated phase (after 20 ns), with BPA-ESR1 at 0.35 ± 0.03 nm, BPA-TNF at 0.26 ± 0.01 nm, and BPA-TP53 at 0.25 ± 0.03 nm. While most proteins and ligands maintained stable RMSD profiles, the BPA ligand in the BPA-TP53 complex displayed more dynamic behavior with fluctuations throughout the simulation (Figure 6B). The radius of gyration (Rg) is an important parameter for measuring the compactness of protein structure. The smaller the Rg value, the more compact the protein structure. In the 100 ns simulation, BPA-TP53, BPA-TNF, and BPA-ESR1 exhibited Rg values of 1.47 ± 0.01 nm, 2.15 ± 0.01 nm, and 2.31 ± 0.01 nm, respectively (Figure 6C). The solvent-accessible surface area (SASA) metrics, which quantify protein-solvent interactions and hydrophobic characteristics, showed that these 3 complexes maintained stable SASA profiles, with values of 219.25 ± 5.15 nm2, 194.78 ± 3.85 nm2, and 74.35 ± 1.55 nm2 for BPA-ESR1, BPA-TNF, and BPA-TP53, respectively (Figure 6D). The number of hydrogen bonds reflects the strength of the protein-ligand interaction, and the 3 complexes exhibited steady hydrogen bond density and strength throughout the simulation: BPA-ESR1 averaged 0.58 ± 0.73, BPA-TNF 1.68 ± 0.73, and BPA-TP53 1.00 ± 0.29 (Figure 6E). Further, the MM/GBSA method was used to calculate the binding free energies (ΔGbind) of the 3 protein-ligand complexes. As a lower ΔGbind value indicates stronger receptor-ligand binding affinity, the ΔGbind values of the BPA-ESR1, BPA-TNF, and BPA-TP53 complexes were −26.47 ± 4.91 kcal/mol, −40.69 ± 3.1 kcal/mol, and −18.4 ± 5.07 kcal/mol, respectively. Molecular dynamics simulation results of BPA with 3 core target proteins (ESR1, TNF, and TP53). (A) Root-mean-square deviation (RMSD) values of the BPA-protein complexes during 100 ns simulation: the complexes showed slight RMSD fluctuations in the initial ∼20 ns, followed by stabilization, indicating overall stability of the BPA-protein complexes. (B) RMSD values of the proteins and ligands during 100 ns simulation: most proteins and ligands maintained stable RMSD profiles, while the BPA ligand in the BPA-TP53 complex exhibited continuous fluctuations, reflecting its relative instability. (C) Radius of gyration (Rg) values of the BPA-protein complexes: the Rg values of the complexes remained within a stable range, indicating compact structures of the target proteins in the complexes. (D) Solvent-accessible surface area (SASA) values of the BPA-protein complexes: the complexes maintained stable SASA profiles, suggesting strong binding interactions between BPA and the 3 core targets. (E) Number of hydrogen bonds in the BPA-protein complexes: the complexes maintained steady hydrogen bond density, indicating the stability of interactions between BPA and the core targets.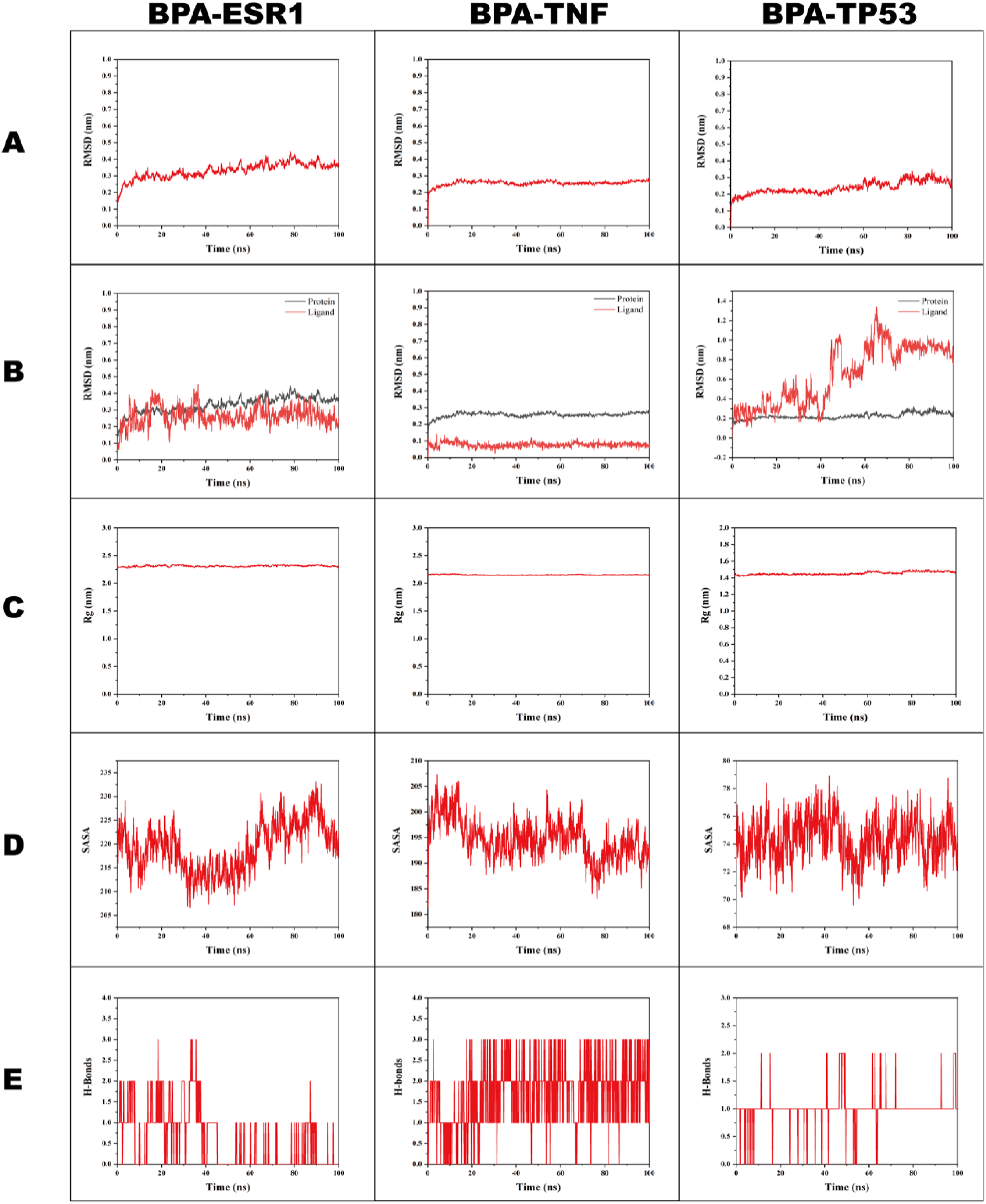
Discussion
This study employed standard computational toxicology methods to systematically explore BPA-induced neurotoxicity and generate testable hypotheses. The investigation identified 52 key molecular targets and demonstrated their functional connections. Identifying these central targets informs on how BPA may cause neurotoxicity. Network topology analysis showed that key regulatory molecules such as TNF, TP53, INS, ESR1, and PTGS2 are in central positions in the network. This suggests that they may play a vital role in BPA-induced neurotoxicity. Further research suggests that these core targets are primarily associated with biological processes, such as responses to external stimuli, synaptic signaling, the regulation of neuroinflammation, and responses to oxidative stress. These processes are thought to contribute to the clinical manifestations of BPA-induced neurotoxicity. These pathways are associated with significant activity in cancer, MAPK signaling, and neuroactive ligand-receptor interaction pathways, suggesting that BPA may cause neurotoxicity by disturbing multiple neuromodulatory pathway networks. Network analysis suggests that BPA may produce neurotoxic effects through a sophisticated mechanism that affects numerous neural signaling pathways simultaneously rather than targeting a single molecule. These findings contribute to understanding potential neurotoxic pathways of BPA and identifying candidate key molecules, which are informative for generating hypotheses in future research on BPA-related neuroprotective strategies.
Network analysis highlights potential primary targets with critical topological roles and profound biological relevance. These primary targets are chiefly engaged in biological processes such as the inflammatory response, apoptosis, and endocrine metabolism, which suggests that their altered function may lead to neurotoxicity. TNF appears to play a key role in BPA-induced neurotoxicity. As a cytokine that promotes inflammation, TNF is closely tied to neurotoxicity and exerts its effects through various mechanisms. These include triggering neuroinflammatory responses, inducing neuronal apoptosis, impairing the integrity of the blood-brain barrier, and disrupting the transmission of synaptic signals. 29 Prolonged TNF activation has been linked to neurodegenerative diseases such as Alzheimer’s and Parkinson’s. 30 Animal studies have shown that subchronic exposure to BPA significantly increases the expression of the pro-inflammatory cytokine genes TNF-α, IL-17, and IL-6 in male mice, suggesting that BPA may induce neuroinflammation and neurotoxicity. 31 In vitro studies indicate that BPA significantly increases the expression of TNF-α genes and the secretion of TNF-α proteins in human trophoblast cells. This suggests that exposure to BPA may exacerbate neuroinflammatory diseases, ultimately impairing cognitive and neural function. 32
As a key regulator of cell cycle and apoptosis, TP53 increases neuronal cell damage by initiating the activity of its downstream pro-apoptotic gene BAX. 33 Once the environmental substance BPA enters the organism, its action seems to disable part of the TP53 pathway, which in turn interferes with the dynamics of cell proliferation and cellular demise patterns. 34 Studies have also revealed that the protein, in response to stress or disease, enhances the expression of genes that produce oxidatively active substances and inhibits the expression of genes with antioxidant functions, ultimately exacerbating oxidative stress in the body. 35 Previous literature also suggests that exposure to BPA not only triggers increased oxidative stress and DNA damage but also triggers the TP53 signaling pathway, leading to mitochondrial dysfunction and further driving cells toward apoptosis or cycle arrest. 36
INS encodes insulin, which is not only a key hormone in the regulation of blood glucose but also crosses the blood-brain barrier and enters the central nervous system, where it is deeply involved in various neurophysiological processes. BPA significantly increased fasting glucose levels and decreased the protein levels of insulin signaling molecules, which reduced glucose uptake and oxidation, suggesting that BPA impairs glucose homeostasis. 37 The observation that BPA-treated mice show markedly reduced insulin sensitivity and significantly decreased levels of the key glucose transport proteins GLUT1 and GLUT3 in brain tissue strongly suggests that BPA exposure may constitute a potential risk factor for neurodegenerative diseases. 38
ESR1 is a component of the steroid hormone receiver superfamily and is crucial for maintaining endocrine homeostasis. ESR1 mediates gene expression and influences cell multiplication and differentiation in target tissues. 39 It is widely accepted that neurotoxicity stems mainly from its demonstrated estrogenic activity, which disrupts the normal pathway of endogenous estrogen signaling and that disruption of this signaling pathway may impair neural development and its plasticity. 40 BPA is chemically similar to natural estrogenic compounds and has estrogen-like activity. It binds to ESR1. BPA interferes with its genomic transcriptional regulation and non-genomic signaling by competitively binding to the ESR1 binding site. 41 Studies suggest that this interference may affect the neural development of zebrafish embryos and cause neurological damage to neurons, partly by interfering with the expression of estrogen receptors. 42 ESR1 abnormalities have been strongly implicated in neurotoxic processes in neurodegenerative diseases. 43
PTGS2 is an enzyme that plays a key role in the conversion of arachidonic acid to prostaglandins. Its encoded product is COX-2. PTGS2 is actively linked to inflammatory responses and is associated with the pathological mechanisms of aging and neurodegeneration. 44 Under normal circumstances, it is lowly expressed in most tissues. BPA can induce its expression, promoting neuroinflammation cascades and neuronal apoptosis. 45
GO and KEGG enrichment analyses of BPA-associated neurotoxicity targets identified not only pathways directly related to neurotoxic mechanisms but also novel associations that warrant in-depth exploration. The neurodegenerative disease pathway and the serotonergic synapse have been identified as core plausible pathways that are highly consistent with the generally acknowledged mechanisms of BPA neurotoxicity. Specifically, Yin et al. demonstrated that BPA induces hyperphosphorylation of tau protein in Neuro-2a cells, 46 while Li et al. found that BPA exposure increases amyloid-β plaques and induces the overexpression of amyloid precursor protein (APP) in mice. 47 The overexpression of APP, hyperphosphorylation of tau, and accumulation of amyloid-β plaques are core pathological hallmarks of AD. 48 For the PD-related pathway, BPA impairs dopaminergic neurotransmission in mouse midbrain cultures 49 and reduces tyrosine hydroxylase activity in rats’ brain, 50 findings closely associated with the core pathological features of PD. Additionally, the neurotoxicity of BPA can be demonstrated by its concentration-dependent inhibitory effects on serotonin. 51 Regarding the MAPK signaling pathway, we focused on the JNK subpathway, which is closely tied to BPA-induced neurotoxicity: studies show that BPA exposure can increase the inflammatory response of BV2 cells and simultaneously upregulate the expression of TNF-α and IL-6, and this effect can be partially reversed by JNK inhibitors. 52 Additionally, our study identified 2 unexpected pathways with potential research value: the linoleic acid metabolism pathway and the arachidonic acid metabolism pathway. Though their link to known BPA mechanisms is unclear, recent studies suggest clues: linoleic acid derivative accumulation induces neuroinflammation, 53 and arachidonic acid oxidation products cause neuronal toxicity. 54 These provide new perspectives for exploring non-classical BPA neurotoxicity mechanisms. Regarding cancer-related pathways, although studies have suggested a potential association between BPA exposure and brain gliomas, among other tumors, 55 with some mechanisms overlapping with the pathways enriched in this study, they were not emphasized in the present research due to their weak direct relevance to neurotoxicity. These specific associations between our in-silico pathway enrichment results and published experimental data enhance the translational relevance of our findings, indicating that the neurotoxic effects of BPA may involve these disease-specific molecular cascades.
To further verify these findings, comprehensive research incorporating molecular docking and MD simulations was performed. Molecular docking analysis revealed that BPA displayed significant binding affinity with the top 5 hub targets, with binding energies ranging from −8.3 to −5.3 kcal/mol. The docking results suggested BPA’s effective binding to the active sites of these hub targets, indicating a potential interaction mechanism that provides in silico evidence for a potential ability to induce neurotoxicity through these molecular interactions. In line with general molecular docking principles, lower scores indicate a more favorable predicted interaction between a ligand and its target. Critically, however, this trend only holds robust biological significance when docking score differences between distinct targets exceed 1 kcal/mol. Given that the score discrepancies among ESR1 (−8.3 kcal/mol), TNF (−8.0 kcal/mol), and TP53 (−7.6 kcal/mol) fall below this threshold, their numerical variations should not be overinterpreted as meaningful differences in binding affinity, but rather as minor fluctuations in energy simulation. To examine the BPA-ESR1, BPA-TNF, and BPA-TP53 interactions in greater detail, we conducted molecular dynamics simulations. Analysis of RMSD and Rg values confirmed system stability and convergence throughout the 100 ns simulation period, validating the structural stability of the BPA-ESR1, BPA-TNF, and BPA-TP53 complexes identified through docking.
As a new method of toxicological prediction, network toxicology emerges as a cutting-edge interdisciplinary field integrating computational approaches, bioinformatics, and systems biology. By leveraging advanced tools such as artificial intelligence, machine learning, and big data analytics, it aims to identify chemical hazards and evaluate associated risks with reduced reliance on traditional animal experiments. 56 Its strengths lie in efficient processing of massive datasets, high-throughput chemical screening, modeling complex biological interactions, 57 and identifying key toxicity nodes to deepen mechanistic understanding, 58 thereby enhancing the accuracy and biological relevance of predictions. 59 However, network toxicology remains inherently predictive or descriptive in nature, as its findings depend on computational models and existing datasets. Although our approach to network toxicology provides valuable insights into the neurotoxicity of BPA, we acknowledge that critical limitations persist, as predictions are constrained by model assumptions, data quality, and potential gaps in biological representation. Thus, the observed trends and mechanistic insights proposed herein require confirmation through subsequent wet-lab experiments and large-scale longitudinal epidemiological studies to validate the validity of the core targets and signaling pathways identified by network toxicology. Such experimental follow-up is essential to bridge the gap between computational predictions and biological reality, providing a more robust empirical basis for developing prevention and treatment strategies targeting BPA-induced neurotoxicity.
This study contributes to BPA-induced neurotoxicity research through integrated application of established computational tools: First, we constructed a network linking BPA exposure to neurotoxicity pathogenesis by integrating 5 multi-source databases to identify 255 potential neurotoxicity-related targets and 52 core regulatory hubs (e.g., TNF, TP53, and ESR1) identified via integrated analysis (Figures 2 and 3). Second, our functional enrichment analysis reveals that these core targets converge on key signaling cascades such as inflammatory responses, neurodegenerative pathways, serotonergic synapse pathway, and MAPK signaling, providing distinctive mechanistic insights into how BPA induces neurotoxicity through multi-pathway crosstalk (Figure 4). Third, we validate the stability of BPA-core target interactions using molecular docking (with binding energies as low as −8.3 kcal/mol for ESR1) and 100 ns molecular dynamics simulations, complemented by MM/GBSA binding free energy calculations to confirm conformational stability (Figures 5 and 6). These core computational approaches add value to the field by (1) providing a systems-level perspective on BPA’s neurotoxic mechanisms (complementing existing single-target studies), (2) identifying high-confidence core targets and pathways for experimental validation, and (3) establishing a reference computational framework for generating testable hypotheses in the neurotoxicity research of other environmental endocrine disruptors. Based on current network toxicology methods, we uniquely integrated multi-database screening, network topology analysis, and structural biology validation to provide some supplementary ideas and references for environmental neurotoxicity research.
Conclusion
In this study, we developed a combined computational strategy that integrates network toxicology, molecular docking, and molecular dynamics simulation. This approach allowed us to probe the potential targets and molecular mechanisms underlying BPA-induced neurotoxicity. We identified a series of potential neurotoxicity-associated targets and their core regulatory molecules by integrating large amounts of data from various databases, supplemented by sophisticated network topology screening analysis. Subsequently, detailed GO and KEGG pathway enrichment investigations characterized the neurotoxicity of BPA as an intricate mechanism involving numerous targets and pathways. Molecular docking and molecular dynamics simulations further provided in silico evidence for stable interactions between BPA and key target proteins, especially ESR1, TNF, and TP53. These in silico findings generate testable hypotheses regarding the neurotoxic mechanisms of BPA and offer a computational reference for guiding further experimental validation of core targets and pathways. Going forward, it is suggested that subsequent research should concentrate on validating key target molecules and signaling pathways, exploring protective strategies rooted in network regulation, and establishing scientific proof of the toxic mechanisms of BPA and the development of protective measures.
Supplemental Material
Supplemental Material - Assessing the Neurotoxicity of Bisphenol A Using Network Toxicology, Molecular Docking, and Molecular Dynamics Simulation
Supplemental Material for Assessing the Neurotoxicity of Bisphenol A Using Network Toxicology, Molecular Docking, and Molecular Dynamics Simulation by Wanyu Huang, Xuanyu Lin, Shuang Li, and Wenzong Zhu in International Journal of Toxicology
Supplemental Material
Supplemental Material - Assessing the Neurotoxicity of Bisphenol A Using Network Toxicology, Molecular Docking, and Molecular Dynamics Simulation
Supplemental Material for Assessing the Neurotoxicity of Bisphenol A Using Network Toxicology, Molecular Docking, and Molecular Dynamics Simulation by Wanyu Huang, Xuanyu Lin, Shuang Li, and Wenzong Zhu in International Journal of Toxicology
Footnotes
Author Contributions
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Zhejiang Province, China (grant number: LY18H270001).
Data Availability Statement
Data will be made available on request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
