Abstract
High-throughput screening (HTS) is the main starting point for hit identification in drug discovery programs. This has led to a rapid increase of available screening data both within pharmaceutical companies and the public domain. We have used the BioAssay Ontology (BAO) 2.0 for assay annotation within AstraZeneca to enable comparison with external HTS methods. The annotated assays have been analyzed to identify technology gaps, evaluate new methods, verify active hits, and compare compound activity between in-house and PubChem assays. As an example, the binding of a fluorescent ligand to formyl peptide receptor 1 (FPR1, involved in inflammation, for example) in an in-house HTS was measured by fluorescence intensity. In total, 155 active compounds were also tested in an external ligand binding flow cytometry assay, a method not used for in-house HTS detection. Twelve percent of the 155 compounds were found active in both assays. By the annotation of assay protocols using BAO terms, internal and external assays can easily be identified and method comparison facilitated. They can be used to evaluate the effectiveness of different assay methods, design appropriate confirmatory and counterassays, and analyze the activity of compounds for identification of technology artifacts.
Introduction
High-throughput screening (HTS) is one of the most important methods to identify starting points for chemical equity in pharmaceutical drug discovery programs. In addition, HTS is rapidly expanding in academic drug discovery. 1 HTS methods and throughput have continuously evolved over almost 30 years and are now a routine strategy for hit identification in early drug discovery. The constant development of new cost-effective technologies, including sophisticated laboratory automation, has enabled screening of increasingly larger compound collections, generating rapidly increasing quantities of data stored in both internal and public databases. In recent years, large amounts of screening data from different HTS laboratories have become publicly available through the Molecular Libraries Program (MLP) funded by the National Institutes of Health (NIH). The assay descriptions and results are deposited into the PubChem database. 2 PubChem is an open screening data repository, divided into three parts (Substance, Compound, and BioAssay), for archiving and retrieval of chemical structures and their biological test results. The BioAssay database includes 6790 bioassays generated via the NIH MLP program, which include more than 4500 different protein targets, plus many more records imported from other sources. This content in PubChem has been generated by HTS, medicinal chemistry studies, chemical biology research, and drug discovery programs. 3 However, mining of these massive screening data sets is often difficult, because assay protocols are written as free text, lacking standardized vocabulary and formal annotations; as a consequence, critical information about assay background or method specifications is often difficult to access in an automated and systematic fashion since text mining of protocols is usually not possible, or the data are not available.
Screening data integration has been a hot topic over the past few years. There have been both proprietary and open public initiatives to improve the integration of HTS data from different sources. One example of a proprietary solution is the ChemistryConnect application, 4 while Open PHACTS (http://openphacts.org) and BARD (http://bard.nih.gov) are examples of two major publicly funded initiatives for better integration of screening data. Content in the BARD system includes data generated via the NIH MLP, which was manually curated and annotated using controlled terminology and organized into projects. 5 Open PHACTS is an Innovative Medicines Initiative (IMI)–funded private-public partnership with a focus on the integration and reuse of large-scale chemical biology data. Open PHACTS has defined a set of scientifically relevant competency questions around compound-target, and compound-target-disease and pathway relationships that the platform will aim to answer. 6 This work is part of the Open PHACTS project with the aim to develop annotation and analysis tools within the pharmaceutical and biotechnology industry, as well as public databases (e.g., PubChem and ChEMBL). The in-house assays are annotated manually from protocols with high granularity to cover specifications that can be used to identify technology artifacts and tool compounds and to support future assay design and screening campaign development.
Early in a drug discovery program, during hit and lead identification, there is a general demand for new methods for evaluating compound-target interactions. For example, the development of computational models to predict compound activity for a specific target or target class requires thorough annotation of the biological assay and the screening outcomes (such as end point and mechanism of action) to ensure that the training data are interpreted correctly and meaningfully. To support assay development and hit activity analysis in early screening projects, the publicly available BioAssay Ontology (BAO) has been used for AstraZeneca HTS annotation. The annotation of assay protocols is annotated using BAO standardized vocabulary for assay design and technology that can be analyzed together with external data in future drug discovery programs. BAO enables classification of screening assays to facilitate data analysis and comparison between different HTS data sets.7,8 The ontology includes information about the assay design and technology, end point (result) and metadata to describe a target, such as protein origin and cell line background. BAO annotation enables the analysis of biological activity based on various relevant characteristics such as the assay format and type (e.g. biochemical binding assay vs. functional cell based assay), as well as the assay stage and end point with different compound actions (e.g., agonist/antagonist activity of a compound and the reliability of the results of a primary single-point assay vs. a confirmation EC50 assay). More than 900 PubChem assays have been annotated using the BAO terminology by the BioAssay Ontology team, 7 which can be viewed through the BAO Search tool (http://baosearch.ccs.miami.edu/). Many more assays are available via BARD, which also leveraged BAO for its core terminology.
Several aspects of HTS are important to successfully identify chemical matter as suitable starting points for lead generation and lead optimization. It is important that the screened collection is diverse with good physicochemical properties and that the assay is robust and produces reliable results. Different organizations might make different choices of assay technology for a given target, target class, and phenotype. It is therefore of interest to systematically compare different available assay technologies, the usage within each target class or for a specific target or phenotype, and the obtained results. BAO provides a foundation to perform such an analysis. During assay development, assay methods are typically evaluated and optimized using well-defined tool compounds if available, ensuring a robust and reliable screening setup. However, besides the true active and inactive compounds, there is typically a set of compounds that are wrongly assigned as actives (false positives) or inactives (false negatives) in any screening assay. These inaccurate activity results depend on both the variability of the screening method and the susceptibility of the technology to detect artifactual activity of test compounds. The technology artifacts vary between different assay methods and are dependent on several factors such as detection technology, incubation times, and factors influencing compound solubility. To be able to correctly identify those undesired actives that should be removed from a hit list—for example, technology frequent hitters (i.e., compounds with high activity in many unrelated assays using the same technology, e.g., autofluorescent compounds), it is critical that the assays used in the investigation are described in a standard and consistent way. 9 BAO provides the tools for such a uniform and formal description. Therefore, in the current study, HTS assays from in-house drug discovery programs have been retrospectively annotated with BAO terms. On the basis of the standardized annotations of assay design and detection method, we were able to analyze and compare our in-house assay setup with the assay methods of HTS data sets published in PubChem. In this article, we demonstrate how the ontology can be applied in an industrial setting and the utility of BAO to analyze screening data from different sources. We evaluated internal assay methods with external assays from PubChem to identify desired hits and undesired artifacts and demonstrate how external assay data can be used in verifying active compounds in in-house screening programs.
Methods
Assay Annotation
HTS assays from several AstraZeneca screening sites, hereafter referred as internal assays, have been studied together with external assays from the PubChem database. BAO has been extended to cover all methods used both in internal and external HTS assays. These modifications have been based on the scientifically relevant questions with respect to our in-house organization as well as with respect to the Open PHACTS consortium. The assay method modifications have been included in BAO 2.0, while an external module will be created to include infection-specific annotations (i.e., screening of virus and bacterial targets [to be published]).
The BAO 2.0 has been used for annotation of the HTS assays used as primary screening tests in in-house drug discovery programs run between 2005 and 2013 (381 assays). The assays have been annotated using BAO terms manually from assay protocols and include HTS assays run in a different assay format, on divergent target classes, and with a broad usage of assay methods (data not shown). Standardized annotation of in-house HTS assays using BAO is demonstrated by previously published in-house screening assay development experiments ( Table 1 ).10–12
Examples of In-House Assay Annotation Using BioAssay Ontology Terms.
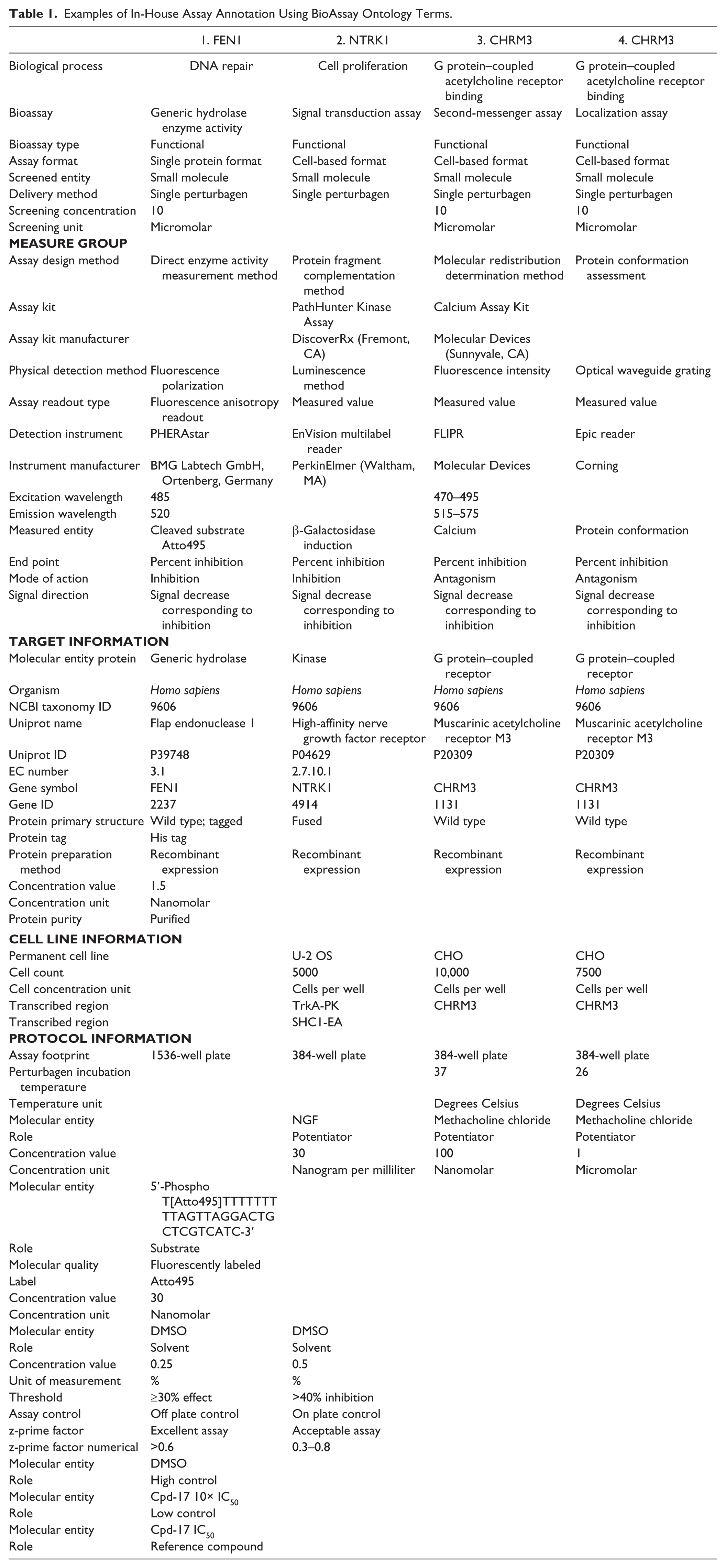
The assay design and detection methods of the in-house HTS assays have been analyzed together with 233 primary PubChem HTS assays. 7 Annotated PubChem assays have been provided by the BAO team and are mainly biochemical assays, G protein–coupled receptor (GPCR) assays, and various assays detected by luminescence. We have also evaluated BARD as an external source of annotated PubChem assay data. Although BARD contains a large subset of PubChem assays annotated using BAO terms, they contain annotations of using only a subset of the BAO concepts. For our analysis, we therefore used assays annotated by the BAO team since we required the capture of information on assay design method and assay stage to allow comparison of assay methods and the identification of confirmatory assays and counterscreens. Mappings of BARD and BAO terms are available at https://bard.nih.gov/BARD/dictionaryTerms/dictionaryTerms.
Compound Activity Analysis
Compound overlap was studied between in-house assays and assays annotated from PubChem. Compound structures were standardized according to AstraZeneca rules 13 and used for compound identification and comparison of compounds tested in both internal and external HTS assays. 4 Compound activity between primary HTS assays screened on the same human-derived protein target (defined by gene symbol) was analyzed using compound activity overlap A1,2 /(A1 + A2 + A1,2), where A1,2 is the number of compounds active in both assay 1 and assay 2, and A1 and A2 are the number of compounds only active in one of the assays and inactive in the other. Where more than two assays were evaluated for the same target, the compound activity was analyzed both in pairs as above and by evaluating the compound activity overlap (also calculated in assay pairs according to the equation above) using only compounds tested in all three assays. By the use of BAO assay stage annotations, both internal and external screening cascades, including confirmation assays and counterscreens, could be identified and used to evaluate the most robust actives identified in the primary HTS assay.
Results
Primary HTS Assay Annotation Using BAO 2.0
The ontology has been extended to facilitate the annotation of HTS assays from pharmaceutical companies. For example, new relationships for thorough annotation of detection wavelengths, restructuring the annotation of target protein primary structure, and the addition of assay kits and instruments used within AstraZeneca have been added to the ontology. BAO 2.0 classes and properties 8 have then been used to represent our in-house drug discovery data annotations as semantic triples. In-house HTS assays are represented as new instances of “bioassay,” “screening campaign,” and “measure group,” which are the core of the screening cascade and assay annotations in BAO 2.0. The remaining assay data around assay design and detection methods, cell line background, and protein origin are then annotated according to the ontology in relation to these instances, summarized in Figure 1 and the following workflow.
An HTS assay corresponds to an instance in the “bioassay” ontology class, which is a concept in BAO.
The bioassay has a defined assay stage and relations (e.g., “is confirmatory assay of” or “has orthogonal assay technology”) to other instances of “bioassay” in the same “screening campaign.”
Each “bioassay” has one or more “measure group” instances, dependent on the “assay readout content,” where the “assay design method” and “physical detection method” are described.
End points are annotated with relations to one or several “measure group(s)” that describes the methods leading to each result type (e.g., “IC50”or “percent inhibition”).
Each “bioassay” is associated with the “assay format” describing the model system of the assay (e.g. “biochemical,” “cell based,” etc.).
For each “bioassay” instance, biological details and screened entities are described by corresponding instances of material entities and roles, such as a protein with the role “target,” a “cell line cell” for a cell-based assay, and small molecules with the “screened entity role.”
In addition, various associated details such as “assay title,” “source,” and so on are described.
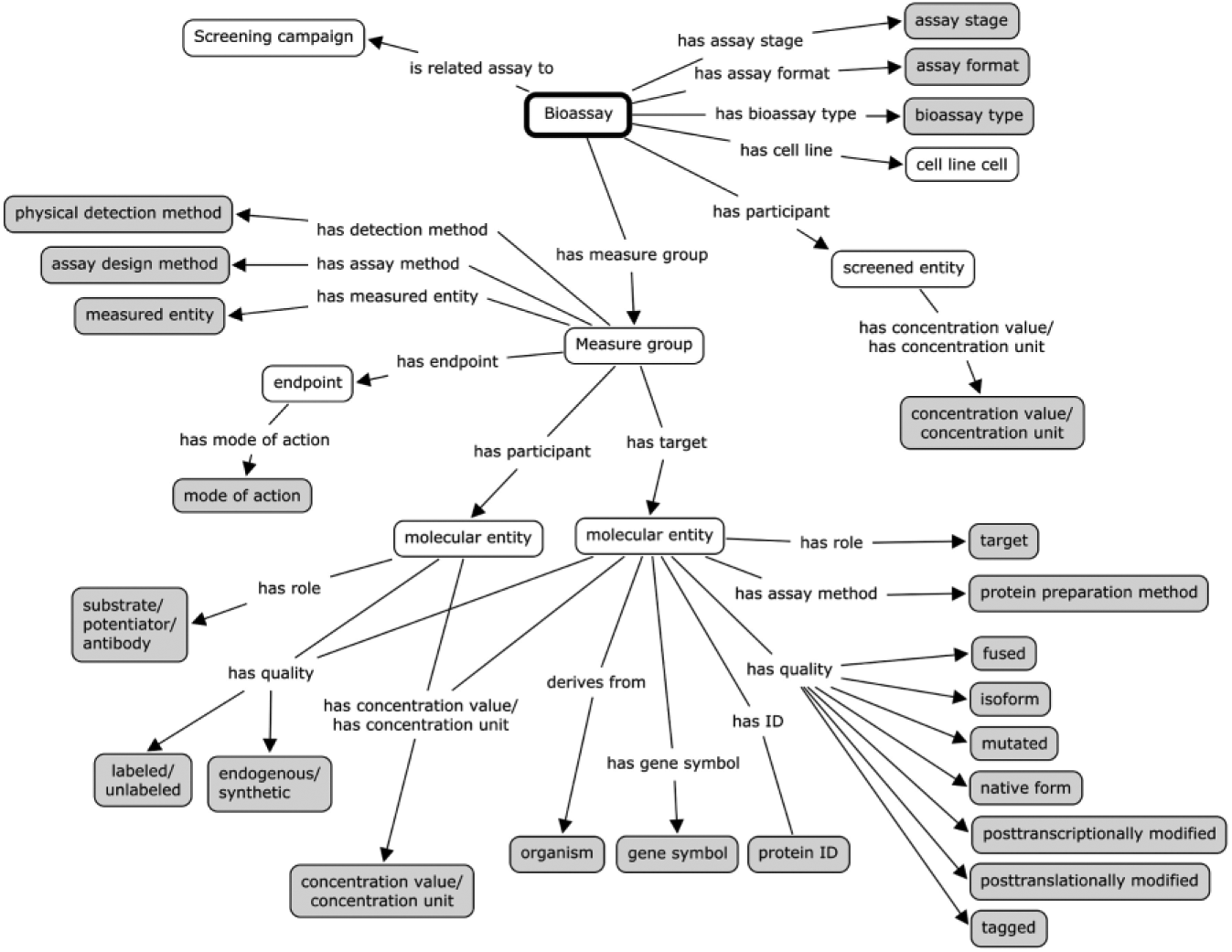
Schematic presentation of BioAssay Ontology classes and properties used for in-house high-throughput screening (HTS) annotation. The data are annotated using already defined classes in BAO 2.0 (in gray) or by new instances that can be further described by properties as has quality and has concentration (in white).
By inserting new instances in these classes, we can assert specifications, for example, concentrations and qualities of the assay target, substrate, and screened compound, in a semantic format ( Fig. 1 , Table 1 ). The setup enables us to define target-specific molecular entities, such as substrates or potentiators, and easily compare many similar assays in depth, for example, with regard to the screening concentration or the labeling of a coupled substrate. A fluorochrome-labeled substrate can be identified by the substrate name (or, in this case, by a DNA sequence) but can also be defined as a “fluorescently labeled” substrate by the object property “has quality” and the property “has part” “Atto495” (see substrate annotation of the FEN1 assay, Table 1 ). BAO 2.0, with our recent extensions, can be used to annotate targets in a similar way. The protein can be described as a wild-type protein with a protein tag or with posttranslational modification. Furthermore, to facilitate activity data analysis from different sources, we have extended the assay quality assessment segment of the ontology. The “z-prime factor” data have been classified into excellent and acceptable assays (Z′ value >0.5 and 0-0.5, respectively14,15) and the “on plate control”/“off plate control” parameters have been included in the ontology since both plate setups are used for in-house HTS screening (FEN1 and NTRK1 assay, Table 1 ).
Assay Design and Detection Technology Usage in Primary HTS
Assay design and detection method of in-house HTS assays and PubChem assays employing human-derived targets have been analyzed as a function of assay format, and the results are presented in Figures 2 and 3 . Several in-house binding assays have used radiometric methods such as scintillation proximity assays (SPAs) and filter-wash methods, neither of which could be found among the annotated PubChem assays. Neither microscopy nor optical waveguide grating, 16 which have been used in a number of in-house screening activities, were used as detection technologies in the annotated external screening assays published in PubChem. SPAs and optical waveguide grating are flexible methods used for detection of enzyme, GPCRs, and other targets in either a biochemical or cell-based format.
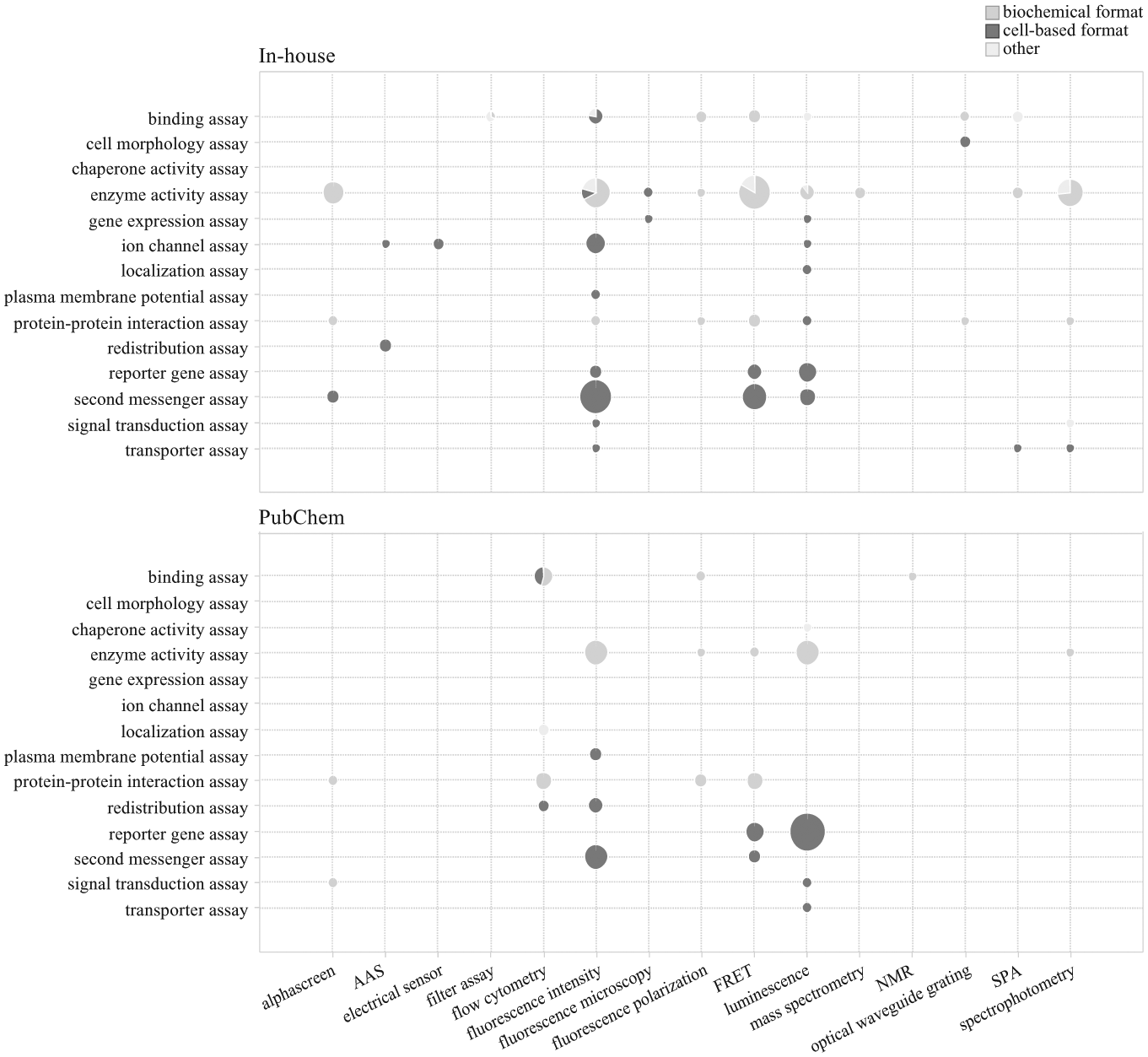
Bioassay type and detection technology of in-house and annotated PubChem high-throughput screening assays on human targets (297 and 190 assays, respectively). Numbers of assays are represented by size of each pie with sectors divided into cell-based assay format (dark gray), biochemical format (gray), and other assay formats (i.e., subcellular, tissue-based, and whole-cell lysate format; light gray). AAS, atomic absorption spectrophotometry; FRET, fluorescence resonance energy transfer; NMR, nuclear magnetic resonance; SPA, scintillation proximity assay.
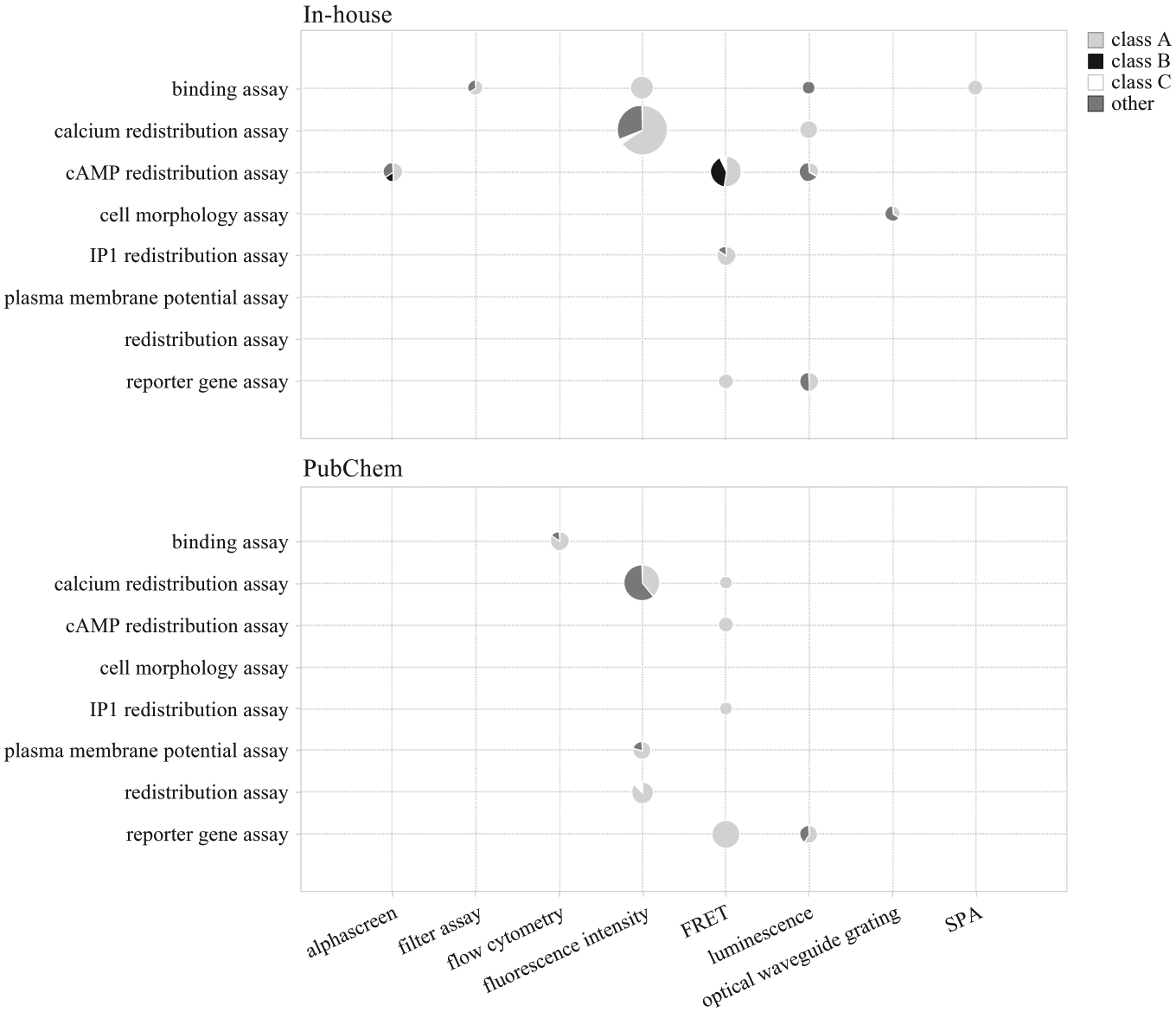
Bioassay type and detection technology of in-house and annotated PubChem G protein–coupled receptor (GPCR) assays on human targets (95 and 60 assays, respectively). Numbers of assays are represented by size of each pie with sectors divided by GPCR subclasses. FRET, fluorescence resonance energy transfer; SPA, scintillation proximity assay.
There were only a limited number of ion channel assays among the annotated PubChem assays, and therefore no assays detected by atomic absorption spectrophotometry (AAS) or electrical sensors were included ( Fig. 2 , data not shown). There are, however, PubChem assays using plasma membrane potential and dye redistribution methods to measure the GPCR activity by fluorescence in an ion channel coupled design ( Fig. 3 ). Furthermore, we found flow cytometry to be a technology used externally (e.g., for GPCR and nuclear hormone receptor assays). Flow cytometry is not present among our in-house HTS assays. In another example, reporter gene assays are much more frequent among PubChem assays, both for GPCRs and other target classes, compared with our in-house assays.
Among the differences between in-house and external assays, a more frequent usage of second-messenger cAMP in GPCR assays could be seen in the in-house HTS data ( Fig. 3 ). One reason for the difference in second-messenger usage can be that no class B GPCRs, which generally are Gs coupled, have been used for screening in the annotated PubChem assays (data not shown). Thus, differences in assay technologies used in different organizations depend on different target classes screened, availability, and cost of detection technologies as well as other factors.
Analysis Using External BAO Annotated Assays for Verification of Active Compounds
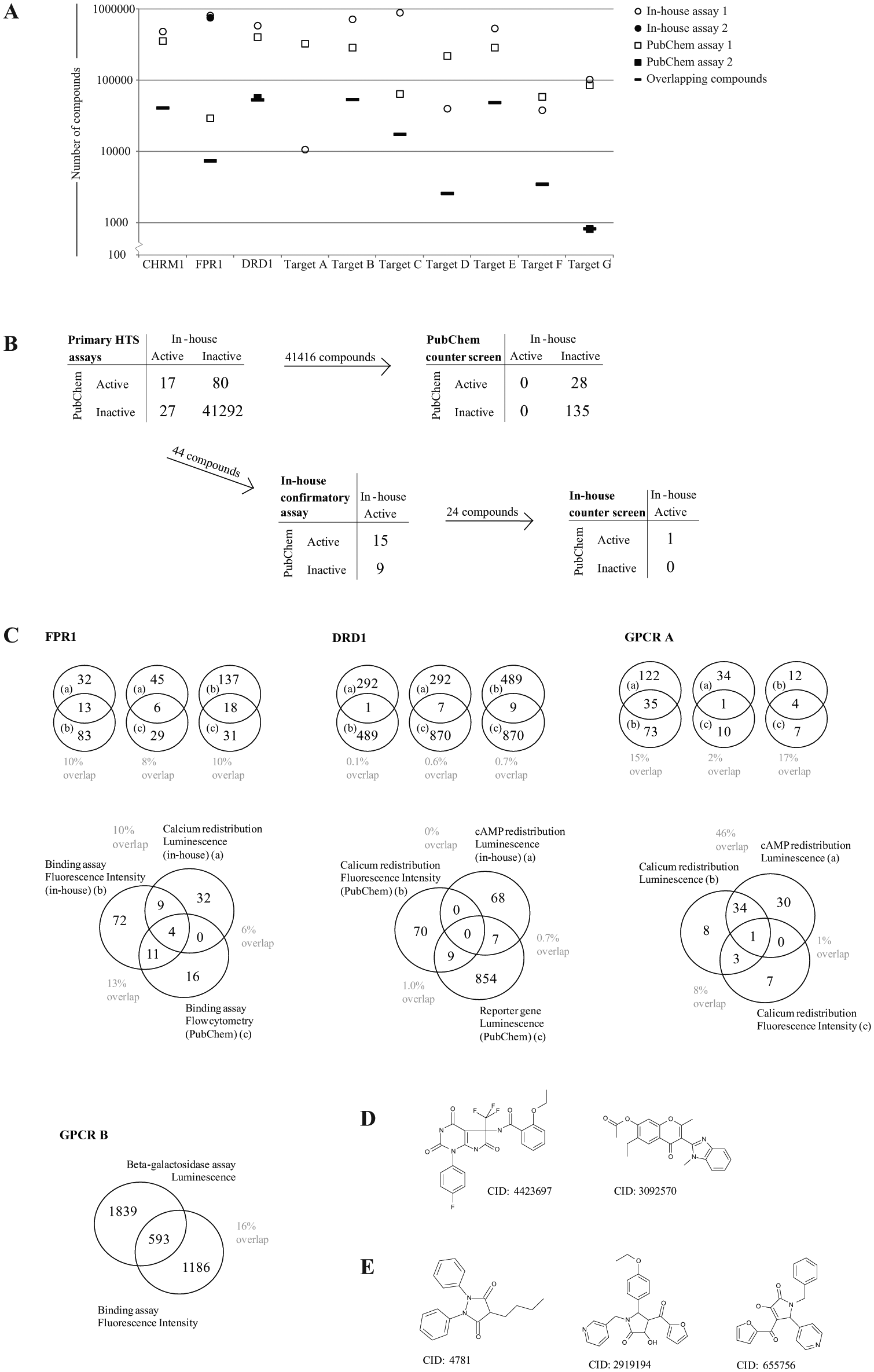
Fourteen human-derived targets in the BAO annotated HTS data set have been screened both in house and externally, where the results have been uploaded to PubChem. Of these, 10 targets were chosen as they have the same mechanism of action; these were used for compound overlap analysis between in-house and PubChem screening ( Fig. 4A ). To further evaluate the possibility of using external data for verification of active compounds in HTS screening programs, compound activities for three GPCR targets were selected—namely, muscarinic acetylcholine receptor M1 (CHRM1), N-formyl peptide receptor (FPR1), and dopamine D1 receptor (DRD1).
(
For CHRM1, two similar assay designs were used in the in-house and external primary HTS assay ( Table 2 , PubChem AID: 588814, BARD EID: 224917,18). The target was expressed in Chinese hamster ovary (CHO) cells in both assays, and the compound activity was detected in a Ca2+ flux assay by FLIPR. The assays did use different detection dyes, but the fluorescence emission wavelength was within the same range. A chemical structure analysis revealed an overlap of over 40,000 compounds between the in-house and external assays, due to the inclusion of commercially available compounds in the in-house compound collection. Among the overlapping compounds, 17 were found active in both assays, while 27 and 80 compounds were found active exclusively in the in-house and external assays, respectively ( Fig. 4B ). The BAO annotated PubChem data were also used to identify a suitable counterscreen (PubChem AID: 602248, BARD EID: 208319,20), identifying false positives that interfere with the assay method, using nontransfected parental cells. Using this assay, 28 of the 80 compounds found active exclusively in the primary PubChem assay were identified as false actives, while none of the compounds active in the in-house primary assay was active in the PubChem counterscreen. In house, the active compounds in the primary assay were screened in a single concentration retest, which confirmed 15 of the 17 compounds (88%) active in both primary (in house and external) assays and 9 of the 27 compounds (33%) not found active in the PubChem primary assay. Thereafter, a counterscreen similar to that used in the PubChem screening cascade identified one additional compound active in both primary assays as falsely active. The active compound overlap in the CHRM1 assays analyzed was around 14%. The difference in compound activity can be explained by the general variability seen between any two assays but also by a possible difference between the assays in parameters not considered in this analysis. The current study does, however, show the usefulness of external assays in building confidence in the hits that have been found. From the compounds found active in the internal CHRM1 primary assay, a higher number of compounds could be confirmed in the in-house confirmatory assay among the compounds already confirmed by the external CHRM1 HTS assay, compared with those that were not active in the external primary HTS assay (88% and 33%, respectively).
Assay Design and Technology of Internal and PubChem Assays Used for Verification Analysis of Active Compounds.
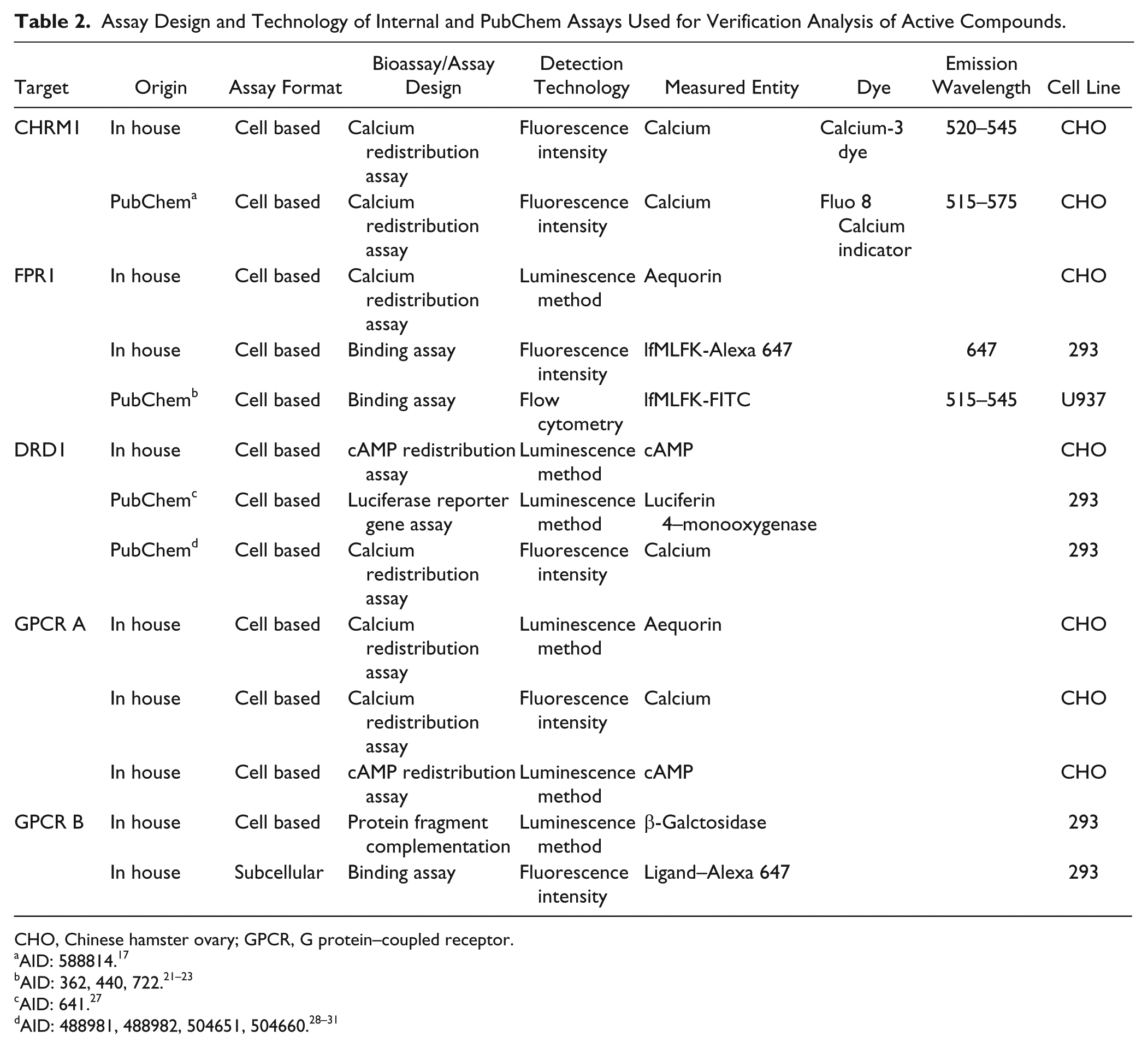
CHO, Chinese hamster ovary; GPCR, G protein–coupled receptor.
AID: 588814. 17
AID: 641. 27
Second, the compound activity in three assays using different detection technologies was analyzed in FPR1 and DRD1 screening. For the FPR1 target, both an external and an in-house competitive binding assay detected by fluorescence have been used ( Table 2 , PubChem AID: 362, 440, 722; BARD EID: 324, 642, 295021–26). These fluorescent ligand binding assays were detected by flow cytometry and fluorescence intensity, respectively, at different wavelength ranges due to the use of different fluorochromes for ligand labeling. In addition, we have included an in-house FPR1 HTS using a functional calcium redistribution assay method detected by luminescence in the analysis ( Table 2 ). The overlap between one of the internal assays and the external assay was only about 7000 compounds, but more than half, approximately 4500 compounds, were analyzed in all three assays. Based on the compounds tested in all three assays, the active compound overlap between the two binding assays was 13%, while the corresponding value was 10% for the two in-house assays and 6% between the external binding assay and the in-house functional assay ( Fig. 4C ). This can be compared with the active compound overlap in the previous analysis using similar assay methods ( Fig. 4C , CHRM1) and the analysis of only in-house HTS assays ( Fig. 4C , GPCR A and B), which both had an active compound overlap of around 15%. Different scaffolds were picked up in all three FPR1 assays (examples presented in Fig. 4D ), while certain scaffolds were found only in the binding assays ( Fig. 4E ). To evaluate the probability of the compounds being active only in the ligand binding assays detected by fluorescence to be technology artifacts, we further studied the activity of these compounds. All of the presented compounds were tested in more than 100 in-house HTS assays with an activity of less than 3% of the assays. In assays using fluorescence for detection, the compounds were not active in more than 6% of the assays (e.g., CID: 4781 was active in 3 of 59 assays using different targets [data not shown]). Since the assays were based on different assay design methods and used various fluorochromes with different emission wavelengths, we concluded the probability of CID: 4781 being a technology frequent hitter to be low.
The three DRD1 assays evaluated were all functional screens but used different assay designs. The in-house HTS was measuring second-messenger cAMP by luminescence, while the two external assays measured calcium redistribution by fluorescence and the luciferase reporter gene method ( Table 2 ; PubChem AID: 641, 488981, 488982, 504651, 504660; BARD EID: 2911, 755, 756, 1654, 166327–35). More than 13,000 compounds have been tested in all three assays, and almost 54,000 compounds have been tested in the in-house assay and at least one external assay. Surprisingly, there was a compound activity overlap of less than 1% between any of the assays, far less than observed for the other targets ( Fig. 4C ).
To evaluate the usage of external assays in the studies verifying active compounds, we further analyzed the overlap of active compounds of two in-house GPCR targets screened with different methods. First, the compound activity in two calcium redistribution assays detected by fluorescence and luminescence, respectively, and a second-messenger cAMP assay detected by luminescence were used for comparison ( Table 2 , GPCR A). The overlap of active compounds was 17% and 15% when assay design or technology was the same (in this case, assays using the calcium redistribution method and assays detected by luminescence respectively) but only 2% between the primary HTS assays in which neither the assay design nor detection method was the same ( Fig. 4C , GPCR A). Furthermore, the compound activity analysis between two other dissimilar in-house assays—namely, β-galactosidase fragment complementation assay detected by luminescence and fluorescent ligand binding assay—showed an active compound overlap of 16% of the more than 20,000 compounds tested in both assays ( Table 2 , Fig. 4C , GPCR B).
All the included examples of active compound analyses illustrate how external assays can be used to verify active compounds in HTS evaluation comparing either assays screened with a similar method (CHRM1) or screening tests using a different assay design and detection technology (FPR1). The compound activity overlap is only slightly lower when external assays are included than what can be seen in the analysis of only internal HTS assays (GPCR A and B). The analysis also shows the usage of assays annotated with BAO terms helps in analyzing assays with high false activity, which could be the case in the DRD1 assay, in which the active compound overlap is low.
Discussion
The BioAssay Ontology has been used to annotate in-house HTS by a common vocabulary for assay design and detection method and other categories that describe and categorize assays and screening results. BAO HTS assay annotations thus enable the comparison and analysis of assays based on various concepts, such as assay design and detection methods, format, biology, and other meta-descriptors. These annotations, for example, enable scientists to relate biological processes or diseases with assay formats or target classes or to identify assays based on criteria such as a specific fluorescent emission wavelength. The muscarinic acetylcholine receptor M3 (CHRM3) example ( Table 1 ) highlights differences in the detection of active compounds between the more conventional GPCR assay—namely, FLIPR and optical waveguide grating, which detect dynamic mass redistribution in the cell upon compound treatment, measured by an Epic instrument. 12 In the annotated data, the assay methods are easily evaluated; for example, differences in incubation temperature and methacholine chloride concentration, which may affect the measured percent effect, in the two assays are evident ( Table 1 , CHRM3 assays 3 and 4). A thorough manual annotation of assay protocols using a standard vocabulary is a laborious assignment but can be used for numerous analyses with external data, both retrospectively and in upcoming drug discovery programs. Using BAO to compare results by concepts such as assay method between various data sources, one should take into account that the depth of annotation, as well as the specific implementation of how each assay is annotated, can vary. However, in a sufficiently descriptive ontology with formal semantics, 36 the meaning of such annotations would be expected to stay the same or at least not be contradictive for more or less specific annotations (subsumption hierarchy). In such cases, one would still be able perform analyses of BAO annotated assays with less granularity. In addition, informatics systems to annotate assays using specific ontology-driven templates, such as BARD, will drive the adoption of well-defined vocabularies and formal ontologies to retrieve data based on specific and well-defined descriptors.
During the development of new screening assays, one often has an option to choose from several screening methods, which can differ substantially in cost, speed, robustness, and the types of artifacts that are expected and consequently the counter and confirmatory assays required for a screening campaign. More cost-effective methods and methods with higher throughput are constantly being developed and marketed. During assay development early in a discovery program, several methods are often compared and evaluated for each target,11,12,37 which leads to the optimization and progression of the method that best meets the criteria of cost, automation, and robust detection of tool compounds. The assay method preference for a specific target or target class is not obvious and differs between screening sites and organizations, since choice of assay method is also influenced by retrospective analysis of success with a particular technology within the organization, the experience and bias of the researchers, and the availability of reagents and instruments. To evaluate such differences and to identify any technology gaps, we compared in-house HTS screening methods with external HTS methods. In house, we use a diverse set of bioassays and detection technologies such as optical waveguide grating, SPA, and AAS, which could not be found in the annotated PubChem assays ( Figs. 2 and 3 ). Fluorescence resonance energy transfer (FRET) is commonly used for the detection of enzyme activity internally but not among the annotated external assays from PubChem. It is also evident that flow cytometry, a method not used for detection in in-house HTS assays, is used externally (e.g., PubChem AID: 440, 759, 761, 1423; BARD EID: 642, 2984, 2986, 358122,25,38–43). Using BAO for assay annotation, assay design and detection method can readily be evaluated for a new target or target class early in drug discovery programs, and new screening methods and useful reagents can be identified.
To evaluate how publicly available data can be leveraged in the verification of active hits from HTS, we have compared, based on BAO annotations, the overlap of compound activity between in-house and PubChem assays. For all targets studied here, except DRD1, the active compound overlap was around 10% to 15% and seemed not to be dependent on the assay source ( Fig. 4B , C ). The low compound activity overlap shows the importance of the choice of assay method for the primary HTS assay and subsequent orthogonal assays for compound hit confirmation. The compound activity overlap between in-house and PubChem assays did not differ significantly from the internal analysis using different methods (GPCR A and B) or from what has been described previously.44–46 Furthermore, and perhaps not surprisingly, the overlap of active compounds is significantly lower in assays using different methods than the noise seen between assays when a set of compounds is screened twice under the same conditions, 45 due to the detection of assay method-specific artifacts. Whether the assay design, detection method, or another factor is the predominant cause for the inconsistency in compound activity is yet to be investigated. In this study, we saw a lower compound activity overlap when both the assay design and detection method differed (assays a and c; GPCR A, Fig. 4C ) than when either the assay design or detection was the same (assays b and c and assays a and b, respectively; GPCR A, Fig. 4C ), suggesting that both factors can have an impact on the result. Further complicating the picture for GPCR A is that measurement of cAMP production and calcium redistribution capture signaling are via different pathways. It is well established that ligands can have a pathway bias, and selection of assay technology therefore needs to take into consideration which pathway to target. The primary assay setup and screening cascade design for a new target might benefit from a similar analysis systematically incorporating external HTS results. Furthermore, the advantage of using two different methods to broaden the identification of new lead series for the target needs to be evaluated.
The overlap of active compounds between the DRD1 assays was surprisingly low, regardless of the fact that only two of three compounds active only in the in-house assay were confirmed in a retest ( Fig. 4C , data not shown). This could be due to the fact that very different assay methods have been used, leading to different sets of false actives ( Table 2 ). It could also be influenced by other factors such as the difference in DNA constructs used for target expression or compound incubation time, and the possibility of a variance between the methods to detect different compound sets (i.e., the loss of hit compounds due to false-negative activity) cannot be ruled out.
A thorough annotation of assay design and detection method enables the identification of internal and external reference assays using a similar method, which can be used for artifact analysis. Identification of technology frequent hitters at an early stage of the screening cascade would improve the active compound overlap between two methods and facilitate the definition of compound hits in drug discovery programs. Standardized vocabularies for assay annotations enable informatics systems to retrieve data based on specific and well-defined descriptors. For example, the annotations of the assays illustrated here revealed two assays with different target classes, enzyme, and GPCR, and thus different assay designs and assay kit/reagents (FEN1 assay and CHRM3 assay 3, Table 1 ), which nevertheless were measured by fluorescence with excitation and emission wavelengths in the same range. Therefore, it can be anticipated that the assays may share common frequent hitters (i.e., compounds active in both assays due to autofluorescence or compounds interfering with the detection technology). Such information has not been captured previously because of the additional burden on assay depositors to explicitly annotate assay protocols and published studies, as well as the lack of established controlled terminologies. However, these challenges are now being addressed by BAO, BARD, and technologies to facilitate annotations based on machine learning and leveraging semantics in ontologies. 47
The utilization of the rapidly increasing amount of external HTS assay results stored in different repositories is impeded by the lack of comprehensive semantically and syntactically uniform and consistent assay annotations. Here, we illustrate the advantage of a standardized vocabulary, clear definitions, and, in many cases, formal descriptions to annotate and analyze HTS assay and screening results. BAO was developed as a foundation to annotate and describe assays, results, and screening campaigns consistently. Upon annotating HTS assays and screening campaigns based on BAO, we are able to analyze screening methods and assay setup both internally and in comparison with external data. The analysis revealed preferential assay methods and technology gaps for specific targets between in-house and external data. The external data also enabled compound activity analysis of several overlapping targets; in some cases, the percentage of overlapping active compounds is in range with what has been previously published, while in other cases, the detection of actives critically depends on the assay method. The analysis highlights the importance of assay choice and selection of orthogonal assays for subsequent cascades. We conclude that a comparative analysis with external data is useful to support assay development in the early stage projects as well as for verifying active compounds in drug discovery programs.
Footnotes
Acknowledgements
We thank Mats Ormö and Péter Várkonyi for their contribution to this work; Tom Plasterer, Plamen Petrov, Simon Rakov, and Isabella Feierberg for useful discussions; and Rurika Oka and Mari Hansson for assisting with the in-house HTS assay annotation.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was supported by Open PHACTS (http://openphacts.org), funded by the Innovative Medicines Initiative of the EU and EFPIA (![]() ). The development of BAO was supported by the National Institutes of Health (NIH) by grants RC2-HG005668 and U01-HL111561 to SCS.
). The development of BAO was supported by the National Institutes of Health (NIH) by grants RC2-HG005668 and U01-HL111561 to SCS.
