Abstract
In this work, a sensitive and convenient protease-based fluorimetric high-throughput screening (HTS) assay for determining peptidyl-prolyl cis-trans isomerase activity was developed. The assay was based on a new intramolecularly quenched substrate, whose fluorescence and structural properties were examined together with kinetic constants and the effects of solvents on its isomerization process. Pilot screens performed using the Library of Pharmacologically Active Compounds (LOPAC) and cyclophilin A (CypA), as isomerase model enzyme, indicated that the assay was robust for HTS, and that comparable results were obtained with a CypA inhibitor tested both manually and automatically. Moreover, a new compound that inhibits CypA activity with an IC50 in the low micromolar range was identified. Molecular docking studies revealed that the molecule shows a notable shape complementarity with the catalytic pocket confirming the experimental observations. Due to its simplicity and precision in the determination of extent of inhibition and reaction rates required for kinetic analysis, this assay offers many advantages over other commonly used assays.
Introduction
Peptidyl-prolyl cis-trans isomerases (PPIases) are ubiquitously expressed and highly conserved proteins found in both prokaryotic and eukaryotic cells. 1
Based on drug specificity and primary sequence homology, PPIases have been divided into three distinct families: (1) the cyclosporin A (CsA) binding proteins, cyclophilins; (2) the FK506 and rapamycin binding proteins, FKBPs; and (3) the parvulins, which do not bind immunosuppressant drugs.1,2
The principal function of PPIases is to participate in protein folding by catalyzing the cis-trans isomerization of X-Pro peptide bonds in polypeptide chains (where X is any amino acid).1,2 The conformational interconversion of the backbones of signaling proteins mediated by peptidyl-prolyl cis-trans isomerization plays a pivotal role in many aspects of cellular processes, including posttranslational modifications such as phosphorylation and glycosylation, whose dysregulation does contribute to the development of many human diseases. 3
For this reason, great efforts are spent on the identification of specific PPIase modulators for therapeutic approaches. However, the discovery of novel inhibitors has been limited by the lack of suitable assays for high-throughput screening (HTS).
In this study, we have developed a sensitive and robust HTS method for monitoring the peptidyl-prolyl cis-trans isomerase activity of human cyclophilin A (CypA) and used the assay to identify prototypical inhibitors from the Library of Pharmacologically Active Compounds (LOPAC), a commercial library of bioactive drugs.
CypA is the most important member of the human cyclophilins family. 4 CypA plays a critical role in a variety of biological processes, including protein folding, trafficking, assembly, immunomodulation, and cell signaling. 4 Moreover, besides its physiological role, the PPIase activity of CypA is also implicated in various pathological conditions and diseases, including viral infection, cancer, and neurodegeneration. 4
Conventionally, the enzymatic activity of PPIases has been determined by UV/Visible spectrophotometry with N-succinylated tetrapeptide-4-nitroanilides (pNA) as standard substrates (succinyl-Ala-X-Pro-Phe-pNA). 5 This assay is based on the conformational specificity of chymotrypsin, which cleaves the pNA moiety from the substrate only when the X-Pro amide bond is in the trans conformation. In the presence of PPIases, the X-Pro bond is more rapidly converted to the trans conformation and the substrate is then more readily cleaved by chymotrypsin, leading to the formation of the colored product pNA. 5 However, this method suffers from low sensitivity of the chromogenic compound, as well as the fast spontaneous cis-trans isomerization, which translates into the rapid processing of the substrate (generally in the range of seconds) by chymotrypsin. 5 Since the spontaneous cis-trans isomerization is too fast for reliable calculation of rate constants at room temperature, the assay is usually carried out at or below 10 °C and in presence of tetrafluoroethylene (TFE) to force the cis conformation of the substrate. Nonetheless, even under these conditions, a typical halftime of the uncatalyzed cis-trans isomerization is about 100 s. 5
In the attempt to develop assays more sensitive and suitable for HTS programs, several new probes and detection methods have been developed. 6 However, to date, no suitable substrates for monitoring PPIase activity in HTS modality have been reported. Indeed, the reaction is always too fast and needs organic solvents, such as TFE, that strongly affect viscosity and impair volume measurements during liquid handling. The main features of HTS-compatible assays include robustness, sensitivity, automation, and miniaturization. Sensitivity, in this instance, is intended as the capability to generate signals sufficiently higher than the noise, but also the ability to detect small inhibition differences. Robustness instead refers to the possibility to have assay readouts stable and reproducible in a time range adequate to the assay timelines. 7
In this framework, we have designed and tested a novel peptide substrate targeting the catalytic site of CypA and useful in HTS-based protease-coupled PPIase assays. To transfer the assay in an HTS format, we have investigated and optimized parameters such as buffer composition, reaction time, enzyme/substrate ratio, and DMSO content. The conditions for achieving optimal signal-to-noise ratio (S/N) and Z′ factor values have been identified. Moreover, using the optimized and automated assay, we have selected a small-molecule inhibitor (<500 Da) of CypA by the screening of the LOPAC. The inhibitor shows an IC50 in the low micromolar range, and molecular docking studies suggest how it interacts and occupies the catalytic groove of CypA.
Materials and Methods
Protected amino acids, coupling agents (HATU), and Fmoc-Rink Amide AM Resin used for peptide synthesis were purchased from IRIS Biotech GmbH (Marktrewitz, DE). Fmoc-Glu(EDANS)-OH and Fmoc-Lys(Dabcyl)-OH were from PolyPeptide Group (Strabourg, France). Solvents, including acetonitrile (CH3CN), dimethylformamide (DMF), trifluoroacetic acid (TFA), and methanol (CH3OH) were purchased from ROMIL (Dublin, Ireland). Other products, such as Sym-collidine, DIPEA, piperidine, CsA, TLCK-treated α-chymotrypsin (hereafter, chymotrypsin) from bovine pancreas, and LOPAC1280 small library were from Sigma-Aldrich (Milan, Italy).
Analytical high-performance liquid chromatography (HPLC) analyses for monitoring the cis-trans transition were performed on an Alliance HT WATERS 2795 system, equipped with a PDA WATERS detector 2996. Preparative purifications were carried out on a WATERS 2545 preparative system (Waters, Milan, Italy) fitted out with a WATERS 2489 UV/Visible detector. Liquid chromatography–mass spectrometry (LC-MS) analyses were performed using an ESI Ion Trap HCT ETD II HC Ultra PTM discovery system Bruker mass spectrometer coupled with an HPLC System Alliance e2695 separation module fitted out with a 2998 PDA detector (Waters). An automated MICROLAB STAR Liquid Handling Workstation from Hamilton Robotics (Bonaduz, Switzerland) was used to develop the screening assay. The workstation includes a set of eight independent pipetting channels, several positions for sample tubes and plates, the barcode identification for samples, microplates, reagents, and carriers. The liquid handler was implemented with the Total Aspiration and Dispense Monitoring (TADM) system, with the Liquid Level Detection system and the Tip Attachment (CO-RE) system. In all procedures, the Monitored Air Displacement (MAD) system was used. All workstation functions and integrated third-party devices were controlled by the Venus software (Hamilton Robotics). An EnSpire Multimode Plate Reader (PerkinElmer) was used to perform fluorescence measurements.
HPLC Analysis
Time- and concentration-dependent isomerase activity of CypA was evaluated by HPLC using the new Fmoc substrates. The conditions used were as follows: Onyx monolithic C18 column (50 × 2 mm) operated at a flow rate of 0.6 mL/min; eluents: H2O + 0.1% TFA and CH3CN + 0.1% TFA; wavelength 265.8 nm. A linear gradient from 15% to 80% of CH3CN + 0.1% TFA in 4 min was applied to elute the products. The differences between CypA-catalyzed and spontaneous cis-trans isomerization rates of the Fmoc-protected substrates were evaluated by following the Fmoc-full-length peptide signal reduction and/or the increase of chymotrypsin-hydrolyzed Fmoc-peptide at 265.8 nm (maximum Fmoc absorption). The assays were carried out at room temperature and in a volume of 200 μL, by using the substrates at 10 ng/μL, commercial chymotrypsin at 100 ng/μL, and CypA at 200 ng/μL. Assays were performed in two different buffers, HEPES 35 mM and LiCl 5 mM or HEPES 35 mM and NaCl 150 mM, to evaluate the effect of positive ions on the population of cis conformers and on the reaction kinetics.
Assay Automatization
The chymotrypsin-coupled enzymatic assay was performed in 384-well black plates (PerkinElmer) in a volume of 50 μL using the automated MICROLAB STAR Liquid Handling Workstation from Hamilton Robotics. For assays, chymotrypsin, CypA, and expanded polypropylene (EPP) substrate solutions in reaction buffer (phosphate-buffered saline [PBS] 1×, pH 7.4) were prepared at a concentration fourfold higher than that of the final concentration used in the assay. Solutions of inhibitors were prepared in 96-well plates (mother plates) at a 4× concentration in the reaction buffer.
The procedure consisted of the following steps:
Dilution of compounds from the stock solution in the reaction buffer in the mother plate
Dispensing 12.5 μL of buffer or inhibitor solution into 384 wells of black OptiPlate-384
Addition of 12.5 μL of FRET substrate (60 ng/μL)
Addition of 12.5 μL of CypA (2.2 μg/μL)
30 min incubation at room temperature, in the dark
Addition of 12.5 μL of chymotrypsin solution (600 ng/μL)
Reading over time of fluorescence (λex = 340 nm and λem = 510 nm)
Experiments were run as quadruplicate and reported as averaged values ± standard deviation (SD).
Determination of the Steady-State Kinetic Parameters for EPP Substrate
To determine the Km/Kcat values, the FRET substrate was serially diluted in PBS 1×, pH 7.4 (from 1 to 250 μM). The peptide was then mixed with CypA (600 ng/μL) and chymotrypsin (150 ng/μL). After 30 min, the fluorescence of EDANS was measured at 510 nm upon excitation at 340 nm.
Assay Parameter Calculations
Signal-to-background ratio (S/B) was defined as the XP/XN ratio of peak areas obtained for the PPcut and EPPcut peptides during the CypA-assisted (XP) and not assisted (XN) chymotrypsin cleavage. The S/N was calculated by using eq 1: 8
Z′ factor values were calculated on 32 independent data points for each condition, as reported in literature. 8 In the assays, chymotrypsin (150 ng/μL) was added in 384-well plates in which the substrate and CypA were already dispensed as previously described, and fluorescence was recorded after 30 min. The assay was performed also in the presence of increasing amounts of DMSO (1%, 2%, and 3%).
IC50 Determination
Dose-dependent assays with the CsA and with hits were performed at concentrations ranging between 0.001 and 25 μM for the CsA and between 0.001 and 50 μM for the selected inhibitors. The FRET substrate, the CypA, and the hits were incubated for 30 min at room temperature. After incubation, chymotrypsin was added, and after 30 min, fluorescence was detected. For all these assays, each data point was in quadruplicate. Experimental data were fitted with GraphPad Prism, v5.00 (GraphPad Software, San Diego, CA).
NMR Spectroscopy
Nuclear magnetic resonance (NMR) experiments were carried out at 298.0 K using an Inova 600 MHz spectrometer (Varian, Inc., Palo Alto, CA), equipped with a cryogenic probe optimized for 1H detection. NMR data were collected on samples of EPP substrate without the Fmoc group to improve solubility. Samples were prepared by dissolving the peptide at a concentration of 600 µM in 20 mM sodium phosphate buffer, pH 7.0, containing 10% 2H2O (500 μL) and 150 mM NaCl or 5 mM LiCl. One-dimensional (1D) 1H spectra and two-dimensional (2D) [1H, 1H] DQF-COSY, TOCSY, NOESY, and ROESY spectra were acquired according to the procedures described in Kaczmarek et al.
9
(see
Molecular Modeling
Molecular docking studies were performed using DockBench platform DockBench 1.01.
10
The procedures used are reported in
Results
Design and testing of substrates
Two new PPIase substrates, hereafter named PP and EPP, were designed in order to force their structures, at room temperature and in the absence of organic solvents, into a predominantly cis conformation, and thus to enhance assay sensitivity. Specifically, besides the consensus sequence Pro-Phe (P-F) present in common PPIase substrates, a second P was inserted before the P-F segment, as suggested by several other studies. 12 Moreover, a lysine (K6 in PP peptide or K8 in EPP peptide) was introduced in both sequences to improve the peptide solubility. A glutamic acid was inserted at the N-terminus of the EPP peptide (E1) to potentially stabilize the cis conformation of the X-Pro bond through a salt bridge with the K8 at the C-terminus. 13 Both peptides bore an Fmoc group on the N-terminus to facilitate identification and detection by HPLC.
Peptides were easily synthesized using the standard solid-phase Fmoc method (see
Time- and concentration-dependent isomerase activity of CypA were evaluated by HPLC using the PP and EPP substrates conjugated with the Fmoc group (see
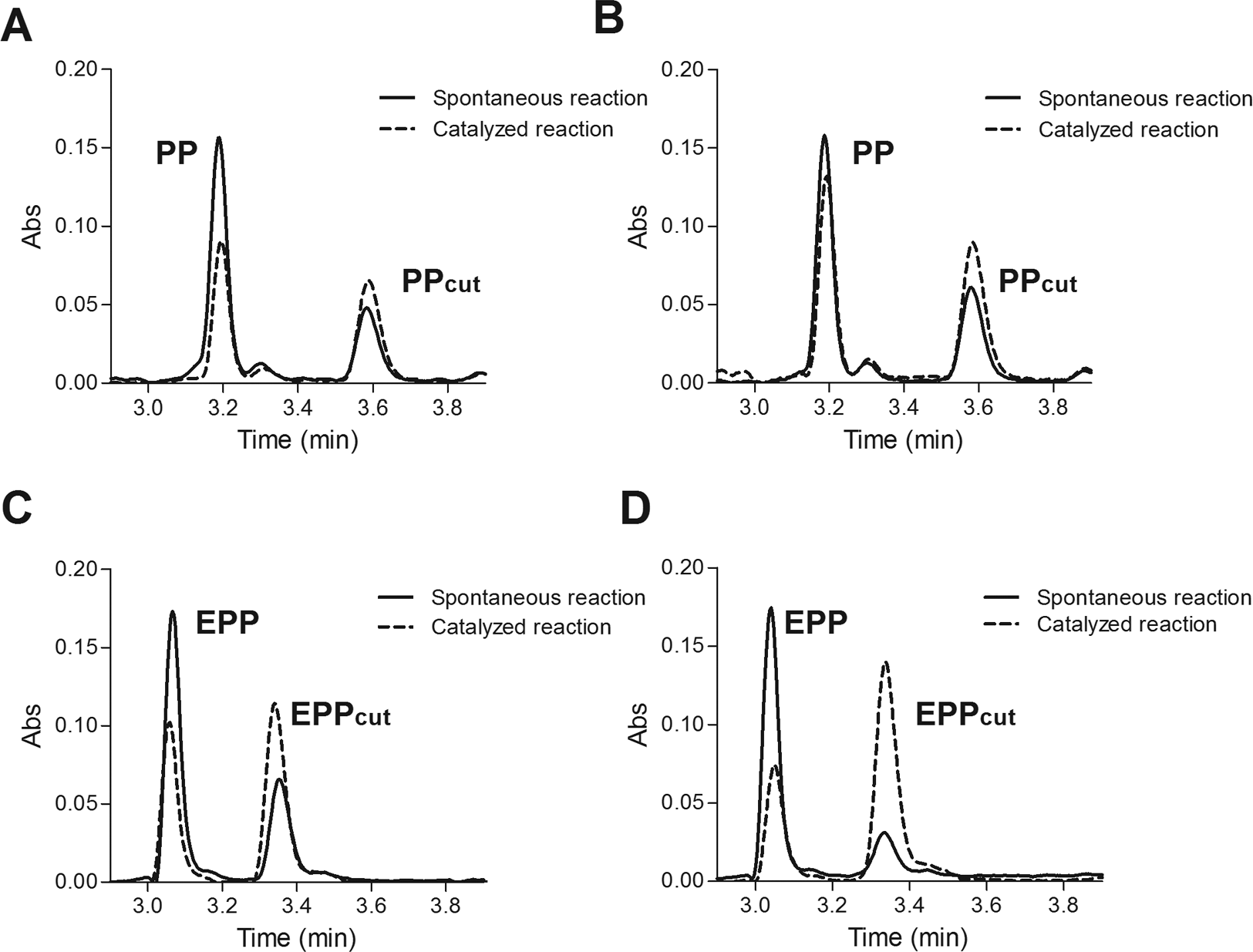
Chymotrypsin-coupled isomer-specific proteolysis HPLC assay. (
Sequences of Peptides Used for Monitoring the CypA Isomerase Activity through HPLC.
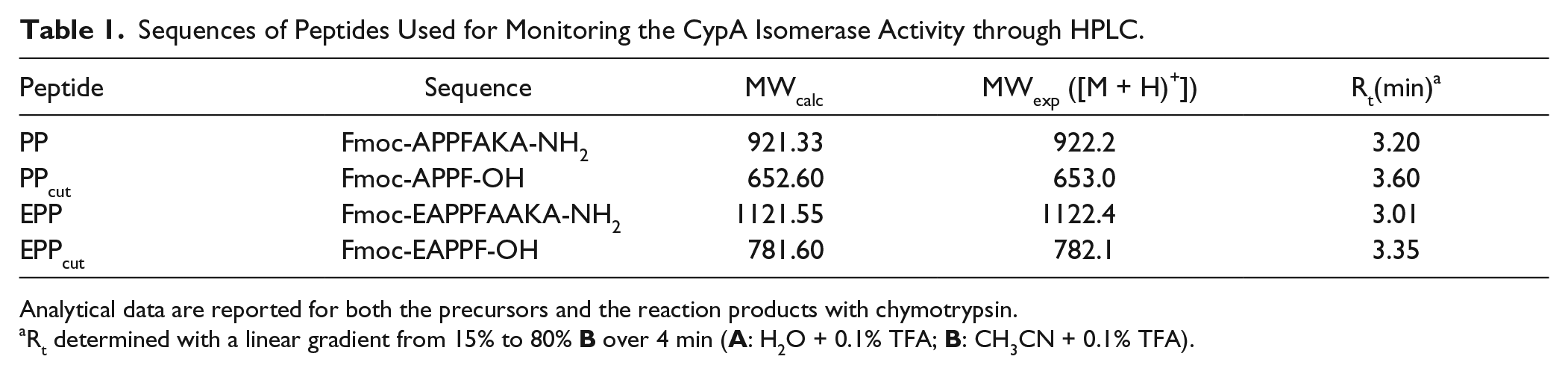

Analytical data are reported for both the precursors and the reaction products with chymotrypsin.
Rt determined with a linear gradient from 15% to 80%
Most bioactive compounds are characterized by very low solubility in aqueous solvents; thereby compounds used in HTS are typically dissolved in DMSO starting stock solutions. For this reason, enzymatic assays are mostly developed in the presence of this organic solvent. Therefore, we carried out time-course experiments using the EPP substrate (10 ng/μL), CypA (200 ng/μL), and chymotrypsin (100 ng/μL) in HEPES 5 mM, NaCl 150 mM, pH 7.4, at increasing concentrations of DMSO (from 0.5% to 5%) and measuring the enzyme kinetic as described above. Assay performances in these conditions were evaluated in terms of S/B values calculated as described in the
We also evaluated the effect of the natural inhibitor CsA
1
on CypA isomerase activity in our conditions. Reactions were carried out as described above in the presence of 1% DMSO, because of the low solubility of CsA. Data show that the presence of CsA interfered with the CypA isomerase activity in a dose-dependent manner, showing a IC50 in the nanomolar range (150 nM), compatible with values reported in the literature (
NMR Structural Characterization of the New CypA Substrate
To provide structural insights into the conformation of the selected EPP substrate, we performed NMR analysis on the Fmoc-free peptide in aqueous buffers containing the different ions. In particular, 1D 1H and 2D [1H, 1H] TOCSY, NOESY, and ROESY spectra of the peptide were acquired in phosphate buffer in the presence of either LiCl or NaCl.
According to the presence of two X-Pro peptide bonds (A2-P3 and P3-P4), four conformational states (numbered from 1 to 4) were observed in a slow-exchange equilibrium. Indeed, as can be seen in Figure 2A , B , the F5 HN, immediately following the P3-P4 segment, exhibited four different chemical shifts in either saline buffer tested (NaCl and LiCl). However, different relative peak intensities, correlating with the relative populations of the four conformers, were observed in the two buffers tested. The F5 HN-Hβ cross-peak volumes from TOCSY spectra were used to determine the relative conformer populations, which are shown in Figure 2C .
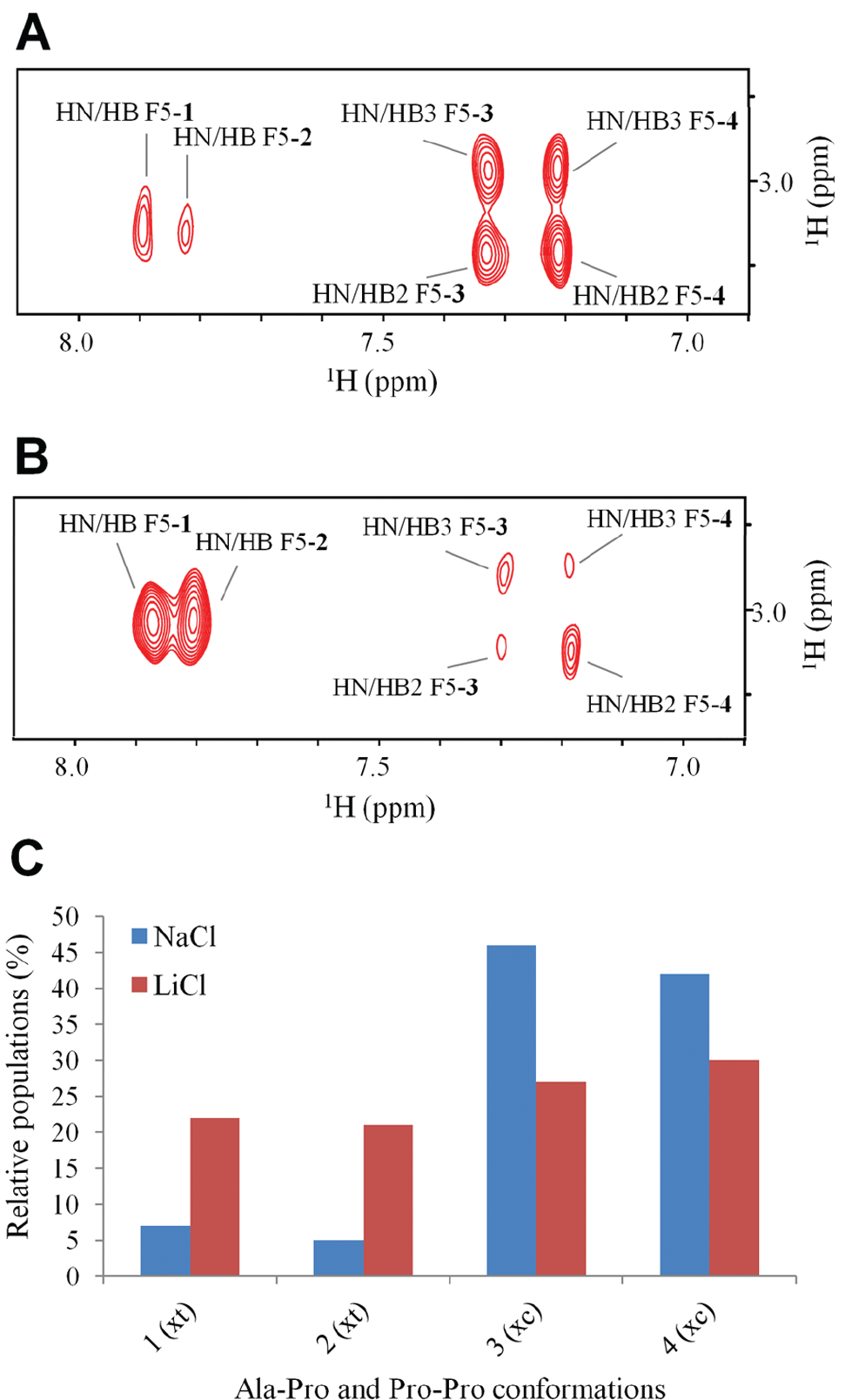
NMR structural characterization of the EPP substrate. Expansion of the HN-aliphatic correlation region of the 2D [1H, 1H] TOCSY spectra of the peptide in phosphate buffer containing NaCl 150 mM (
Due to the low number of signals in both the NOESY and ROESY spectra, there was some ambiguity in assigning the trans-cis states of the two X-Pro bonds from the NOE patterns. Nonetheless, considerations on the influence of proximal and distal proline isomerization on the chemical shifts were used to gain insight into the proline isomerization in the peptide. 15 Specifically, since the F5 HN was separated from the P3-P4 peptide bond by one covalent bond, greater chemical shift changes (Δδ) were expected following isomerization of the proximal proline (i.e., P4), while minor changes were expected upon distal proline isomerization (i.e., P3).
On this basis, the two F5 HNs at 7.88 and 7.82 ppm (conformers 1 and 2, respectively) and those at 7.33 and 7.21 ppm (conformers 3 and 4, respectively), which showed small differences of chemical shift (ΔδHN1–2 = 0.06 ppm and ΔδHN3–4 = 0.12 ppm), result from the isomerization of the distal A2-P3 bond. On the other hand, the two sets of peaks (1–2 and 3–4), showing larger differences for the HN chemical shifts (ΔδHNav1–2/3–4 = 0.58 ppm), arise from the isomerization of the proximal P3-P4 peptide bond (
Fig. 3A
,
B
and
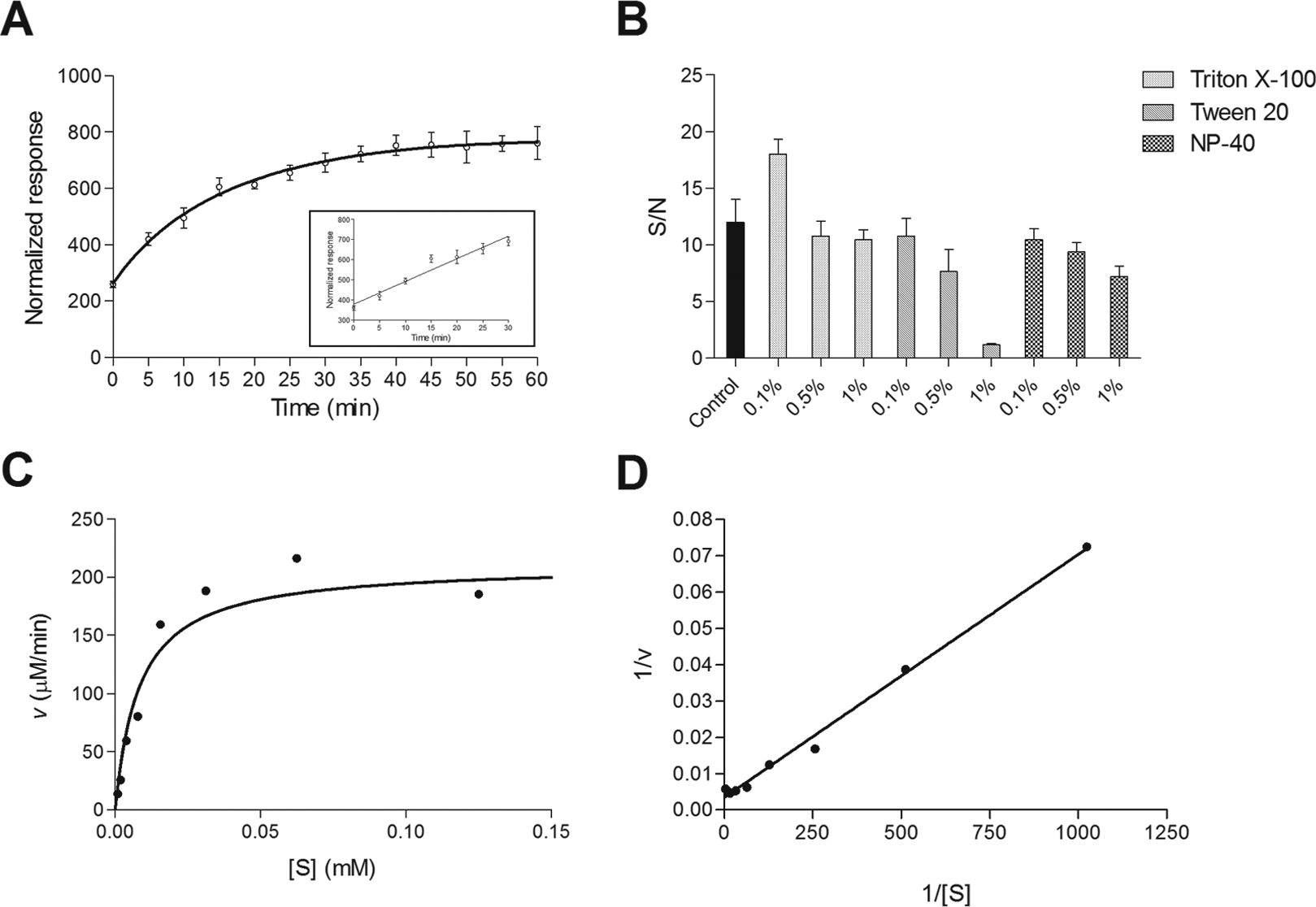
Assay setup. (
Taking into account that the P3-P4 peptide bond in cis conformation seems to be more favored in NaCl than in LiCl, as indicated by the chymotrypsin-coupled assay, conformers 1 and 2 reasonably have the P3-P4 peptide bond in trans conformation (conformers xt). Conversely, conformers 3 and 4 should correspond to the P3-P4 peptide bond in cis conformation (conformers xc). In this way, the cis P3-P4 population in NaCl is 88%, whereas in LiCl it is 57%. Moreover, the A2-P3 bond appears to have the same cis and trans contents of about 50% in both buffers ( Fig. 3C ). It is worth noting that in the case of conformers 3 and 4, the F5 HN chemical shift is upfielded and the two germinal Hβs appear as distinct signals, exhibiting their magnetic nonequivalence and suggesting a more restricted rotation of the F5 ring. Moreover, the four cis-trans states also gave rise to multiple spin systems for the aliphatic protons of the proline residues. Although the peaks’ assignment result was intricate, we could observe a proline Hα proton highly upfielded (3.77 ppm), likely due to the close proximity of the P3-F5 side chains, as found in some proteins and peptide containing the segment P-P-F, 12 which gives rise to the ring current effect on Hα protons of the first proline. Closely interacting P3-F5 side chains are supposed to be at the basis of the observed stability of the proline–proline cis conformers in the segment P-P-F, when present in a short peptide, devoid of any tertiary interaction.
Development of a FRET Assay
In order to develop a FRET assay, the selected EPP substrate was endowed with the FRET pair EDANS–Dabcyl.16,17
EDANS–Dabcyl is a widely used donor–quencher pair. The optimal absorbance and emission wavelengths of EDANS are λabs = 336 nm and λem = 490 nm, respectively, and for Dabcyl, the maximum absorbance wavelength is λabs = 472 nm, which, to a large extent, overlaps with the emission spectrum of EDANS. When the two fluorophores are in the range 10–100 Å, the energy emitted by EDANS is quenched by Dabcyl. The intact molecule is thereby internally quenched, while EDANS fluorescence is readily restored upon protease cleavage within the peptide chain and the intensity change can be detected continuously and directly.16,17
The substrate Ac-EK(Dabcyl)PPFAE(EDANS)KA-NH2 was efficiently synthesized, purified with a high yield (~40%), and characterized by mass spectrometry (
The incubation of recombinant CypA at room temperature with the substrate resulted in the specific enzymatic cleavage by chymotrypsin and a time-dependent increase of fluorescence intensity (λex/λem = 340 nm/510 nm) that was linearly related to the extent of CypA-mediated substrate isomerization.
Assay conditions were standardized manually and subsequently validated in an HTS format using a liquid handler and 384-well black plates. In particular, we investigated the influence of substrate and chymotrypsin concentration on the kinetics of EDANS release from the substrate in the absence of the isomerase. Then, time-course experiments were optimized using four substrate concentrations (15, 25, 50, and 100 ng/μL) and four different substrate/chymotrypsin ratios (1:1, 1:5, 1:10, and 1:20 w/w) in a total volume of 50 μL in PBS 1× buffer at pH 7.4. PBS 1× was chosen as reaction buffer because of the higher stability of CypA over time in this buffer, as assessed by determining protein concentration at each step of protein preparation.
As expected, a higher fluorescence emission was generated at increasing substrate concentrations and substrate/chymotrypsin ratios (
We next optimized the concentration of CypA. As shown in
Km and Kcat Determination
The enzyme reaction optimized followed the Michaelis–Menten kinetics with the reaction rate (v) increasing with substrate concentration (S) ( Fig. 3C ). A first-order rate kinetic was observed and a double reciprocal Lineweaver–Burke plot of 1/v against 1/[S] gave values of Kcat/Km of 2.5 × 107 M–1 s–1 at room temperature in physiological conditions (PBS 1×, pH 7.4) ( Fig. 3D ). The kinetic constant here determined is in agreement with the value reported in literature (1.4 × 107 M–1 s–1), calculated with the chymotrypsin-coupled PPIase assay, using the recombinant wild-type CypA and the substrate N-succinyl-AAPF-pNA dissolved in HEPES 35 mM buffer (pH 8.0) and 66% (vol/vol) aqueous DMSO at 10 °C. 18
Assay Procedure Automation and Validation
The assay procedure described for the manual experiments was next automated using an integrated platform comprising an 8-channel Starlet station for liquid handling, a robotic arm, and a multiwell plate reader (EnSpire Multimode Plate Readers, PerkinElmer). The assay was also performed in the presence of increasing amounts of DMSO (1%, 2%, and 3%) to simulate the experimental conditions, whereby compounds from stock solutions in this solvent are submitted to screening. Optimal assay performances were evaluated in terms of S/N and Z′ factor values.
As can be seen in Figure 4A , excellent S/N values of 11.6, 11.1, and 10.9 were determined for the assays without DMSO and in the presence of 1% and 2% solvent, respectively; the S/N value decreased to 9.1 at 3% DMSO. The Z′ factor values determined on 32 points for each condition tested (0 ÷ 3% DMSO) ( Fig. 4B ) were 0.80, 0.75, 0.68, and 0.56 for 0%, 1%, 2%, and 3% of solvent, respectively. Altogether, data suggest that Z′ factor and S/N values are still sufficiently high for using the assay in screening campaigns at the highest solvent concentration tested.
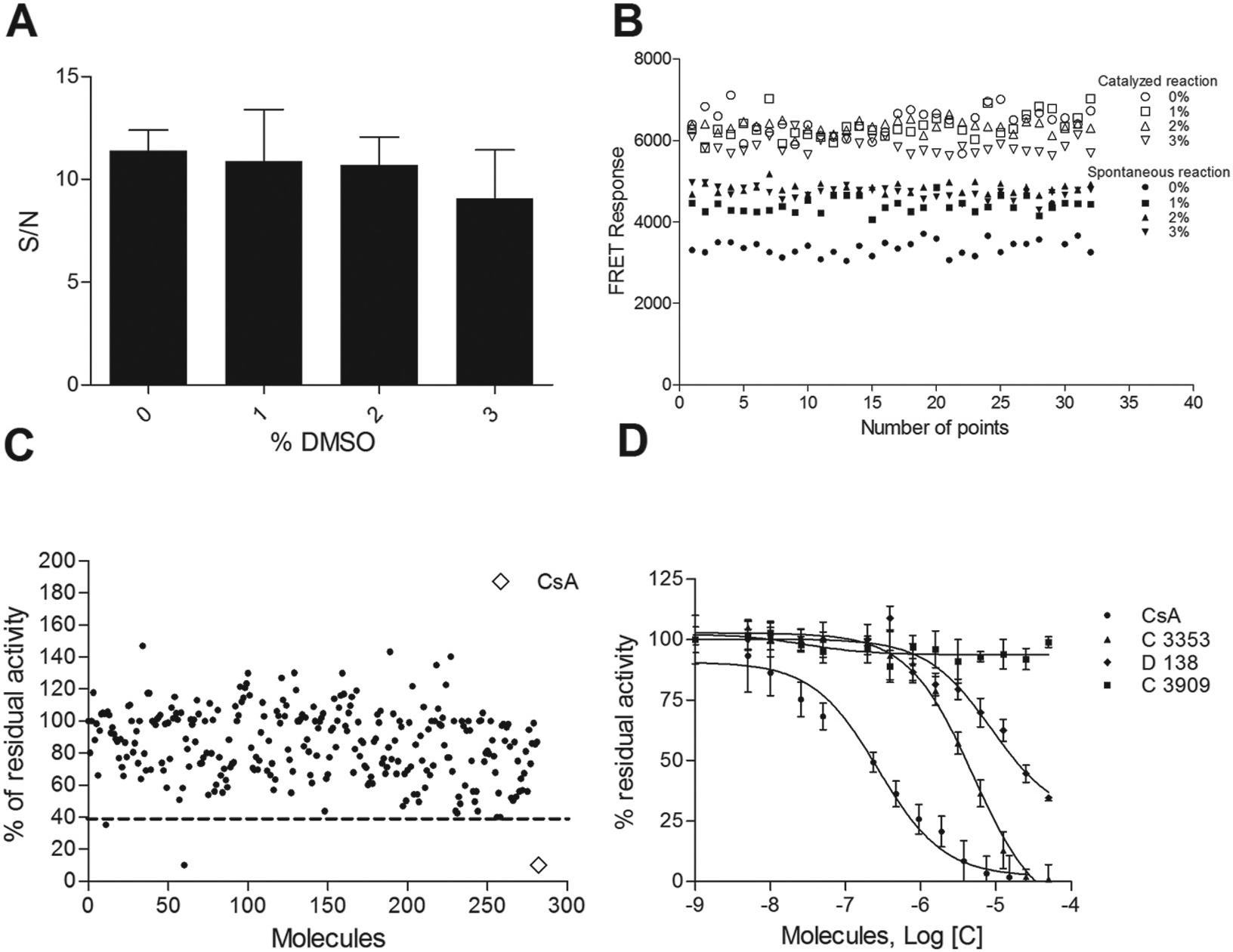
Evaluation of FRET assay performance. Thirty-two wells of a 384-well plate were used to determine the suitability of the assay for HTS in terms of S/N (
In order to estimate the assay suitability for HTS, a pilot screen against the LOPAC library (n = 1280 compounds) was performed. LOPAC compounds were tested at 50 μM in quadruplicate in PBS 1× at pH 7.4, supplemented with DMSO 1%. From a set of 292 compounds (
Fig. 4C
), three were selected for further characterization. They were N2-(cis-2-aminocyclohexyl)-N6-(3-chlorophenyl)-9-ethyl-9H-purine-2,6-diamine hydrochloride (LOPAC code C3353), 5,7-dichlorokynurenic acid (LOPAC code D138), and as expected, the CsA, present within the set. D-Cycloserine (LOPAC code C3909), instead, was used in the next assays as negative control. The testing was performed in a 14-point dose–response format in quadruplicate at a 1:2 serial dilution (see
Molecular Docking Studies of C3353 and D138 Molecules
Molecular docking simulations were carried out to investigate the binding modes of the novel inhibitors identified in the HTS and with the aim to interpret the experimental binding data. To identify which docking protocol and which protein conformation were more suitable for our compounds, we performed a self-docking benchmark based on the CypA complexes available in the protein data bank (PDB). We selected four complexes in which a unique binding mode was present (PDB codes: 3RDD, 4N1N, 4N1Q, and 4N1R). In the benchmark performed with DockBench software,
10
we compared the ability of 17 different protocols in reproducing the four complex conformations. Except for 4N1N, all the complex geometries were nicely reproduced by several protocols (
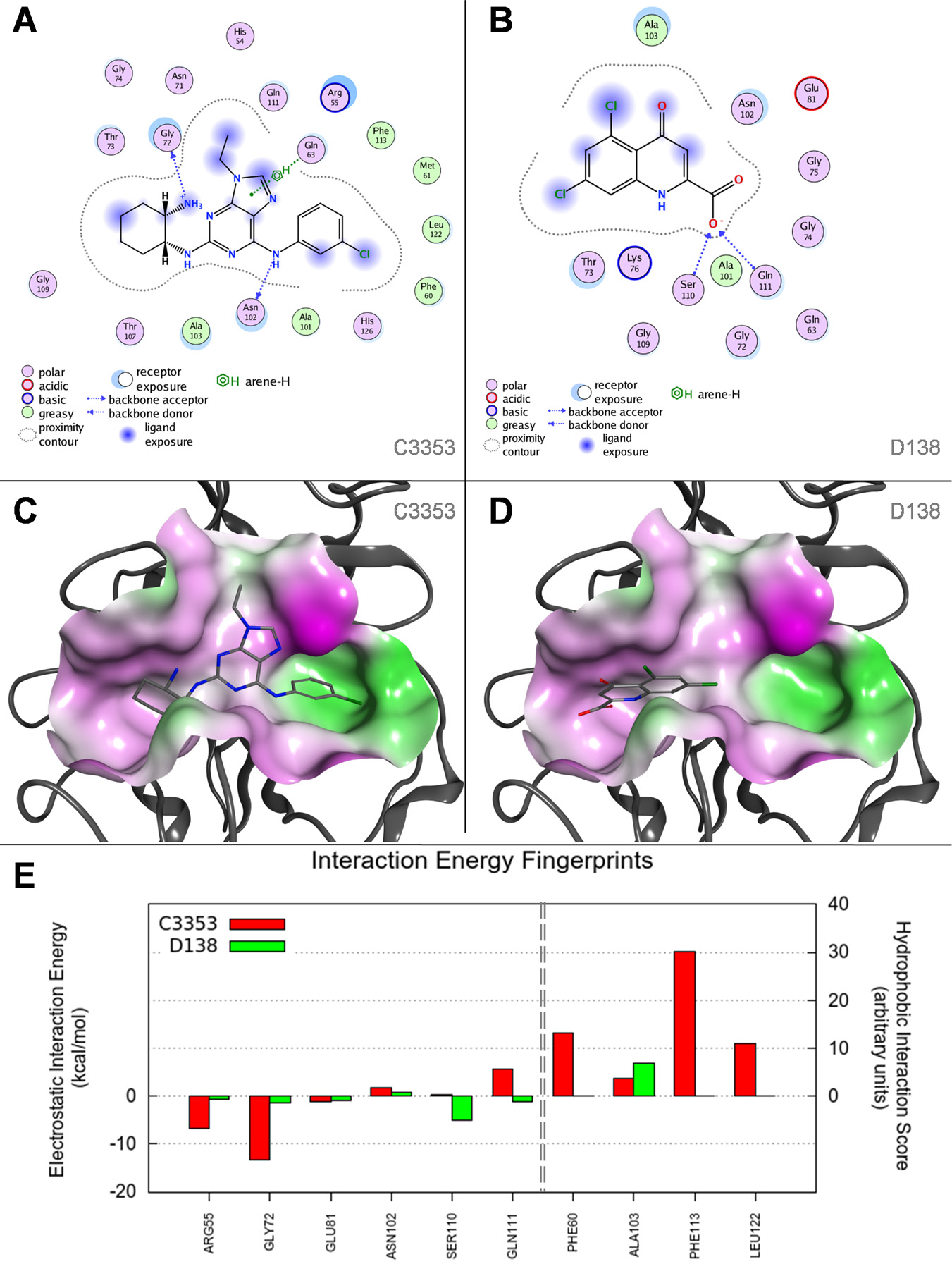
Molecular docking studies of C3353 and D138 compounds. In
The 5,7-dichlorokynurenic acid (D138) showed a clear propensity for the hydrophilic groove of the CypA catalytic site. The carboxylic moiety establishes two hydrogen bonds with S110 and Q111, while the condensed ring lies in front of the methyl group of A103 ( Fig. 5B , D ). The molecular basis explaining the different abilities to inhibit the CypA enzymatic activity can be ascribed to the different sizes of the two compounds and to the number of interactions they are able to establish with the catalytic site. In particular, C3353 is able to occupy most of the catalytic cleft, and the final score returned by Goldscore function was clearly higher than that for the molecule D138, 76.8 and 48.6, respectively. These observations are more evident when the contributions to the estimated interaction energy of the two compounds are plotted in a per-residue histogram, as reported in Figure 5E . In this graph, the electrostatic interaction energy and a hydrophobic score are computed for each residue of the binding site and the most relevant ones are reported. While the electrostatic component of the predicted interaction energy is relevant for both ligands, the hydrophobic score is notably weaker in the D138 molecule that interacts only with A103.
Discussion
The PPIases represent a large class of biological proteins, such as cyclophilins, with broad physiological functions. Alteration of their activity may lead to several pathological conditions, making them valid targets for therapeutic intervention. 19 For example, cyclophilin inhibitors have shown promise as replication inhibitors of several RNA viruses, including HIV, hepatitis C virus (HCV), severe acute respiratory syndrome (SARS) corona, and influenza virus. Nonimmunosuppressive cyclosporin derivatives have reached clinical development for the treatment of hepatitis C infection. 20 Moreover, formulations of cyclosporin have also shown some promise in new therapeutic applications, for example, in traumatic brain injury, 21 muscular dystrophy, 22 respiratory diseases, 23 cardiovascular diseases, 24 and Alzheimer’s disease. 25 Much effort is spent on selecting specific PPIase modulators for therapeutic approaches, and in this framework, the discovery of novel inhibitors may largely benefit from suitable assays for HTS.
The biochemical reaction catalyzed by PPIases interconverts cis and trans isomers of a proline–peptide bond. However, this reaction also occurs spontaneously in the absence of catalysis within a few minutes, especially in short synthetic peptide substrates. Inhibitor screens therefore have to distinguish between the catalyzed and spontaneous reactions, which are separated by a time window of a few minutes. Such screens have been developed, but they require low temperatures and elaborated experimental protocols in the presence of organic solvent; they therefore are invariably of low throughput and are unsuitable for large-scale screens.
Here, we have developed and optimized a simple and robust homogenous fluorescent HTS assay for the discovery of cyclophilin inhibitors in 384-well plates, using a new FRET-based substrate probe useful for chymotrypsin-coupled isomerase assays. The probe, easily prepared by a straightforward solid-phase approach in a highly soluble form, contains the well-known FRET pair Dabcyl–EDANS.16,17 The donor (EDANS) is separated from the quencher (Dabcyl) by a short peptide linker containing a modified chymotrypsin cleavage site (Pro-Phe) and residues of glutamic acids and lysine at the N- and C-termini, respectively, to potentially increase the cis conformation in solution. Noteworthy, the NMR structural characterization of the probe showed that the cis conformation of the substrate was highly favored, as hypothesized in the design. Treatment of the substrate with recombinant CypA in the absence of organic solvents promoted the rapid proteolytic cleavage by chymotrypsin and an increase of fluorescence intensity that is linearly related to the extent of CypA-mediated substrate isomerization.
Technical parameters of the HTS assay (Z′ factor, S/N, IC50 values of pharmacological control) were optimized; therefore, the test appears to be ready for large-scale screenings. As an exemplary screening, we used here the commercial LOPAC1280 library. From such screening, some selective CypA inhibitors were identified, including the well-known CsA, present within the library as a positive control. Among the other compounds showing activity, two were tested in dose–response experiments. The purine-based compound C3353, known as a Cdk inhibitor selective for cyclin-dependent kinase-1 (Cdk1), 26 showed a 6 μM IC50 inhibition, which is a good starting point for developing new and more selective inhibitors. To gain insights into the structure–activity relationship of the two compounds, they were subjected to a comparative computational study that confirmed the predilection of the ligands for the CypA active site. Moreover, the different IC50 values detected were realistically explained by the higher number of interactions established by C3353 with the hydrophobic groove present in the catalytic site.
In conclusion, we have set up an HTS screening assay for the selection of CypA cis-trans isomerase activity inhibitors. The assay is based on the use of a new fluorescent peptide substrate designed in order to have a high proportion of cis conformers and to allow the detection of fluorescence intensity increase or decrease, depending on enzyme activation or inhibition. This very robust assay has been optimized on 384-well plates and can be used in screening campaigns of large compound libraries.
Footnotes
Acknowledgements
The authors thank L. Zona (IBB-CNR, Napoli, Italy) for NMR technical support.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The project was partially funded by Progetto PON01_01434 - REACT and Progetto N PON 03 PE_00155_2 - OPTIMA to MR. The computational work of MS has been supported by University of Padova, Italy (N° 79122, Progetto Giovani Studiosi 2013 - Area 4).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

