Abstract
Cellular signaling is in part regulated by the composition and subcellular localization of a series of protein interactions that collectively form a signaling complex. Using the α7 nicotinic acetylcholine receptor (α7nAChR) as a proof-of-concept target, we developed a platform to identify functional modulators (or auxiliary proteins) of α7nAChR signaling. The Broad cDNA library was transiently cotransfected with α7nAChR cDNA in HEK293T cells in a high-throughput fashion. Using this approach in combination with a functional assay, we identified positive modulators of α7nAChR activity. We identified known positive modulators/auxiliary proteins present in the cDNA library that regulate α7nAChR signaling, in addition to identifying novel modulators of α7nAChR signaling. These included NACHO, SPDYE11, TCF4, and ZC3H12A, all of which increased PNU-120596-mediated nicotine-dependent calcium flux. Importantly, these auxiliary proteins did not modulate GluR1(o)-mediated Ca flux. To elucidate a possible mechanism of action, we employed an α7nAChR-HA surface staining assay. NACHO enhanced α7nAChR surface expression; however, the mechanism responsible for the SPDYE11-, TCF4-, and ZC3H12A-dependent modulation of α7nAChR has yet to be defined. This report describes the development and validation of a high-throughput, genome-wide cDNA screening platform coupled to FLIPR functional assays in order to identify functional modulators of α7nAChR signaling.
Introduction
G protein–coupled receptors (GPCRs) and ion channels rarely act in isolation; instead, they function in concert with a multitude of signaling/effector proteins that constitute a signaling complex. 1 Signaling complexes may contain proteins that aid in folding to appropriate conformation, trafficking to site of action, tethering to subcellular location, and modulating pharmacology. Indeed, components of the signaling complex are likely dynamic at any given stage of a particular cascade. Understanding the critical components of the signaling complex and how they relate to disease pathology has ramifications for drug discovery and may provide novel drug targets.
Ion channels and GPCRs are two large classes of targets that the pharmaceutical industry has embarked on in drug discovery programs in order to address a plethora of diseases. However, in many instances, identifying a compound that displays receptor/ion channel selectivity and elicits the desirable response in select tissues has proved challenging. Moreover, for diseases such as psychiatric disorders, targeting specific tissue subregions (i.e., the hippocampus in brain) is a high hurdle. Indeed, targeting allosteric sites of GPCR or ion channels can help achieve subtype selectivity. 2 α-Amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptors (AMPARs) are positively and negatively modulated by a collection of auxiliary proteins that include transmembrane AMPAR proteins (TARPs), cornichons, and porcupine, respectively.3,4 In this scenario, the auxiliary proteins have been shown to modulate endoplasmic reticulum (ER) export, trafficking, pharmacology, and the kinetics of AMPARs.4,5
Exploiting the key components of the signaling complex responsible for regulating the activities of a specific GPCR or ion channel target for drug discovery is challenging, especially because not all protein components have been identified. We sought to develop and implement a screening platform that would enable the identification of functional modulators of an ion channel. Several desirable criteria were established. The platform had to be amenable to automation with medium to high throughput, the target should be expressed in a cellular environment in order to recapitulate a physiologically relevant conformation, and the cellular readout should be a direct measure of target activation in order to avoid downstream reporter systems. We chose the α7 nicotinic acetylcholine receptor (α7nAChR) as a case study. The α7nAChRs are ligand-gated, calcium-permeable homopentamers. The α7nAChR is expressed in discrete regions of the brain (i.e., hippocampus and cortex) and contribute to learning and memory, a deficit presented by Alzheimer’s disease and schizophrenic patients. 6 However, drug discovery efforts surrounding α7nAChR have proved challenging due to the poor α7nAChR functional response in many mammalian cell lines transfected with α7nAChR. 7 The poor α7nAChR activity can be partially circumvented by cotransfecting α7nAChR with the molecular chaperone, RIC3. 7 RIC3 has been described as a chaperone contributing to the proper folding/assembly of α7 receptors. 8 However, the mammalian RIC3 modestly increases α7nAChR activity and is neither necessary nor sufficient for efficient assembly of mammalian α7nAChR.7,9 In a recent publication, RIC3 did not increase α7nAChR surface expression when coexpressed in HEK293 cells. 10
Using an automated high-throughput platform and the Broad genome-wide cDNA library, we transiently cotransfected each Broad cDNA in an arrayed 384-well format with the cDNA encoding the α7nAChR and measured the propensity for each expressed cDNA to modulate α7nAChR-dependent calcium flux using FLIPRTETRA. We identified several known α7nAChR-interacting proteins validating the platform and approach. Importantly, we identified novel α7nAChR-modulating proteins, including the recently published NACHO. 10 Identifying and understanding the key auxiliary proteins that regulate α7nAChR activity will contribute to our understanding of α7nAChR function and may unveil a novel target or point of interception for the development of novel therapies. Moreover, the genome-wide cDNA screening platform described in this report can be applied to other targets, including ion channels, GPCRs, transporters, and enzymes, whose activities can be monitored in high-throughput screening.
Materials and Methods
Materials
The HEK293T cells were purchased from the American Type Culture Collection (ATCC, Manassas, VA). Dulbecco’s modified Eagle’s medium (DMEM) high glucose and PenStrep were obtained from Corning (Corning, NY). Detachin was purchased from Genlantis (San Diego, CA). Defined fetal bovine serum (FBS) was purchased from PAA (Pittsburgh, PA). OptiPRO serum-free media was purchased from Thermo Fisher Scientific (Waltham, MA). Ca5 calcium dye and probenecid were purchased from Molecular Devices (Sunnyvale, CA) and Biotium (Haywad, CA), respectively. FuGENEHD was purchased from Promega (Madison, WI). The Broad cDNA library was purchased from the Broad Institute at the Massachusetts Institute of Technology (MIT). Hank’s Balanced Salt Solution (HBSS) was custom-made by HyClone (Logan, UT) and contains the following: 137 mM NaCl, 2 mM KCl, 0.5 mM MgCl2, 10 mM HEPES, 5 mM D-glucose, pH 7.4, osmolarity 300 mOsm/L. (–) Nicotine ditartrate, AR-R17779, and 1-(5-chloro-2,4-dimethoxy-phenyl)-3-(5-methyl-isoxazol-3-yl)urea (PNU-120596) were purchased from Tocris (Pittsburgh, PA). Black, clear-bottom, 384-well, poly-D-lysine-coated plates were purchased from Corning.
Broad cDNA Library and Control Constructs
The Broad cDNA library contains 17,255 cDNA clones. Each cDNA was cloned into the pLX-317 vector and transcription driven by the EF1a promoter. Each cDNA was engineered to express a V5 tag at the C-terminus of the translated protein. The library was arrayed into a 384-well format. For screening experiments, the following constructs were used: pcDNA3.1-α7nAChR and pCMV6-XL5-RIC3. For immunofluorescence studies, the following constructs were used: pcDNA3.1-α7nAChR-HA, pCMV6-XL5-RIC3, pCMV6-XL5-NACHO, pLX317-TCF4, pLX317-SPDYE11, and pLX317-ZC3H12A.
Cell Culture and Cotransfections
HEK293T cells were maintained in DMEM high glucose supplemented with 10% defined FBS and 1× PenStrep in a humidified incubator at 37 °C/5% CO2. Cells were maintained at ~80% confluency. Cells were detached using detachin, centrifuged at 1000 rpm/5 min, and resuspended in DMEM supplemented with 10% FBS. Cells were dispensed into 384-well poly-D-lysine-coated plates at 10,000 cells/well. Each well of 384-well plates was cotransfected with pcDNA3.1-α7nAChR cDNA and a single Broad library cDNA at a ratio of 4:1. Cotransfections were performed using FuGENEHD and diluted in OptiPRO serum-free media. Wells assigned to high or low controls were cotransfected with α7nAChR+RIC3 or α7nAChR+vector cDNAs, respectively. To ascertain nonspecific calcium flux, the α7nAChR cDNA was replaced with the pcDNA3.1 vector. Cells were incubated for 24 h at 37 °C/5%CO2/95% humidity, followed by 24 h at 30 °C/5%CO2/95% humidity. The flow scheme for automated transient transfection is illustrated in
Calcium Flux Assay
Cells were washed four times with assay buffer (custom HBSS buffer supplemented with 2 mM CaCl2 and 0.5 mM MgCl2), and an equal volume of the Ca5 dye was added. Ca5 dye was prepared according to the manufacturer’s instructions using assay buffer supplemented with probenecid (final concentration in well 1.25 mM). Cells were incubated at room temperature for 1 h, followed by four series of washes with assay buffer. Plates were immediately loaded into the FLIPRTETRA, with the first addition of (–) nicotine ditartrate followed by a second addition of PNU-120596. FLIPR traces were analyzed using ScreenWorks 3.2 (Molecular Devices). The output statistic was defined as the maximum relative light units (RLU) during the kinetic read. Assay development data were exported to GraphPad Prism 5 software (GraphPad, San Diego, CA). Screening data were exported and analyzed using in-house software. The fluorescence change from baseline for each well (F) was used to calculate the response (R) relative to controls (H = high control; L = low control) as follows: R = 100((F – L)/(H – L)).
HA Epitope Immunofluorescent Staining
HEK293T cells were cotransfected using an α7nAChR C-terminal HA-tagged construct (α7nAChR-HA) as described previously. 10 Briefly, HEK293T cells were cotransfected as follows: α7nAChR-HA+vector, α7nAChR-HA+RIC3, α7nAChR-HA+SPDYE11, α7nAChR-HA+NACHO, α7nAChR-HA+TCF4, or α7nAChR-HA+ZC3H12A. The ratio of each construct for the cotransfections was 1:1 (α7nAChR-HA: library cDNA/vector). Cotransfections were performed using FuGENEHD and diluted in OptiPRO serum-free media as described above. For surface staining, cells were assayed 48 h posttransfection and incubated with AF 555 florescence-conjugated HA tag antibodies (Thermo Scientific) in culture medium for 30 min at 37 °C. Cells were washed three times with ice-cold PBS and fixed with 4% paraformaldehyde for 15 min. Cells were permeabilized using 0.3% Triton X-100 and blocked by 1% BSA. Cells were incubated with AF 488 florescence-conjugated HA tag antibodies for 1 h at RT. After three washes, cells were mounted with DAPI-containing media and images captured using a Zeiss M2 Imager. DAPI was present in the fixative to ensure viability (data not shown). Images were processed using Zeiss Zen 2011 software.
Subcellular and Expression Profiles of α7nAChR Interacting Proteins
Subcellular and expression profiles were obtained from GeneCards Human Gene Database (www.genecards.com).
Results
α7nAChR-Dependent Calcium Flux in HEK293T Cells
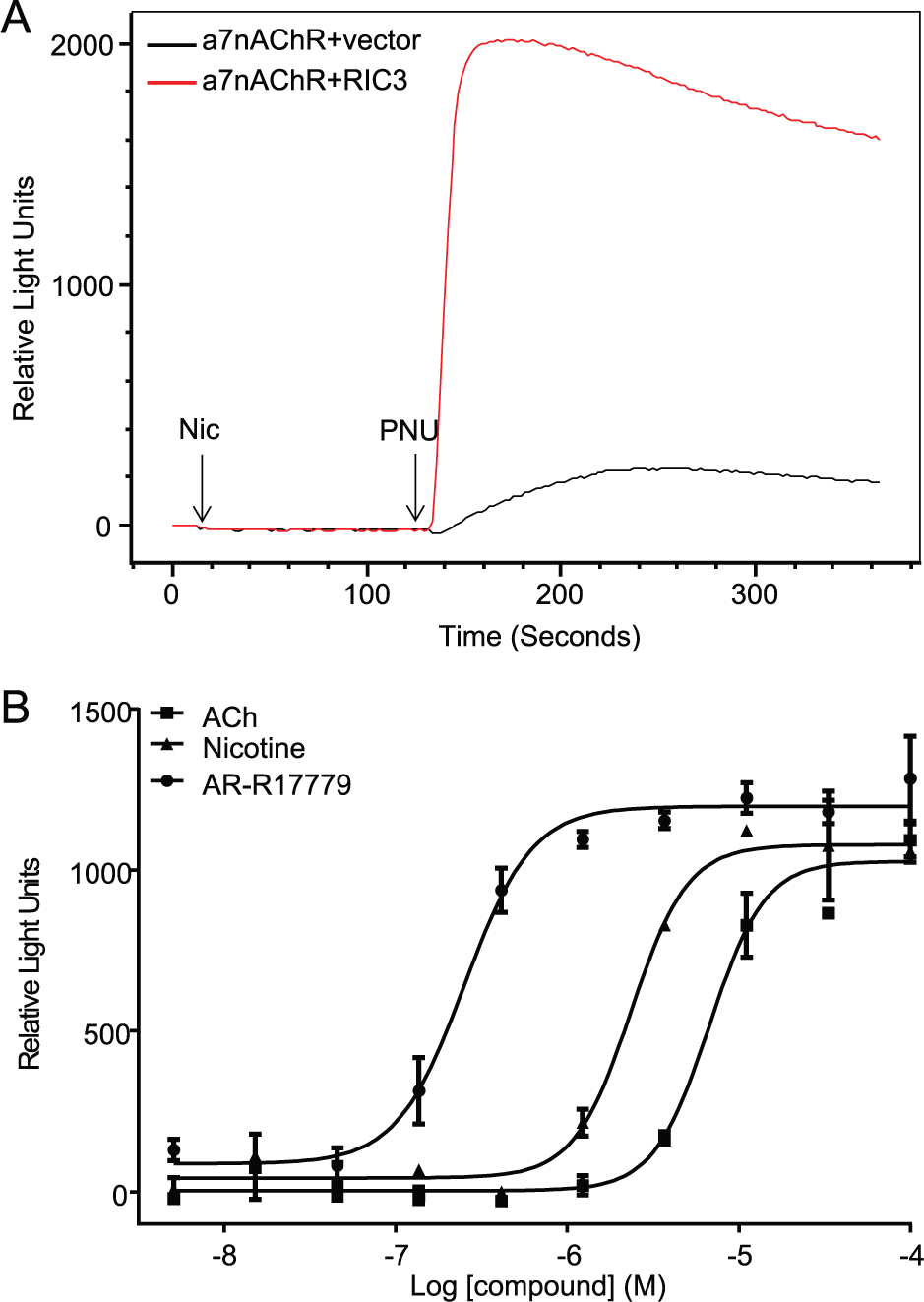
In HEK293T cells transfected with α7nAChR alone, nicotine did not elicit a calcium response in the FLIPR (
Fig. 1A
). The kinetics of the α7nAChR is rapid, and the channel desensitizes faster than the FLIPR can record changes in calcium flux.
11
Type 2 positive allosteric modulators (PAMs), such as PNU-120593s, are reported to affect the peak current, evoke a secondary weak decaying current, and reactivate desensitized currents.12,13 Upon addition of PNU-120596 to HEK293T cells transfected with α7nAChR, a nicotine-dependent calcium response, albeit small, was measured in the FLIPR (
Fig. 1A
). However, robust nicotine-dependent calcium flux was measured in the presence of PNU-120596 using HEK293T cells cotransfected with α7nAChR and the molecular chaperone RIC3 (
Fig. 1A
). To develop and establish an α7nAChR screening platform to identify auxiliary proteins that may regulate/enhance α7nAChR function, we used RIC3 cDNA as the control α7nAChR modulator protein in combination with PNU-120596 treatment to optimize the assay conditions. We titrated the ratio of α7nAChR/RIC3 cDNA transiently transfected into HEK293T cells. To conserve the Broad cDNA library, we sought a robust calcium flux with a minimal ratio of α7nAChR/RIC3. A series of α7nAChR/RIC3 titrations were performed (1:1, 2:1, and 4:1) while maintaining the α7nAChR constant. The α7nAChR/RIC3 ratio of 4:1 corresponding to 30:7.5 ng cDNA/well did not significantly decrease the nicotine/PNU-dependent calcium flux response window when compared with an α7nAChR/RIC3 ratio of 30:30 ng/well (
(
Pharmacology of HEK293T Cells Expressing α7nAChR+RIC3
To validate the approach, we examined the pharmacology of three α7nAChR agonists using the transiently transfected HEK293T cells expressing α7nAChR+RIC3. The agonist pharmacology of HEK293T cells coexpressing α7nAChR and RIC3 was consistent with the published literature.14–16 The rank order of functional potency is AR-R17779 > nicotine > Ach, a profile that matched the calcium flux data shown in Figure 1B . AR-R17779 is a potent and selective full agonist for α7nAChR. 17 These data suggest that the transiently transfected HEK293T cells cotransfected with α7nAChR+RIC3 form functional channels with a pharmacological profile similar to that of other expression systems and native tissue.
Examining Assay Robustness
We examined the intrawell, intraplate and interday variability and signal window. For a screening-ready assay, we sought Z′ > 0.5 and a signal–background ratio (s/b) with less than 10% standard deviation (SD). The assay met these criteria (
Fig. 2
). Candidate α7nAChR modulator proteins (hits) were determined based on 3 SD from the mean.
Figure 2B
illustrates the mean for a given day (solid lines) and 3 SD from the mean (dotted line), and a clear distinction between the α7nAChR+RIC3 (high control) and α7nAChR+vector (low control) was achieved. Moreover, the Z′ value remained >0.5 from plate to plate and day to day (
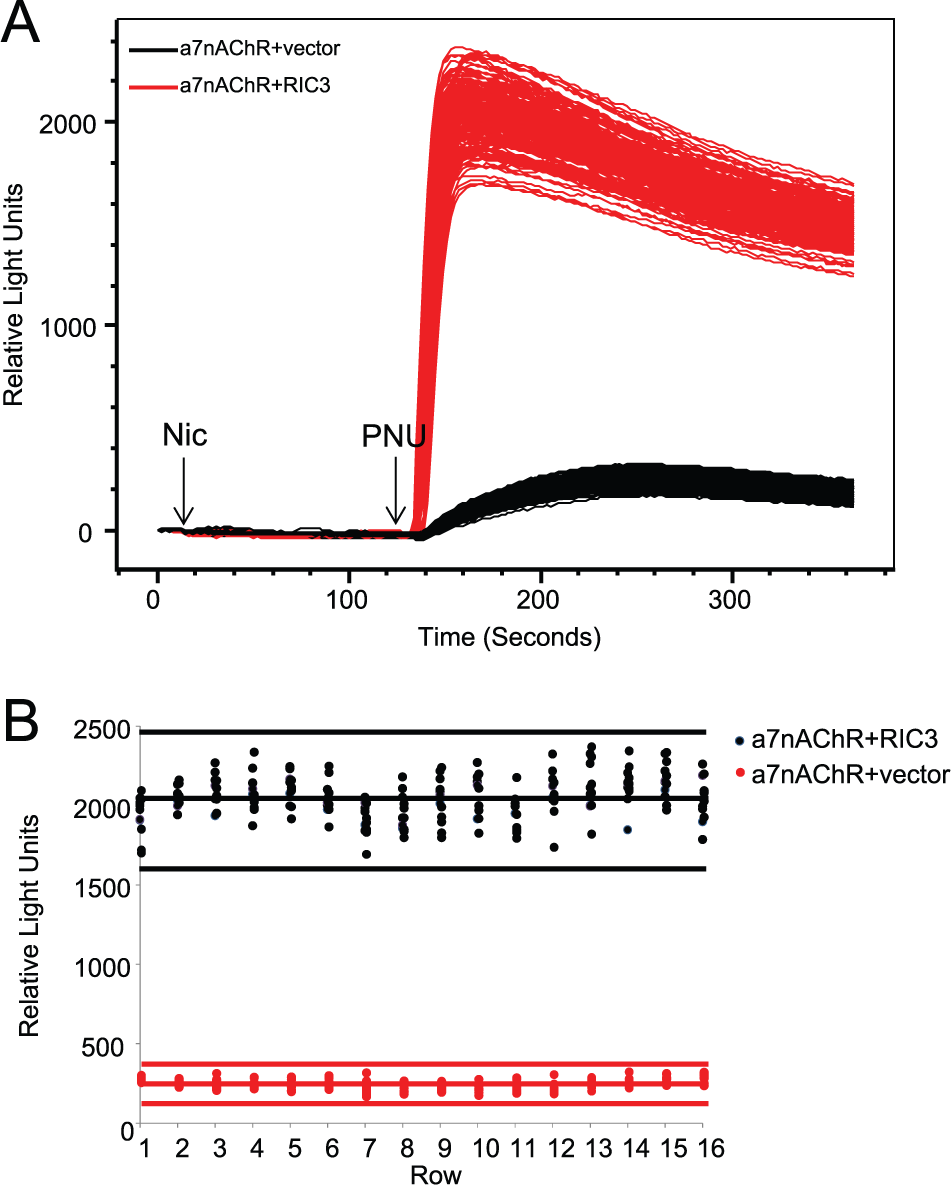
Assessing robustness and reproducibility of screening assay. (
Assay Validation
To validate that the platform was robust enough to identify novel α7nAChR modulator proteins, we ran the protocol using the Broad library plate containing RIC3 cDNAs. In this scenario, we conducted a mock run and employed all screening settings and equipment that would be used for screening the entire Broad library. Indeed, the two RIC3 cDNAs present in the Broad library (full length and truncated variants) elicited a robust calcium response (
Fig. 3
). Moreover, the Broad library RIC3 clones elicited a calcium response of similar magnitude to the RIC3 cDNA control clone when coexpressed with α7nAChR (
Fig. 3B
). In the absence of α7nAChR, RIC3 did not elicit calcium flux (
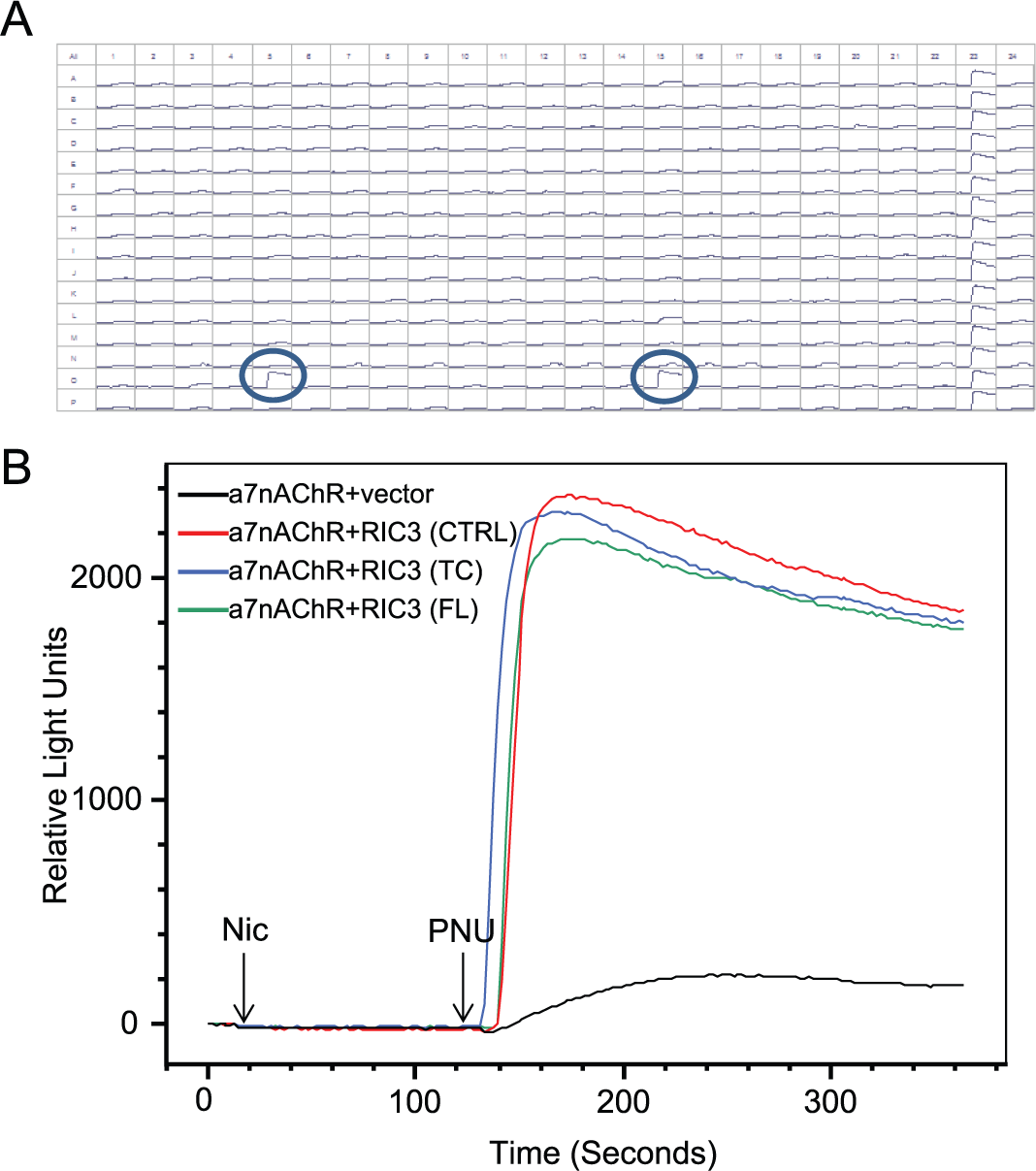
Screening of a pilot library plate to assess library performance. (
Screening Broad Genome-Wide cDNA Library to Identify Functional Modulators of α7nAChR Calcium Flux
Using the assay protocol and screening paradigm described in Materials and Methods and
A cDNA encoding a candidate functional α7nAChR modulator was classified as a hit if the α7nAChR calcium flux response reproduced in duplicate wells. Based on the cutoff criteria of 2 SD from the mean, the hit rate was 0.18%. cDNAs that elicited greater than 10.6% activity of the controls (defined by columns 23 and 24 corresponding to cells transfected with α7nAChR+RIC3 or α7nAChR+vector, respectively) in the calcium flux assay were progressed to confirmation studies. Importantly, both RIC3 variants, full length and truncated, were identified as hits in the screening campaign. A total of 32 cDNAs were reprepped from glycerol stocks, sequenced, and tested in the confirmation assay. The confirmation assay format was designed to address several questions. First, the reprepped cDNA was tested to ensure that modulation of the α7nAChR calcium flux was reproducible. Second, the cDNAs were individually cotransfected with vector and in the absence of α7nAChR in order to assess off-target, α7nAChR-independent calcium flux. The controls for the confirmation assay were the same as those for the primary screen. Columns 23 and 24 were reserved for the high- or low-control α7nAChR+RIC3 or α7nAChR+vector, respectively. The confirmation rate was 50% using the cutoff defined in the primary screen. The Z′ for the confirmation screen was 0.7. Of the confirmed hits, three cDNAs displayed α7nAChR-independent calcium flux and were not pursued further.
Functional Modulators of α7nAChR: A Data Mining Exercise
The confirmed hits were further triaged based on a data mining exercise using the Human GeneCards database to investigate the expression and biological functions (if known) of the putative α7nAChR modulator proteins. The GeneCards database integrates genomic, transcriptomic, proteomic, clinical, and functional information from a multitude of web sources. Based on microarray and RNAseq data, α7nAChR modulator transcripts that appeared to be expressed in the brain were pursued further. Subcellular localization predictions were based on COMPARTMENTS data analysis within the GeneCards database. Briefly, subcellular localization information was based on high-throughput microscopy-based screens and predictions from primary sequence and literature/text mining. 19 The predicted subcellular localization of the α7nAChR modulator proteins aligned with their functional description in most instances. For example, the control α7nAChR modulator protein, RIC3, is a molecular chaperone and localized within the ER and Golgi apparatus and was accurately predicted by COMPARTMENTS. We identified two transmembrane-containing proteins (NACHO and CCDC107) and proteins that regulate transcription or translation (EIF2AK, TCF4, CREB1, and ZC3H12A). No functional or expression information was available for the cDNA encoding SPDYE11 ( Table 1 ).
Confirmed cDNAs That Modulate α7nAChR Calcium Flux in HEK293T Cells.
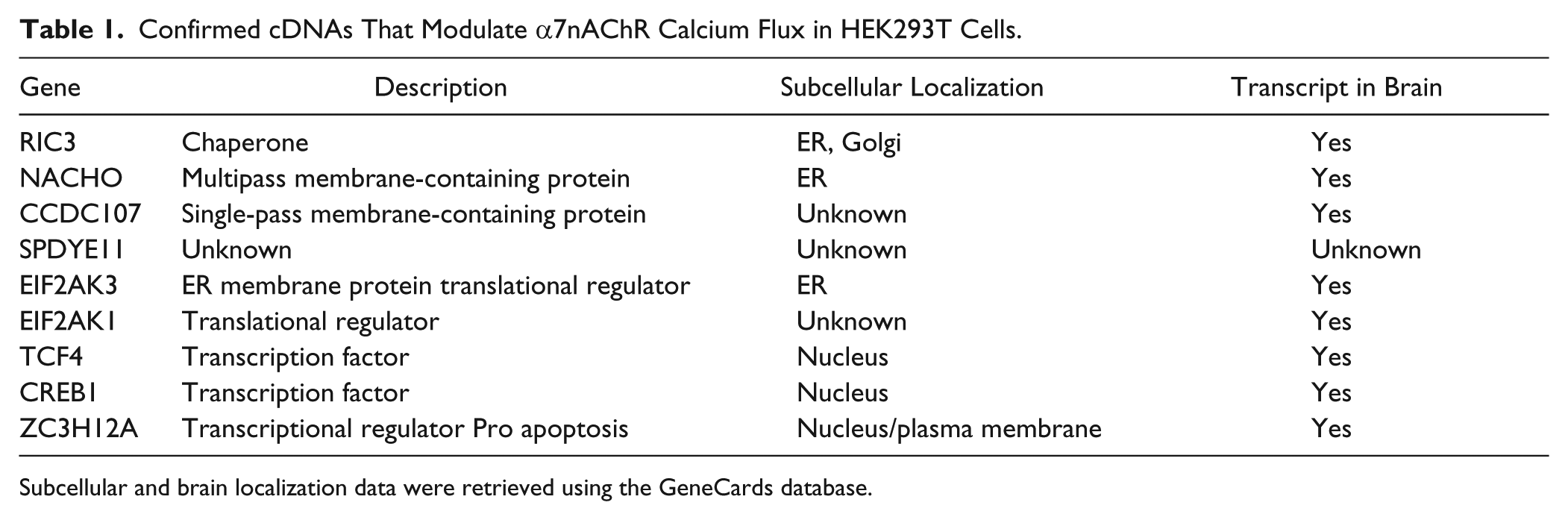
Subcellular and brain localization data were retrieved using the GeneCards database.
NACHO, SPDYE11, TCF4, and ZC3H12A Are Functional Modulators of the α7nAChR-Mediated Calcium Flux
We studied four α7nAChR modulator proteins in further detail. We selected NACHO, SPDYE11, TCF4, and ZC3H12A based on their apparent localization in the brain, predicted differential cellular/subcellular localization, and inferred functions. NACHO elicited a robust α7nAChR-mediated calcium flux when compared with vector alone (
Fig. 4A
). NACHO is a transmembrane protein that localizes in neuronal ER.
10
Interestingly, the protein with no known functional or localization data, SPEDYE11, elicited an α7nAChR-dependent calcium flux that was as equally robust as NACHO (
Fig. 4A
). The magnitude of calcium flux elicited by NACHO or SPDYE11 was comparable to that of RIC3 (
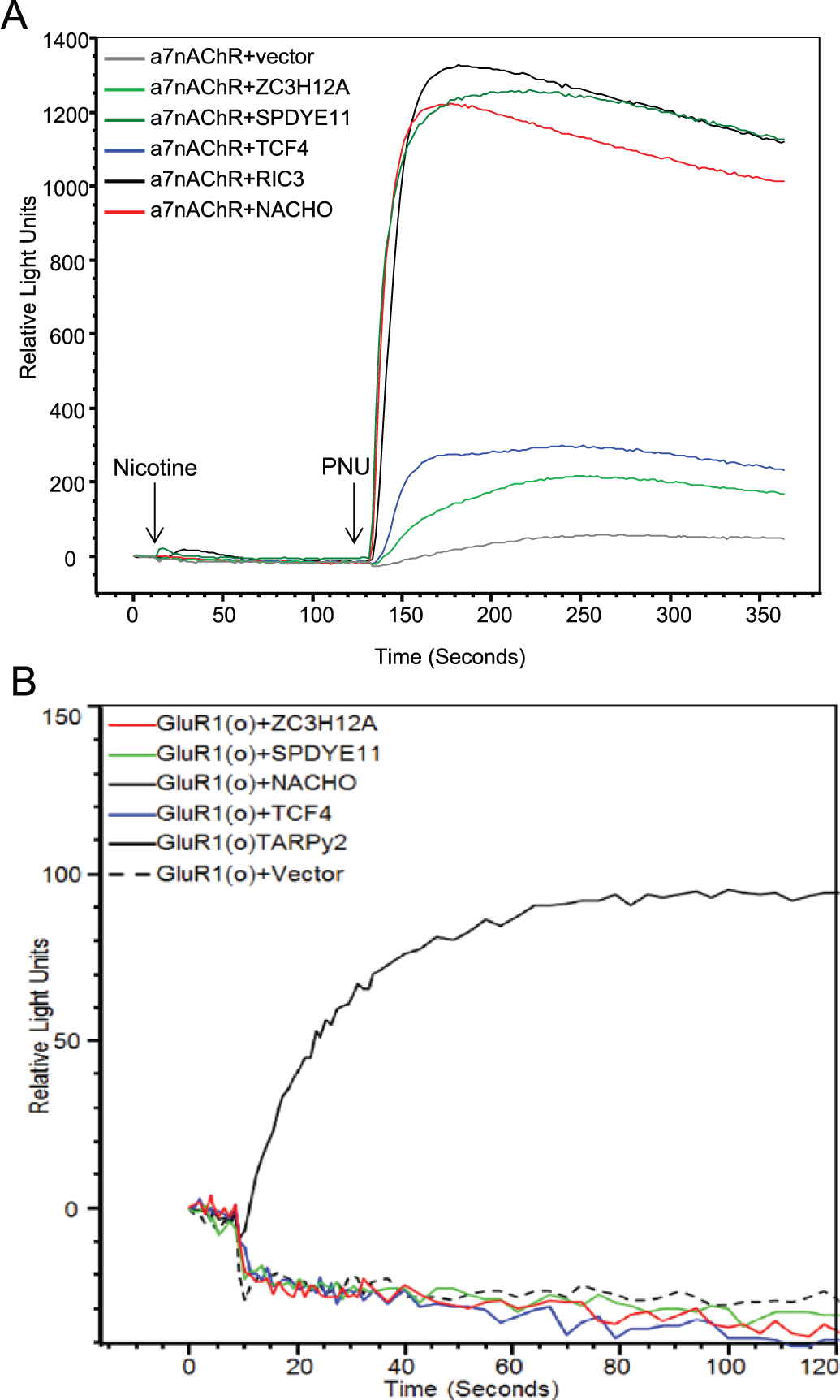
Selective modulation of α7nAChR-dependent calcium flux. (
NACHO, SPDYE11, TCF4, and ZC3H12A Do Not Elicit AMPAR GluR1(o) Calcium Flux
To begin addressing whether NACHO, SPDYE11, TCF4, and ZC3H12A are general ion channel functional modulators, we examined their propensity to modulate the AMPAR GluR1(o). GluR1(o) calcium flux is regulated by several auxiliary/modulator proteins, including TARP γ2. The regulation of GluR1(o) by TARPγ2 has been well documented in the literature and was chosen as a nonnicotinic ion channel that elicits calcium flux. In the absence of TARPγ2, a calcium response cannot be measured in HEK293T cells expressing GluR1(o)+vector (
Fig. 4B
). However, HEK293T cells cotransfected with GluR1(o) and TARPγ2 elicit a measurable calcium response (
Fig. 4B
). Significantly, none of the α7nAChR modulators, NACHO, SPDYE11, TCF4, and ZC3H12A, modulated Glu-dependent GluR1(o) calcium flux (
Fig. 4B
,
NACHO but Not SPDYE11, TCF4, or ZC3H12A Enhanced α7nAChR Surface Expression
We hypothesized that the α7nAChR modulators may enhance calcium flux by increasing the number of functional α7nAChRs expressed at the cell surface. To visualize α7nAChR surface expression, we cotransfected α7nAChR-HA-tagged cDNA with cDNAs encoding NACHO, SPDYE11, TCF4, ZC3H12A, or RIC3. The α7nAChR was engineered to express an HA tag at the extracellular C-terminus and, in combination with a fluorescently labeled HA antibody, permitted the visualization of α7nAChR expressed at the cell surface (as described in Materials and Methods). 10 As previously published, α7nAChR-HA cell surface expression was increased in HEK293T cells cotransfected with α7nAChR+NACHO. 10 Interestingly, α7nAChR-HA surface expression was not increased by SPDYE11, TCF4, ZC3H12, or RIC3. In fact, little α7nAChR was observed at the plasma membrane in HEK293T cells unless coexpressed with NACHO ( Fig. 5 ). In each of these scenarios, cells were permeabilized to confirm α7nAChR expression ( Fig. 5 ).
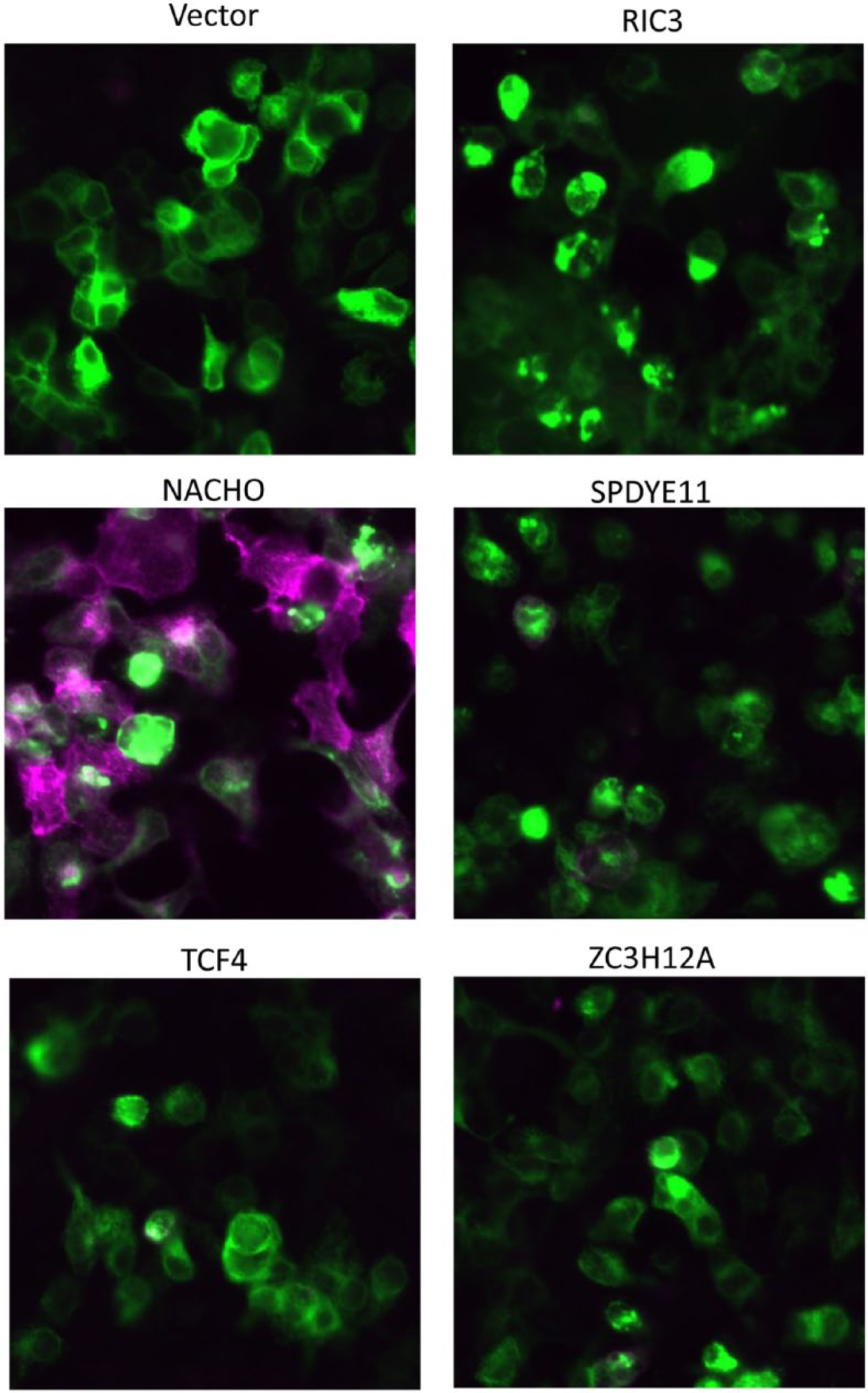
Cell surface expression of α7nAChR-HA in HEK293T cells coexpressing select cDNAs. HEK293T cells were cotransfected with α7nAChR-HA and cDNA’s encoding vector, RIC3, NACHO, SPDYE11, TCF4, or ZC3H12A. α7nAChR-HA was labeled with and without cellular permeabilization using fluorescence-conjugated HA antibodies. Purple illustrates α7nAChR-HA surface expression. Green illustrates intracellular α7nAChR-HA expression.
Discussion
In this report, we describe the development and validation of a genome-wide cDNA screening platform to identify functional modulators of α7nAChR. The α7nAChR was an attractive case study for several reasons. As mentioned earlier, the α7nAChRs are proposed therapeutic targets for treating cognitive impairment, including schizophrenia and Alzheimer’s disease. The α7nAChRs are prominently expressed in regions of the brain associated with cognitive function, such as the hippocampus, cortex, and some subcortical limbic structures, including the ventral tegmentum and substantia nigra. 6 The α7nAChRs are expressed both presynaptically and postsynaptically to modulate neurotransmitter release and excitatory neurotransmission, respectively. Nicotine-dependent activation of α7nAChR has been shown to enhance hippocampal long-term potentiation and regulate attention and memory. However, little is known about the signaling complex associated with α7nAChR. Indeed, simply transfecting HEK293T cells with α7nAChR was not sufficient to elicit a calcium response ( Fig. 1 ). However, cotransfection with the molecular chaperone, RIC3, robustly enhanced nicotine-mediated calcium flux in the presence of PNU-120596. These data suggest that the function of the α7nAChR was dependent on auxiliary proteins that may play a role in channel assembly and shuttle to the plasma membrane to elicit a response upon agonist challenge.
We sought to develop a platform that would identify proteins that functionally modulate the α7nAChR and be amenable to high-throughput screening. There are several approaches that can be employed to identify components of a signaling complex, all of which have their pros and cons. One method includes a target-specific coimmunoprecipitation approach coupled to mass spectrometry (MS) sequence analysis.20,21 In this scenario, the target is immunopreciptated from cellular lysates and subjected to two-dimensional gel electrophoresis. Unique spots or bands were excised for MS sequence analysis. An advantage to this approach is that the target of interest is expressed in a cellular environment, preserving its conformation and propensity to form protein interactions with components of the signaling complex. Ideally, the target of interest should be immunoprecipitated from a physiologically relevant cell type to increase the chance of identifying relevant modulator/auxiliary proteins and reduce artifact. The ability to immunopreciptate the target is largely dictated by the quality of the antibody and the level of expression in the cellular environment. Unfortunately, identifying quality antibodies for many integral membrane proteins can be challenging. To circumvent this caveat, the target of interest can be tagged (i.e., HA tag) to enable immunoprecipitate with HA-specific antibodies. The assumption is that the tag and/or antibody immunoprecipitating the target does not induce artificial conformational changes in the target of interest that may engage or disengage protein associations. Moreover, proteins that are less abundant may be below the levels of detection and/or migrate with highly abundant proteins during the electrophoresis procedure, and therefore may be masked. In the case of the AMPAR, the protein constituents of the AMPAR complex were identified using multiple epitope affinity purifications with 10 different antibodies specific for the AMPAR subunits to overcome the pitfalls associated with individual antibody immunoprecipitations. 20 The precipitates were subsequently analyzed by high-resolution nanoflow liquid chromatography tandem MS. It is noteworthy that associated proteins identified using these approaches bear no information on their propensity to functionally modulate the target of interest and require further studies to address this. Furthermore, this approach is not high throughput or automatable.
In this report, we describe a screening platform that utilized the Broad cDNA library in an arrayed format containing an annotated version of the human ORFeome. 18 HEK293T cells were cotransfected with the α7nAChR cDNA and each individual cDNA of the Broad library in an automated 384-well format. The HEK293 cell line was chosen for several reasons. First, the cell line was not expected to contain many regulatory proteins of receptors specifically expressed in neurons, providing the opportunity to identify modulator proteins using overexpression studies. Additionally, HEK293 cells can be transiently transfected in a robust, reproducible fashion required for an automated high-throughput screen of an arrayed cDNA library. We assessed the propensity for each cDNA from the library to modulate α7nAChR-dependent calcium flux, a direct measure of α7nAChR activity. Previously, Abujarour et al. employed a similar approach to identify regulators of pluripotency by cotransfecting cells with the Nanog luciferase reporter and individual library cDNAs. 22 Wnt signaling activators were identified using an arrayed cDNA library in combination with the b-catenin/TCF/LEF-responsive TOPflash reporter, a downstream readout of wnt signaling. 23 Ma et al. screened 409 cDNAs in a 96-well format and assayed for upregulation of a thymidine kinase–luciferase reporter construct as a readout for cell proliferation. 24 Importantly, the platform we describe in this manuscript is the first report utilizing a high-throughput genome-wide cDNA library to successfully identify functional modulators of an ion channel whereby the readout is a direct measure of the target’s activity, in this case calcium flux. As mentioned previously, the α7nAChR was a suitable case study due to the potential role in psychiatric disorders and the previously identified α7nAChR modulator protein RIC3 that served as a positive control to optimize the assay platform. In proof-of-concept experiments, we identified RIC3 present in the library as a modulator of α7nAChR. We subsequently implemented several deconvolution parameters post–primary screening. The first level of deconvolution was to confirm hit cDNAs that modulated calcium flux in HEK293T cells cotransfected with α7nAChR and not HEK293T cotransfected with vector. We hypothesized that cDNAs eliciting a calcium flux in cells cotransfected with vector (and not α7nAChR) are likely modulating endogenous receptors. The second tier of deconvolution was to prioritize α7nAChR functional modulators by performing a data mining exercise to establish their expression profile based on mRNA levels and/or function where possible ( Table 1 ). We selected four α7nAChR modulators, NACHO, SPDYE11, TCF4, and ZC3H12A, to proceed into subsequent deconvolution/characterization assays based on apparent expression in the brain and predicted differential compartment localization. NACHO and SPDYE11 were selected based on their propensities to robustly potentiate α7nAChR-dependent calcium flux when compared with other α7nAChR modulator proteins identified in the screen. The role of NACHO on α7nAChR function and regulation has been characterized and is the focus of a recent publication. 10 NACHO was shown to be pivotal for α7nAChR assembly and function in the brain. NACHO localizes to the ER and mediates correct folding and surface expression of α7nAChRs. 10 Importantly, these findings further substantiate and validate the platform described in this paper. SPDYE11 (speedy/RINGO cell cycle regulator family E11) is encoded by 265 amino acids. Based on sequence analysis, SPDYE11 contains a spy1 domain (also known as RINGO 1) that has been implicated in cell cycle regulation. However, nothing is known about the function or expression profile of this gene, and it may therefore be a truly novel finding if SPDYE11 does in fact regulate α7nAChR signaling via a novel mechanism. TCF4 encodes transcription factor 4 protein and has been implicated in psychiatric disorders. For example, single-nucleotide polymorphisms (SNPs) associated with the genome locus for TCF4 have been associated with schizophrenia based on genome-wide association studies.25,26 Furthermore, TCF4 mutations that correspond to decreased TCF levels were directly associated with the neurodevelopmental disorder Pitt–Hopkins syndrome. 27 Interestingly, TCF4 was shown to modulate the excitability of rat prefrontal neurons by a mechanism that may involve repression of the expression of SCN10a and KCNQ1 encoding sodium and potassium voltage-gated channels, respectively. 27 ZC3H12A (also known as MCPIP) is a transcriptional activator that is induced by the monocyte chemoattractant protein 1 (MCP1). MCP1 binds to its receptor, CCR2, and induces the expression of ZC3H12A. 28 Gene-based analyses have associated ZC3H12A with Alzheimer’s pathogenesis. 29 Furthermore, amyloid precursor protein (APP), the hallmark of Alzheimer’s disease, was found to increase the expression of ZC3H12A (MCPIP1) in NT2 neuroprogenitor cells. The increased levels of MCPIP induced glial differentiation and cell death. 30
All four α7nAChR functional modulator proteins, NACHO, SPDYE11, TCF4, and ZC3H12A, were evaluated in the GluR1 AMPAR FLIPR assay to explore ion channel selectivity. None of the α7nAChR modulators potentiated Glu-stimulated GluR1-dependent calcium flux when compared with the GluR1 auxiliary protein TARPγ2. These data suggest that the α7nAChR modulators exhibit some degree of selectivity toward α7nAChR overexpressed in the HEK293T cellular system. We performed α7nAChR cell surface staining in the presence of each modulator, hypothesizing that they may function by increasing the cell surface expression of α7nAChR. NACHO was the only α7nAChR modulator that increased cell surface expression. Neither SPDYE11, TCF4, nor ZC3H12 appeared to increase α7nAChR cell surface expression in these studies. How these proteins functionally modulate α7nAChR is unclear. We speculate that they may elicit intracellular calcium release that is synergistic with ion channel opening. For example, α7nAChRs in microglia have been shown to increase intracellular Ca2+ levels by the activation of phospholipase C and Ca2+ release from intracellular stores. 31 Alternatively, the modulator proteins may promote an ion channel conformation that supports the increased probability of channel open time and/or reduced desensitization. In-depth electrophysiology studies will be required to address these possibilities and are outside the scope of this manuscript.
In summary, we describe a high-throughput cell-based screening platform to identify functional modulators of the α7nAChR. The platform employs the Broad genome-wide cDNA library in conjunction with calcium flux recordings using FLIPRTETRA. Importantly, the screening platform directly measures modulation of α7nAChR activation and does not rely on artificial reporter constructs. Understanding the repertoire of proteins that constitute the α7nAChR signaling complex may unveil new opportunities for the design of novel therapeutics. Moreover, this approach can be applied to other ion channels, as well as other classes of targets, such as GPCRs and transporters.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
