Abstract
Condylomata acuminata (CA), induced by low-risk human papillomaviruses (HPVs), is one of the most common sexually transmitted diseases. The increasing incidence and the high recurrence rate of CA have significantly contributed to public health problems around the world. Because HPVs cannot be cultured in vitro for a long time, there has been little progress in the development of HPV-specific antiviral agents. In this study, we established an HPV11.HaCaT system by introducing the recircularized genome of HPV-11 into HaCaT keratinocytes with transfection techniques and cultured them in a special medium. The existence and replication of HPV-11 DNA were positively detected in established HPV11.HaCaT cells. The HPV-11 DNA in HPV11.HaCaT cells has been stably replicated in definite passages of cells. We preliminarily studied the anti–HPV-11 effects of recombinant human interferon α1b (rhIFN-α) and 13-hexyl-palmatine hydrochloride (HP-13) in HPV11.HaCaT cells. The results suggest that HP-13 significantly inhibited the proliferation of HPV11.HaCaT cells in a dose-dependent manner, whereas rhIFN-α did not. HP-13 and rhIFN-α inhibited the replication of HPV-11 DNA and the expression of E1∧E4 mRNA in HPV11.HaCaT cells. In conclusion, the established HPV11.HaCaT cells can provide us with a convenient and relatively stable tool for screening anti–HPV-11 agents.
Introduction
Human papillomaviruses (HPVs) are a group of small, double-stranded DNA viruses. More than 180 types of HPV DNA have been identified, which can be divided into two categories: the high-risk and low-risk genotypes. 1 The high-risk types (e.g., HPV-16, HPV-18, and HPV-31) are the etiologic agents of cervical cancers, whereas the low-risk types (e.g., HPV-6 and HPV-11) predominantly induce benign genital warts. Until now, most research attention on HPV has been focused on high-risk HPVs because of their carcinogenic capabilities. Although the low-risk HPVs rarely promote carcinogenesis, some of them are commonly known to mediate sexually transmitted diseases (e.g., genital warts). Genital warts [also known as condylomata acuminata (CA)] induced by HPV-6 and HPV-11 are one of the most widespread sexually transmitted diseases. In recent years, studies have shown that the prevalence of CA has increased throughout the world. 2 CA is especially prevalent among the young and sexually active population, and its incidence peaks before 24 years of age in females and between 25 and 29 years of age among males. 3 Although few persisting forms of CA cause mortality like cervical cancer, the increasing incidence and high recurrence rate of CA have contributed significantly to public health problems, heavy economic burdens, productivity loss, and considerable psychological stress such as fear of disfigurement, cancer, and infertility. All of these factors influence the quality of life of the patients and prompt them to take active treatments for CA.
At present, a number of treatment strategies of CA are available, including topical application of drugs (e.g., imiquimod, podophyllin, interferon, and green tea extract) and physiotherapy (e.g., cryotherapy, surgical removal, and carbon dioxide laser).3,4 Effective antiviral medicines (especially specific anti-HPV agents), however, are not available as first-line treatments for CA. Thus, there is an urgent need to develop novel anti-HPV agents.
One of the major obstacles in the development of specific anti-HPV agents is the lack of effective screening systems for testing anti-HPV efficacy, because the papillomavirus cannot be cultured in vitro and cannot be successfully inoculated in an animal body. Although the human epithelium xenograft–immunocompromised mouse system has been successfully developed to study HPV-11 production and the molecular events associated with the pathogenesis of wart formation, there are some limitations in applying this animal model to screen and evaluate anti-HPV agents due to its complexity in operation and time-consuming process.5–7
The complete life cycles of all HPVs are strictly regulated by the differentiation program of infected epithelial cells, which are divided into early stages and virus-productive stages. 8 After HPVs invade the epithelium through skin wounds, the early stage of the life cycle of HPVs occurs in the proliferating basal-layer cells. With virus migration toward the epithelial cell surface, the virus-productive stage of HPVs occurs in differentiating epithelial cells. To date, many attempts have been made to mimic HPV-11 infection in tissues in vitro. Several types of mammalian cells, such as SCC-4 cells (derived from a squamous cell carcinoma), primary human foreskin keratinocytes (HFKs), and immortalized human keratinocytes (N-Tert cells with the catalytic subunit of human telomerase and HaCaT cells),9–12 have been used to establish HPV-11 DNA–containing cells by introducing gene segments, from the linear or circularized genome of HPV-11, with electroporation or chemical transfection techniques. In these cells, the replication of HPV-11 DNA in transient- or definite-passage cell cultures has been observed. In a few studies, HPV-11 transfectants exhibited altered differentiation patterns in organotypic raft cultures. For an in vitro model for screening antiviral activities, however, some important factors have to be considered. First, the in vitro model should be as close as possible to the natural infection status. Second, the in vitro model should be repeatable, and the established cells can be conveniently stored for future use. In addition, HPV-11 genomes in the established host cells should be stably maintained and replicated in long-term cultures or in multiple passages. It is very important to select reliable host cells for establishing in vitro screening models. The HFK is not an ideal candidate because of the lack of homogeneity and the finite lifespan. Immortalized keratinocytes (e.g., HaCaT) exhibit many characteristics similar to primary or early-passage HFKs. 13 Our previous studies have shown that HaCaT keratinocytes appeared differentiated in the organotypic raft culture system. 14 Because they are nontumorous and easy to maintain in cell culture for a long time, HaCaT keratinocytes could be a suitable host cell line for establishing an in vitro screening model for anti–HPV-11 agents.
In this study, we developed an in vitro screening system for anti–HPV-11 activities. We established HPV11.HaCaT by transfecting a circularized HPV-11 genome into HaCaT cells. To assess the stability of cells containing the HPV-11 genome, we monitored the replication of HPV-11 DNA and the expression of E1∧E4 viral gene in different passages of fresh and thawed cells after frozen storage in liquid nitrogen. We also used HPV11.HaCaT to preliminarily study the anti–HPV-11 activities of recombinant human interferon α1b (rhIFN-α), which has been used in the clinical treatment of CA. We also explored the potential anti–HPV-11 effects of two compounds, 13-hexyl-berberine hydrochloride (HB-13) 15 and 13-hexyl-palmatine hydrochloride (HP-13), which have shown obvious anti–herpes simplex virus effects in our previous studies (data not shown).
Materials and Methods
Circularized HPV-11 DNA Preparation for Transfection
The pBR322.HPV11 plasmid [12,294 base pairs (bp)] from Escherichia coli (ATCC No. 45151, ATCC, Manassas, VA, USA) contains the full-length HPV-11 genome inserted into the BamHI site of pBR322. Cultivation of E. coli with the pBR322.HPV11 plasmid was performed in Luria–Bertani (LB) culture medium supplemented with 50 mg/L ampicillin to ensure plasmid stability. Plasmid DNA was extracted and purified by the PureLink™ HiPure Plasimid DNA Purification Kits (Invitrogen, Carlsbad, CA, USA). Then, the purified plasmid DNA was digested with BamHI enzyme (Promega, Madison, WI, USA) to release viral genome from the vector at the BamHI site of pBR322. The linear full-length HPV-11 DNA was collected from the low-melting-point agarose gel by electrophoresis and was purified by the Biospin Gel Extraction Kit (Bio Flux, Tokyo, Japan). The linear genome was self-circulated with T4 DNA ligase (Promega). All the target DNA products in the experiments were evaluated by 0.6% agarose gel electrophoresis and ethidium bromide staining with a DNA ladder.
Cell Culture and DNA Transfection
The HaCaT cells, a spontaneously transformed nontumorigenic human keratinocyte cell line, 16 were routinely cultured in fresh Dulbecco’s Modified Eagle Medium (DMEM; Gibco Laboratories, Grand Island, NY, USA) supplemented with heat-inactivated 10% (v/v) fetal bovine serum (FBS), 100 U/mL penicillin, and 100 μg/mL streptomycin (pen/strep) at 37 °C in a humidified atmosphere at 5% CO2. Both 3 μg of the circularized HPV-11 DNA and 1 µg of pTK-neo DNA (Invitrogen) were transfected into HaCaT cells on a six-well plate using Lipofectamine™ 2000 reagent (Invitrogen) following the manufacturer’s instructions. After selection with 500 μg/mL G418 (Amersco, Solon, OH, USA) for about one week after transfection and then growing in media containing 250 μg/mL G418, the resulting G418 colonies (HPV11.HaCaT) were pooled as a cell population.
HPV11.HaCaT Monolayer Culture
After the pooled cell populations (HPV11.HaCaT) were expanded, cells were grown to about 80% confluence in fresh DMEM supplemented with 10% FBS, calcium chloride at 0.03 mM final concentration, and without pen/strep at 37 °C in a humidified atmosphere at 5% CO2 for subsequent analysis.
HPV-11 DNA Examination by Oligonucleotide Primed in Situ Labeling Test
Cell suspension of HPV11.HaCaT in phosphate-buffered saline and the cell smears were freshly prepared. The cells on the slides were fixed with 4% paraformaldehyde for 30 min and air-dried at room temperature. The reaction mixture for the primed in situ labeling (PRINS) technique (total volume: 100 µL) consisted of 1 mM of each of dATP, dCTP, dGTP, dTTP, and DIG-11-dUTP (Roche, Mannheim, Germany); 50 µM of primer HPV-11 (5′-ATG CCT CCA CGT CTG CAA C-3′); 5 U/µL of Taq DNA polymerase (Promega); 10 µL 10× polymerase chain reaction (PCR) buffer; 25 mM MgCl2; and distilled water. The slides were heated for 1 min at 94 °C before application of 15 µL of the PRINS mixture onto each slide. Then, the slides were heated for 5 min at 94 °C and incubated for 5 min at 65 °C. The reaction was stopped by washing slides for 3 to 5 min in stop buffer at 65 °C. The slides were incubated in blocking buffer for 30 min at 37 °C and incubated with biotin–SP-conjugated mouse antidigoxin antibody (Boster Co., Ltd. Wuhan, China) for 2 h at room temperature, then they were washed four times for 5 min with 0.5× tris-buffered saline (TBS buffer) and incubated with SABC-AP (Boster Co., Ltd.) at 37 °C for 30 min. After washing again with 0.5× TBS buffer, the slides were stained with BCIP/NBT (Boster Co., Ltd.) for 20–30 min at 37 °C. The results were examined with the microscope.
Fluorogenic Quantitative PCR Analysis for HPV-11 DNA
The HPV11.HaCaT cells (about 2×104 cells in each sample) were washed three times with phosphate-buffered saline, and total DNA was extracted. Fluorescent quantitative PCR (FQ-PCR) was performed using the HPV type 6 and 11 kits (Da’an Gene, Guangzhou, China) in accordance with the manufacturer’s instructions. The primer sequences for HPV-11 were shown as follows: 5′-ATG CCT CCA CGT CTG CAA C-3′ (forward) and 5′-CTC TGC CGG TGG TCA GTG CAT-3′ (reverse). An HPV-6/-11 positive quantitative control sample (1×106 copies/μL) was obtained from the HPV type 6 and 11 kits. Serially diluted samples (dilution factors of 100, 1000, 10,000, and 100,000) were submitted to an FQ-PCR. Two microliters of total DNA was used in a 40 μL reaction mixture, and conditions were 93 °C for 2 min at stage 1 and 40 cycles of 93 °C for 45 s and 55 °C for 120 s at stage 2. There existed a linear relationship between the CT values and logarithms of the starting numbers of target-specific products in the standard curves ( Fig. 1A ). Cells were counted with a hematocytometer, and the tested samples were prepared from the cell suspension. The resulting CT values of the tested samples were used to calculate the starting numbers of target-specific products according to the standard curves. Then, the averaged copies of HPV-11 in each cell were calculated by absolute quantification using the standard curves and the cell count results. Normal HaCaT cells without HPV-11 DNA were used as a blank control.
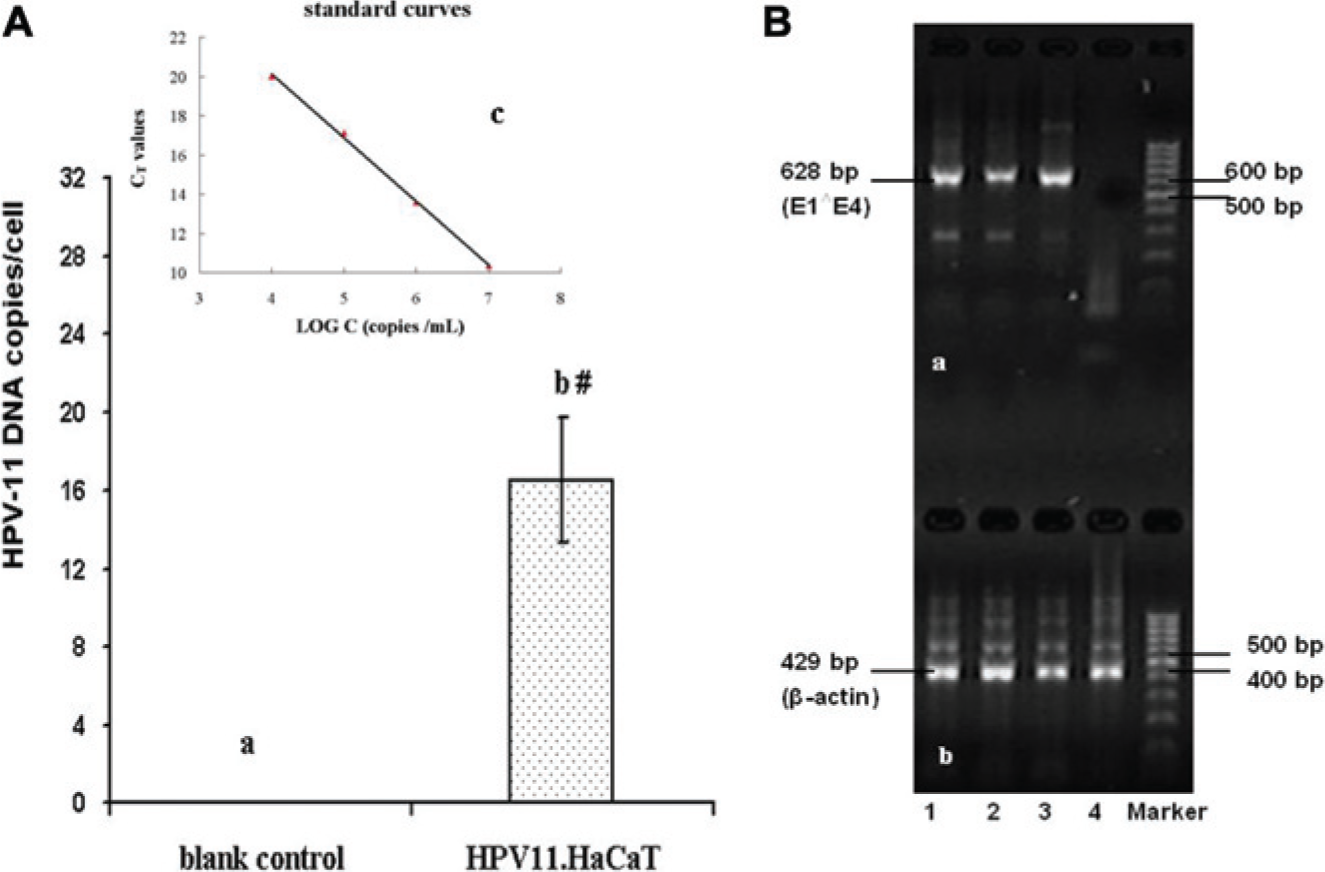
Detection of human papillomavirus (HPV)-11 DNA or HPV-11 E1∧E4 transcripts from the established HPV11.HaCaT cells. Total DNA or total cellular RNA, extracted from three separate expanded HPV11.HaCaT cell populations, was subjected to fluorescent quantitative polymerase chain reaction (FQ-PCR) detection or nested reverse transcription PCR (RT-PCR) analysis. (
Total RNA Isolation and Nested RT-PCR for HPV-11 E1∧E4
HPV11.HaCaT cells were seeded in six-well plates (1×106 cells each well) and grown to about 80% confluence. Total cellular RNA was extracted from the HPV11.HaCaT samples using TRIZOL reagent (Invitrogen), and cDNA was synthesized using the Reverse Transcription System (Promega). The products were amplified by nested primer sets and two 30-cycle PCR rounds with the outer primer and inner primer, respectively, to detect the HPV-11 E1∧E4 spliced transcript. The primers are described in Table 1 . Ten microliters of cDNA were used for the first 30-cycle reaction, and 2 µL of the first PCR product was used for a second 30-cycle round of PCR. The temperature file is as follows: first, 2 min at 94 °C, then 30 cycles of 30 s at 94 °C, 30 s at 60 °C, and 55 s at 72 °C, with a final extension of 10 min at 72 °C. Normal HaCaT cells without HPV-11 DNA were used as a blank control.
Oligonucleotide Primer Used in Nested PCR Assays.
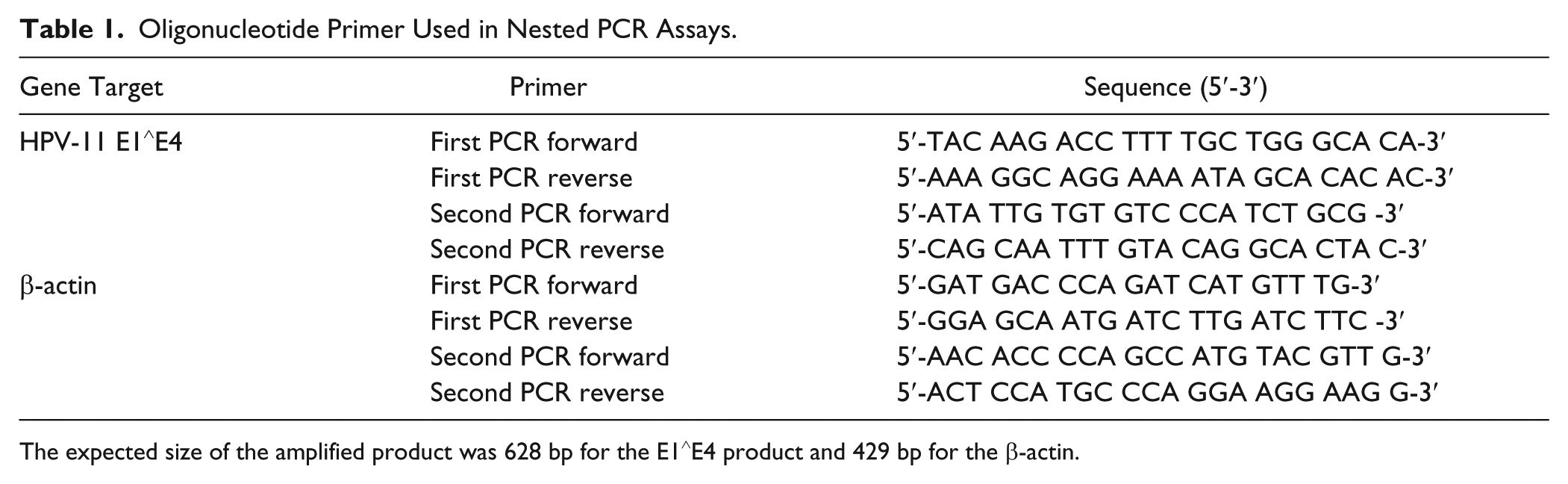
The expected size of the amplified product was 628 bp for the E1∧E4 product and 429 bp for the β-actin.
Detection of Stabilities of HPV-11 DNA Replication and E1∧E4 mRNA Expression of HPV11.HaCaT Cells in Different Passages and Thawed Cells from Liquid Nitrogen Storage
The HPV11.HaCaT cells were passaged repeatedly after being grown to 80–90% confluence until passage 15 (P15). In addition, the HPV11.HaCaT cells were stored in liquid nitrogen for one month, and then were thawed and followed passages until passage 18 (P18). Extraction of total DNA and RNA of indicated passage cells (P1, P3, P5, P10, and P15) or the thawed and followed passage cells (P2, P5, P10, P15, and P18), detection of HPV-11 DNA replication by FQ-PCR, and E1∧E4 mRNA expression by nested RT-PCR were performed using the protocol described in this article.
Cytotoxicity Analysis
HB-13 (purity of 99.36%) and HP-13 (purity of 97%) were kindly provided by Professor Nai-san Li from China Pharmaceutical University (Nanjing, China). Chemical structures of HB-13 and HP-13 are shown in Figure 2 . RhIFN-α was purchased from Kexing Co., Ltd. (Beijing, China). The HPV11.HaCaT cells were seeded in triplicate onto 96-well plates at a density of 5×104 cells per well. After 24 h, the medium was replaced with medium containing various concentrations of tested agents. After incubating for 24 h at 37 °C, 5% CO2, the cell viability was determined by an MTT assay.
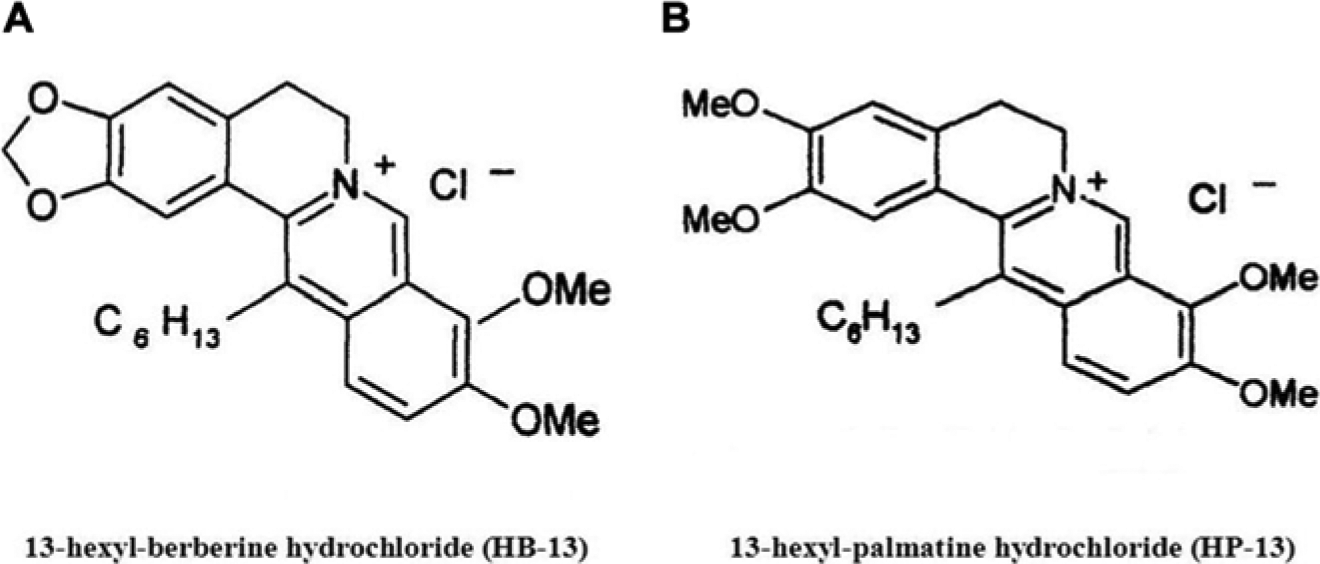
The chemical structures of HB-13 and HP-13. (
Antiviral Agent Treatment Assays
Detection of the Effects on HPV-11 DNA Replication and E1∧E4 mRNA Expression in HPV11. HaCaT Cells Treated by Tested Agents
The HPV11.HaCaT cells were seeded onto six-well plates at a density of 1×106 per well and incubated for 24 h at 37 °C, 5% CO2. After removing the supernatant, the cells were treated with indicated concentrations of tested agents for 16 h. Extraction of total DNA and RNA, detection of HPV-11 DNA replication by FQ-PCR, and E1∧E4 mRNA expression by nested RT-PCR were performed by protocol as described in this article. Normal HaCaT cells without HPV-11 DNA were used as a blank control.
Statistical Analysis
Results were expressed as mean ± standard deviation and analyzed using a Student t test. P values lower than 0.05 were considered statistically significant.
Results
Construction of HPV-11 Episomal DNA Containing HaCaT Cells
We first prepared circular HPV-11 DNA for transfection. The pBR322.HPV-11 plasmid was extracted and purified from E. coli after culturing and amplifying ( Fig. 3A1 , lanes 1, 2, and 3; 12,294 bp). The linear HPV-11 genome was released from pBR322 by digestion with the BamHI enzyme ( Fig. 3A2 , lane 3; 4363 bp for pBR322 and 7931 bp for linear full-length HPV-11 DNA) and purified with low-melting-point agarose gel electrophoresis and gel extraction ( Fig. 3A2 , lane 2; 7931 bp for purified linear full-length HPV-11 DNA). The linear full-length HPV-11 DNA was collected and recircularized with T4 DNA ligase ( Fig. 3A2 , lane 1; 7931 bp for circular HPV-11 DNA). HaCaT cells were transfected, together with pTK-neo DNA encoding a neomycin resistance gene, by using lipofectamine with circular HPV-11 DNA as described in the Materials and Methods section. Following selection for neomycin resistance, the resulting positive colonies were pooled and expanded as a cell population.
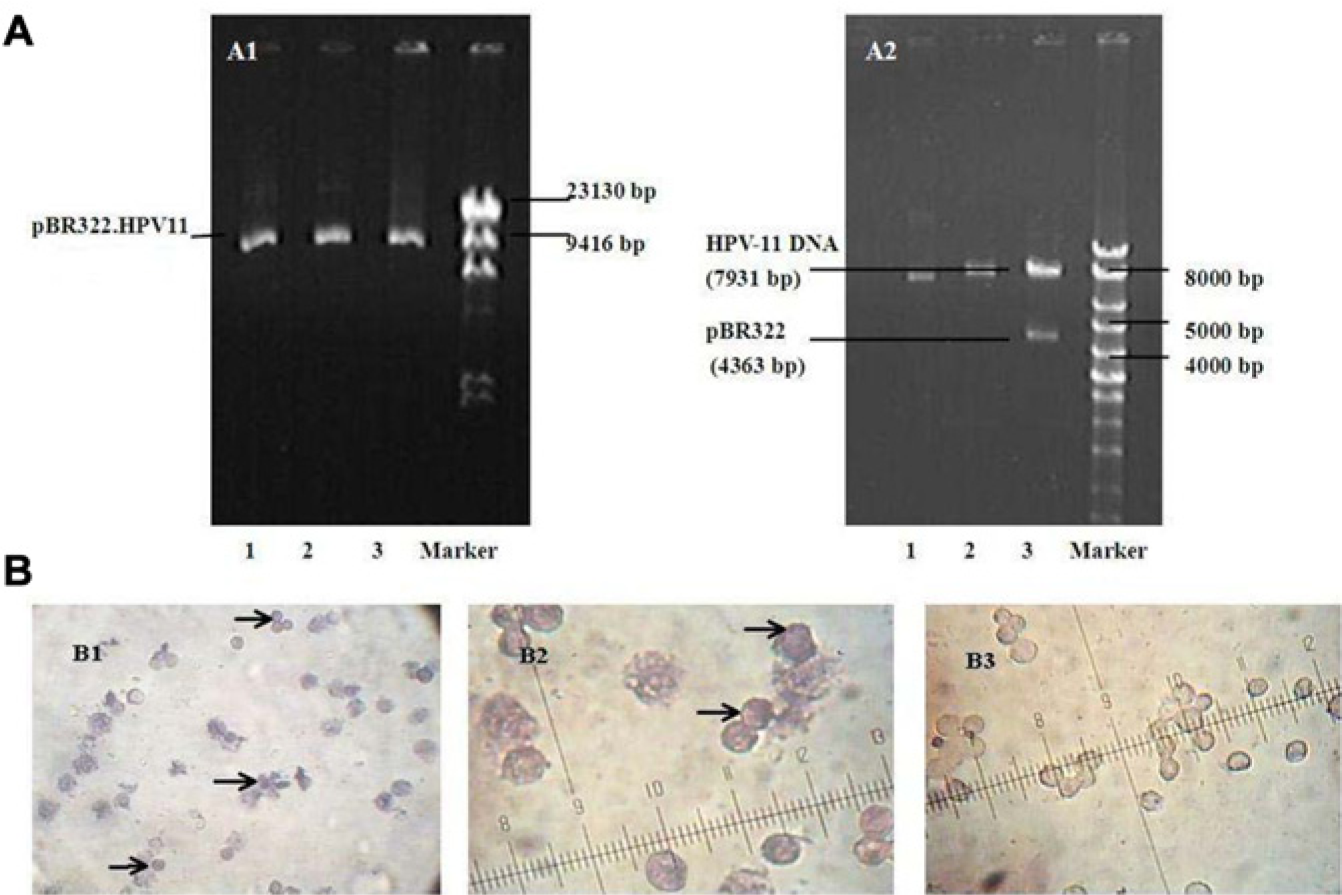
(
Detection of HPV-11 DNA from the Established HPV11.HaCaT Cells
Analysis of HPV-11 DNA Replication in the Expanded HPV11.HaCaT Cell Populations
The PRINS technique was applied to determine HPV-11 DNA in expanded cell populations. HPV-11–positive signals, shown by blue-black nuclear staining, were detected in HPV11.HaCaT smears ( Fig. 3B1 , 3B2 ), whereas the nuclear signals for HPV-11 exhibited negative staining in the normal HaCaT cells ( Fig. 3B3 ).
In addition, FQ-PCR analysis of three separate expanded HPV11.HaCaT cell populations showed a significant increase (p < 0.05, Student t test) in HPV-11 DNA copy numbers in each HPV11.HaCaT cell (16.5±3.2, Fig. 1A , column b) compared to blank controls (normal HaCaT cells without HPV-11 DNA).
Expression of HPV-11 E1∧E4 mRNA in Established HPV11.HaCaT Cells
To confirm the expression of HPV-11 E1∧E4 mRNA in established HPV11.HaCaT cells, total RNA samples isolated from three separate expanded HPV11.HaCaT cell populations were subjected to nested RT-PCR analysis. Results showed that the spliced E1∧E4 transcripts were detectable in the populations ( Fig. 1B , lanes 1, 2, and 3; 628 bp).
HPV11.HaCaT Cells Can Stably Maintain HPV-11 Episomes for Multiple Passages
We assessed the stability of HPV11.HaCaT cells in maintaining HPV-11 episomes for various passages, including in thawed cells following passages, by detecting copy numbers of HPV-11 DNA per cell and the expression of E1∧E4 mRNA. As shown in Figure 4A , FQ-PCR analysis revealed that the average copy number declined gradually during passage 3 (P3) to P10 and maintained at approximately 15–30 copies per cell over the first 10 passages. There was a sharp decline between P10 and P15. In thawed and followed passage HPV11.HaCaT cells, copy numbers (per cell) of HPV-11 DNA were slightly reduced compared with those of unfrozen cells during P2 to P10. After P10, there was a sharp decline pattern in the copy number of HPV-11 DNA similar to that seen in unfrozen cells ( Fig. 4B ).
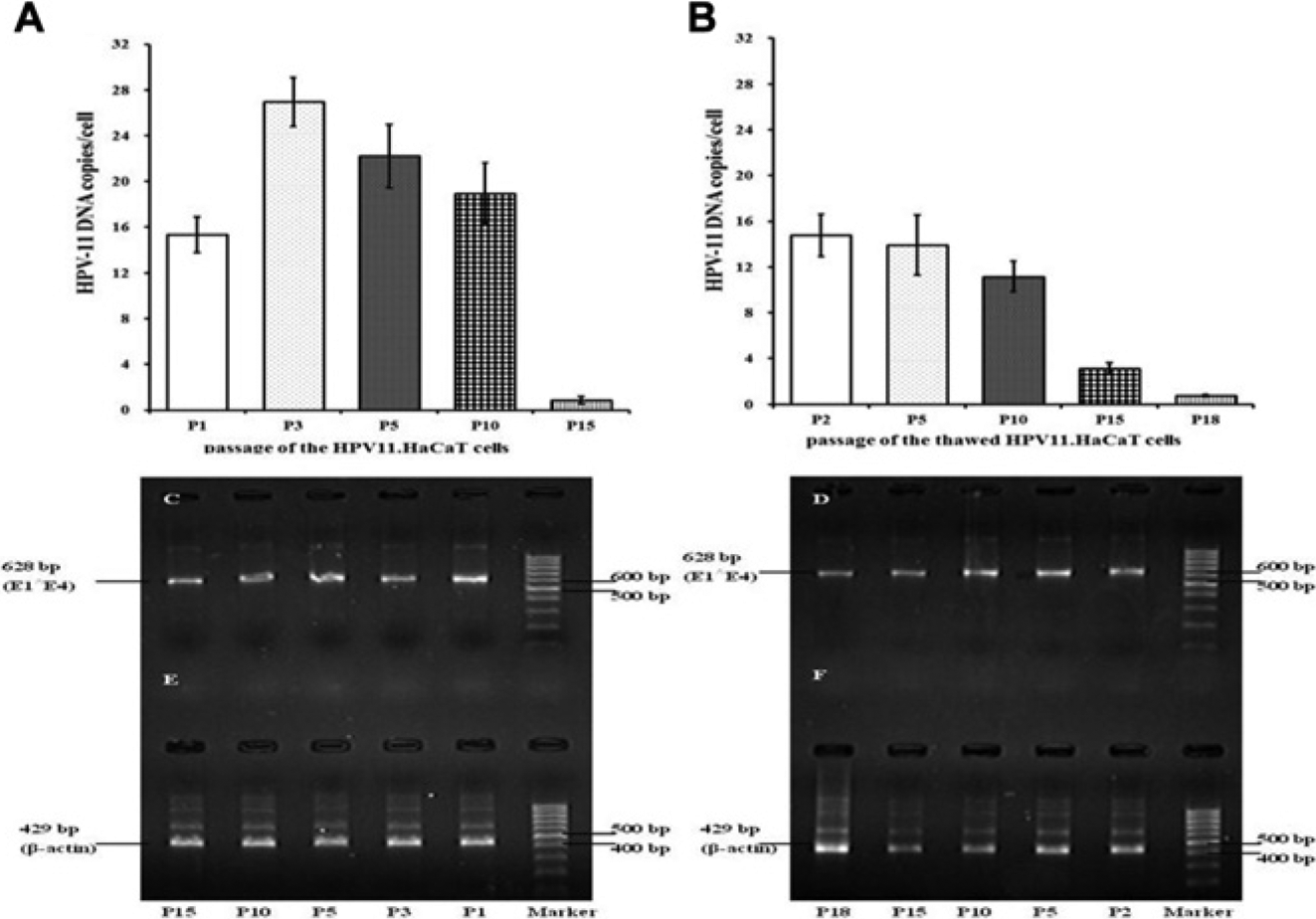
Detection of human papilloma virus (HPV)-11 DNA copy number by fluorescent quantitative polymerase chain reaction (FQ-PCR) or HPV-11 E1∧E4 transcripts by nested reverse transcription (RT)-PCR analysis in established HPV11.HaCaT cells after serial passages. Total DNA or total RNA extracted from the various passages of HPV11.HaCaT cells was subjected to FQ-PCR detection or nested RT-PCR analysis. (
The examination of HPV-11 E1∧E4 mRNA expression in consecutive passages revealed relatively stable transcriptional activities in all samples. As shown in Figure 4C , the spliced E1∧E4 transcripts (628 bp) were detectable from different passages of the HPV11.HaCaT cells (lanes P1, P3, P5, P10, and P15), at least up to passage 15, without obvious loss of HPV-11 E1∧E4 mRNA expression. After preservation in liquid nitrogen, HPV-11 E1∧E4 mRNA (628 bp) in the thawed HPV11.HaCaT cells was still steadily expressed following many passages ( Fig. 4D , lanes P2, P5, P10, and P15), except for only a small decrease in passage 18 ( Fig. 4D , lane P18).
Both the analysis of FQ-PCR and RT-PCR showed that established HPV11.HaCaT cells stably maintained HPV-11 episomes after many passages.
Effects of Proliferation of HPV11.HaCaT Cells Treated by Tested Agents
An MTT assay was used to analyze metabolic activity in proliferating cells. MTT assay results of HPV11.HaCaT cells treated by HB-13, HP-13, and rhIFN-α are shown in Figure 5 (A , B ). HB-13 and HP-13 significantly inhibited proliferation of HPV11.HaCaT cells in a dose-dependent manner at the concentration range of 0.78–6.25 μg/mL. Furthermore, HB-13 showed 60% cell survival at 0.78 μg/mL, which suggested its higher cytotoxicity at concentrations of 0.78–6.25 μg/mL, and it could not be used to subject cellular DNA and RNA tests. RhIFN-α at the concentration range of 0.312×106−2.5×106 U/mL had a mild effect on the proliferation of HPV11.HaCaT cells with more than 80% cell survival.
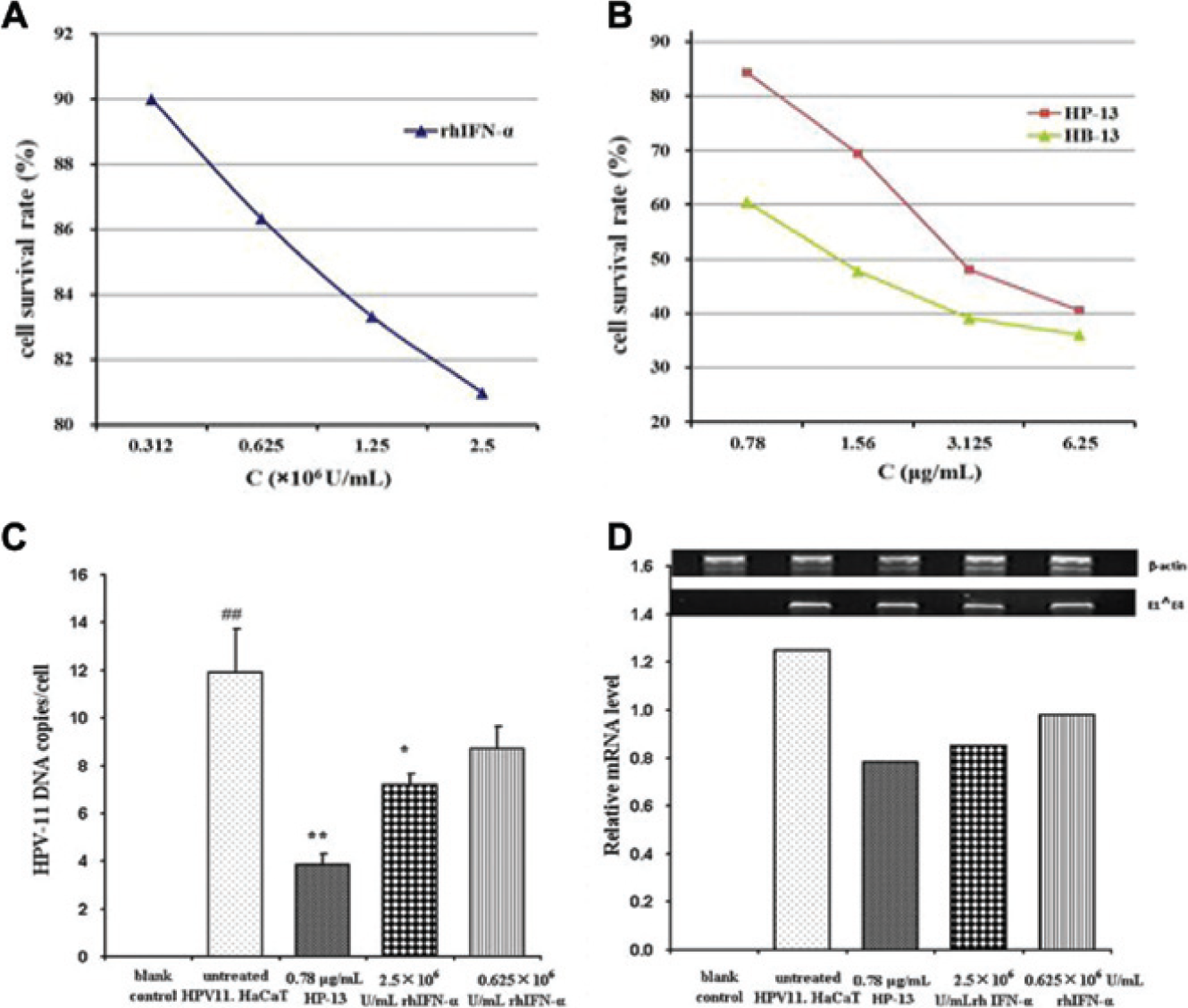
Effects of proliferation, human papillomavirus (HPV)-11 DNA replication, and E1∧E4 mRNA expression of HPV11.HaCaT cells treated by tested agents. (
Inhibition of HPV-11 DNA Replication and E1∧E4 mRNA Expression by Tested Agents
Based on the results of MTT tests, we chose HP-13 and rhIFN-α at the concentration without significant effects on the proliferation of HPV11.HaCaT cells for HPV-11 DNA replication and E1∧E4 gene expression tests. FQ-PCR results showed that HP-13 at 0.78 µg/mL and rhIFN-α at 2.5×106 U/mL significantly decreased the averaged copy number of HPV-11 DNA in each HPV11.HaCaT cell (68% and 39% inhibition, respectively) as compared to untreated HPV11.HaCaT cells (p < 0.01 and p < 0.05, respectively). RhIFN-α at 0.625×106 U/mL had no significant effects on the averaged copy numbers of HPV-11 DNA in each HPV11.HaCaT cell ( Fig. 5C ). Similarly, treatments of HP-13 at 0.78 μg/mL and rhIFN-α at 2.5×106 and 0.625×106 U/mL induced a general reduction in HPV-11 E1∧E4 mRNA expression ( Fig. 5D ).
Discussion
In this article, we developed a convenient, relatively stable, and reproducible cell culture system for screening or evaluating anti–HPV-11 activities in vitro. Our approach has shown significant improvements as compared with those currently used in the field.
Here, we chose HaCaT cells with nearly normal keratinocyte phenotypes, rather than HFKs, as the host because of some clear advantages, such as easiness of propagation, high homogeneity, and the extended lifespan. 13 Recent data also showed that the basal level of autophagy in HaCaT keratinocytes was much lower than that in primary keratinocytes, which might contribute to a better survival of HPV in HaCaT. 17 In addition, our previous work also provided evidence for the virus-productive stages of the HPV-11 life cycle in HPV11.HaCaT cells grown on the raft cultures. In a reconstructed three-dimensional structure of the epithelium, we observed L1 capsid protein expression of HPV-11. 14 Given the advantages and the ability of HaCaT cells to support the HPV-11 life cycle, the cells may be suitable host cells as an additional alternative for investigating the life cycle of HPV-11 and evaluating anti–HPV-11 effects.
HPV-11 is a double-stranded and circular DNA virus that infects stratified epithelial keratinocytes and replicates as an episome without integrating into chromosomes of its host. 18 To closely mimic the natural status of HPV-11 infection, we recircularized HPV-11 DNA and introduced it into HaCaT keratinocytes with pTK-neo DNA encoding a neomycin resistance gene, which made it easy to select cells harboring HPV-11 DNA. The results of PRINS and FQ-PCR indicated that HPV-11 DNAs could be partitioned to daughter cells during mitosis. This result was also supported by the evidence of expression of HPV-11 E1∧E4 gene from three separate expanded HPV11.HaCaT cell populations. We cannot entirely rule out the possibility, however, that these positive cultures also contained cells without HPV-11 DNA and with only pTK-neo DNA, which is a question requiring further clarification.
In the early stage of the life cycle of HPV, infection occurs in the mitotically active basal layer of epithelia, where the HPV-11 genome establishes long-term persistent infections. This phase is characterized by the continuous and latent presence of viral DNAs at a low copy number but the absence of viral production. 19 This stable maintenance of the HPV genome in host cells is thought to be achieved by replicating its DNA like the cellular DNA. 20 It is thought that the persistent and latent infections lead to an effective evasion of defensive system of the host cells, which results in the difficulty in thoroughly curing CA and the high recurrence rate after the initial remission following treatments.9,21 Thus, the inhibition of the replication of HPV-11 DNA by anti-HPV agents at this stage may help the clearance of virus from the infected epithelial cells. To develop an effective in vitro screening system for anti–HPV-11 drugs, the critical step depends on the establishment of the replication and amplification of viral DNA in host cells. To achieve this goal, we have chosen a culture medium with a low calcium concentration to culture the HPV11.HaCaT cells because HaCaT could be stably maintained in a basal-like state for long periods in this medium. 13 Next, we have chosen reliable parameters or indicators to determine the replication and amplification of viral DNA in infected cells. In this article, we have examined the averaged number of HPV-11 DNA copies in each cell as a direct measurement of the replication and amplification of HPV-11 DNA. In addition, the E1∧E4 transcript, expressed early and throughout in the HPV-11-infected epithelium, correlates with the onset of viral-genome amplification. 22 Therefore, the expression of E1∧E4 mRNA can be another indicator. Both the averaged number of HPV-11 DNA copies and E1∧E4 mRNA expression levels can be easily detected with quantitative or semiquantitative FQ-PCR or nested RT-PCR techniques, which are rapid, sensitive, and specific for anti-HPV activity screening. It is noteworthy that a significant difference in the growth rate between HPV11.HaCaT cells and normal HaCaT cells is detected with the cell growth curve assay and cell cycle analysis using flow cytometry (data not shown). We speculate that this difference may be due to changes in cell cycles resulting from the HPV-11 DNA.
The replication activities of HPV-11 DNA may also contribute to the stability of HPV-11 episomes in HPV11.HaCaT cells, just as the virus establishes long-term persistent infections through HPV DNA replication. In this report, the slight changes in the copy number in each cell during the first 10 passages indicate that the HPV-11 DNA could establish continuous replication to achieve a relatively steady-state level during a definite number of passages of HPV11.HaCaT. Although the replication of HPV-11 DNA in the thawed and subsequent passages of HPV11.HaCaT were slightly reduced as compared with unfrozen cells, HPV-11 DNA was still maintained at a relatively stable level (about 11–15 copies per cell) during P2–P10. This stability was also supported by the results on the expression of HPV-11 E1∧E4 mRNA in various passages of HPV11.HaCaT cells and the thawed ones. HPV11.HaCaT cells stably carrying HPV-11 episomes in definite passages could provide us with a quick and convenient tool for anti–HPV-11 drug screening.
Meanwhile, the effectiveness of HPV11.HaCaT cells as a screening tool for antiviral agents should be verified with clinically approved anti-HPV drugs. There are, however, no first-line anti-HPV drugs for CA to date. In this article, we preliminarily examined the anti–HPV-11 effects of HB-13, HP-13, and rhIFN-α with the HPV11.HaCaT cells.
Berberine is a quaternary ammonium salt from the protoberberine group of isoquinoline alkaloids, which is usually found in the roots, rhizome stems, and bark of such plants as Berberis, Berberis vulgaris, Coptis chinensis, and so on. Some studies with berberine have shown that berberine could inhibit the growth of certain tumor-derived cell lines, some bacteria, and several viruses.23–25 Recent research has shown that berberine can block the expression of oncoproteins E6 and E7 and result in a concomitant increase in p53 and pRb expression in the HPV-16–positive and HPV-18–positive cervical cancer cell lines. 26 HB-13 is a derivative with a 13-hexyl group on berberine parent structure. HP-13 is a derivative with a 13-hexyl group on palmatine parent structure, and its molecular structural formula is similar to that of HB-13. It has been reported that the addition of a 13-hexyl group on berberine parent structure showed better anti-HIV activities. 27 Our previous works have shown that HB-13 15 and HP-13 have anti–herpes simplex virus activities (data not shown). In this article, we found that HB-13 showed a higher cytotoxicity in HPV11.HaCaT cells at the concentration range of 0.78–6.25 μg/mL. HP-13 significantly inhibited the proliferation of HPV11.HaCaT cells in a dose-dependent manner at the concentration range of 0.78–6.25 μg/mL, and the results suggested that HP-13 might suppress HPV-11–containing epithelial cells in CA treatment. FQ-PCR and nested RT-PCR results further demonstrated that HP-13 was efficient in inhibiting the replication of HPV-11 DNA and expression of E1∧E4 mRNA, which indicates the potential therapeutic value of HP-13 in the treatment of CA.
IFN-α is a cytokine that exhibits potent antiviral, antiproliferative, and immunomodulatory properties. IFNs elicits broad antiviral activities, including blocking de novo protein synthesis, activating proteolysis of abnormal proteins by inducing the IFN-stimulated gene ubiquitin-like protein, and causing activation of RNase L and disaggregation of polysomes. 28 RhIFN-α has been used to treat CA and obtained mixed results, showing unsatisfactory clinical efficacy. 29 Previous findings have revealed that IFN-α inhibits the proliferation of HPV-16 immortalized cervical epithelial cells by potently suppressing the expression of viral E6 and E7 oncoproteins and preserving p53 and pRB activities.30,31 Our results have shown that rhIFN-α at a dose of 2.5×106 U/mL had a mild effect on the viability of HPV11.HaCaT, which suggested rhIFN-α did not inhibit the proliferation of HPV-11–containing epithelial cells. RhIFN-α significantly reduced the copy number of HPV-11 DNA per cell and the expression of E1∧E4 mRNA with inhibition ratios of 39% and 32%, respectively. These results suggested that rhIFN-α treatment has some antiviral effects on the latently infected cells in. These data also suggested that the clinical efficacy of rhIFN-α in CA treatment could be due to its combined actions of antiviral activity, antiproliferative activity, immunoregulation, and so on. Although a study by others has shown only marginal efficacy of α-IFN in another culture system established with SCC-4 cells, this discrepancy may be due to the use of a different IFN-α and cell line. 9
In conclusion, we have developed a convenient and relatively stable in vitro cell model that may be used to screen anti–HPV-11 drugs. Our data have demonstrated that this system is easy to operate, and convenient to store and reproduce. The constructed HPV11.HaCaT also provides valuable cytological materials for the study of the etiology of HPV-11–induced infection. Nevertheless, further investigations are required to confirm the stability and practical application value of the model.
Footnotes
Acknowledgements
The authors acknowledge Professor Nai-san Li at China Pharmaceutical University for kindly providing the tested compounds and Associate Professor Xiao-Bing Gao at Yale University, United States, for his valuable modifying of English grammar in the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Funds of China (Grant no. 30671881) and the Natural Science Funds of Jiangsu Province, China (Grant no. BK2006016).
