Abstract
The nitramine explosive, hexahydro-1,3,5-trinitro-1,3,5-triazine (RDX) is associated with acute and chronic toxicity in mammals and targets both the central nervous system and liver. After a single oral dose of RDX in male rats, the systemic distribution of RDX and the toxicodynamic response was measured using clinical chemistry and Affymetrix Rat Genome® 230 2.0 gene expression arrays, respectively. Nominal doses of 0, 9 and 36 mg/kg pure RDX were administered to animals followed by liver, cerebral cortex, and hippocampus gene expression analysis at 0, 3.5, 24, and 48 hours. RDX quickly entered the liver and brain, increasing up to 24 hours. For the 36 mg/kg dose, RDX was still measurable in liver and brain at 48 hours, but was non-detectible for the 9 mg/kg dose. At 3.5 hours, the time within which most convulsions reportedly occur after RDX ingestion, the hippocampus displayed the highest response for both gene expression and pathways, while the cortex was relatively non-responsive. The top 2 impacted pathways, primarily involved in neurotransmission, were the GABAergic and glutamatergic pathways. High numbers of genes also responded to RDX in the liver with P450 metabolism pathways significantly involved. Compared to the liver, the hippocampus displayed more consistent biological effects across dose and time with neurotransmission pathways predominating. Overall, based on gene expression data, RDX responses were high in both the hippocampus and liver, but were minimal in the cerebral cortex. These results identify the hippocampus as an important target for RDX based on gene expression.
Keywords
Introduction
The nitramine explosive, hexahydro-1,3,5-trinitro-1,3,5-triazine, hexagen, also known as cyclonite (RDX) has been in use since World War II and its toxicology has been well described.1-3 At acutely high doses, electrogenic seizures leading to convulsions have been verified by EEG in animal studies.4,5 Human exposure from inadvertent or deliberate ingestion of RDX or inadequate industrial hygiene has led to transient convulsions at high doses.6-8 Furthermore, animal studies show a positive association between observed convulsion and the internal doses of RDX in the blood and brain. 5 The mechanism of action (MOA) of RDX was identified by Williams et al (2011), where apparent binding to the picrotoxin site of the ionotropic γ-aminobutyric acid receptor (GABAAR) causes temporary inhibition of the chloride channel.5,9 This inhibition in turn leads to derepression of glutamatergic neurons by inhibitory GABAA neurons, resulting in seizures and convulsions. 10 In this article we make a distinction between convulsions, which refers to observable seizures, and seizure activity which can be observed in electroencephalograms (EEGs) without manifestation as convulsions.
For chronic studies, oral RDX exposures in rats and mice have been equivocal for cancer, though low incidences of liver and lung cancers have been observed in some studies.11-13 Critical toxicological effects for risk assessment applications have therefore relied on central nervous system convulsions and liver cancer from animal studies. 13 However, due to its infrequent occurrence in groundwater at concentrations that might raise public concern, the U.S. Environmental Protection Agency (USEPA) decided not to regulate RDX in drinking water. Instead, the USEPA developed a 2 µg/L lifetime health advisory, whereas state regulations range from .3 to 9.7 ug/L in drinking water. 14 These values are 1000-fold lower than those that could cause convulsions.
Two key determinants of RDX toxicology in mammals are target organ concentration levels in the liver and brain. 15 RDX is rapidly absorbed from the gastrointestinal tract which is likely followed by liver metabolism via P450 enzymes. 16 Mammalian study results however, indicate low excretion levels at low doses of the parent compound so observed systemic levels and toxicological effects are presumably attenuated through liver metabolism, which subsequently lowers the internal dose delivered to the brain. 17 While the intermediate products are still in question, it can be assumed that metabolism and/or excretion attenuates the neurotoxic effects (convulsions) after RDX ingestion at high doses. Depending on aerobic status, metabolic products of RDX may differ.16,18 However, it is generally agreed upon that 3 nitrate reductants 19 and 2 open ring products 20 occur. While these 5 products have been found in the environment, 21 not all have been identified in mammals. 17
We have previously shown that RDX dramatically increases gene expression once maximal internal dose in tissues was reached. 15 Our current work retained the same time points as prior studies, 15 but separated the rat cortex and hippocampus, showing that male Sprague-Dawley rats have a higher responsiveness to oral RDX in the hippocampus compared to the cerebral cortex. These results imply there are important neurotransmission pathways that are significantly impacted by RDX exposure in specific brain areas.
Materials and Methods
Chemicals
Hexahydro-1,3,5-trinitro-1,3,5-triazine, hexagen, cyclonite (RDX) (98% pure) was obtained from the Department of the Navy, Indian Head MD. RDX was characterized using gas chromatography with electron capture detector (GC-ECD, see Analytical Methods) with no chemical impurities detected. RDX was handled according to in-house protocols for hazardous materials/explosives. Dosing capsules were filled by the Army Public Health Center Directorate of Laboratory Sciences by weighing pure RDX into size 9 gel capsules (Torpac®, Fairfield, NJ). Capsules were then individually stored in dark glass sample bottles for up to 2 days in a refrigerator until dosing.
Animals
Male Sprague-Dawley rats (Charles River Laboratories, Wilmington, MA) were pair-housed and had a 1 week acclimatization period. A 12/12 light/dark cycle with ad libitum, food and water was also provided. Animals weighing 300-400 g were administered a one-time oral dose of either 9 or 36 mg/kg RDX in a gel capsule (Torpac®) via a capsule dispenser per the manufacturer’s instructions. After dosing, animals were then returned to their home cages. Doses used in this study were calculated based on previous work, 15 so as not to induce convulsions. Animals were observed during the first hour to confirm no convulsions had occurred. At 0, 3.5, 24, and 48 hours after dosing, animals (n = 3 for each dose and time) were euthanized using decapitation and the cerebral cortex as well as the hippocampus were removed from the brain and flash frozen in liquid nitrogen for RNA extraction and microarray analysis. The remainder of the brain was used for RDX analysis. An additional 5 minute time point, collected at necropsy, was also included in the analysis. Blood samples for RDX analysis were placed into tubes with EDTA, which served as an anticoagulant. A segment of the median liver lobe was immediately flash frozen in liquid nitrogen for gene expression analysis. All liver samples were stored at −80°C until RNA extraction, while the remainder of the brain and liver samples were stored at −40°C for RDX analysis.
The animal protocol for this study was reviewed and approved by the Institutional Animal Care and Use Committee and the work was carried out in an AAALAC accredited facility. Guidelines for the Guide for the Care and Use of Laboratory Animals were followed (http://www.nap.edu/catalog.php?record_id=12910#toc). The study was carried out in accordance with the standards found in Title 40 Code of Federal Regulations, Part 792, Good Laboratory Practices (https://www.ecfr.gov/current/title-40/chapter-I/subchapter-R/part-792).
Analytical Methods
Samples were analyzed using Gas Chromatography - Electron Capture Detection (GC-ECD) based on a previously published method. 15 For whole blood samples, erythrocytes were lysed by adding 100 μL of mixed whole blood into 100 mL of deionized (DI) water in a volumetric flask. To extract RDX, 5 mL of isoamyl acetate (Pesticide Residue Grade, anhydrous, Sigma-Aldrich, St Louis, MO, USA) was added to the 100 mL flask, followed by liquid/liquid extraction at room temperature for 1 hour. Standards were made from pure RDX dissolved in acetonitrile (ACN) that were then spiked into DI water at 0, .01, .02, .1, .2, .5, 1.0 and 2.0 μg RDX/mL. These standards were extracted using the same procedure as the blood and tissue samples. An aliquot of the isoamyl acetate extract was then used for analysis. The above method extracts both RDX and other organic molecules into the organic phase, with water soluble molecules separating into the aqueous phase. This extraction method also decreases gas chromatogram signal interference seen in other methods used for explosives analysis.
Tissue samples were treated differently than blood samples, with flash frozen tissue being partially defrosted and then blended to an even consistency in a tissue macerator with 20 mL DI water. Five milliliters of isoamyl acetate was added to this mixture, where sample extraction occurred on a flat-bed shaker for up to 3 hours; time duration depended on the tissue mass measured. The mixture was then centrifuged for 20 minutes to separate the isoamyl acetate extract from the tissue and water. An aliquot of the supernatant was then used for analysis.
Prior to extraction, a surrogate chemical, 3,4 dinitrotoluene (3,4-DNT) was added as an external running standard; this surrogate served as an additional quality control measurement added to each sample prior to extraction. Since DNT elutes before RDX, this surrogate compound was used to monitor extraction efficiency, accuracy, and reproducibility for each sample. Blind controls were also used for each sample extraction batch. Spiked blood, brain, and liver samples had the following reporting limits respectively: .10 μg/mL and .12, .25 μg/g. Tissue recovery of spiked 3,4-DNT was between 80 and 120%.
Microarray Analysis
All experiments were performed using the Affymetrix Rat Genome® 230 2.0 whole genome oligonucleotide arrays (Santa Clara, CA). Frozen tissues were homogenized in Tri Reagent (Sigma Chemical Company, St Louis, MO) and RNA was extracted according to the manufacturer’s protocol. RNA purification, microarray probe synthesis, hybridization, and microarray processing were previously described. 15
Data Analysis
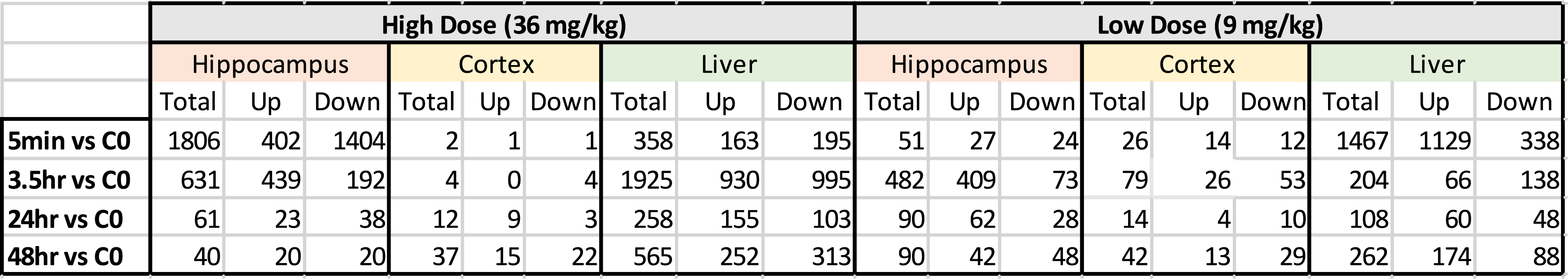
Summary of Total Significant Genes.

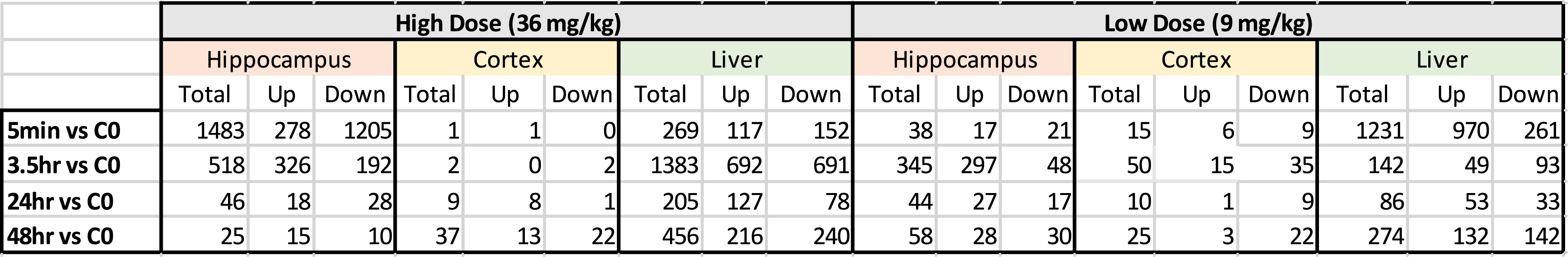
Number of Genes for Pathway Analysis.

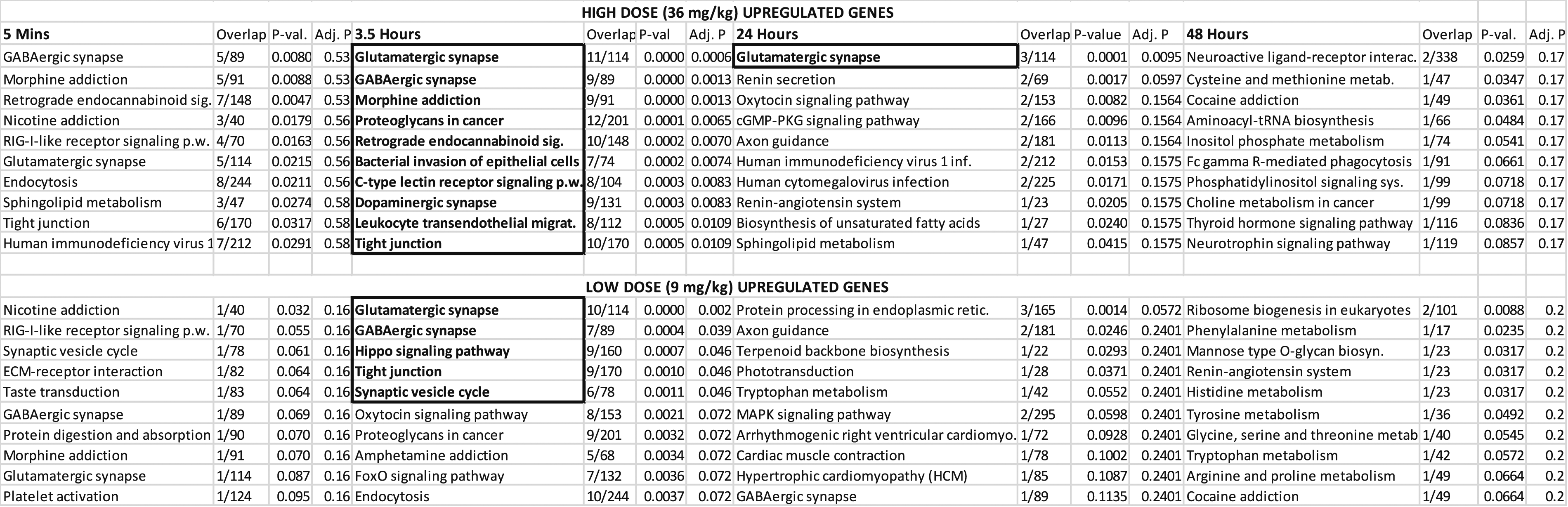
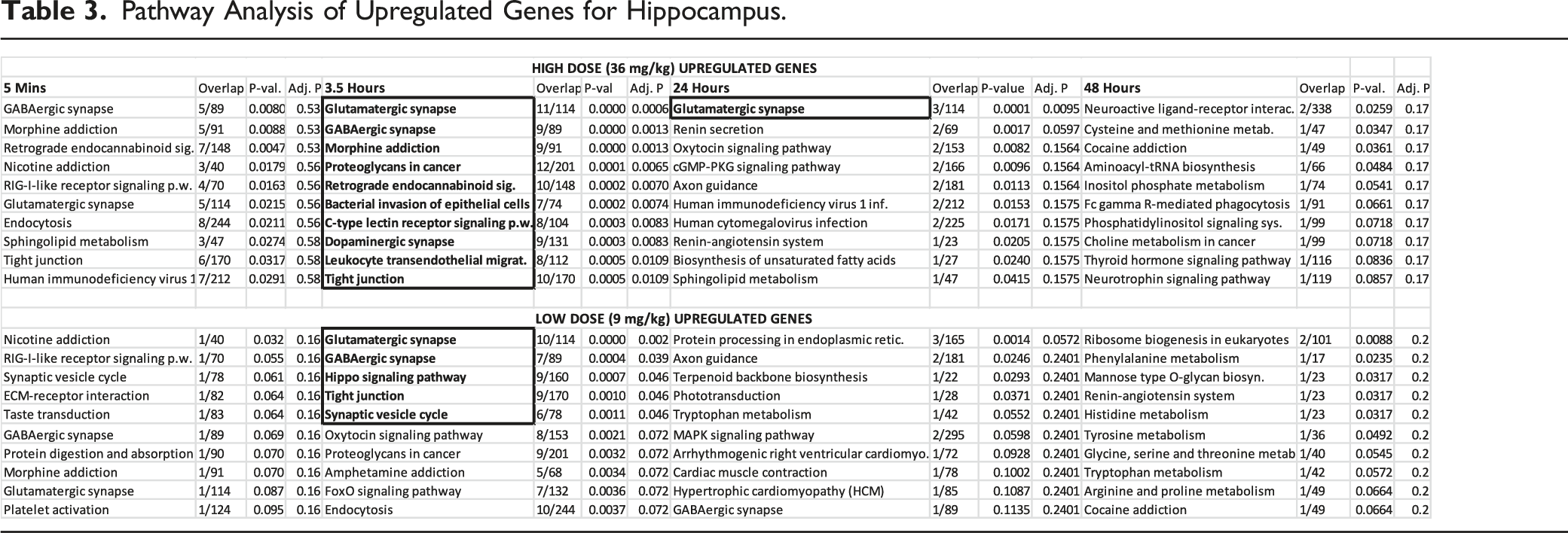
Pathway Analysis of Upregulated Genes for Hippocampus.
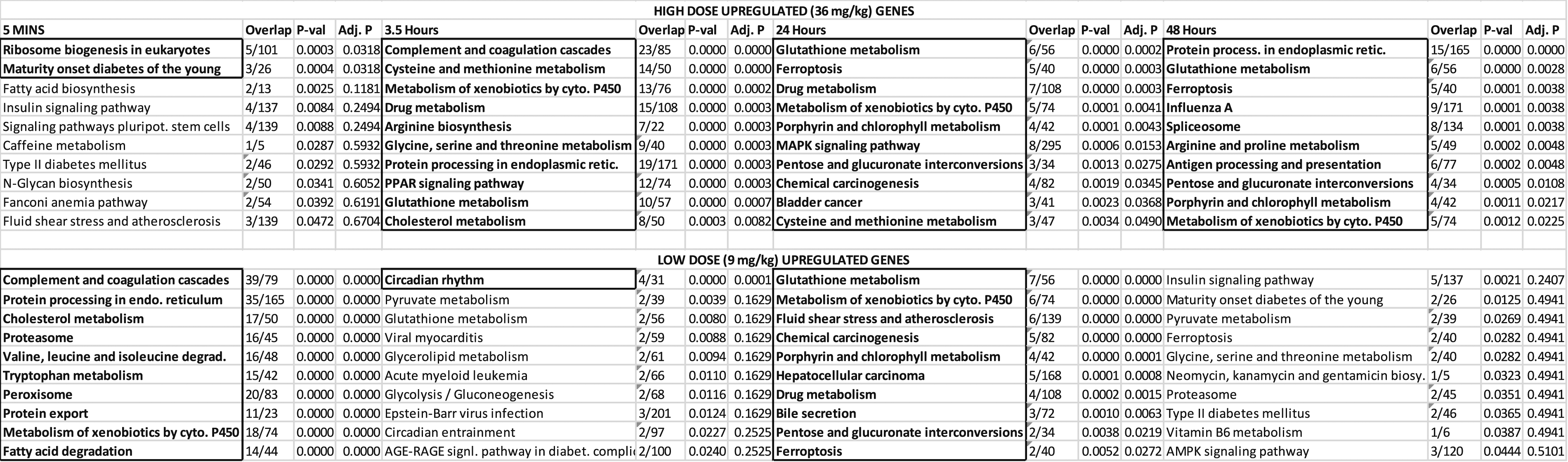
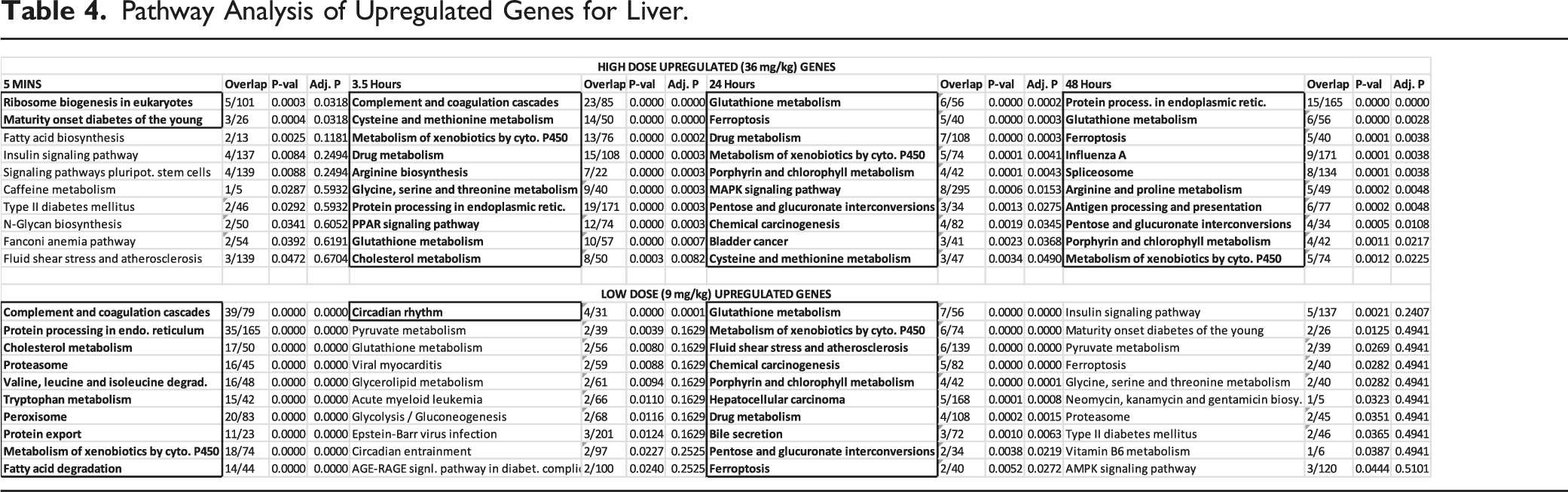
Pathway Analysis of Upregulated Genes for Liver.
Results
Blood samples from rats were analyzed using Gas Chromatography - Electron Capture Detection (GC-ECD) as described in the methods section. Scatterplots of RDX concentrations by dose and tissue are shown in Supplementary Figure 1 where the highest correlations were between blood and brain as well as the blood and liver values. A regression analysis shows RDX Liver = −.05 + 1.95 x Blood RDX (r2 = .915) and RDX Brain = .20 + 2.45 x Blood RDX (r2 = .979). Figure 1 shows the same data as a time course of RDX concentrations in blood and tissues for up to 48 hours. While blood samples contained relatively lower RDX concentrations than brain and liver, the temporal profiles for all 3 were similar. Plots of RDX levels. Concentrations in blood (1A), liver (1B), and brain (1C) showing measurements in individual animal tissues after a single oral dose of RDX. Controls are green boxes, 9 mg/kg doses is blue, and 36 mg/kg doses is red. RDX was measured using gas chromatography – electron capture detection as described in Analytical Methods. All graphs on the same scale. Results depicted as mean and s.d. (n = 3) of RDX measurements.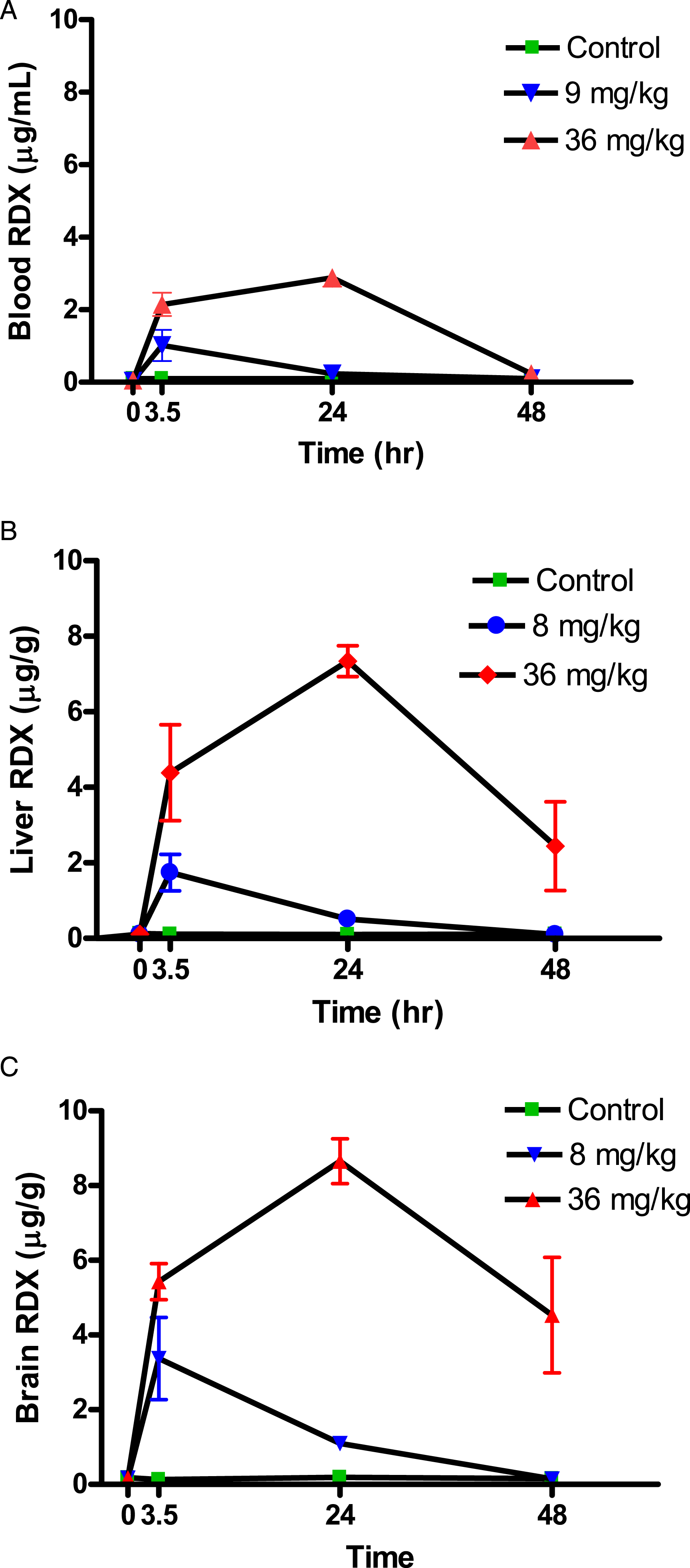
Principal component analysis (PCA) of the expression data (Figure 2) shows distinct separation by tissue type for the liver, cortex, and hippocampus. Note also that the liver was spatially separated from both the hippocampal and cerebral cortex tissues. Outlier samples removed prior to ANOVA analysis are shown on the PCA (Figure 2); they comprise of 2 liver samples (LV 621, LV 601), 1 cortex sample (CT 621), and 1 hippocampus sample (HP 601). Table 1 shows the total number of statistically significant transcripts (putative expressed genes) from the ANOVA for the liver, cortex, and hippocampus. In contrast, Table 2 shows the actual number of genes that were used for pathway analysis. The lower numbers in Table 2 were due to numerous expressed sequence tags (ESTs) being removed from the dataset as well as removal of genes that may not yet have any functional significance in pathway analysis. Principal component analysis. PCA analysis of liver, cortex, and hippocampus for all probe sets in the study. Outliers indicated were removed before analysis, including Liver 621 and 601(control and 3.5 hours) and Hippocampus 601 and Cortex 621 (control and 3.5 hours). LV = Liver, HP = Hippocampus, CT = Cerebral Cortex: 3.5 = 3.5 hour timepoint: cont = control.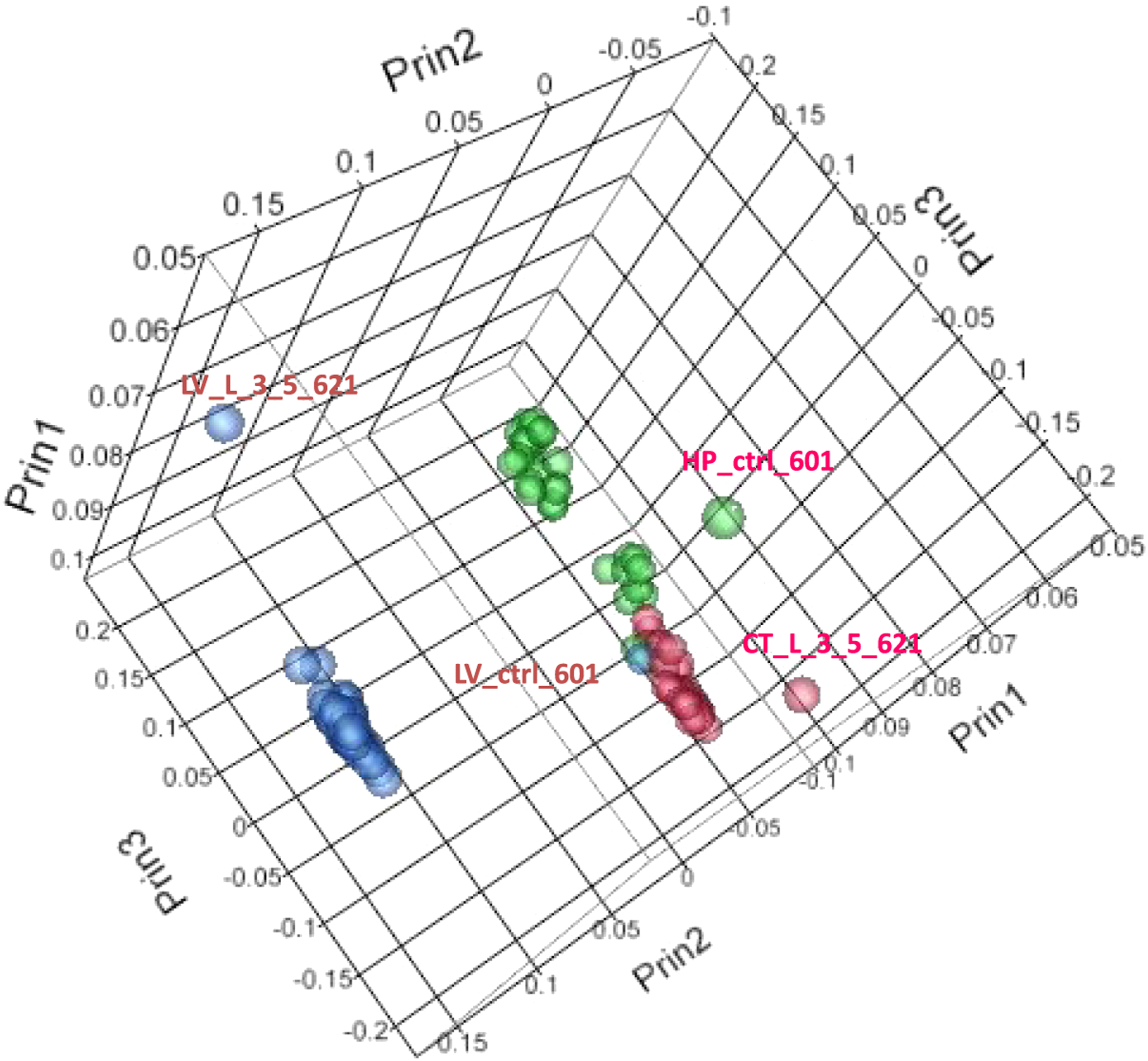
Brain cortex gene expression counts were lower at nearly all time-points or doses, which implies the cortex was insufficient for pathway analysis (Table 2). In contrast, the hippocampal and liver gene expression levels were high (Table 2) and pathway analysis was viable at both the high and low doses across all time-points (Supplementary Hippocampus Tables and Supplementary Liver Tables). Summary results are shown in Table 3 (hippocampus) and 2b (liver) for upregulated genes. This group of genes are more informative than the results for downregulated genes (see Supplementary Hippocampus and Supplementary Liver Tables for downregulated results). Overall, results show that the highest proportion of significant pathways (after FDR correction) appeared at 3.5 hours for hippocampus and at 24 hours for liver, while generally there was a higher number of significant pathways for the high vs the low doses. Below is a separate analysis for liver and hippocampus.
Liver: After FDR correction, a wide variety of pathways are impacted in liver compared to hippocampus. The peak number for both the high and low doses appear at the 24-hour time point. These 2 doses show a high consistency in their pathway terms. Metabolic pathways (Table 4) predominately affected by RDX exposure include Cytochrome P450 Metabolism of Xenobiotics and Glutathione Metabolism. Both of these pathways share the Glutathione S-Transferase M4 (GSTM4) gene (Supplementary Liver Table). Other pathways that are of significance include the chemical carcinogenesis and bladder cancer pathways; both of these pathways implicate the role of the proto-oncogene MYC. Across time, the Protein Processing in Endoplasmic Reticulum pathway and Ferroptosis are considered significant.
Hippocampus: After FDR correction, the GABAergic Synapse and Glutamatergic Synapse were the 2 main significant neurotransmission pathways across doses. Other neurotransmission pathways with related genes content include the Morphine Addiction and Retrograde Endocannabinoid Signaling (Table 3 and Supplementary Hippocampus Tables). Since genes can be used in several pathways, common genes for all these pathways include several GABA genes (GABRB3; GABRA2) as well as guanine nucleotide-binding protein (GNAI1), a second messenger protein (Supplementary Hippocampus Tables). When time was considered, the 5 minute and 3.5 hour time points had nearly identical pathway analyses; however, the pathways at the 5 minute time point were non-significant after post-hoc FDR correction (Table 4)
Given RDX’s known neurological effects, the relative importance of the hippocampal pathways were further visualized. Figure 3 shows a volcano plot of the odds ratio vs pathway P-values. This figure illustrates the GABAergic and glutamatergic pathways, both of which are involved in neurotransmission, are the highest ranking pathways, in addition to the morphine addiction pathway. As with other neurotransmission pathways, the most significant pathways shared similar groups of genes (Supplementary Hippocampus Tables). For example, the Hippocampus Supplementary Data sheet shows 7 out of a total of 9 genes significant in each pathway were common to both GABAergic and Morphine Addiction pathways (3.5 hour time point). Relative pathway importance. Plot of fold-change vs P-value for pathways. Graph was generated in EnrichR. Results show the relative significance of pathways, with Glutamatergic Synapse and GABAergic Synapses being the top pathways. Morphine addiction was driven by the same set of genes as GABAergic Synapse (see Supplementary Hippocampus Table at 3.5 hour). While D-Glutamate and D-glutamine had a high effect size, they did not reach significance (-log (.05) < 1.3).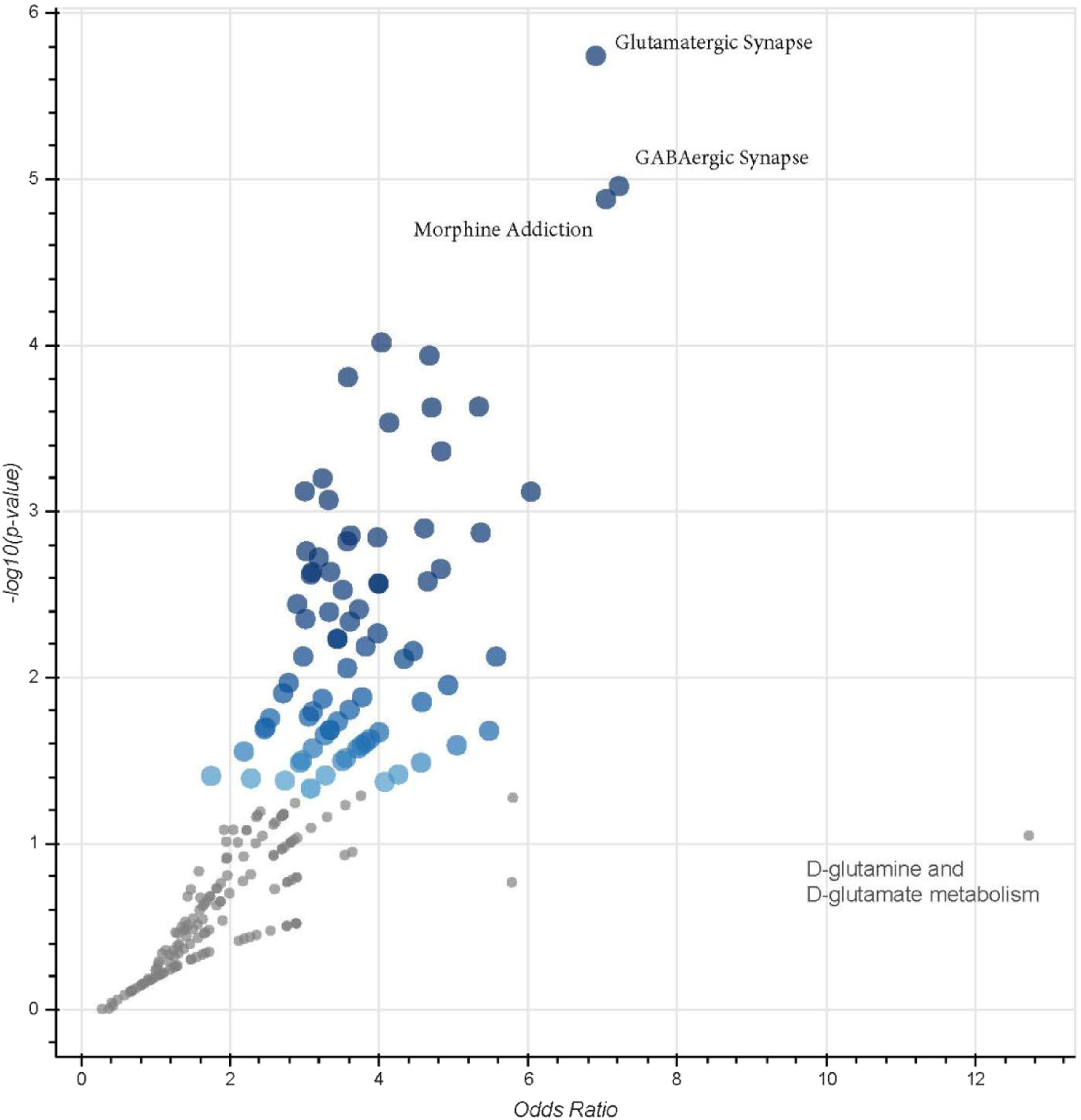
Discussion
A single sub-convulsive oral dose of RDX was rapidly absorbed in rats, showing high systemic correlations between blood, brain, and liver. Peaks occurred in brain, liver, and blood at the 3.5 hour for a 9 mg/kg dose and 24 hour time points for a 36 mg/kg dose (Figure 1). The total brain concentration at the high dose was close to a previously identified threshold (8 µg/g) for convulsions, 5 though this threshold was derived from a single 75 mg/kg RDX dose, which was 2-fold greater than our highest dose. Moreover, the single 75 mg/kg RDX dose was dissolved in a mixture of methylcellulose and tween 5 which could also influence absorption, whereas our pure RDX powder is likely similar to occupational exposures. Past work has also shown that convulsions predictably occur within the first 4 hours after RDX oral doses in animals; although a few convulsions can reoccur in some animals at later time points.24,25 Our concentration profiles show that the brain and liver RDX levels were still high even where blood values fell below the limit of detection, indicating there is some residual RDX accumulation in cells, which would be expected to disappear as RDX is metabolized and/or excreted. However, these levels were at or below the threshold for seizures (Figure 1C, Brain RDX)
Pathway analysis showed that RDX preferentially targeted the hippocampus, while the cerebral cortex appeared to be minimally disrupted. It has previously been shown that regional variations in the brain exist for GABA metabolism during chemically induced status epilepticus, with cortex being the lowest, 26 though brain region gene expression after RDX exposure has not been explored. For hippocampus, the maximum biological response occurred at 3.5 hours - where there was the greatest number of significant pathways (Table 3). However, even at the highest RDX concentration in the brain, no frank convulsions were observed at the 36 mg/kg dose. Johnson (2015) found that convulsions generally appear in rodents within minutes to hours after RDX administration and were dependent on the dose’s magnitude. 25 Once a threshold of approximately 8 µg/g in brain is reached, 5 the level of RDX presumably exceeds the effective dose at the molecular targets (GABAA receptors), which then triggers seizure activity in the brain. Therefore, the initial short-term cumulative dose and extent of occupancy of the GABAA receptor’s picrotoxin convulsant sites may be more important than later peak brain concentrations. 27
The GABAergic and glutamatergic pathways in the hippocampus indicate that neurotransmission is the dominant MOA in all susceptible species. 28 This is consistent with the GABA disinhibition hypothesis, where fast acting GABAergic and glutamatergic processes work in a balanced interplay. Then, once the GABAA receptor is blocked, depression of inhibition occurs and can result in seizures and convulsions due to fewer inhibited post-synaptic glutamatergic neurons. 29 Prior work from this laboratory using binding assays to screen a battery of 25 neurotransmitters demonstrated that 33 µM RDX (∼7.3 ug/mL) inhibited >75% of the GABAA chloride channel’s picrotoxin convulsant site compared to the control; this dose is close to the threshold for seizures (Williams 2011b). Alternatively, the binding screen showed the glutamate AMPA or NMDA binding sites remained unaffected 5 by RDX. More recent work has shown that RDX preferentially inhibits the α1β2γ2 GABAAR subtype configurations of the pentameric GABAAR channel 27 though not all subtypes were tested in that work. The current work shows that α2, α4, and γ4 subunits were significantly upregulated in response to RDX (Hippocampus H3.5 vs C0 Up) so it’s possible that RDX is targeting a specific subtype of GABAAR. Kim showed that changes in GABAA receptor channel subunit mRNA expression can occur with GABA treatment alone 30 and our also results indicate a differential GABAAR subunit expression response (compared to control) in the hippocampus.
In contrast to the brain tissues, the liver responded to the high dose beyond the 3.5 hour time point and there were significant pathways at the 24 and 48 hour time periods. At the 24 hour time point, both the high and low doses had similar pathways highlighted (Table 4). The glutathione metabolism, P450 metabolism of xenobiotics, and drug metabolism pathways in particular were driven by several common genes that involve Phase I metabolism of xenobiotics; these include CYP1A2 and CYP2E1 (Supplementary Liver; 3.5 H3.5 vs C0 UP). Phase II enzymes, such as GSTM4, GSTA5, and UGT1A1, were also implicated; the latter enzyme being 31 involved in glucoronidation of small molecules. 32 Bhushan 2003 suggested that RDX is metabolized in the liver by P450 reduction and end products can depend on low or normal oxygen conditions 19 and several potential RDX metabolites were also previously identified in swine. 17 While this work further suggests that CYP2E1, CYP1A2, and GSTM4 may be involved in human metabolism and detoxification, evidence using radiolabeled RDX in mice has shown that up to 40% of administered RDX is exhaled as CO2 (Schneider 1978). These results indicate the complete mineralization of a large fraction of oral RDX doses 33 with our work implicating Phase I metabolism in particular.
While evidence from cancer studies using oral RDX in animals is equivocal, 11 2 cancer pathways, Chemical Carcinogenesis and Bladder Cancer, were identified as significant in this study. In particular, the proto-oncogene MYC, which drives the cell cycle, underpins both of these cancer pathways (Supplementary Table Liver) as well as oncogenic c-MYC, is associated with hepatic tumors. 34 However, since MYC is a part of the normal cell cycle, moving beyond association to causation is not possible with the current work. Other genes (such as VEGFA) shown in these cancer pathways are likely associated with tumor development rather than serving as initial drivers of carcinogenicity. Overall, while the MYC pathway is significant, it cannot be determined from this work if oncogenic c-MYC is activated. The cancer pathways therefore provide weak evidence.
In summary, this study examined the relationship between the toxicokinetics and toxicodynamics of a single oral dose of pure RDX. Two neurotransmission pathways, GABAergic and glutamatergic, predominated in the hippocampus. By contrast, liver pathways suggested there are specific P450s involved in RDX metabolism. In addition to GABAergic mechanisms, we also identified the role of the Glutamatergic pathway, which would be downstream of the inhibitory GABA neurons. Our work further validates the prior use of convulsions as the critical effect for the development of an RDX oral reference dose 13 rather than the somewhat inconclusive evidence implicating possible carcinogenic effects of RDX on liver. The work demonstrates that with careful experimental design, expression data can be useful in the examination of neurotoxicity.
Supplemental Material
Supplemental Material - Gene Expression and Pathway Analysis in Rat Brain and Liver After Exposure to Royal Demolition Explosive (Hexahydro-1,3,5-Trinitro-1,3,5-Triazine)
Supplemental Material for Gene Expression and Pathway Analysis in Rat Brain and Liver After Exposure to Royal Demolition Explosive (Hexahydro-1,3,5-Trinitro-1,3,5-Triazine) by Desmond I. Bannon, Wenjun Bao, James F. Dillman, Russ Wolfinger, Christopher S. Phillips, and Edward J. Perkins in International Journal of Toxicology
Supplemental Material
Supplemental Material - Gene Expression and Pathway Analysis in Rat Brain and Liver After Exposure to Royal Demolition Explosive (Hexahydro-1,3,5-Trinitro-1,3,5-Triazine)
Supplemental Material for Gene Expression and Pathway Analysis in Rat Brain and Liver After Exposure to Royal Demolition Explosive (Hexahydro-1,3,5-Trinitro-1,3,5-Triazine) by Desmond I. Bannon, Wenjun Bao, James F. Dillman, Russ Wolfinger, Christopher S. Phillips, and Edward J. Perkins in International Journal of Toxicology
Supplemental Material
Supplemental Material - Gene Expression and Pathway Analysis in Rat Brain and Liver After Exposure to Royal Demolition Explosive (Hexahydro-1,3,5-Trinitro-1,3,5-Triazine)
Supplemental Material for Gene Expression and Pathway Analysis in Rat Brain and Liver After Exposure to Royal Demolition Explosive (Hexahydro-1,3,5-Trinitro-1,3,5-Triazine) by Desmond I. Bannon, Wenjun Bao, James F. Dillman, Russ Wolfinger, Christopher S. Phillips, and Edward J. Perkins in International Journal of Toxicology
Footnotes
Acknowledgments
The authors acknowledge the contribution of Krisa Camargo, Mark Johnson, and Lindsay Holden for helpful technical review of the manuscript. We acknowledge the late Mike Hable at APHC for developing methods for tissue analysis of RDX.
Author Contributions
DB designed the study, interpreted the data, and wrote the MS. JD and CP carried out RNA extraction and microarray analysis. WB and RW provided genomic data analysis. EP provided genomic analytical expertise.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was funded by the Department of Defense.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.