Abstract
In vitro models that accurately and rapidly assess hepatotoxicity and the effects of hepatic metabolism on nonliver cell types are needed by the U.S. Department of Defense and the pharmaceutical industry to screen compound libraries. Here, we report the first use of high content analysis on the Integrated Discrete Multiple Organ Co-Culture (IdMOC) system, a high-throughput method for such studies. We cultured 3T3-L1 cells in the presence and absence of primary human hepatocytes, and exposed the cultures to 4-aminophenol and cyclophosphamide, model toxicants that are respectively detoxified and activated by the liver. Following staining with calcein-AM, ethidium homodimer-1, and Hoechst 33342, high content analysis of the cultures revealed four cytotoxic endpoints: fluorescence intensities of calcein-AM and ethidium homodimer-1, nuclear area, and cell density. Using these endpoints, we observed that the cytotoxicity of 4-aminophenol in 3T3-L1 cells in co-culture was less than that observed for 3T3-L1 monocultures, consistent with the known detoxification of 4-aminophenol by hepatocytes. Conversely, cyclophosphamide cytotoxicity for 3T3-L1 cells was enhanced by co-culturing with hepatocytes, consistent with the known metabolic activation of this toxicant. The use of IdMOC plates combined with high content analysis is therefore a multi-endpoint, high-throughput capability for measuring the effects of metabolism on toxicity.
Introduction
In vitro toxicity experiments attempt to replicate the toxicological effects of a given compound as they occur inside a living organism, but the complex physiology that influences toxicity in vivo is difficult to replicate in vitro, and improved models are needed to reduce the dependence on animal testing. 1 One limitation of standard in vitro models is that many do not assess the potential for hepatotoxicity, or the effects of metabolism on the toxicity of a particular drug or toxicant. In vivo xenobiotic metabolism of a given toxicant by the liver can activate or deactivate a parent compound, thus influencing the pharmacokinetics and pharmacodynamics for that toxicant. In vitro systems that include an appropriate metabolic component are thus more biologically relevant than plate-based experiments that use a single cell type, and several clever and intricate systems have been described (for a review, see Ref. 2 ). Although these systems recapitulate in vivo physiology better than many simpler in vitro models, they are often too cumbersome to allow for the high-throughput screening of large chemical libraries for drug development and toxicity testing as required by pharmaceutical companies and the U.S. Department of Defense, respectively. As such, a biologically relevant in vitro liver model that allows for rapid screening and detects multiple parameters for toxicity evaluation is needed.
High content analysis is a quantitative, multiparametric tool that is powerful for the evaluation of toxicity in vitro 3 and has been used to predict drug-induced liver injury in both primary human hepatocyte and HepG2 models.4,5 To simultaneously evaluate both hepatotoxicity and metabolism, a high content model is needed that can evaluate multiple cells types at once to determine the effects of metabolism on the toxicity of compounds for both hepatocytes and other cell types in co-culture. The IdMOC system 6 provides such a cell culture system but has previously not been combined with high content analysis.
The IdMOC system is a simple tool that allows for the incorporation of a metabolic component for in vitro toxicity screening. In this method, cells are cultured independently in short inner wells that are housed together in the same outer chamber (
We report herein the development of a high content–based multiparametric method for the evaluation of toxicity in co-cultures of primary human hepatocytes and 3T3-L1 cells using IdMOC plates. The mouse-derived 3T3-L1 cell line is a commonly used model in cytotoxicity testing 7 that was chosen here due to its demonstrated lack of xenobiotic metabolism. 8 The model toxicants, 4-aminophenol and cyclophosphamide, were used in these experiments because they are known to be respectively detoxified 9 and activated 10 by hepatic metabolism in vivo, effects that are demonstrable in our system. This method provides researchers with a rapid, informative, and physiologically relevant means for the screening of large compound libraries for both hepatotoxicity and metabolic effects.
Materials and Methods
Materials
The LIVE/DEAD viability–cytotoxicity kit for mammalian cells and Hoechst 33342 dye were purchased from Invitrogen (Carlsbad, CA). Cyclophosphamide, 4-aminophenol, and 1-aminobenzotriazole were obtained from Sigma-Aldrich (St. Louis, MO). Collagen-coated 96-well IdMOC plates, Universal Cryo Recovery Medium, Universal Primary Cell Plating Medium, and Hepatocyte Induction Medium (HIM) were purchased from In Vitro ADMET Laboratories (Columbia, MD).
Cells and Cell Culture
Cryopreserved human primary hepatocytes were obtained from In Vitro ADMET Laboratories (lot #HH1020). These hepatocytes were obtained from a female donor, and they were selected for their viability and robust expression of many P450 isoforms in vitro. 11 The mouse 3T3-L1 cell line was obtained from ATCC (Manassas, VA). The 3T3-L1 cells were subcultured in RPMI 1640 medium (Invitrogen) supplemented with 10% fetal bovine serum and 5% penicillin in a humidified incubator with 5% CO2 at 37 °C. The cells were expanded, frozen at −80 °C in complete medium containing 10% dimethyl sulfoxide (DMSO), and then stored in liquid nitrogen. They were used experimentally at passages 9 to 11.
IdMOC Experiments
IdMOC experiments were conducted essentially as described. 8 Briefly, cryopreserved primary human hepatocytes from lot #HH1020 were thawed at 37 °C and placed into 50 mL warm Universal Cryo Recovery Medium, whereas cryopreserved 3T3-L1 cells were similarly thawed and placed into 50 mL warm RPMI 1640 medium supplemented with 10% fetal bovine serum and 5% penicillin. After centrifugation at 100× g for 10 min at 4 °C, the cell pellets were resuspended in Universal Primary Cell Plating medium to a final density of 106 cells/mL of hepatocytes and 2.0×105 cells/mL of 3T3-L1 cells. Thirty-five microliters of cell suspension or Universal Primary Cell Plating medium were added to each well of a 96-well collagen-coated IdMOC plate. The plates were kept in the cell culture hood for 30 min to allow the cells to attach, and then they were moved to a 37 °C humidified incubator with 5% CO2 for 4 h. After incubation, 390 µL of HIM was added to each outer chamber, followed by 600 µL of 2× toxicant diluted in HIM for a total volume of 1.2 mL of media per chamber. Both cyclophosphamide and 1-aminobenzotriazole stocks were dissolved in HIM, whereas stock 4-aminophenol was dissolved in DMSO. In the 4-aminophenol experiments, the DMSO concentration was maintained at 0.25% in all wells. Toxicant concentrations were based on previous experiments that used cyclophosphamide 8 and 4-aminophenol. 12 After exposure to the toxicants, the IdMOC plates were placed in a humidified incubator with 5% CO2 at 37 °C for 24 h.
Cell Staining and High Content Analysis
After the 24-h exposure, the media and toxicant were removed, and each chamber was washed with 1.2 mL of phosphate-buffered saline (PBS). Then, 1.2 mL of PBS containing 2 µM ethidium homodimer-1 (dead stain), 1 µM calcein AM (live stain), and 2 µM Hoechst 33342 (nuclear stain) was added to each chamber, and the plates were incubated at room temperature for 1 h.
Image analysis of the IdMOC plates was conducted using the Target Activation BioApplication on a Cellomics Arrayscan VTI HCS Reader (ThermoFisher, Pittsburgh, PA). Three channel measurements were used with 10× magnification and XF93-Hoechst, XF93-FITC, and XF93-TRITC filters for the Hoechst 33342, calcein-AM, and ethidium homodimer-1 stains, respectively. Cells were identified using the nuclear Hoechst 33342 stain, and the cytoplasm was measured using a wider mask relative to the nucleus. Five hundred cells per well were analyzed for calcein-AM and ethidium homodimer-1 fluorescence intensity in addition to nuclear area based on Hoechst 33342 staining. The mean total fluorescence intensity per well, the mean area of Hoechst 33342 stain per well, and the mean number of cells per field were reported in the vHCS:ViewTM software, from which the data were exported and analyzed further in Microsoft Excel 2007 (Microsoft Corporation, Redmond, WA).
Data Analysis
The relative fluorescence units of the live and dead stains were determined by normalizing the average fluorescence intensity of the experimental wells to that of the vehicle control wells. To compare mean responses between data sets, two-way ANOVA was performed followed by Sidak’s multiple comparisons posttest using GraphPad Prism version 6.0 for Windows (GraphPad Software, La Jolla, CA). IC50 values were calculated using nonlinear regression analysis in GraphPad Prism.
Results and Discussion
Multiparametric Analysis of Hepatocyte and 3T3-L1 Co-Cultures for Metabolism and Toxicity Assessment
IdMOC experiments have previously been conducted using endpoints that are evaluated on traditional plate readers. High content analysis, however, vastly improves on the amount and quality of data that are accessible on standard plate-based cell culture experiments by quickly imaging and quantifying multiple parameters on a cell-by-cell basis, and it has been used previously to evaluate hepatotoxicity in monocultures of liver-derived cells.4,5 Because the IdMOC system increases the physiological relevance of typical in vitro experiments by evaluating metabolism and toxicity for multiple cell types in co-culture, we chose to further increase the value of the information generated by the IdMOC system by combining it with high content analysis.
To demonstrate both metabolic detoxification and activation in the IdMOC system, we chose to use the model toxicants 4-aminophenol and cyclophosphamide in the following experiments. The toxic industrial chemical 4-aminophenol has previously been shown to be detoxified in rat hepatocytes and a liver-derived cell line via conjugation to glutathione, acetylcysteine, and glucouronic acid by phase II metabolic enzymes.9,13 We thus expect to observe decreased toxicity in 3T3-L1 cells cultured in the presence of hepatocytes compared to cells in monoculture due to the effects of hepatic metabolism. In contrast, cyclophosphamide is a pro-drug used in chemotherapy that, once metabolized by the phase I cytochrome P450 enzymes of the liver, forms the cytotoxic metabolites 4-hydroxycyclophosphamide and phosphoramide mustard.10,14 In this case, increased toxicity is expected on cyclophosphamide exposure for 3T3-L1 cells cultured in the presence of hepatocytes relative to 3T3-L1 cells cultured alone.
We cultured 3T3-L1 cells alone and in combination with primary human hepatocytes in collagen-coated 96-well IdMOC plates. After 24 h of exposure to various concentrations of 4-aminophenol or cyclophosphamide, the cells were stained with calcein-AM (live stain), ethidium homodimer-1 (dead stain), and Hoechst 33342 (nuclear stain). High content analysis was subsequently performed on the plates using a ThermoFisher Cellomics Arrayscan VTI HCS Reader. We chose a Target Activation Bioapplication method in which the nuclear stain was used to select primary objects and determine the area of the nuclei, and the fluorescence intensities of the live and dead stains were determined for 500 cells per well. In our assay, this analysis allowed for the study of four toxic endpoints: fluorescence intensity of the live and dead stains, nuclear area, and the number of cells per field (cell density). These experiments were repeated for three identical IdMOC plates, with nine wells total per toxicant per concentration.
Generally, as cytotoxicity increases, the fluorescence intensity of the live stain decreases, whereas the intensity of the dead stain increases. In our experiments, the mean fluorescence intensity of the 3T3-L1 cells for both the live and dead stains (
Figs. 1A
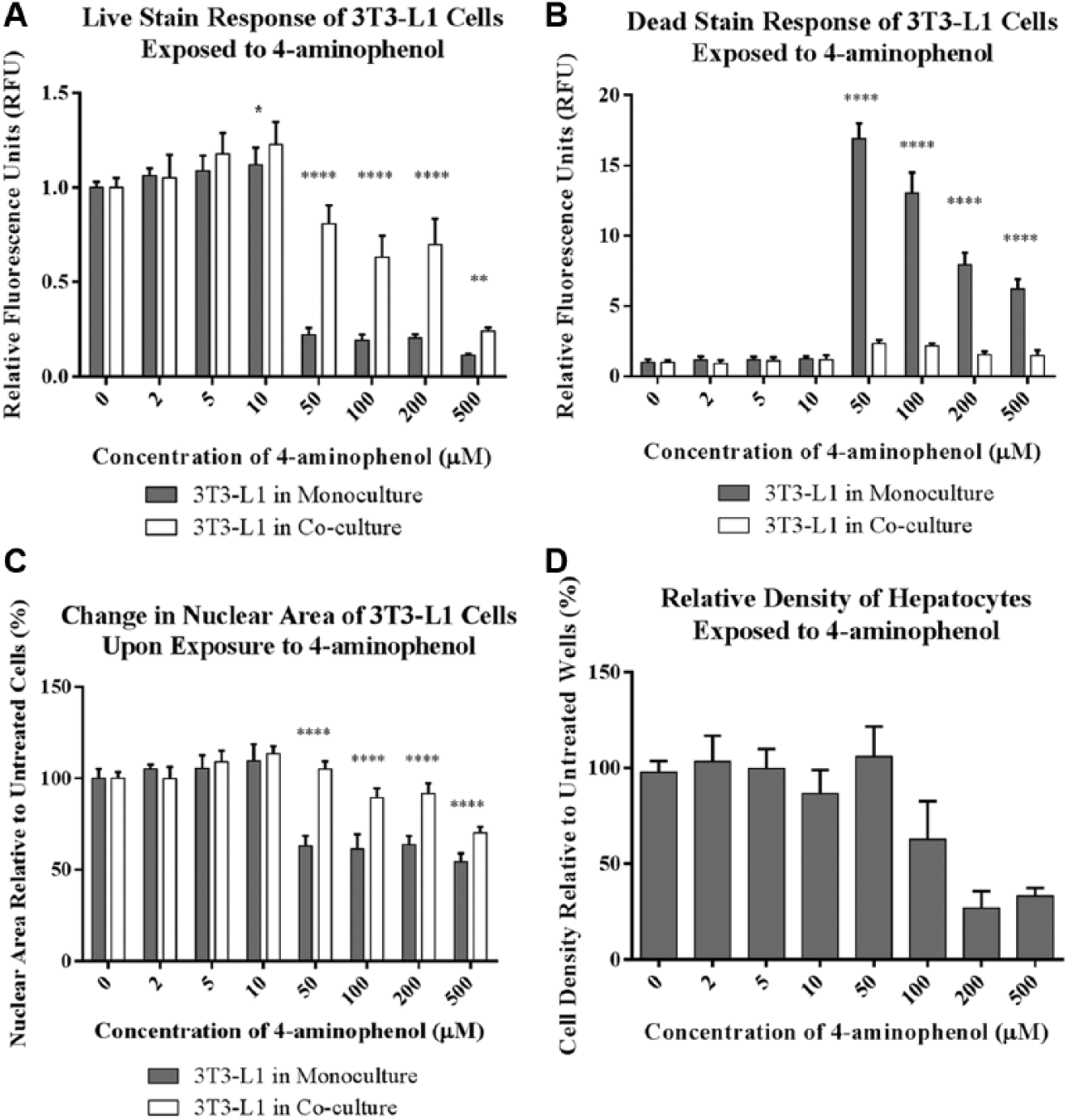
Cytotoxicity measurements conducted using high content analysis of 3T3-L1 cells and primary human hepatocytes exposed to 4-aminophenol in 96-well Integrated Discrete Multiple Organ Co-Culture (IdMOC) plates. The relative fluorescence intensity of the (
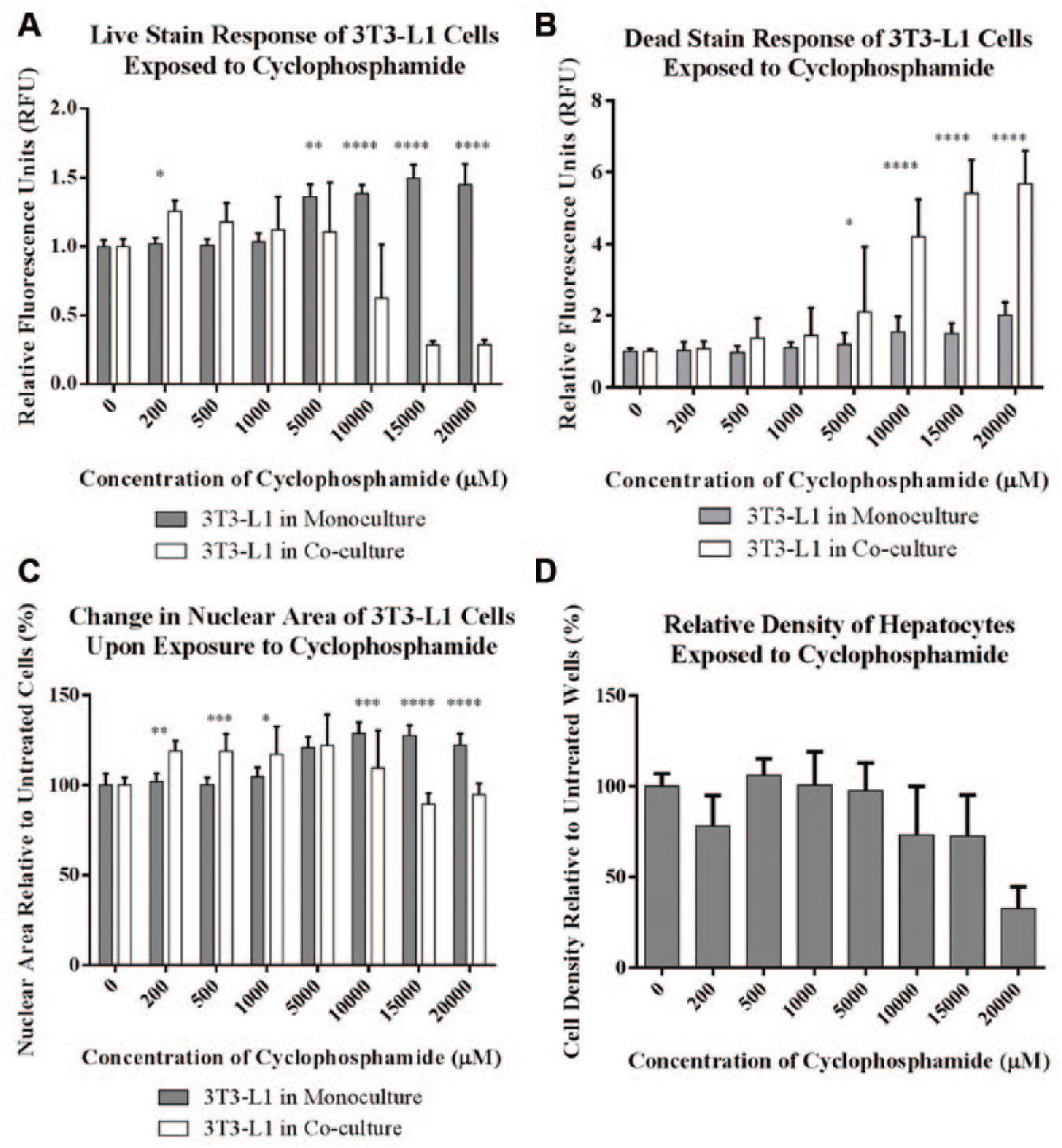
Cytotoxicity measurements conducted using high content analysis of 3T3-L1 cells and primary human hepatocytes exposed to cyclophosphamide in 96-well Integrated Discrete Multiple Organ Co-Culture (IdMOC) plates. The relative fluorescence intensity of the (
IC50 values obtained from the 4-aminophenol and cyclophosphamide dose–response curves of 3T3-L1 cells in monoculture and in co-culture with hepatocytes. The reported values are derived from the live- and dead-stain fluorescence intensities, changes in the nuclear area, and previously published work for comparison. Dash marks indicate the absence of a value, and this is attributable to either the lack of response for an endpoint under the specified conditions or the absence of comparable conditions reported in the literature. Although cyclophosphamide was previously studied under conditions similar to those reported here, 4-aminophenol was previously studied in the pig-derived LLC-PK1 cell line as opposed to the mouse-derived 3T3-L1 cell line.
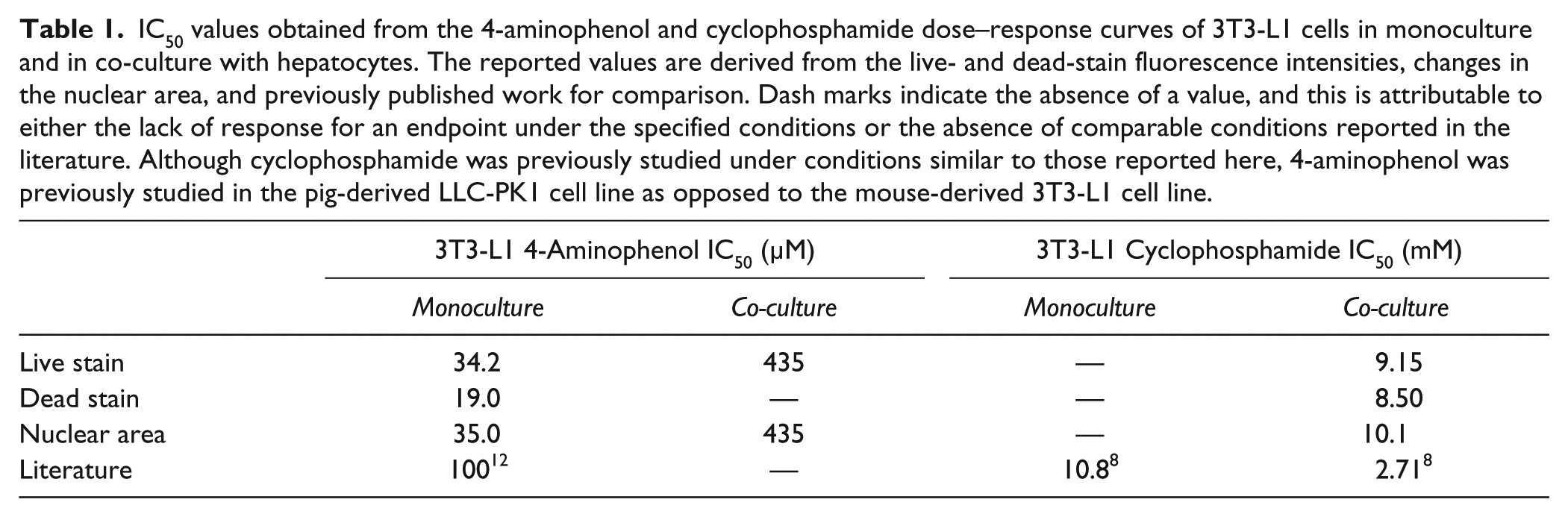

In addition to the fluorescence intensities of the live and dead stains, nuclear area was a valuable endpoint in measuring the cytotoxicity of 3T3-L1 cells in response to 4-aminophenol ( Fig. 1C ), whereas only a slight decrease in the area of the nucleus was observed in response to cyclophosphamide ( Fig. 2C ). Chromatin condensation and nuclear collapse are hallmarks of apoptosis 15 that, in high content analysis and other types of microscopy, are measured as a decrease in the area of the nucleus. We found that the area of the nuclei of 3T3-L1 cells decreased on exposure to 4-aminophenol, and that the IC50 values generated using this endpoint were consistent with those obtained using the fluorescence intensities of the live and dead stains ( Table 1 ). Nuclear size decreased at lower concentrations of 4-aminophenol for 3T3-L1 cells cultured in the absence of hepatocytes compared to cells cultured in the presence of hepatocytes, suggesting that hepatic metabolism decreased the toxicity of 4-aminophenol. Conversely, nuclear size did not change substantially with cyclophosphamide exposure, and, because this morphological effect is dependent on the mechanism of cell death, its utility as a high content endpoint may be limited by the cell type and toxicant under investigation.
Beyond measuring cell death in 3T3-L1 cells cultured with and without hepatocytes, the IdMOC system allows for the evaluation of hepatotoxicity. We found that for the hepatocytes, the intensity of the live stain in response to 4-aminophenol exposure fluctuated from cell to cell, and as such, the average fluorescence response of each well varied considerably (data not shown). We predict that this variability could be attributed to the metabolism of the live stain by the hepatocytes. The area of the nuclei of the hepatocytes decreased on exposure to 4-aminophenol (data not shown), but there was a high degree of variability in these data, likely as a result of the polyploidy that is present in a significant portion of adult primary hepatocytes, or the heterogeneous nature of the hepatocytes that potentially arises from isolation from different zones within the liver. We found that a better endpoint for hepatotoxicity in our system was cell density, or the number of cells per field, as determined in high content analysis by the primary object identification feature of the nuclear channel (
Figs. 1D
Verification of the Effects of Hepatic Metabolism on Toxicity
To verify that the differences in cytotoxicity that we observed in 3T3-L1 cells cultured in the presence or absence of hepatocytes are due to metabolism by the hepatocytes and not caused by some other effect, we applied high content analysis to cytotoxicity studies conducted in the presence of the cytochrome P450 inhibitor, 1-aminobenzotriazole (ABT). In these experiments, we cultured 3T3-L1 cells in 96-well IdMOC plates in the presence of hepatocytes, and we exposed the co-cultures to either 4-aminophenol or cyclophosphamide. Cell staining and high content analysis were performed as described in this article. ABT did not appear to affect the health of the hepatocytes, because the cell density and live staining fluorescence intensity of these cells were not altered by the addition of ABT in control wells (data not shown). Using the fluorescence intensity of the live stain as an endpoint, we observed no difference in the cytotoxicity of 3T3-L1 cells in the presence or absence of ABT on exposure to 4-aminophenol ( Fig. 3A ). This effect is consistent with previous work using primary rat hepatocytes, and it is likely due to the fact that 4-aminophenol is metabolized by phase II–dependent conjugation to glutathione and acetylcysteine instead of oxidation by the cytochrome P450 enzymes. 9
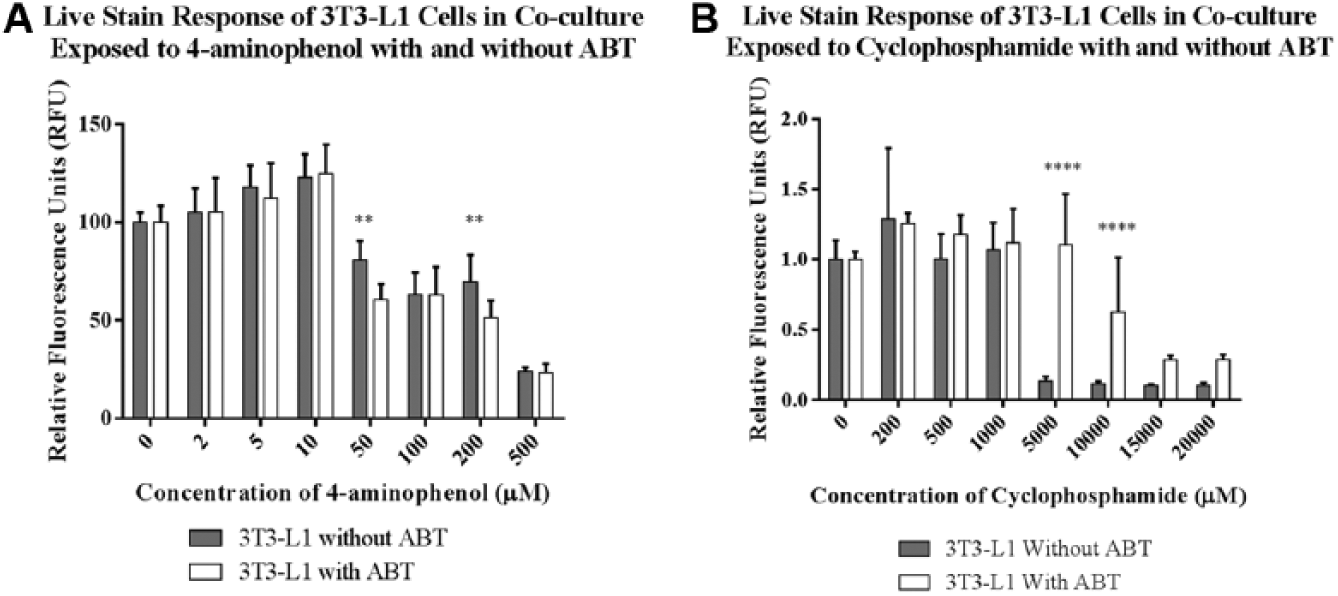
Effects of 1-aminobenzotriazole (ABT) on the cytotoxicity of 4-aminophenol and cyclophosphamide to 3T3-L1 cells co-cultured with primary human hepatocytes in 96-well Integrated Discrete Multiple Organ Co-Culture (IdMOC) plates. The mean fluorescence intensity of the live stain in 3T3-L1 cells, measured using high content analysis, shows that (
To confirm that the hepatocytes activate cyclophosphamide in a P450-dependent manner in our cell culture system, we repeated the cyclophosphamide exposure of the co-cultures with and without ABT. In the absence of ABT, cyclophosphamide exposure to the co-cultures elicited cytotoxicity in the 3T3-L1 cells ( Fig. 3B ). When ABT was present, however, cyclophosphamide was toxic to 3T3-L1 cells only at higher toxicant concentrations than that observed in the absence of ABT, indicating that ABT inhibited cytochrome P450–dependent activation of the pro-drug.
In conclusion, we found that high content analysis of IdMOC plates provides a rapid and multiparametric capability for the study of the effects of metabolism on toxicity in vitro. This method replicated the known in vivo metabolic effects of the model toxicants 4-aminophenol and cyclophosphamide. We found that 4-aminophenol was less toxic to 3T3-L1 cells cultured in the presence versus the absence of hepatocytes, indicating metabolic detoxification of this compound. This detoxification was unaffected by the addition of the P450 enzyme inhibitor ABT, as expected. 9 In contrast, cyclophosphamide was more toxic to 3T3-L1 cells in co-culture than 3T3-L1 cells in monoculture, which is consistent with the in vivo metabolic activation of this pro-drug. The metabolic activation of cyclophosphamide is known to be dependent on the P450 enzymes of the liver, 10 and this was demonstrated in our system by the inhibition of metabolic activation and toxicity via the addition of ABT.
The IC50 values generated from the dose–response curves of 4-aminophenol and cyclophosphamide based on the live stain, dead stain, and nuclear area endpoints were internally consistent, and they fell within threefold of the values reported previously in the literature ( Table 1 ). The dead stain produced an approximately twofold decreased IC50 value for 4-aminophenol in 3T3-L1 monocultures compared to the values obtained using the live stain and nuclear area endpoints. As such, the dead stain may be inherently less reliable than the other endpoints studied, possibly resulting from the fact that dead cells often detach from the cell culture dish and are subsequently washed away during the staining process before image analysis takes place. General discrepancies between the IC50 values reported here and those from the literature may be contributing to differences in the cell type under examination (previously, LLC-PK1 cells were used to study 4-aminophenol exposure in vitro, 12 as opposed to the 3T3-L1 cells used here), differences in the composition of the medium during exposure, or inherent differences in the sensitivities of the endpoints chosen for study.
Although we chose a relatively simple approach in the experiments described herein, using live, dead, and nuclear staining as endpoints, a plethora of other cell type–specific dyes that report toxicity are also appropriate for this type of analysis, including the evaluation of reactive oxygen species, mitochondrial membrane potential, and glutathione content in hepatocytes. In addition, the use of human-derived cell types, particularly primary cells, in co-culture with human hepatocytes in the IdMOC system and analyzed using high content assays will be useful for making human in vivo estimations of toxicity. This first demonstration of the utility of using multiparametric high content analysis in conjunction with the IdMOC cell culture system provides the government and the pharmaceutical industry with the capability to quickly analyze the metabolic and toxicological effects of a large number of compounds in a simple co-culture system with multiple descriptive endpoints appropriate for the effects under consideration.
Footnotes
Acknowledgements
The authors gratefully acknowledge Donna Hoffman for administrative help throughout the course of this work, Dr. Jonathan Oyler for assistance with chemical preparations, Dr. Utkarsh Doshi for providing reagents, and Drs. Crystal Randall and Robert Kristovich for comments on the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the National Research Council Research (NRC) and the Defense Threat Reduction Agency’s Chemical and Biological Defense Program while Stephanie Cole held an NRC Research Associateship Award at the Edgewood Chemical Biological Center.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
