Abstract
Catechol-O-methyltransferase (COMT) plays an important role in the deactivation of catecholamine neurotransmitters and hormones. Inhibitors of COMT, such as tolcapone and entacapone, are used clinically in the treatment of Parkinson’s disease. Discovery of novel inhibitors has been hampered by a lack of suitable assays for high-throughput screening (HTS). Although assays using esculetin have been developed, these are affected by fluorescence, a common property of catechol-type compounds. We have therefore evaluated a new homogenous time-resolved fluorescence (HTRF)–based assay from CisBio (Codolet, France), which measures the production of S-adenosyl-L-homocysteine (SAH). The assay has been run in both HTS and medium-throughput screening (MTS) modes. The assay was established using membranes expressing human membrane-bound COMT and was optimized for protein and time to give an acceptable signal window, good potency for tolcapone, and a high degree of translation between data in fluorescence ratio and data in terms of [SAH] produced. pIC50 values for the hits from the HTS mode were determined in the MTS mode. The assay also proved suitable for kinetic studies such as Km,app determination.
Introduction
Catechol-O-methyltransferase (COMT) is a catabolic enzyme that uses S-adenosyl methionine to transfer a methyl group to a variety of substrates. COMT plays an important role in the deactivation of catecholamine neurotransmitters, as well as influencing the metabolism of other substrates, such as catechol estrogens and dietary polyphenols. 1 COMT exists in two forms: a soluble form (S-COMT) and a membrane-bound form (MB-COMT). The membrane-bound form has an additional 50 amino acids at the N-terminus for membrane anchoring. 2 Even though the catalytic sites are nearly identical, the two forms differ in their kinetic properties for substrates such as dopamine and S-adenosyl methionine (SAM). 3 The COMT enzyme operates with an ordered mechanism where SAM binds first, followed by magnesium and catechol substrate. 2 The sources of COMT used in studies have been from tissues and recombinant systems (Escherichia coli and Sf9 insect cell). Each has its own set of optimal substrates and kinetic parameters, and this must be taken into account when comparing activity data. 4
COMT’s role in dopamine metabolism in the brain has made this enzyme an area of great interest. 5 This role includes its ability to metabolize L-DOPA, a precursor to dopamine that is given to Parkinson’s disease patients to boost dopamine levels in the brain since dopamine itself cannot cross the blood–brain barrier but L-DOPA can. Even though L-DOPA is capable of crossing the barrier, when exogenously administered, only about 1% of it makes it into the brain, and high doses have peripheral side effects. 6 Coadministration of inhibitors of L-DOPA metabolism has been shown to reduce the overall dose of L-DOPA and prolong its effects, and two COMT inhibitors (tolcapone and entacapone) have been developed for this purpose. 7 Both compounds are potent COMT inhibitors but contain a potentially problematic nitrocatechol pharmacophore. 8 Tolcapone has greater bioavailability than entacapone, as well as a longer half-life, and both compounds have the desired effect of reducing the required dose of L-DOPA. Tolcapone, however, has a black box warning for severe liver toxicity. 8 There have been attempts to improve entacapone since it does not have the liver toxicity of tolcapone, but no new agent has yet been approved for clinical use. 5 We have endeavored to discover COMT inhibitors lacking the nitrocatechol moiety, which may be responsible for some of these undesirable effects.
A fluorescent assay had been developed based on the principle that COMT interacts with esculetin and, in the presence of magnesium and SAM, transfers the methyl group to create S-adenosyl homocysteine (SAH) and scopoletin, a fluorescent product. Unfortunately, the fluorescence wavelengths required to read the assay (355 nm excitation, 460 nm emission) are within the UV range and can cause background problems for some catechol compounds. 4
We encountered some success in using a fluorescence polarization (FP)–based assay that measures the reaction product SAH with a SAH-specific antibody in order to alleviate some of the fluorescence issues. 9 Some of the test compounds were still found to fluoresce in the wavelengths used to read the assay (480 nm excitation, 535 nm emission). An additional limitation to this assay was the use of an antibody and its specificity for SAH, as opposed to SAM. The antibody is procured as part of a large kit (IMX Homocysteine Reagent Pack, Abbott Laboratories, Chicago, IL), its stability was not fully determined for our purposes, and maintaining a sufficient assay signal window was found to be challenging. The assay was modified in order to increase the wavelength even further, 10 but it still had some fluorescence issues. It was necessary to find a way to detect activity of the enzyme without the complication of a fluorescence background while still maintaining a good signal window.
To this end, we evaluated a new homogenous time-resolved fluorescence (HTRF) assay from Cisbio, which measures production of SAH. The assay principle is as follows: The HTRF assay is a competitive binding assay where SAH, produced by demethylation of SAM, competes for the binding of a labeled SAH (SAH-red) to an antibody. The antibody is tagged with Lumi4-tb cryptate that acts as the donor. When the SAH-red acceptor is bound and the complex excited at 337 nm, there is energy transfer resulting in emission from the SAH-red acceptor at 650 nm. This assay allowed us to complete not only a high-throughput screen (HTS), but also a medium-throughput screen (MTS) for pIC50 values, in addition to mechanistic data, such as Ki values.
Materials and Methods
Assay Procedure
An HTRF kit purchased from CisBio was used: EPIgeneous methyltransferase assay, 10,000 tests, cat. 62SAHPEH. Assay reagents were kept frozen at −80 °C, thawed, and used immediately. Stock solutions were made and dispensed into single-use aliquots. No stability information is available longer than 1 day after freeze–thaw.
HTS Format
All reagents were dispensed using a Tempest liquid handler manufactured by Formulatrix (Bedford, MA) and were made fresh each day. The dopamine solutions were kept on ice until use.
One microliter of compound at 100 µM in 5% DMSO final (from prediluted plate: 2 µL at 2 mM 100% DMSO into 38 µL assay buffer) was added to 2 µL MB-COMT 0.1 µg/well and allowed to be incubated at room temperature (RT) for 30 min. Next, 2 µL [SAM 5 µM final + dopamine 10 µM final] was added to start the reaction and allowed to go for an additional 30 min at RT.
As a no-activity control, a 10 µM concentration of tolcapone and a reaction without enzyme were evaluated. Both reactions resulted in the same output, so 10 µM solution of tolcapone was selected as the contro, and this was used for the Z′ calculation along with DMSO control.
One microliter of stop buffer was then added, and 1 µL of DB1 buffer was added. This was mixed and allowed to go to 30 min at RT. Lastly, 3 µL [SAH-red + anti SAH-cryptate Ab] was added and allowed to incubate at RT for 60 min. The plate was then read on the Envision plate reader per kit instructions.
MTS Format
Compounds were diluted in two steps, the first in 100% DMSO (1:32 dilution of 10 mM liquid stock) and then by 20× into assay buffer (5 µL into 95 µL assay buffer using a mosquito dispenser, TTP LabTech, Cambridge, MA), resulting in a final DMSO concentration of 1% (1 µL in 5 µL reaction). Two microliters of a mixture of 10 ng (final concentration per well) of lysate preparation of MB-COMT, premixed with SAM (final concentration of 20 µM, 2 mM stock solution provided by kit), was added to the plate and allowed to incubate with compound for 30 min at 37 °C. Freshly prepared dopamine solution (Sigma H8507) was added to give a reaction concentration of 4 µM, and the reaction was allowed to proceed for 40 min at 37 °C. The DB1, cryptate antibody, and detector were added (no stop buffer used in the MTS format) and incubated according to the kit insert (50× dilution of antibody and 8× dilution of SAH-red based on the 20 µM SAM in the reaction) using a Formulatrix Tempest liquid handler. The results were read on a Tecan Infinite Pro or PerkinElmer Envision and analyzed using Prism GraphPad software.
Standard curves
A standard curve was run on every plate. Using the HTRF ratio (665 nM/620 nM × 104), a standard curve is determined according to the manufacturer’s instructions using a mixture of SAM and SAH, with the final concentration of SAM being equivalent to that used in the enzyme reaction mixture.
Results
The assay was initially optimized on time and enzyme level to be in the linear range of the SAH standard curve and for tolcapone potency. In order to determine if the HTRF assay was capable of handling a high-throughput compound screen, its robustness was evaluated. It was also evaluated on an MTS level in anticipation of later studies. To that end, 159,003 compounds were tested in single-point format, and 413 showed >50% inhibition and 1323 showed >30% inhibition. Of those compounds, 1317 were retested in MTS format. A 3 SD cutoff using DMSO controls was determined to be 23% ( Fig. 1 ). The signal window was reproducible with a 2.5- to 3-fold difference and the Z′ (calculated in GraphPad Prism by standard methods) was 0.72 ( Fig. 1 ). Tolcapone was used as the benchmark for maximum signal window and consistency of signal, to which other compounds in the screen were compared.
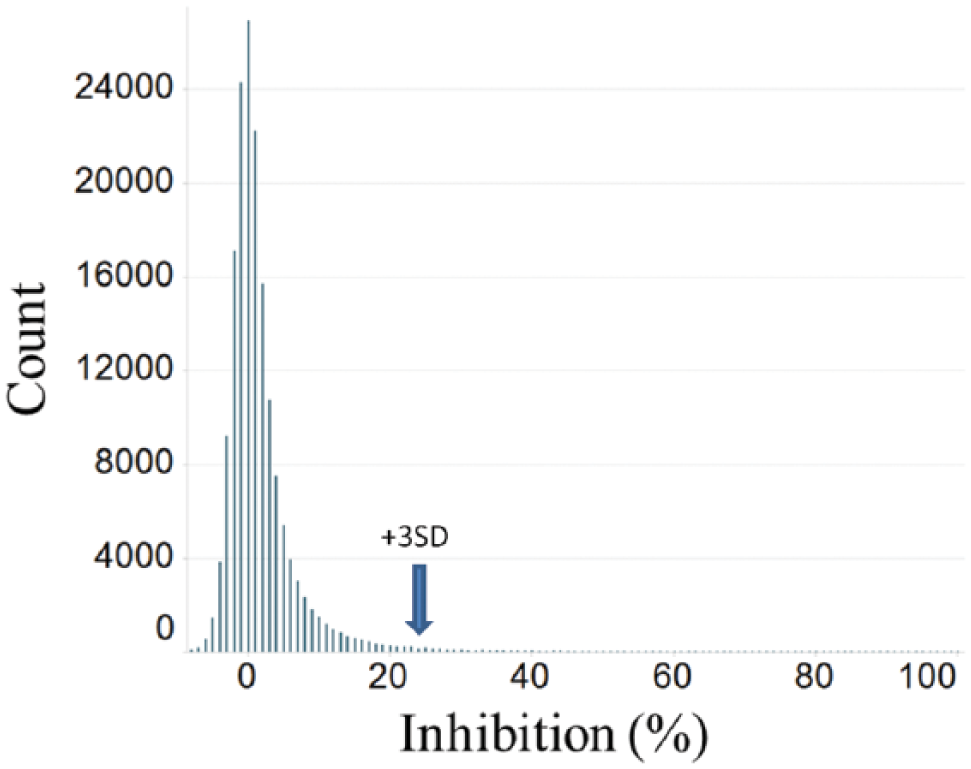
HTS assay results histogram. Histogram of the HTS assay results for 159,003 compounds. The 3 SD cutoff was determined to be 23%.
Km,app Values
We determined Km,app values for the MB-COMT lysate for two substrates: dopamine (DA) and S-adenosyl methionine (SAM). In order to determine the Km,app for DA, the reaction was run in MTS format on a 384-well plate. The reaction time, DA concentration, and enzyme concentration were varied. We used a high concentration of SAM (20 µM) and varied the concentration of DA around its literature Km value.2,11 The resulting data were graphed as a function of DA concentration ( Fig. 2A ) and of time ( Fig. 2B ), and it was calculated that the shared Km,app was 4 µM for DA. The shared Km,app was calculated in GraphPad Prism by plotting all of the curves from a single plate and then applying the shared Km,app constraint to the calculation. The DA Km,app value was then used as a constant in determining the Km,app value for SAM. The amount of enzyme was kept constant in the presence of a high concentration of dopamine (20 µM), and the concentration of SAM and reaction time were varied ( Fig. 2C ). The shared Km was determined to be 20 µM. The Km,app values for both substrates were comparable to those reported in the literature,2,11 so compound screening was initiated. The time data allowed for determination of the optimal reaction length for the assay.
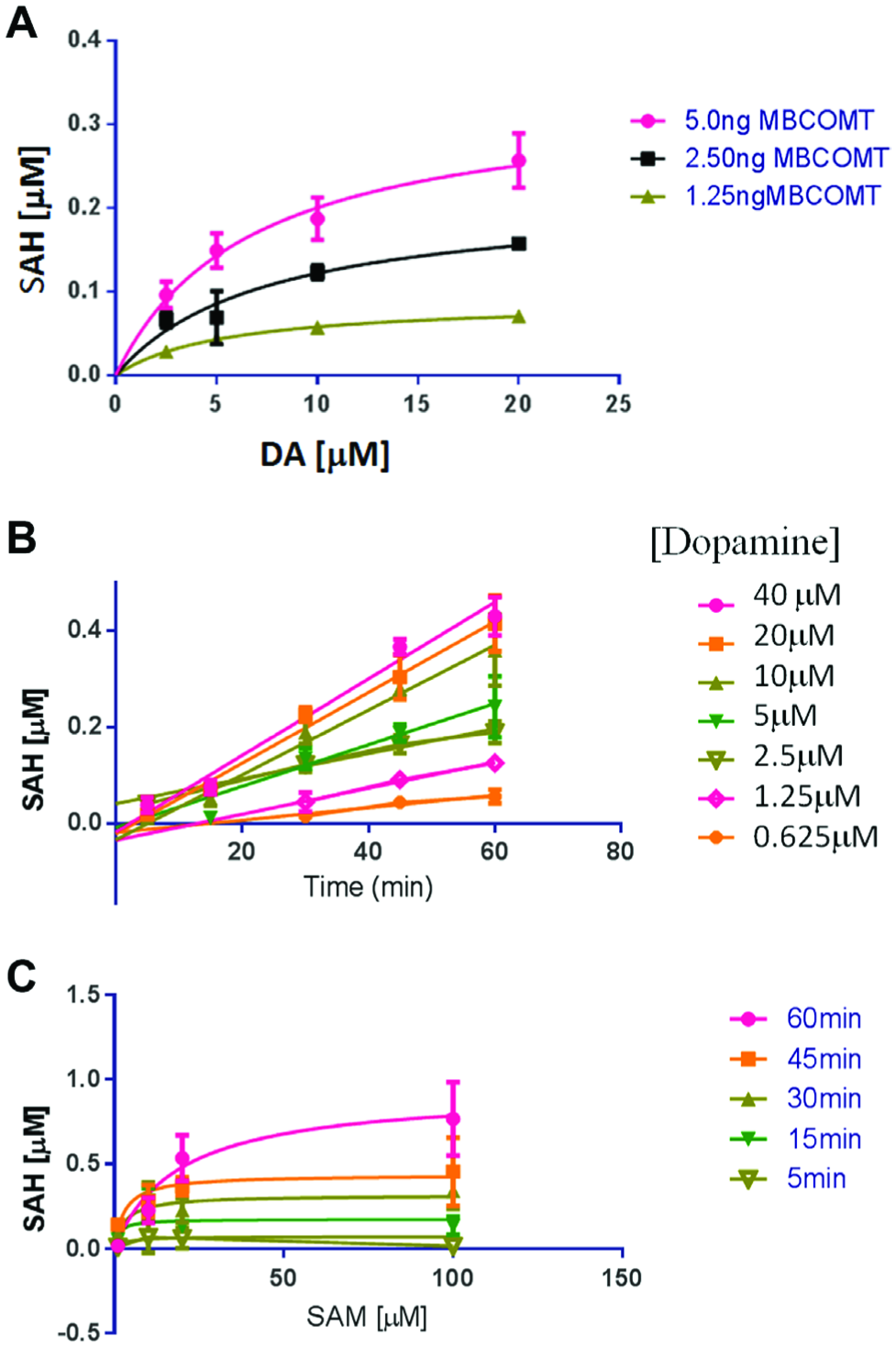
Km,app value for dopamine.
Inhibition Detection
Both formats of the assay were used to screen for inhibitors and for the pIC50 of tolcapone. Tolcapone is known to be a potent inhibitor of COMT 12 and, as such, was used as a benchmark for assay performance. The assays were run under substrate Km,app conditions; the compound was allowed to preincubate with the enzyme/SAM mixture for 30 min prior to initiating the reaction with dopamine. The MTS assay was designed to be as physiological as possible, so it was run at 37 °C as opposed to the HTS, which was run at room temperature. For the HTS assay, room temperature was necessary so as not to compromise throughput and automation. These changes caused the substrate and enzyme concentrations to be different in the HTS versus MTS assays as well.
In the HTS format, tolcapone displayed an IC50 of 18 nM ± 3 (mean ± SD, n = 4, pIC50 7.74) and gave an assay window ratio of 2.4 ( Fig. 3B ). In the MTS format, tolcapone was more potent with an IC50 of 1.9 nM ± 1 (n = 4, pIC50 8.7), and a similar assay window was observed ( Fig. 3A ). All were run with a 10-point dose–response curve starting at 32 µM. Inhibition of subsequently tested compounds was compared to tolcapone at 10 µM, which was set as the 100% inhibition value.
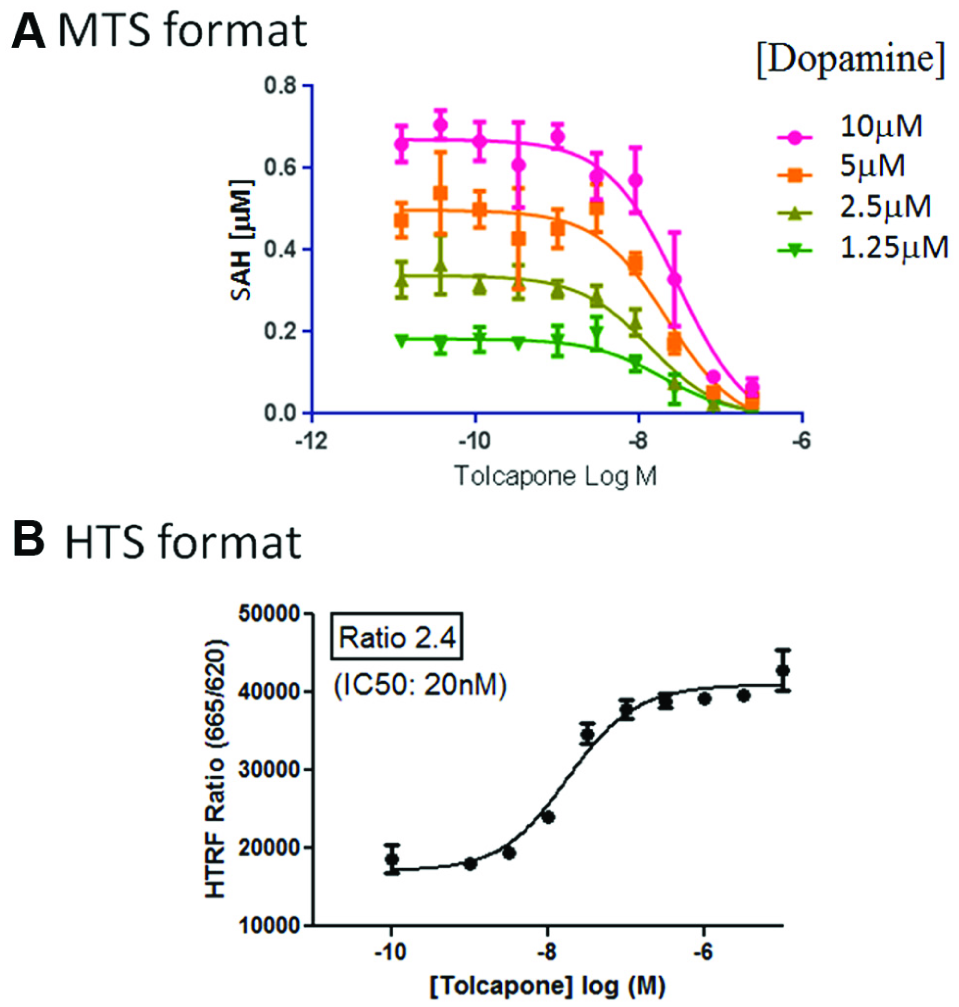
Assay detects potent inhibitors. Both formats of the assay were used to determine the IC50 for tolcapone. Tolcapone is known to be a potent inhibitor of COMT and, as such, was used as the benchmark. The assays were run under Km,app substrate conditions, and the compound was allowed to preincubate with the MB-COMT/SAM mixture for 30 min prior to initiating the reaction with dopamine.
Ki,app Values
Being able to determine the pIC50 at even low concentrations of DA allowed us to proceed with the next step, determining the Ki,app value. The reaction was performed with the standard amount of enzyme, SAM, and for the same length of time as screening. Five concentrations of compound were chosen based upon the pIC50 values. For tolcapone, the values chosen were 50, 25, 10, 5, and 1 nM. The compound was allowed to preincubate with the enzyme/SAM mixture for 30 min prior to starting the reaction with various amounts of DA. The highest concentration of DA was chosen to be 10 times higher than the Km,app value. The reaction time was kept as close to that of the screening protocol as possible to ensure that the output was still an initial rate of the reaction. Using these data, Prism GraphPad software was used to determine the Ki,app value and the possible competitive/noncompetitive binding mode of the compound. In this instance ( Fig. 4 ), we determined that the pKi,app value for tolcapone in the assay is 9 (1 nM), and that it is a competitive binder. This is in good agreement with the literature reports for tolcapone.2,13
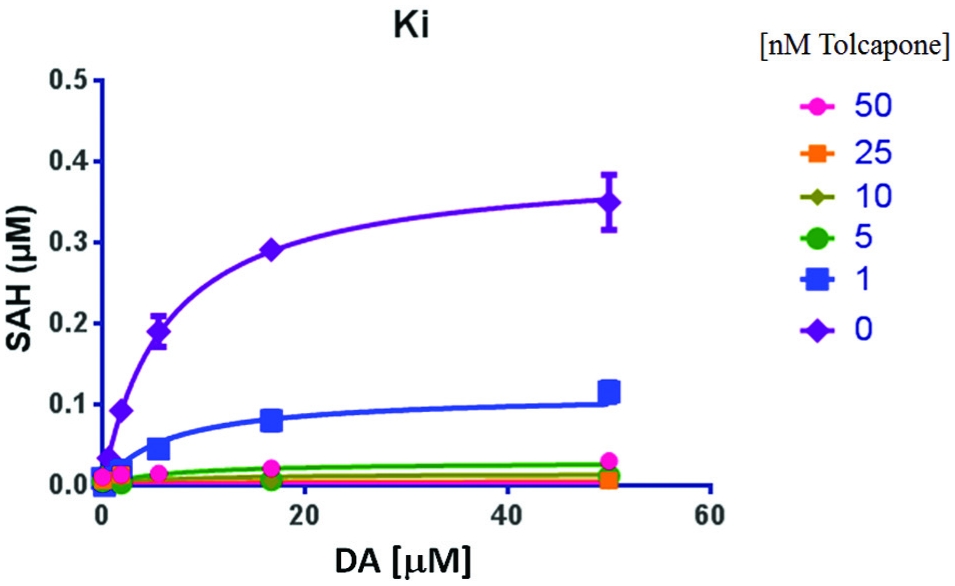
Ki value determination for tolcapone. The reaction was performed with 10 ng of enzyme, 20 µM SAM, and varying micromolar amounts of DA. The tolcapone and the MB-COMT/SAM mixture were allowed to preincubate for 30 min at 37 °C, and then the DA was added to start the reaction. It was allowed to proceed for 40 min. The tolcapone concentrations are nanomolar. A prism was used to determine the Ki via a nonlinear curve-fitting analysis (GraphPad Prism, f test compared to noncompetitive and uncompetitive models).
Discussion
We successfully implemented a new assay format for the use of high- and medium-throughput screening of COMT inhibitors. Our results show that the assay is robust (Z′ = 0.7), has a broad signal window, and it was used to determine the enzymatic properties as well as perform some initial mechanism of action studies for our compounds.
This assay compares favorably to the others previously used in the literature in that it poses less fluorescence background issues, although we have no data from the HTS screen that only examines donor quenching. However, in principle, the HTRF format minimizes (but does not eliminate) this possibility. The assay was also sensitive enough to detect as little as a 0.03 µM SAH output. This was not possible when attempting to use the esculetin or the FP formats described in the literature (data not shown). When testing these assays, we found that many known catechol and noncatechol COMT inhibitors interfered with the assay readout and gave values above those of the background and zero enzyme reaction wells (data not shown). A modification of the FP assay was used by a group at Merck 10 by adjusting the tracer in the assay to the far-red region, but that still did not alleviate the issues of time and antibody sensitivity. The HTRF version of the assay has a longer overall assay run time than the esculetin assay, but shorter than the FP version, as the FP detection reaction requires a significant amount of time.
Unlike the esculetin fluorescent assay, HTRF also allows us to use dopamine (a natural substrate) and can add advantages in picking up novel types of inhibitors, such as allosteric modulators. It also allows for a side-by-side comparison of both isoforms of the enzyme (soluble and membrane bound) where there are differences in substrate affinity, as well as a comparison on selectivity against other methyltransferases. Lastly, this screen has been run in 384-well format and should be amenable to further miniaturization.
We have performed a high-throughput screen on a library of almost 160 K compounds at 20 µM. From this, more than 1300 compounds were selected for confirmation, and along with further informatics filtering (such as structure and molecular weight), approximately 400 compounds were selected for IC50 determination on MB-COMT and a closely related methyltransferase, phenylethanolamine-N-methyltransferase (PNMT), using the same assay format. Although HTRF assays are less sensitive to interference from fluorescent compounds, they can be affected by quenching and compound aggregation. However, we found only 19 compounds that displayed similar activity in both assays. No follow-up was done on these hits to determine if they were assay artifacts or nonselective methyltransferase inhibitors. The overall result of the screening effort identified more than 30 compounds with MB-COMT IC50 < 1 µM and which were essentially inactive on PNMT.
In the future, it may be possible to use this assay to test for enzymatic activity in tissue or blood samples before and after treatment with compound, although it remains to be determined whether the assay is sensitive enough for such ex vivo applications. If this is possible, then it would allow for some of the drug metabolism and pharmacokinetic (DMPK) studies to be performed in the assay instead of via the traditional mass spectrometer.14,15 This could increase the speed at which data are obtained and analyzed for decision making.
In conclusion, we have shown that the SAH HTRF assay can be used to measure the enzymatic properties of MB-COMT and its inhibition. We show that this assay is sensitive and robust. Using this assay, we have run a high-throughput screen that identified inhibitors of MB-COMT. Such compounds may prove useful in both elucidating the role of MB-COMT in the CNS and as novel therapeutic agents in cognitive and movement disorders.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
