Abstract
Macrophages are highly plastic cells, which serve as sentinels of the host immune system due to their ability to recognize and respond to microbial products rapidly and dynamically. Appropriate regulation of macrophage activation is essential for pathogen clearance or preventing autoimmune diseases. However, regularly used endpoint assays for analyzing macrophage functions have the limitations of being static and non–high throughput. In this study, we introduced a real-time and convenient method based on changes in cellular impedance that are detected by microelectronic biosensors. This new method can record the time/dose-dependent cell response profiles (TCRPs) of macrophages in real time and generates physiologically relevant data. The TCRPs generated from classically interferon-γ/lipopolysaccharide-activated macrophages showed considerable consistency with the data generated from standard endpoint assays. We further explored this approach by using it for global screening of a library of protein tyrosine kinase/phosphatase (PTK/PTP) inhibitors to investigate their impact on macrophage activation. Collectively, our findings suggest that the cellular impedance-based assay provides a promising approach for dynamically monitoring macrophage functions in a convenient and high-throughput manner.
Introduction
Macrophages are first-line defenders of the host innate immune system against invading pathogens. They recognize pathogen-associated molecular patterns (PAMPs) in microorganisms through pattern recognition receptor (PRRs), such as Toll-like receptors (TLRs). 1 Classical activation of macrophages, induced by interferon-γ (IFN-γ) and the TLR4 ligand, lipopolysaccharide (LPS), is characterized by secretion of high levels of proinflammatory cytokines (interleukin [IL]–1β, tumor necrosis factor [TNF]–α, IL-6, CCL2), production of bactericidal intermediates (nitric oxide, reactive oxygen species), and enhanced antigen-presenting capacity. 2 All these features contribute to pathogen clearance, initiating type I immune response and boosting antitumor immunity. However, uncontrolled overactivation of macrophages may cause detrimental outcomes, such as pulmonary inflammation, chronic obstructive pulmonary disease (COPD), inflammatory bowel disease (IBD), or cancer, underscoring the importance of appropriate regulation of macrophage activation.3–6
Macrophages respond in a highly plastic manner to external perturbations and consequently differentiate into phenotypically distinct subpopulations. Thus, macrophage activation in response to external stimuli is dynamically regulated, temporally as well as spatially, which is essential for appropriate modulation of immune response. 7 However, the factors that influence the modulation of macrophage activation have not been elucidated yet. This is partially due to the lack of compatible analytical tools that are convenient to use and at the same time generate physiologically relevant data. The most commonly used in vitro methods for studying macrophage activation are endpoint assays, which use a single or limited time course to monitor macrophage response upon stimulation. The readouts of these methods are relatively static and require seeding multiple wells of cells for a single treatment to assess the activation status of macrophages. Moreover, some antigen-based assays, such as immunofluorescence, require tedious and time-consuming experimental manipulations, making them unsuitable for high-throughput analysis. In our study, we introduce a real-time, label-free method for monitoring the kinetic behavior of classically activated macrophages, using a cellular impedance-based system. 8 Activated macrophages undergo cytoskeleton-associated morphological changes, 9 resulting in alteration in the cellular impedance (CI) index, which is continuously captured by microelectronic biosensors to generate time/dose-dependent cell response profile (TCRP) readouts. We propose that in vitro cell-based manipulation correlates these kinetic TCRPs with macrophage polarization in a more physiologically relevant manner and also appears to be highly convenient and can be used in a high-throughput manner. In the present study, we report that in macrophages, IFN-γ/LPS stimulation causes an immediate increase in the CI values followed by a decline after maximal CI values were reached. Morphologically, IFN-γ/LPS stimulation leads to an increase in cell size, which is then gradually declined, consistent with the featured TCRPs produced by macrophages. In contrast, inhibition of macrophage activation by transforming growth factor–β (TGF-β) partly reversed the increase of CI values and decreased peak CI values. Interestingly, disruption of the rearrangement of microtubules but not actin filaments was shown to be effective in altering IFN-γ/LPS–induced TCRPs in macrophages. We further screened a small compound library containing 153 protein tyrosine kinase/phosphatase (PTK/PTP) inhibitors based on the TCRP values for the purpose of investigating the impact of these PTKs/PTPs on classic macrophage activation.
In this study, we introduce a cellular impedance-based method for dynamically monitoring macrophage response after classic activation. Besides its real-time advantage, this system is particularly valuable in screening chemical compounds that can potentially affect macrophage functions in a high-throughput and unbiased manner, providing a convenient tool for discovering novel regulatory molecules involved in innate immune responses.
Materials and Methods
Chemicals and Reagents
Dulbecco’s modified Eagle’s medium (DMEM) and fetal bovine serum (FBS) were purchased from HyClone (Beijing, China). LPS, cytochalasin D, and colchicine were purchased from Sigma-Aldrich (St. Louis, MO). IFN-γ and TGF-β were purchased from PeproTech (Rocky Hill, NJ). InhibitorSelect 96-Well Tyrosine Kinase and Phosphatase Inhibitor Library IV were purchased from Merck Millipore (Billerica, MA).
Cell Culture
Murine macrophage cell line Raw264.7 was purchased from American Type Culture Collection (Manassas, VA). Cells were cultured in a humidified incubator at 37 °C in 5% CO2 in DMEM containing 50 U/mL penicillin/streptomycin (Gibco, Grand Island, NY) and 10% heat-inactivated FBS. Medium containing 2% FBS was used for macrophage activation experiments.
Real-Time Cell Analyzer System
The detailed procedures of Real-Time Cell Analyzer (ACEA Biosciences, San Diego, CA) manipulation for TCRP measurement and the calculation of CI values have been described previously.10,11 Briefly, background readings were obtained by adding 50 µL DMEM containing 2% FBS to 96-well E-plates (Roche, Mannheim, Germany). Certain numbers of Raw264.7 cells were added to E-plates and incubated at room temperature for 30 min, followed by being placed onto the reader for continuous reading of CI values. Cells were allowed to attach for 5 h to get stable baseline readings before IFN-γ (10 ng/mL) and LPS (100 ng/mL) administration. In some experiments, cells were pretreated with chemical regents or inhibitors for 2 h prior to IFN-γ stimulation, and cells were monitored every 2 min for a total of 80 h to obtain TCRPs.
Immunofluorescence Microscopy
Cells were seeded into a 24-well culture plate with glass coverslips, then treated with IFN-γ/LPS for the indicated time. Following washing with phosphate-buffered saline (PBS), cells were fixed in 4% paraformaldehyde for 20 min at room temperature. Fixed cells were washed with PBS for three times and then permeabilized with 0.1% Triton X-100 (v/v) for 20 min. Permeabilized cells were washed with PBS and then blocked with 1% bovine serum albumin for 1 h at room temperature. The cells were then stained with F-actin and α-tubulin antibodies (Beyotime, Shanghai, China) overnight at 4 °C, washed three times with PBS, and counterstained with DAPI for 5 min. Stained cells were photographed with a Zeiss LSM 510 confocal fluorescence microscope (Zeiss, Oberkochen, Germany).
Real-Time PCR
Total RNA was extracted from cells using an RNA extraction kit (Life Technologies, Shanghai, China) and reverse-transcribed using the PrimeScript II first-strand complementary DNA (cDNA) synthesis kit (Takara, Kusatsu, Japan) to generate cDNA. Real-time PCR was performed with the FastStart Universal SYBR Green Master Kit (Roche, Mannheim, Germany) on the LightCycler 480 Real-Time PCR Detection System (Roche). Primer sequences are available upon request.
Statistical Analysis
Data are presented as means ± SEM. Statistical analysis was performed in Prism 5.0 (GraphPad Software, La Jolla, CA) using one-way analysis of variance (ANOVA) or two-way ANOVA followed by Dunnett’s post hoc tests. p < 0.05 was considered statistically significant.
Results
Dynamic Monitoring of TCRPs Produced by Macrophages in Response to IFN-γ/LPS Stimulation
Electrode impedance-based technology has been described in diverse kinds of cellular behavior assays.10–12 We first addressed the feasibility of using this technology for analyzing macrophage response using a Real-Time Cell Analyzer system. To induce the activation of macrophages, different numbers of Raw264.7 cells were stimulated with IFN-γ and LPS sequentially with CI being continuously recorded. LPS stimulation induced an immediate increase in CI values, followed by peaking and then a decline of the CI values, and finally the CI values maintained at relatively low levels. Different numbers of cells generated similar but shifted CI curves ( Fig. 1A ). CI values peaked earlier when higher numbers of cells were seeded. Since peak CI values no longer increased when more than 4 × 104 cells were seeded, we selected 2 × 104 cells to perform all of the following experiments. We then stimulated fixed numbers (2 × 104) of Raw264.7 cells with increasing concentrations of IFN-γ/LPS. A dose-dependent effect of IFN-γ/LPS on peak CI values was observed, which all peaked approximately 40 h after IFN-γ treatment (18 h after LPS treatment) and gradually declined to levels comparable to that of resting macrophages ( Fig. 1B ).
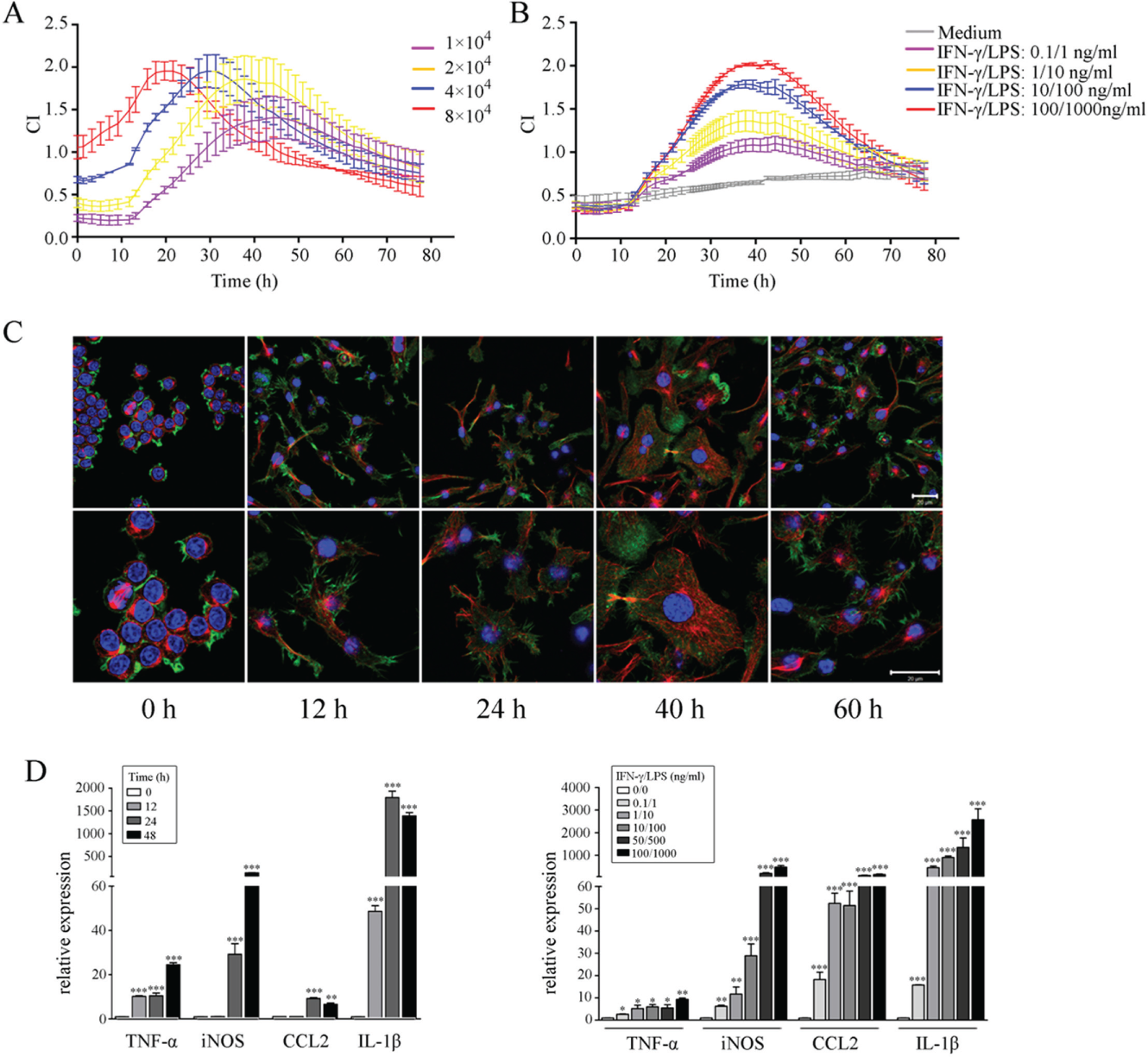
Dynamic monitoring of macrophage responses to interferon-γ (IFN-γ)/lipopolysaccharide (LPS) by time/dose-dependent cell response profile (TCRP)–based method and regular assays. (
Upon being activated, macrophages undergo cytoskeletal rearrangements and morphological changes, and they secrete high amounts of proinflammatory cytokines. We observed membrane ruffling and pseudopodia extension, which were indicative of cytoskeletal rearrangement in macrophages upon activation. A gradual increase in cell size was observed along with time; the most notable morphological changes appeared 40 h after activation, followed by a decline at 60 h, corresponding to the rise, peak, and decline phases of the TCRP, respectively ( Fig. 1C ). Moreover, these visual morphological changes were accompanied with dramatically increased production of the transcripts of proinflammatory cytokines TNF-α, IL-1β, CCL2, and arginine-metabolic enzyme inducible nitric oxide synthase (iNOS), all of which are hallmarks of classically activated macrophages ( Fig. 1D ). These findings suggest that cellular impedance-based TCRPs could serve as an alternative of regular endpoint assays in evaluating the activation status of macrophages.
Different Inhibitors of Macrophage Activation Produce Distinct TCPRs
Since activated macrophages can produce characteristic TCRPs mirroring their activation status, we wanted to investigate how the inhibition of macrophage activation would affect TCRPs. Raw264.7 cells were pretreated with increasing concentrations of TGF-β, a well-known anti-inflammatory cytokine that dampens macrophage response to Toll-like receptor ligands, prior to IFN-γ/LPS stimulation. Macrophage activation induces a marked increase of CI values as we have already shown herein. However, this increase was relatively milder when cells were pretreated with TGF-β, even at concentrations as low as 0.016 ng/mL, demonstrating the high sensitivity of our TCRP system in detecting subtle cellular changes ( Fig. 2A ). TGF-β affected TCRPs by reducing the height of CI curves, without causing a horizontal shift along the X-axis (time axis). This finding suggested that TGF-β “suppressed” rather than “delayed” the classic activation of macrophages.
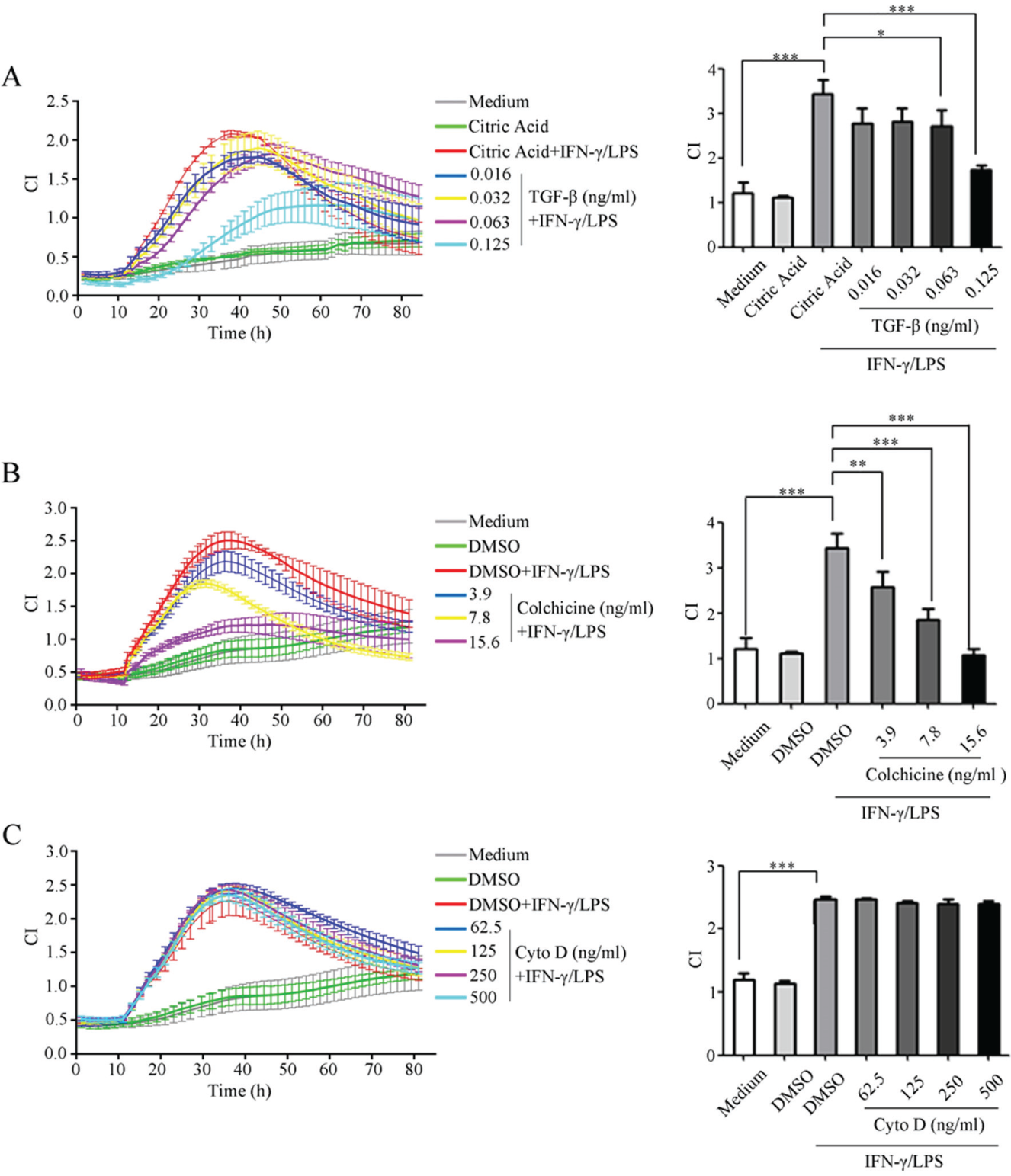
Measurements of time/dose-dependent cell response profiles (TCRPs) produced by macrophages activated in the presence of different inhibitors. Raw264.7 cells were pretreated with different concentrations of transforming growth factor–β (TGF-β) (
TCRPs generated by macrophage activation were associated with cytoskeletal rearrangements; therefore, we were interested in finding out how the CI profiles would be affected when the cytoskeleton was disorganized. We found that colchicine-induced microtubule depolymerization significantly inhibited macrophage activation in a dose-dependent manner, as suggested by a milder CI increase as well as lower peak CI values, compared with DMSO-treated macrophages (
Fig. 2B
). Remarkably, the inhibition of actin filament polymerization by cytochalasin D failed to change CI patterns even at high concentrations (
Fig. 2C
), indicating that cellular impedance may be differentially influenced by the organization of actin filaments and microtubules. TGF-β, colchicine, or cytochalasin D alone did not obviously change CI values in the absence of IFN-γ/LPS stimulation (
Global Screening and Functional Analysis of Kinetic Macrophage Responses to PTK/PTP Inhibitors
High throughput is another advantage of the TCRP-based assay. Hence, we applied this method to the measurement of kinetic cell responses to a number of PTK/PTP inhibitors. PTKs/PTPs are critical regulators of cellular signaling and behavior,
13
and different types of chemical inhibitors have been developed for investigating their impact on cellular functions. A TCRP-based heatmap in terms of macrophage responses to an inhibitor library containing 153 PTK/PTP inhibitors is shown in
Figure 3
and
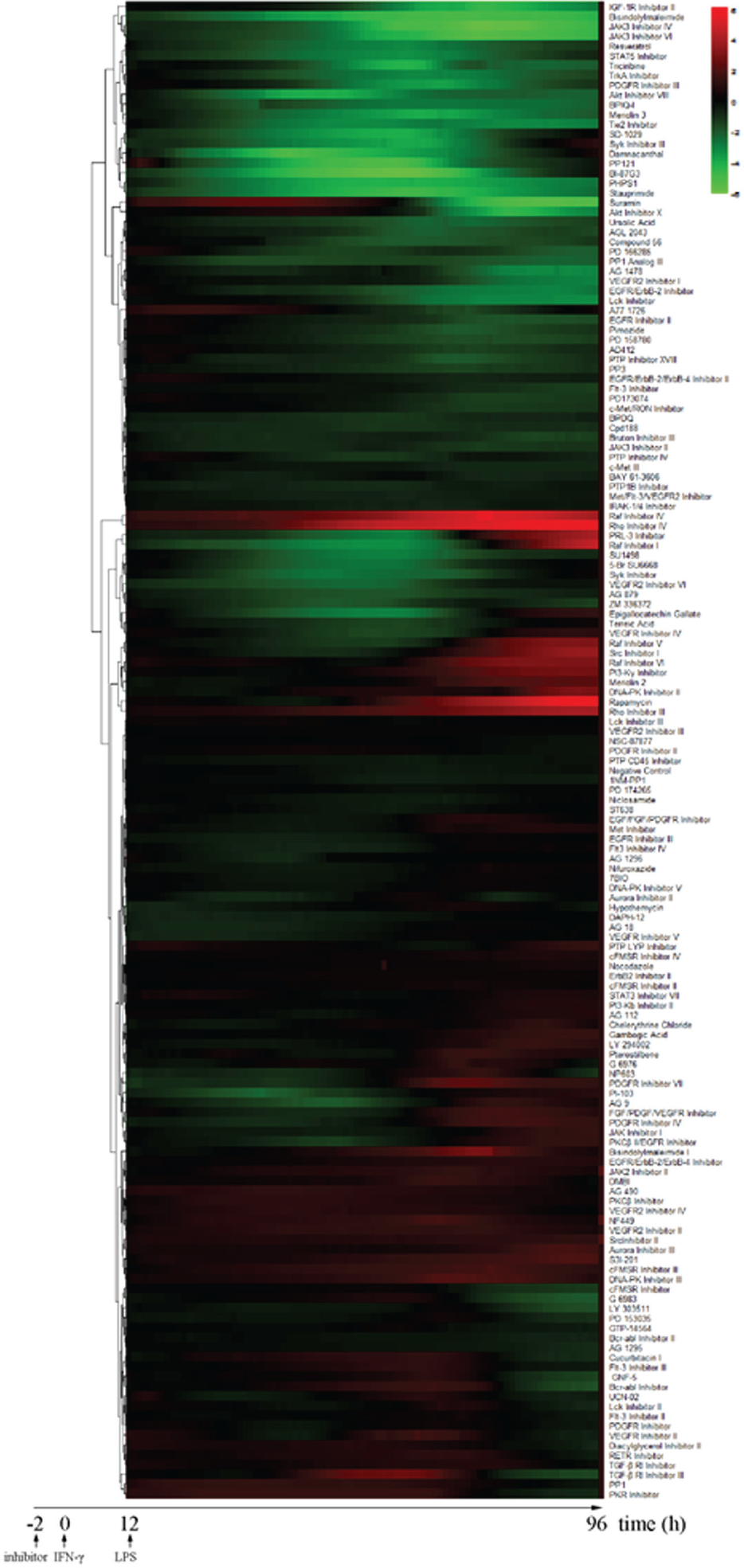

Agglomerative hierarchical clustering analysis of dynamic macrophage responses to interferon-γ (IFN-γ)/lipopolysaccharide (LPS) stimulation. Relative cellular impedance (CI) values were mapped to colors from green (z = −6.0) to red (z = 6.0) and plotted as a function of time from 12 h to 80 h. Inhibitors that had similar functions were subgrouped together. Relative CI values were calculated as CIinhibitor – CIDMSO at each time point.
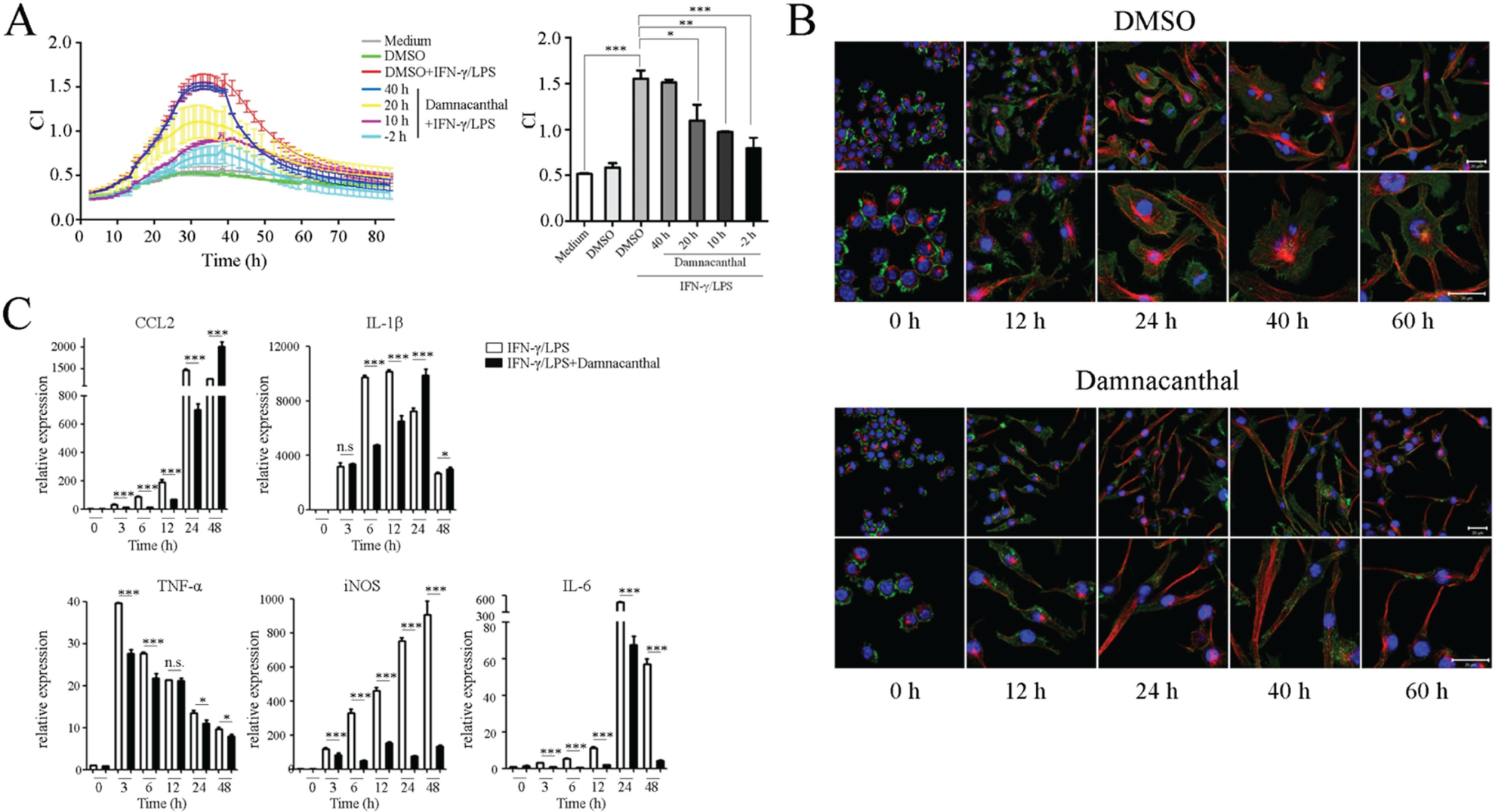
Monitoring the impact of damnacanthal on macrophage activation by time/dose-dependent cell response profile (TCRP)–based method and regular assays. (
Discussion
Appropriate macrophage responses to external stimuli are important for maintaining immune homeostasis in host-microbiome interactions. Understanding the processes of such interactions helps to control infectious diseases and hyperactivated inflammatory responses. Many technologies, such as real-time PCR, Western blot, enzyme-linked immunosorbent assay, flow cytometry, and immunofluorescent imaging have been developed to study macrophage functions. These methods are all endpoint assays that collect and study cells at a single or few time points. However, the responses of macrophages to most stimuli are rapid, dynamic, and even reversible. Therefore, endpoint assays are disadvantageous for real-time monitoring of cell behavior and for tracing subtle cellular responses. Besides, endpoint assays always involve tedious and time-consuming experimental procedures, and at least one well of cells is required for analysis of one time point. These limitations render the currently available assays unsuitable for high-throughput study of cell functions.
In this report, we introduce a cellular impedance-based method for studying macrophage activation by taking advantage of the Real-Time Cell Analyzer system, which has been applied to a wide range of analyses, including study of epithelial barrier function, 11 mast cell function, 15 cell receptor kinase activation,16,17 cell proliferation and cytotoxicity assays, 18 cell adhesion and migration assays, 19 and cell responses to Salmonella infection, 12 or for studying biologically active compounds. 10 The combinatorial treatment of IFN-γ and gram-negative bacteria-derived LPS has been extensively used for in vitro studies to induce classic macrophage activation, which has crucial physiological roles in pathogen clearance. In addition to cytokine production, IFN-γ/LPS–activated macrophages are characterized by the formation of lamellipodia, filopodia, and membrane ruffles, leading to altered cellular impedance. Using CI values as readouts, we observed considerable consistency between TCRP-based and endpoint-based assays when both were used for analyses of classic macrophage activation. Different numbers of cells produce similar but shifted CI curves, while the enhanced and suppressed activation of macrophages can be respectively reflected by the relative rise and decline of CI values in an immediate, dynamic, and unbiased manner. Macrophage activation is accompanied with cytoskeletal changes that are thought to be mediated by the rearrangement of actin filaments and microtubules. We found that the inhibition of tubulin polymerization by colchicine induces a dose-dependent decline of CI values. Surprisingly, cytochalasin D–mediated actin depolymerization had no impact on the CI pattern. This unexpected phenomenon suggests that actin filaments and microtubules may have a different impact on cellular impedance, at least in classic IFN-γ/LPS–activated macrophages.
The convenient and dynamic nature of the TCRP-based method makes it particularly suitable for high-throughput screening purposes; we reported the use of this system for globally analyzing cell responses to 146 herbal plant species in our previous study. 10 In the present study, we have profiled TCRPs generated by IFN-γ/LPS–activated macrophages in response to 153 PTK/PTP inhibitors. PTKs/PTPs are involved in almost all cellular signal transduction pathways, and aberrant PTK/PTP expression or activity is responsible for many diseases such as inflammation, cancer, and diabetes.20–23 Our TCRP-based method provides a promising tool for investigating the role of PTKs/PTPs or other kinds of molecules in the process of macrophage activation.
Although IFN-γ/LPS stimulation is extensively used for mimicking bacterial infection, macrophages can respond to a wide range of pathogen-associated molecular patterns, for example, gram-positive bacterial cell wall components (peptidoglycan, lipoteichoic acid), fungal cell wall components (zymosan), nucleic acid components derived from the genomes of bacteria or viruses (double-stranded RNA/DNA, single-stranded RNA), bacterial flagellin, or helminth-derived products. On the other hand, dendritic cells, neutrophils, and other nonhematopoietic cells such as epithelial cells, endothelial cells, and fibroblasts also express PRRs and participate in host innate immunity. 24 Therefore, our present study provides a promising tool for investigating the regulation of innate immune responses. However, the inherent heterogeneity of different cell types should be taken into account when designing experimental procedures; hence, factors including cell number, growth rate, morphology, and adhesive quality should be considered.
In conclusion, we introduce herein for the first time a TCRP-based method for studying macrophage function. Due to the advantages of being real time, label free, and high throughput, this method can be used either in combination with or instead of regular assays in studying cellular responses against pathogens and has the potential of being developed for broader applications in the future.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (31371415 and 81530001 to Y.K.), (81370506 to Q.C.), and Natural Science Foundation of Zhejiang Province (LY16H030009 to C.Q.).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
