Abstract
Gene knockdown by small interfering RNA (siRNA) has been used extensively to investigate the function of genes in targeted and genome-wide studies. One of the primary challenges of siRNA studies of any scale is to achieve sufficient gene knockdown to produce the biological changes that lead to measurable phenotypes. Reverse, lipid-based transfection efficiency minimally requires the optimization of the following parameters: cell number, knockdown duration, siRNA oligonucleotide concentration, type/brand of transfection lipid, and transfection lipid concentration. In this study, we describe a methodology to utilize the flexibility and low-volume range of the Echo acoustic liquid handler to rapidly screen a matrix of transfection conditions. The matrix includes six different transfection lipids from three separate vendors across a broad range of concentrations. Our results validate acoustic liquid transfer for the delivery of siRNAs and transfection reagents. Finally, this methodology is applied to rapidly optimize transfection conditions across many tissue culture cell lines derived from various originating tissues.
Introduction
The knockdown of mRNA transcripts by small interfering RNAs (siRNAs) is a significant tool for probing cell biology for drug discovery and exploratory biological research into gene function.1–3 siRNA carries the advantages of providing fairly rapid and robust gene knockdown in many cell lines with commercially available reagents. In addition, siRNA libraries can be formatted as whole genome or targeted libraries, allowing for flexibility in how much of the genome is interrogated. These advantages have led to the use of siRNA for the discovery and validation of new drug discovery targets. 4 Likewise, we and others have successfully combined siRNA-mediated gene knockdown with compound treatment to investigate compound mechanism of action.5–9
Despite the strengths of siRNA for gene silencing, a challenge that remains for the technology is the optimization of transfection efficiency, which can vary significantly across individual cell lines. Incomplete knockdown can lead to false-negatives and confounding results in some cell lines, whereas other cell lines are generally resistant to transfection. In addition, the transfection process itself might lead to undesired secondary effects in addition to the off-target effects that already arise from siRNA specificity issues. For example, at high concentrations, transfection lipids become toxic—a factor that needs to be considered in experiments in which cell viability is being measured. In addition, fusion of the transfection lipid with the plasma membrane could alter biological responses depending on the nature of the screen. For each cell line, the optimized transfection condition must be determined to maximize knockdown while minimizing off-target effects.10–12 Unfortunately, comprehensive transfection optimization requires optimizing minimally four parameters for each cell line: cell number, transfection lipid, transfection lipid concentration, and duration of knockdown. In addition, plate type and siRNA concentration may also affect transfection efficiency, but in our experience, these parameters do not require reoptimization for each individual cell line.
To assist with the liquid handling required to evaluate multiple lipids at multiple concentrations, we have taken advantage of the versatility and low-volume capabilities of acoustic liquid handling. This technology applies focused sonic energy at the meniscus of the liquid in a source well, resulting in the delivery of a nanoliter-range volume droplet. 13 With repeated pulses, volumes can be delivered from the low-nanoliter to low-microliter range. In our studies, we used the Labcyte Echo, which dispenses multiples of 2.5 nL droplets. This instrument can be used to perform 1:1 plate replications, but more importantly, for this application, the Echo can perform customized liquid transfers in which source well, destination well, and volume transferred are specific to the experimental design. Such versatility makes this platform ideal for optimizing the parameters required for efficient siRNA transfection. Here, we describe our validation of acoustic dispense for the delivery of siRNAs and transfection lipids as well as our application of this approach to the transfection optimization of several tumor cell lines.
Materials and Methods
Cell Lines and Tissue Culture
Cells were incubated at 6% CO2 and at 37 °C in monolayer culture under humidified conditions. Human lung lines NCI-H520 and NCI-H596, pancreatic line BxPC3, breast line BT-20, colon line HCT116, and liver line HEPG2 were sourced from the American Type Culture Collection (ATCC; Manassas, VA). Lung line LUDLU-1 was purchased from the European Collection of Cell Cultures (Salisbury, UK). SMMC-7721, a liver cell line, was sourced from Shanghai Medicilon, Inc. (Shanghai, China). Human lung lines LC-1/sq and LK-2 were obtained from the Japanese Collection of Research Bioresources Cell Bank (Osaka, Japan). NCI-H520, NCI-H596, LUDLU-1, SMMC-7721, and LK-2 were cultured in RPMI supplemented with 10% fetal bovine serum (FBS; ATCC). BxPC3 was cultured with RPMI with 10% FBS (ATCC), 10 mM HEPES, and 10 mM sodium pyruvate. HEPG2 and BT-20 were maintained with minimum essential media, 10% FBS (ATCC), nonessential amino acids, and 10 mM sodium pyruvate. Lung line LC-1/sq was grown in a 1:1 RPMI/Ham’s F12 medium supplemented with 10% FBS (ATCC). The HCT116 colon line was cultured with McCoy’s 5A media with 10% FBS (ATCC). All cell lines tested mycoplasma negative prior to their experimental use. All cell culture media and supplements, except as noted above, were from Life Technologies (Carlsbad, CA).
Nonbiological Testing of Acoustic Transfer
Assay plates with 384 wells (781948, Greiner, Monroe, NC) were filled with 25 µL/well Dulbecco’s phosphate-buffered saline (PBS; Life Technologies) and loaded into the Echo 555 Liquid Handler (Labcyte, Sunnyvale, CA) as the destination plate. The source plate was prepared by diluting a 10 mM fluorescein isothiocyanate (FITC; Sigma-Aldrich, St. Louis, MO) stock into lipid or lipid-buffer solutions (1 mM final concentration). The transfection lipids tested were Lipofectamine RNAiMAX (Life Technologies); HiPerFect (Qiagen, Valencia, CA); DharmaFECT 1, 2, 3, and 4 (GE Healthcare Dharmacon, Lafayette, CO); and X-tremeGENE (Roche Diagnostics, Indianapolis, IN). X-tremeGENE was removed from the study because it was not amenable to acoustic dispense using the aqueous setting on the Echo (data not shown). Lipid-buffer solution was made by diluting the transfection lipids 1:1 with siRNA buffer (GE Healthcare Dharmacon). The FITC solutions were then pipetted into an Echo qualified, low-dead-volume microtiter plate (LP-0220, Labcyte) and centrifuged for 1 min at 1000 rpm to remove bubbles prior to being loaded into the Echo as the source plate. Custom cherry-pick files were used to deliver specific volumes to specific wells depending on the experimental design. The Echo was operated in 384LDV_AQ_B mode for the source plate. Following dispense, fluorescence intensity was measured by a PHERAstar FS plate reader (BMG Labtech, Cary, NC). Volumes in Figure 1A are the actual volumes delivered, whereas volumes in all other figures are theoretical (based on the assumption of 2.5 nL/droplet delivery).
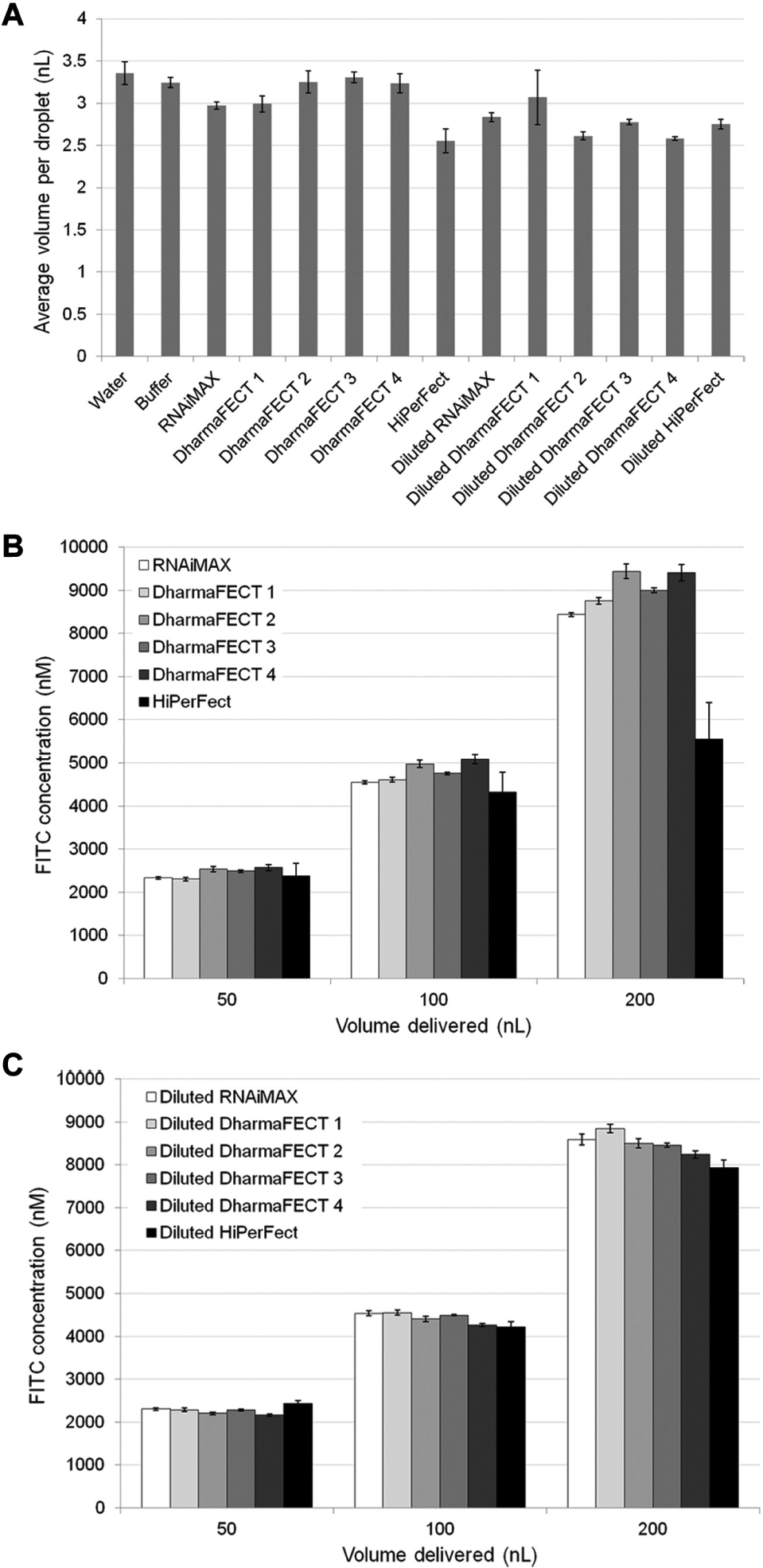
Volumetric validation of transfection reagents delivered by acoustic dispense. (
siRNA Transfection Protocol
Microtiter source plates with 384 wells (LP-0220, Labcyte) were prepared by pipetting the transfection reagents Lipo-fectamine RNAiMAX; DharmaFECT 1, 2, 3, 4; and HiPerFect and the PLK1 and GL2 siGENOME SMARTpool siRNAs (GE Healthcare Dharmacon). Transfection reagents were prediluted 1:1 with siRNA buffer, and the siRNAs were diluted to 20 µM. Before transfer on the Echo, the source plate was centrifuged at 1000 rpm for 1 min. Opti-MEM reduced serum medium (Life Technologies) was dispensed into 384-well destination plates (781948; Greiner, Monroe, NC). The Echo transferred transfection reagents into the plate at 200, 160, 120, 80, 40, and 0 nL per well. PLK1 and GL2 siRNAs were also dispensed by Echo (52.5 nL/well). The source plate setting on the Echo was set to aqueous buffer mode according to the Labcyte protocol. The plate was incubated at room temperature for 20 min to allow siRNA-lipid complex formation. Cells from various cancer lines were trypsinized, counted, and diluted prior to being dispensed on top of the transfection mixture (50 µL/well), and the plates were incubated until viability was assayed.
For experiments in which lipids were hand pipetted, the lipids were first diluted 1:1 with siRNA buffer and then further diluted into Opti-MEM (to achieve the same final volume and concentration as the Echo experiments) prior to being pipetted into an assay plate. siRNAs were delivered by acoustic transfer, and the plates were otherwise handled as described above.
Viability Assay
Cell viability was determined using CellTiter-Glo (Promega, Madison, WI). After 4, 5, or 6 d following transfection, plates were partially aspirated to leave the cells undisturbed and a residual volume of 25 µL using a Biotek 406 plate washer (Biotek, Winooski, VT). CellTiter-Glo reagent was added (25 µL/well), and luminescence was measured using a LEADseeker plate reader (GE Healthcare, Piscataway, NJ) after a 10 min incubation at room temperature. GL2 viability data were normalized to 0 nL/well lipid data. PLK1 viability data were normalized to the GL2 data at the same lipid volume rather than the 0 nL/well lipid data. This more accurately reflects the cell killing that is due specifically to PLK1 knockdown.
Results and Discussion
Nonbiological Validation of Acoustic Dispense of Transfection Reagents
Before using acoustic dispense to optimize siRNA transfection conditions, its ability to deliver both siRNAs and transfection reagents was evaluated. The composition of transfection reagents are not disclosed by their manufacturers, which raises the question of whether they would be compatible with acoustic dispense. By default, the Echo is calibrated to deliver solutions containing at least 70% DMSO, which would not be suitable for dispensing siRNA duplexes nor transfection lipids. With the addition of Labcyte’s OMICS package, two additional modes are available for aqueous solutions: dispensing salt-based buffers and dispensing salt-based buffers containing protein. We tested the accuracy, scalability, and consistency of transfection lipid dispense using the OMICS setting designed for salt-based buffers.
We first examined the accuracy of dispensing undiluted transfection lipids and lipids that were diluted 1:1 in an aqueous buffer (see the Materials and Methods section). For the aqueous buffer, we chose siRNA buffer because it would be compatible with later transfection experiments. FITC was diluted into either pure lipid or 1:1 lipid-buffer mixture (1 mM FITC final) for detection. Assay plates were prefilled with 25 µL/well PBS, and a single droplet of FITC-lipid solution was transferred to each of nine destination wells. Following acoustic dispense, fluorescence intensity was measured. A standard curve prepared from the same solutions ensured that the fluorescence readings were in the linear range and allowed for the determination of the actual volume dispensed (
Fig. 1A
;
Accurately measuring nanoliter volumes is not trivial, and we considered whether the addition of FITC to either undiluted or diluted lipid mixtures could itself have an impact on the acoustic transfer of those solutions. As an orthogonal approach to volume determination, we performed a similarly designed experiment using adenosine triphosphate (ATP) prepared in water. ATP concentration was determined by quantifying the luminescence generated by a luciferase-luciferin reaction. Using a standard curve, the average volume dispensed was 2.8 nL/droplet (n = 20, SD = 0.08;
With the droplet volume accuracy established, we determined the performance over the range of volumes relevant to our transfection optimization experiments. To validate this volume range, we again measured the fluorescence intensity of FITC diluted into undiluted lipid or 1:1 lipid:buffer mixtures acoustically transferred to an assay plate prefilled with PBS. Undiluted and diluted lipids dispensed as expected across this range, with the exception of undiluted HiPerFect ( Fig. 1B, C ). This suggests that the physical properties of HiPerFect differ enough from a typical aqueous buffer that it could not be consistently delivered. Based on this result, we decided to dilute all lipids 1:1 with aqueous buffer prior to acoustic dispense as a standard practice. This necessitates dispensing twice as much reagent, which lowers throughput but increases consistency in volume delivered across all transfection lipids.
Biological Validation of Transfection Reagent Acoustic Dispense
To further validate the utility of acoustic dispense for transfection optimization, we compared acoustic delivery of lipid with the traditional, pipet-based method in a biological setting. The determination of transfection efficiency has been carried out using various methods including quantitative reverse transcriptase polymerase chain reaction, quantitative imaging of fluorescently labeled siRNAs, and Western blot of proteins encoded by the genes targeted for knockdown.14–16 These methods come with their own tradeoffs with respect to ease, susceptibility to artifacts, and biological relevance. Because our primary research interest is to measure the effects of gene knockdown on cell viability, we chose to use the decrease in cell viability following the transfection of siRNA targeting the essential PLK1 gene as a metric for transfection efficiency. Cell death is a downstream consequence of successful PLK1 knockdown, rather than a direct indicator of PLK1 knockdown, but carries the advantage of being amenable to high-throughput experimentation. Using ATP-dependent luminescence assays as an indicator of cell viability, we have been able to infer knockdown efficiency by decreases in viability. In addition, we used siRNA targeting the firefly luciferase gene GL2 as a negative control to determine the effects of off-target toxicity. This can arise from off-target nonspecific siRNA interactions or the transfection lipid itself.
We have successfully used A375 and HCT116 cells in previous genome-wide and targeted siRNA screens.5,6 Based on this experience, these cell lines were transfected in parallel using acoustic dispense or hand pipetting to deliver transfection lipids. In both cases, PLK1 and GL2 siRNAs were dispensed by acoustic dispense, and cell viability was determined after 96 h of knockdown. There were only modest effects on cell viability following GL2 transfection whether lipid was delivered by Echo or pipet, and these effects were seen at higher lipid volumes ( Fig. 2A, B ). This result suggests that neither method results in increased lipid toxicity. As expected, there were severe viability effects following transfection of the PLK1 siRNA with increasing volumes of transfection lipid ( Fig. 2C, D ). Echo and pipet methods tracked closely to each other, with the Echo method appearing slightly more efficient for A375 cells and the pipet method slightly more efficient for HCT116 cells. Although the results from the two methods do not match perfectly, these results show that the results from the Echo method are predictive of what would be obtained by pipet-based transfection.
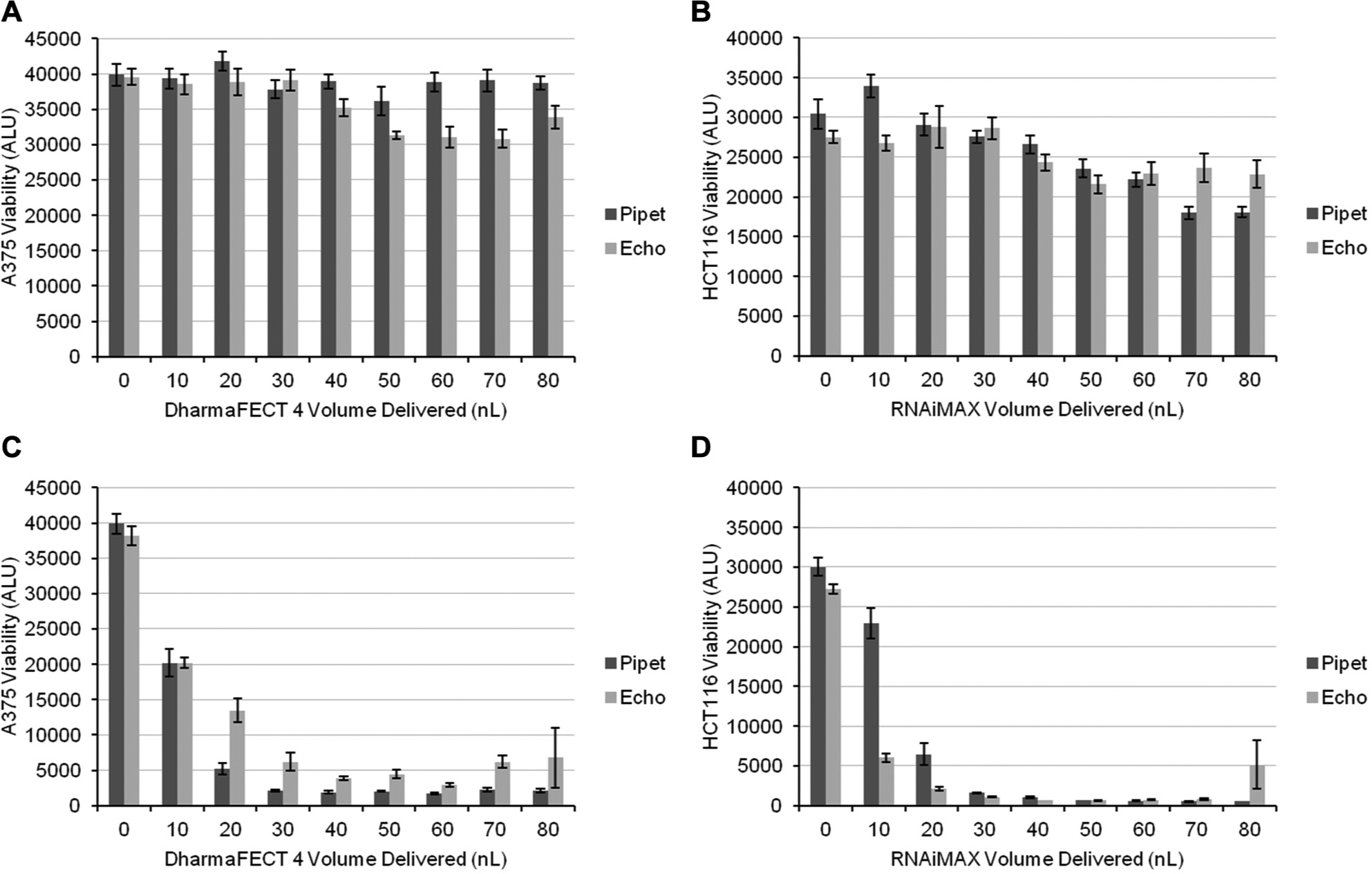
Comparison of pipet-based and Echo-based handling of transfection lipids in A375 and HCT116 cells. (
Interestingly, in both cell lines, we noticed unusually large error bars at 80 nL/well when delivered by acoustic dispense. When the values from the individual wells were examined, it became apparent that there were one or two outlying data points in each case ( Fig. 3A, B ). These relatively high outliers indicate reduced cell killing that is most likely due to incomplete transfection of PLK1 siRNA. As both the siRNA and transfection lipids were delivered by Echo, failure of either reagent to dispense would result in failed PLK1 knockdown. Similar defects were not seen in wells in which lipid was dispensed by pipet and siRNA was dispensed by Echo, suggesting that the sporadic liquid-handling failure was in the delivery of transfection lipid and not siRNAs ( Fig. 2C, D ).
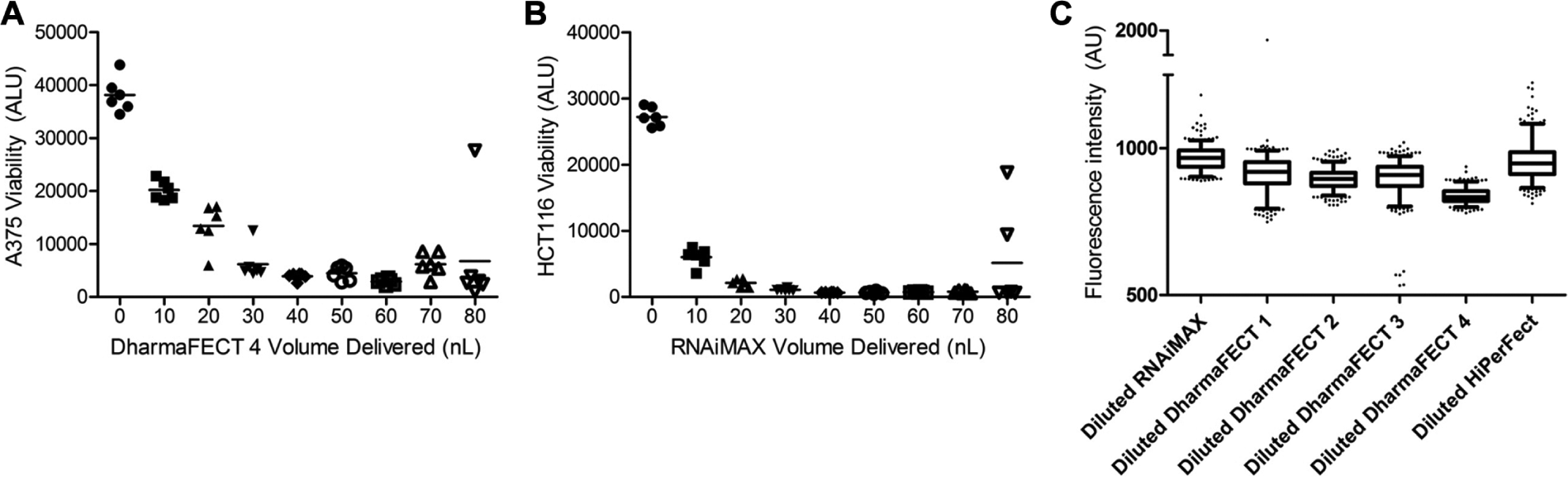
Analysis of acoustic dispense consistency. (
To further test the performance of the Echo to deliver consistently over many wells, we again delivered FITC diluted in the various lipid-buffer mixtures. A single droplet was delivered to each well in a 384-well plate to assess the reliability of delivery. Overall, the Echo dispensed consistently with relatively few outlier wells ( Fig. 3C ). Because only a single droplet was delivered, a missed dispense would have resulted in no detectable fluorescence, but no missed dispenses were detected. A single well from diluted DharmaFECT 1 was twice as fluorescent as other wells, suggesting that it might have received two droplets. The variation observed in other wells is most likely due to differences in droplet size. For instance, there were five wells from diluted DharmaFECT 3 that were roughly half as fluorescent as the other wells, indicating that the droplets delivered to these wells were atypically small.
Throughout our experience using acoustic dispense to deliver transfection lipids, we have consistently observed occasional outlier wells ( Fig. 3A–C , and data not shown). Outliers were caused by underdelivery or overdelivery of reagent. From the six plates dispensed in Figure 3C , 15 wells were at least three standard deviations away from the mean for that lipid. From this, we estimate the frequency of dispense errors to be approximately 1 in 150. Because we were unable to correlate outliers with any definitive cause, we chose to address this factor by performing all experiments with replicate data points. Although no outliers were removed from the data presented in this study, we recommend using median rather than mean in calculations or removing outliers as practical solutions for dealing with wells occasionally missed by acoustic dispense. These approaches minimize the effect of occasional, outlying data points.
Application of Acoustic Dispense for Transfection Optimization across Multiple Cell Lines
siRNA transfection optimization can be affected by the cell number, choice of transfection lipid, transfection lipid concentration, siRNA concentration, plate type, and duration of knockdown. To create a thorough six-dimensional matrix covering the above factors would prevent rapidly optimizing a single cell line, and the optimization of multiple cell lines would be proportionately more difficult and time-consuming. Based on data collected over several years, we have streamlined the optimization matrix by reducing the number of variables considered. First, we have previously found with multiple cell lines that poly-D-lysine–coated microplates consistently give better transfection efficiency than comparable tissue culture–treated microplates (data not shown). Second, we have found no significant increase in knockdown when siRNA concentration exceeds 16 nM for SMARTpools or 8 nM for individual duplexes (data not shown). Third, we have chosen to use starting cell numbers already established for 96-h viability assays. These cell numbers have been chosen to ensure cells are still proliferating 4 d after cells are plated. By removing those three factors, we created a three-dimensional matrix focused on optimizing the remaining factors.
Although there are many commercially available transfection lipids, we limited our matrix to six of the most widely used reagents. Lipofectamine RNAiMAX, HiPerFect, and DharmaFECT 1, 2, 3, and 4 were tested. For each lipid, a range of volumes was tested, from 0 nL/well to 200 nL/well ( Fig. 4A ). Because the lipids were diluted 1:1 with buffer, this represents an actual lipid volume from 0 to 100 nL/well. Importantly, the viability effects following knockdown of the mitotic kinase PLK1 are cell cycle dependent. Based on this, the duration of knockdown required to register a viability effect is linked to the doubling time of the cell line. To address this significant factor, cell viability was assayed following 4, 5, and 6 d of knockdown. An advantage of this approach is that knockdown can be achieved even for stable proteins, because each doubling dilutes the protein twofold once the mRNA is depleted. We recognize that siRNA-mediated gene knockdown is transient, which might allow mRNA levels to recover at later time points. By measuring viability at all three time points, this effect can be detected by following the cell killing following PLK1 knockdown.
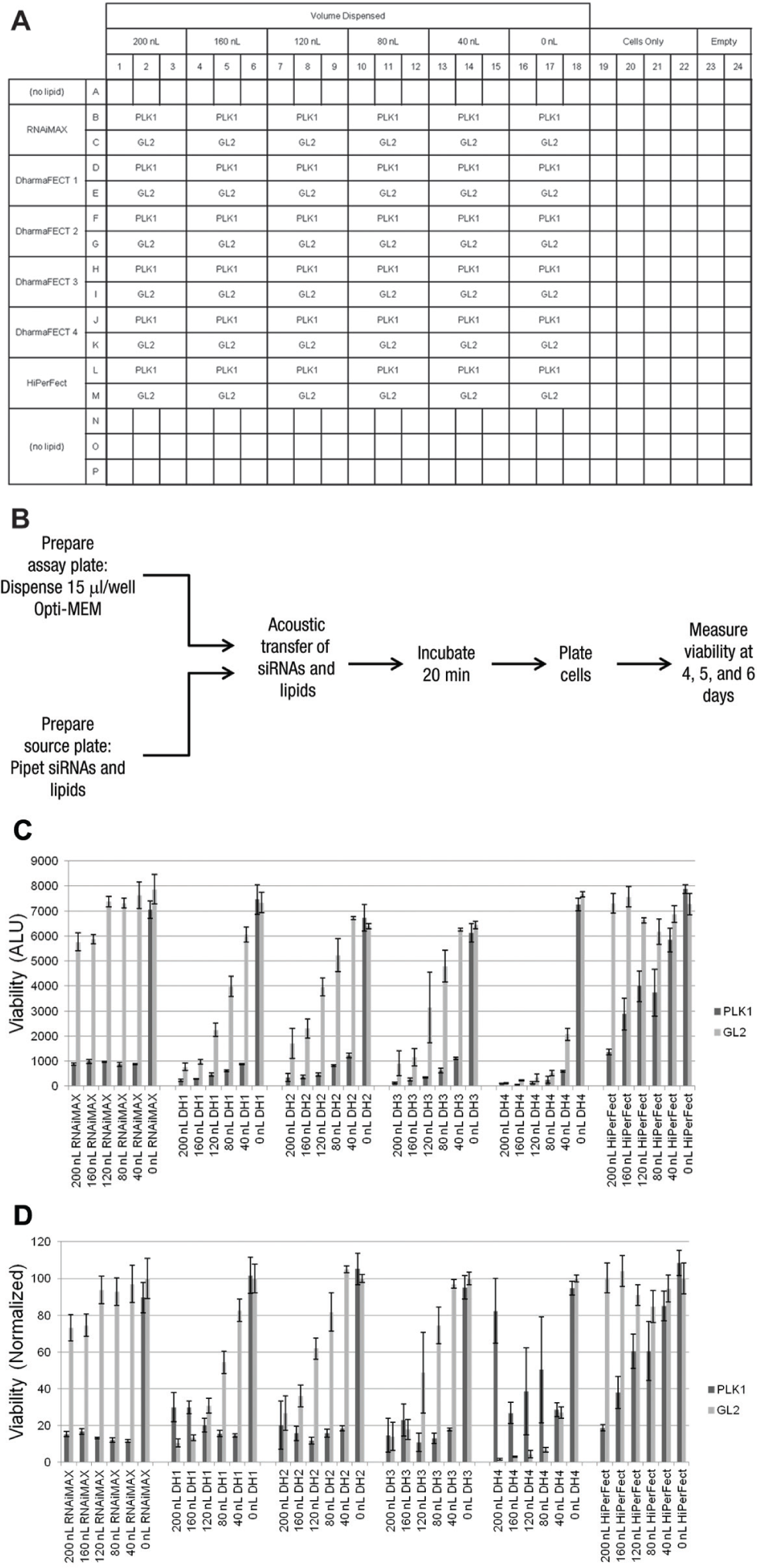
Transfection optimization matrix, workflow, and representative data from BT-20 cells. (
In addition to the determination of knockdown efficiency using PLK1, we assayed the toxicity of the transfection lipids themselves using siRNA targeting the firefly luciferase GL2 gene. GL2 was transfected under the same conditions as PLK1 to give a complete data set to optimize the targeted effect (i.e., cell lethality following PLK1 siRNA transfection) while minimizing off-target toxicity (i.e., cell lethality following GL2 siRNA transfection).
The transfection optimization matrix was applied to cultured, cancer-derived cell lines covering a wide range of originating tissues differing in cell morphology, size, and growth rate. Assay plates were prefilled with 15 µL/well Opti-MEM, and then siRNA and lipid were delivered by acoustic transfer ( Fig. 4B ). After 20 min incubation, cells were plated on top of the transfection mixture. Viability was assessed at 4, 5, and 6 d following transfection.
As an example of the data generated by this approach, viability results after 6 days of knockdown in the BT-20 cell line are shown (
Fig. 4C
). Viability data from 4, 5, and 6 d of knockdown demonstrated that the cells were still proliferating at these time points (
Overall, we found suitable transfection conditions using our criteria for transfectability in 90% of the cell lines tested (
In addition to developing a comprehensive approach to optimizing lipid type, lipid concentration, and duration of knockdown using acoustic dispense, we also wanted to develop a rapid, simple method. Hand pipetting of extensive optimization matrices is time intensive and tedious. In addition, when evaluating multiple reagents at nanoliter volumes, multiple dilutions are necessary, with each dilution introducing an additional source of error. Acoustic dispense possesses two capabilities that make it well suited for siRNA transfection optimization. First, nanoliter volumes can be dispensed, which removes the requirement to predilute reagents. Second, dispenses can be made with fully automated control of the source well, destination well, and volume delivered for each transfer. Once a matrix pattern has been established, it can be repeated without any additional effort. To dispense the matrix described ( Fig. 4A ), the Echo required approximately 3 min per plate. To prepare the dilutions and pipet the same matrix by hand would require at least 10-fold more time.
One complicating factor found when using acoustic dispense for transfection lipids and siRNAs was that not all lipids transfer effectively without being prediluted in an aqueous buffer. In addition, we found replicate data to be necessary in at least triplicate to ensure an assessment could be made in the event of dispense errors. Despite these minor limitations, we successfully applied this methodology to a number of cell lines to rapidly optimize the lipid type, lipid volume, and duration of knockdown to ensure effective knockdown while limiting off-target toxicity due to the transfection itself.
Footnotes
Acknowledgements
The authors express their appreciation to Saurabh Menon, Mike Bembenek, and Mike Kuranda for their helpful discussions and critical review of this manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
