Abstract
Understanding the structure–activity relationships (SARs) of small molecules is important for developing probes and novel therapeutic agents in chemical biology and drug discovery. Increasingly, multiplexed small-molecule profiling assays allow simultaneous measurement of many biological response parameters for the same compound (e.g., expression levels for many genes or binding constants against many proteins). Although such methods promise to capture SARs with high granularity, few computational methods are available to support SAR analyses of high-dimensional compound activity profiles. Many of these methods are not generally applicable or reduce the activity space to scalar summary statistics before establishing SARs. In this article, we present a versatile computational method that automatically extracts interpretable SAR rules from high-dimensional profiling data. The rules connect chemical structural features of compounds to patterns in their biological activity profiles. We applied our method to data from novel cell-based gene-expression and imaging assays collected on more than 30,000 small molecules. Based on the rules identified for this data set, we prioritized groups of compounds for further study, including a novel set of putative histone deacetylase inhibitors.
Keywords
Introduction
Small-molecule profiling (i.e., the characterization of compounds by multiple biological activity measurements) has been shown to capture detailed information about biological effects and mechanisms of action of small molecules. 1 This level of granularity holds great promise for a comprehensive understanding of compound structure–activity relationships (SARs),2,3 which in turn allows optimizing compounds simultaneously against multiple biological endpoints. 4 To overcome the limitations of existing approaches, we developed a computational method to automatically mine small-molecule profiling data for SAR rules.
Profiles can be obtained in parallel by combining separately acquired assay results such as binding constants against different purified proteins, 4 the drug sensitivity of different cell lines,5,6 or high-throughput screening results from different assays. 7 By contrast, multiplexed profiling assays capture complex cell states by simultaneously measuring many features in a single well format as a “signature” (e.g., gene expression, protein levels and modifications, or cell-morphology descriptors).1,4
Growing public bioactivity databases8,9 and novel high-throughput experimental methods10,11 enable researchers to apply both types of profiling data in the early stages of drug- and probe-discovery projects when the number of compounds to consider is still very large.1,4 This application, in turn, requires robust and scalable computational analysis methods to help interpret the large numbers of complex activity profiles. In addition to the inherently multidimensional space of chemical structures, the number of possible profiles grows exponentially with each biological measurement dimension.
Most available SAR analysis methods, however, were designed to handle between one and five bioactivity annotations and are not easily extensible.12 –14 One strategy is therefore to reduce high-dimensional bioactivity data to summary statistics, such as the number of shared targets between two compounds, before connecting this information to chemical structure. 15
Here, we present a computational approach for directly deriving interpretable SAR rules from large amounts of biological profiling data. We used frequent-pattern mining (FPM; also called frequent-itemset mining) and association-rule mining (ARM) to find combinations of substructures that are associated with characteristic biological profiles. 16
FPM was originally developed for market-basket analysis to find combinations of products that are frequently bought together. ARM builds on these “frequent patterns” (or “frequent itemsets”) to find interesting associations between products. These associations are formulated as rules. For example, the rule {
FPM has been used to find frequent substructures in sets of compounds and to identify single contiguous fragments that distinguish active and inactive compounds in a bioactivity assay.17,18 Fragment
We present here a general approach to analyzing SARs for large numbers of high-dimensional biological profiles using FPM and, in addition, ARM to automatically formulate SAR rules. Rules that connect chemical features to patterns in biological profiles are automatically identified and ranked by interestingness. We evaluated our method on gene-expression and cell-morphology profiles for more than 30,000 compounds. The compound collection contains subsets representative of common screening libraries assembled from various sources as well as planned synthetic libraries of compounds with well-defined structural relationships. We used different chemical and biological descriptors to tailor the general approach to specific requirements of the compound library.
Materials and Methods
Compound Sets
We assembled three distinct compound sets with varying structural properties and levels of biological activity annotation. We first selected a collection of 19,637 structurally diverse compounds derived from diversity-oriented synthesis (DOS).
20
The library was synthesized by the Center for the Science of Therapeutics at the Broad Institute using a build–couple–pair strategy.
21
The compounds were built around 23 novel chiral core structures by systematically varying the configuration of core stereocenters and decorating them with various side chains (see
Gene-Expression (GE) Profiles
We followed the protocol of Peck et al. (see
Multiplexed Cytological (MC) Morphology Profiles
We followed the protocol of Gustafsdottir et al. (see
Chemical Descriptors
DOS Fingerprints (DOS-FPs)
We used synthetic history information to define DOS compound core structures. Based on these cores, appendages were determined by R-group decomposition. Information about the attachment position of each appendage to the core was recorded. In addition, the absolute configuration of all stereocenters in the core was determined. This determination was performed on core structures without appendages to keep absolute stereochemistry consistent irrespective of substituent priority changes with different appendages. Concatenating core, appendage, and stereochemistry features into a feature string generated the DOS-FP for each compound.
Extended-Connectivity Fingerprints (ECFPs)
ECFPs 22 with atom types were calculated using a bond distance of 4 (ECFP4). Structural fragments that correspond to each bit were combined into a feature string for each compound. Only features with a bond diameter of 2 and higher were considered to exclude noninformative single atom fragments. All calculations were performed in SciTegic Pipeline Pilot 8.5.
Biological Descriptors
Cluster Attributes
We hierarchically clustered the data based on biological profiles (GE or MC) using a complete linkage method on pairwise correlation distances. All nodes in the cluster dendrogram were indexed. Each compound was assigned a set of indices based on cluster membership.
Signatures
GE signatures were obtained from GE profiles by applying a threshold of 2 (−2) to
Frequent Pattern-Mining and Association Rules
Frequent itemsets were determined using a publicly available implementation (http://mahout.apache.org) of the Parallel FPgrowth algorithm 23 on a Hadoop platform (version 1.0.4; http://hadoop.apache.org). Association rules were generated from the list of frequent patterns in R (version 3.0) using the arules package. 24
SAR Score
We calculated confidence, purity, and
Replicate-Based Hit Selection
To determine which compounds led to stable profile changes compared to negative controls (DMSO), we used replicate correlation and connectivity (i.e., the similarity of replicates compared to all other wells on the same set of replicate assay plates). Replicate correlation was calculated as the Pearson correlation coefficient between their GE (or MC) profiles. Replicate connectivity was calculated for each pair of replicates (e.g., R1 and R2) by ranking all wells on the same set of replicate plates by their profile similarity against R1 and identifying the fraction of unrelated wells that rank higher than R2. The reverse calculation (R2 vs. R1) was also performed because connectivity values are not symmetric. To allow individual replicates to fail, we only considered the top 50% of replicate correlation and connectivity values for each compound. We then calculated the two-dimensional distribution of negative-control replicate correlation and connectivity. Compounds were considered hits if they exceeded the 97% confidence interval of this negative-control distribution.
Results
Background
We applied FPM and ARM to small-molecule gene-expression and cell-morphology profiles to derive SAR rules of the form {
We derived biological attributes from two profiling assays, gene expression (GE) and cell morphology (MC). For GE profiles, the expression levels of 978 protein-coding transcripts were determined by ligation-mediated amplification and Luminex bead-based detection. 10 For MC profiles, changes in 812 cell-morphology features were captured by automated microscopy and computational image analysis. We used a “cell-painting” assay with six fluorescent dyes in five channels to distinguish cellular compartments and organelles. 11
In the following, we follow the notation of Tan, Steinbach, and Kumar
16
to introduce FPM and ARM. FPM and ARM operate on collections of binary attribute vectors (“objects”). Let
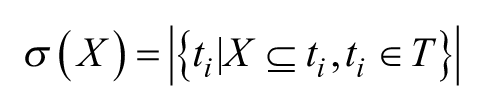
Given a minimum support count threshold
For the purpose of SAR rule mining, objects
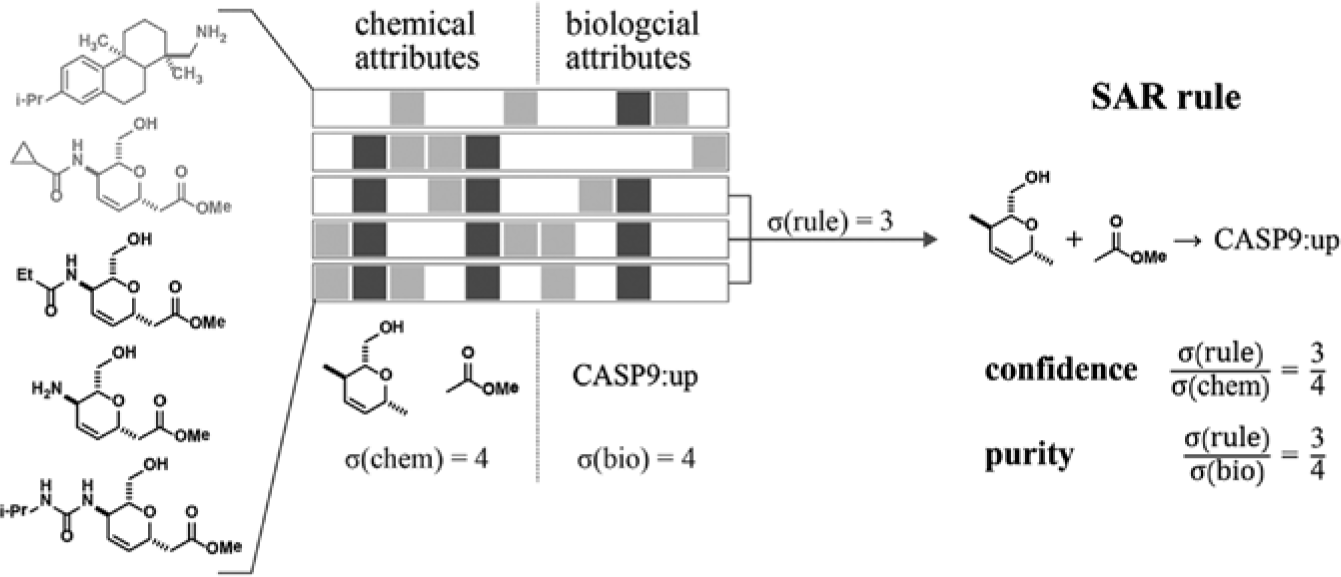
Association-rule mining (ARM) identifies and scores rules that relate the chemical features of compounds to biological profile attributes. In this schematic example, three out of five compounds match a rule that relates the occurrence of two structural features to the upregulation of a gene (CASP9). Confidence and purity measure the overlap between compounds that match the structural rule and compounds that elicit the biological phenotype. This information is used to quantify the quality of a rule.
From these patterns, ARM derives rules of the form
Three measures of rule quality or “interestingness” are then used to filter and prioritize rules. First, the confidence of a rule is defined as the fraction of objects with LHS attributes
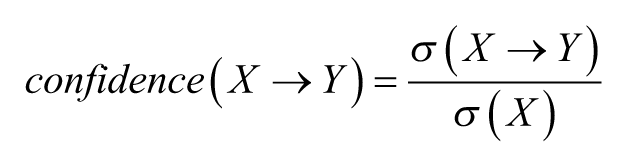
Second, we defined the purity of a rule as the reverse measure (i.e., the fraction of RHS attributes
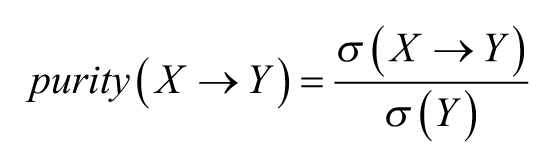
Third, we calculated a
We only kept rules with
We tested our method on GE and MC profiles for two distinct compound libraries. In the following sections, we discuss examples of rules obtained with different chemical structural and biological attributes.
ARM with Custom Substructure Descriptors on Planned Synthetic Libraries Leads to Intuitive and Testable SAR Hypotheses
SAR analyses take advantage of structural similarity between compounds. Although analog series in advanced projects are tailored to meet this requirement, many screening libraries do not. Early-stage SAR exploration is hence especially useful for planned screening libraries that were designed to include groups of compounds with defined structural relationships. 4
We therefore included 19,637 compounds in our experiment that were derived from DOS. 20 The compounds represent systematic combinations of three basic structural elements: core structures, appendages, and stereochemistry. Compounds were built by modular synthesis, attaching different appendages to 23 chiral core structures and systematically varying the configuration of core stereocenters. 21 The resulting collection is chemically diverse but, at the same time, contains related structures: groups of compounds that share common cores, stereochemistry, or appendages can be directly compared in SAR analysis. Importantly, this modular design also allows quickly synthesizing structural analogs of identified hits for follow-up studies.
We used as chemical attributes a custom structural fingerprint (DOS-FP) that reflects the modular design of DOS compounds to obtain rules that directly relate to compound synthesis. Based on the three DOS diversity elements, DOS-FP specifies for each compound (1) the core structure, (2) appendages attached to the core (as a simple occurrence or as “R-groups” that include information about their attachment position in the core), and (3) the configuration of core stereocenters. Stereoisomers of appendages were treated as distinct structures. Note that R-group and stereochemistry features refer to specific positions in the core and hence can only occur in combination with a core feature.
We added a reference set of 2222 compounds with known bioactivities (BIO) to the DOS library to support the generation of hypotheses about the biological effects of novel DOS compounds. The BIO collection was assembled to contain structurally diverse compounds that cover a wide range of biological activities and often have known direct targets. We measured both GE and MC profiles for most compounds. Exact compound numbers measured with each method differ slightly due to quality-control filters on the experimental data (GE: 17,553 DOS compounds + 1935 known bioactives; MC: 17,805 DOS compounds + 2211 known bioactives).
We first aimed to identify SARs on global phenotypic effects. We hierarchically clustered GE and MC profiles to find groups of compounds with similar biological activities. Each node of the resulting dendrogram was considered one cluster. Each compound could therefore be a member of multiple clusters (characterized by superset–subset relationships). Importantly, both DOS and BIO compounds were clustered, but only DOS compounds were subject to ARM for SAR rule mining. Although BIO compounds cannot be represented with DOS-FP, this allowed us to evaluate structural rules for DOS compounds in the context of biologically annotated reference molecules.
We found a total of 6206 rules for GE and 7098 rules for MC (2752 and 2861 for minimal rule sets, respectively). About 70% of the rules for both GE and MC were combinations of core and stereo features, and another 17% specified core features alone ( Table 1 ). This distribution is likely to reflect the fact that the DOS library contains many more compounds that match on these two features than on appendages. Rules that include R-group information are, however, about five- to sixfold enriched among the top 10% of rules ( Table 1 ), indicating their importance for describing specific small-molecule effects.
Gene-Expression (GE) and Multiplexed Cytological (MC) Profiles Show Comparable Distributions of Rule Types.
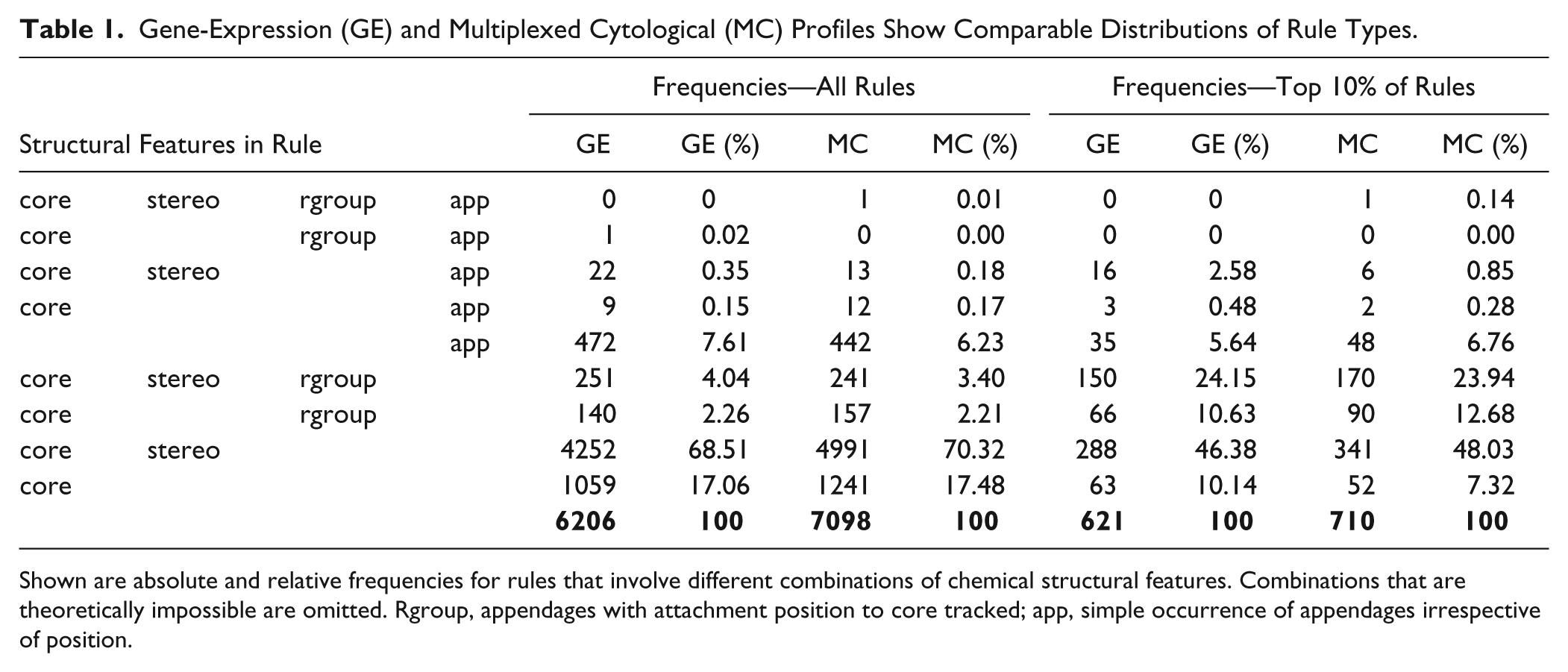
Shown are absolute and relative frequencies for rules that involve different combinations of chemical structural features. Combinations that are theoretically impossible are omitted. Rgroup, appendages with attachment position to core tracked; app, simple occurrence of appendages irrespective of position.
The majority of rules that include R-group information specify analog series, that is, compounds that vary in one R-group (70% for GE and 80% for MC). Hence, these rules lead to directly testable hypotheses that are easily accessible through DOS. Analogously, 11% (GE) and 14% (MC) of rules with R-group information specify all R-groups and hence apply to series of stereoisomers. The remaining fraction consists of rules that allow two of three R-groups to vary. Overall, chemical feature combinations occur with highly similar frequencies in the rules for both profiling experiments ( Table 1 ). These frequencies did not change appreciably when only profiles with very good replicate agreement were used (see the Methods section).
We next examined rules for DOS compounds that co-clustered with known bioactive molecules to find rules with interpretable structural and biological parts. The highest-ranking rule identified a cluster that was strongly enriched for compounds carrying an ortho-aminoanilide (OAA) appendage ( Fig. 2 ). 50% of all OAA DOS compounds are contained in this cluster (confidence: 0.5) and hence have similar GE profiles. The OAA motif is known as a biasing element for inhibitory activity against class I histone deacetylases (HDACs) due to its ability to interact with zinc and active-site residue chains required for hydrolysis of the acetyl moiety on lysine side chains. 27 Indeed, the DOS compounds co-cluster with two known HDAC inhibitors that also contain an OAA residue, mocetinostat (MGCD0103) and tacedinaline (CI-994). Furthermore, the only DOS cluster member that does not match the structural rule, BRD4805, has recently been identified as a low micromolar HDAC inhibitor. 28
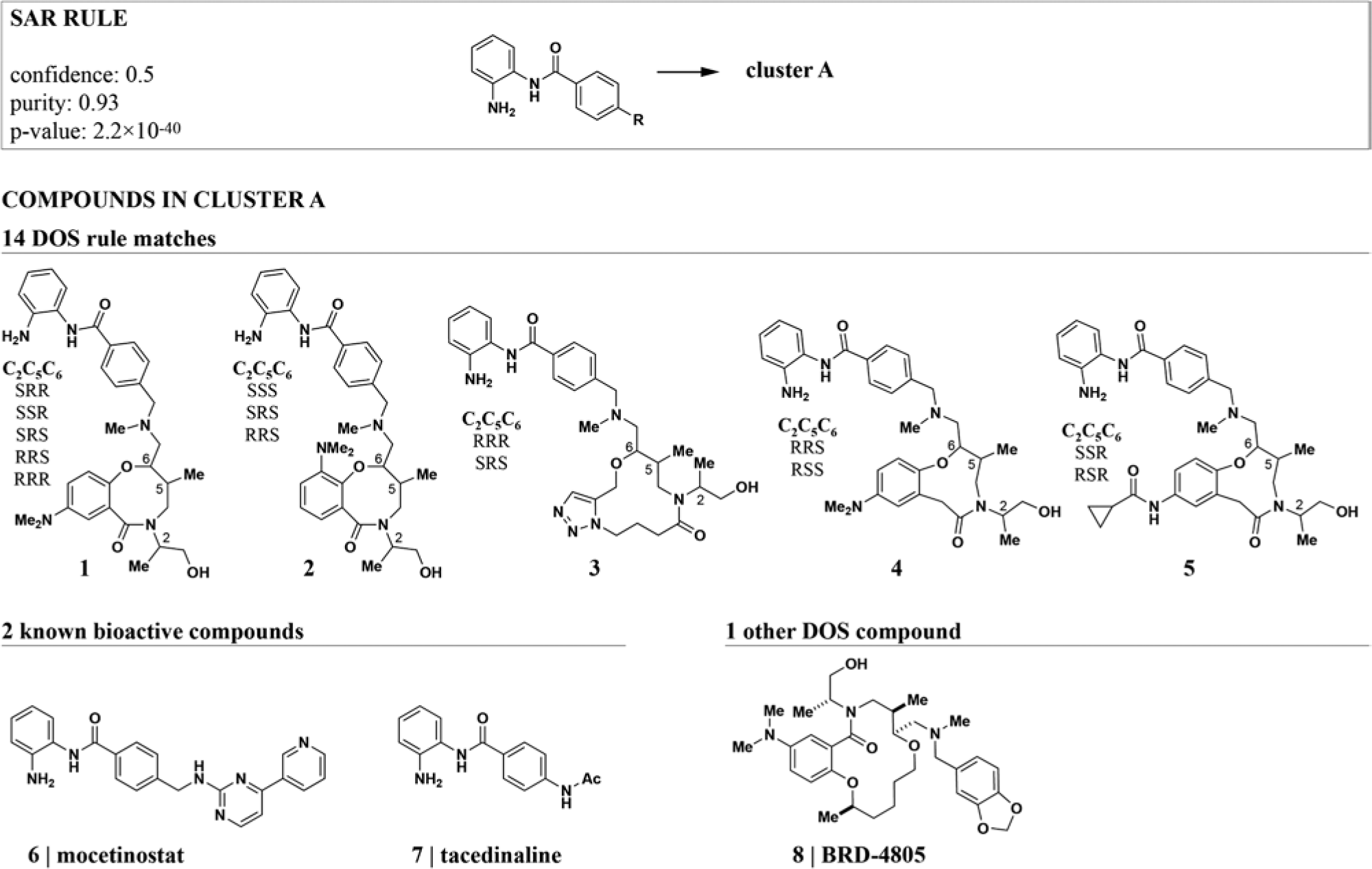
The best-scoring rule for gene-expression (GE) profiles identified a known biasing element for histone deacetylase (HDAC) inhibitory activity. The quality measures for this rule indicate that compounds with an ortho-aminoanilide (OAA) residue are strongly enriched in cluster A. All compounds in cluster A are shown. The cluster contains two known OAA HDAC inhibitors (6 and 7), and all diversity-oriented synthesis (DOS) compounds except one carry the OAA residue. The remaining DOS compound, BRD4805, has recently been identified as an HDAC inhibitor. 28 For simplicity, stereoisomers are shown as combinations of generic structures and configuration tables.
Mocetinostat, tacedinaline, and BRD4805 exhibit selectivity for HDAC1, 2, and 3 over other isoforms.28,29 Interestingly, less selective inhibitors
29
contained in our compound library—trichostatin A (TSA), vorinostat (SAHA), and belinostat (PXD-101)—formed a separate cluster, indicating that GE profiling can distinguish compounds with distinct HDAC isoform selectivity patterns. We therefore hypothesize that the clustered OAA DOS compounds show some degree of specificity for HDACs 1–3. Supporting this hypothesis, stereoisomers of structures 1, 2, and 4 have recently been identified as HDAC1–3 selective inhibitors.
30
Members of this structural class shared the highest profile similarity with the known HDAC inhibitors in our experiment (
Several of the OAA DOS compounds have been tested further in a biochemical assay measuring inhibition of HDAC2 deacetylase activity (
Taking SAR information into account can increase our confidence in profiling assay results. Only six out of the 14 OAA DOS compounds in the cluster had sufficient replicate agreement to be considered “hits” in the GE assay (i.e., compounds that induce a reproducible GE change) (see the Methods section). Relying on replicate agreement as an activity criterion alone would hence disregard the remaining eight compounds and thus 55% of the HDAC2 hits in the cluster (five out of nine). This result provides an example of the overall utility of our method for guiding the selection of initial hits for follow-up biological studies and chemical synthesis.
The OAA rule is an example of a single structural motif whose presence is sufficient to elicit a relatively defined bioactivity. Although in principle one could have predicted such compounds to have HDAC inhibitory activity by structure alone, these results help validate the method for the prediction of less well-known features. More commonly, combinations of features determine a compound’s biological effect. One such example is a set of related DOS compounds that co-cluster with known microtubule destabilizers ( Fig. 3 ). The corresponding rule specifies the core, one of two R-groups, and, importantly, all stereocenters for this compound class, effectively describing a series of analog structures. Stereochemistry is a major determinant of specificity for this compound series. Omitting stereochemistry from the rule causes the confidence to drop from 0.5 (three rule matches out of six structure matches) to 0.065 (three out of 46). The validity of this rule is further supported by the fact that it occurs at the top of both GE (rank 6) and MC (rank 12) rule lists. Microtubule inhibitors co-cluster with the DOS compounds in both the GE and MC rules ( Fig. 3 ). Unlike the OAA case, the known bioactive compounds are not structurally related to the DOS compounds, making this an interesting rule to study for lead-hopping purposes. Using structural features that directly match physical DOS building blocks allows rapidly testing such hypotheses as analogs, and stereoisomers can be synthesized quickly (if they do not already exist).
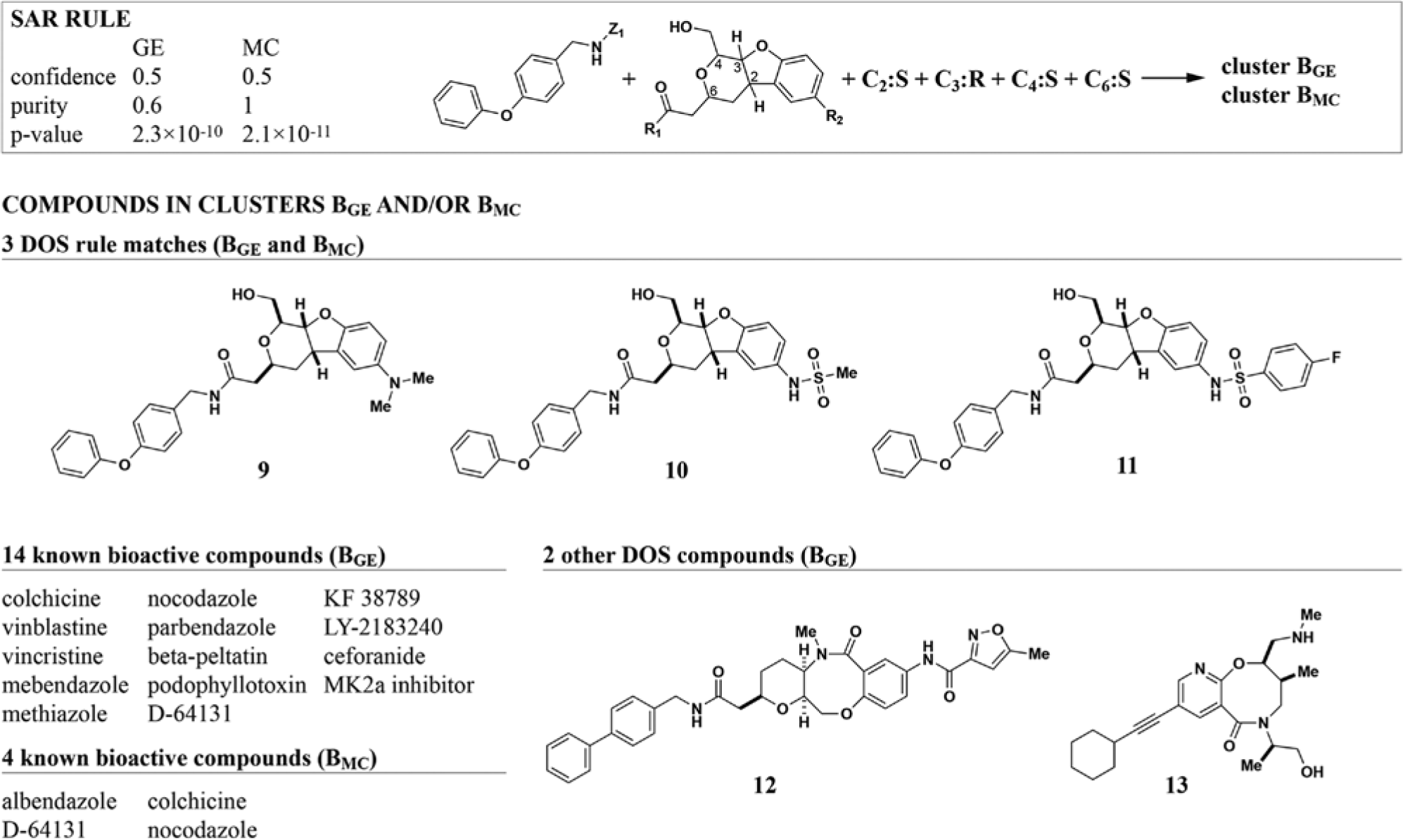
Both gene-expression (GE) and multiplexed cytological (MC) profiles identify the same rule for diversity-oriented synthesis (DOS) compounds that co-cluster with microtubule inhibitors. A series of three DOS analogs was identified based on both GE and MC profile clusters, yielding high-scoring rules in both cases. Interestingly, the rule is highly stereo specific, constraining all four stereocenters in these compounds to one configuration. Known microtubule inhibitors were present in both clusters, although the exact set of bioactives differed. For clarity of presentation, structures for bioactive compounds are omitted (see
General Structure Descriptors Allow Rule Generation for Arbitrary Compound Collections
Not all libraries are designed to include compounds with defined structural relationships like the DOS collection. Arbitrary libraries can, however, be mined by our method by replacing the chemical structural descriptors. We used this approach to analyze MC profile data for a collection of 10,226 compounds from the Molecular Libraries Small Molecule Repository (MLSMR). We used ECFPs (ECFP4 22 ) as a generic structure descriptor that can be used with any compound collection. Although any binary fingerprint can be used readily with our method, we chose ECFPs as a reference for this study because they are widely applied, allow direct mapping of features to structural fragments, and were specifically designed for SAR studies. 22 They achieve high structural resolution by encoding a very large number of features and thus present a good test for the ability of ARM to mine large feature spaces.
We used clusters based on MC profiles as biological attributes. Due to the versatility of ECFP4, we were able to include our set of known bioactive compounds in the rule-mining step. We found 14,198 rules (7328 after redundancy filtering), of which 2176 contained known bioactive compounds. Unlike the DOS-FP features, ECFP4 substructures can describe arbitrary parts of the molecule through combinations of overlapping substructures ( Fig. 4 ). This description allows a more fine-grained and unbiased mapping of relevant substructures. The downside is a reduction of interpretability because rules cannot be directly mapped to existing analogs or straightforward synthesis strategies. In addition, a large number of features are usually needed to achieve high resolution and generality. Even though ARM can handle such large data sets, the number of rules can increase as a result and complicate downstream analyses. Therefore, each compound collection and project require balancing generality, complexity, and interpretability when choosing a chemical descriptor.
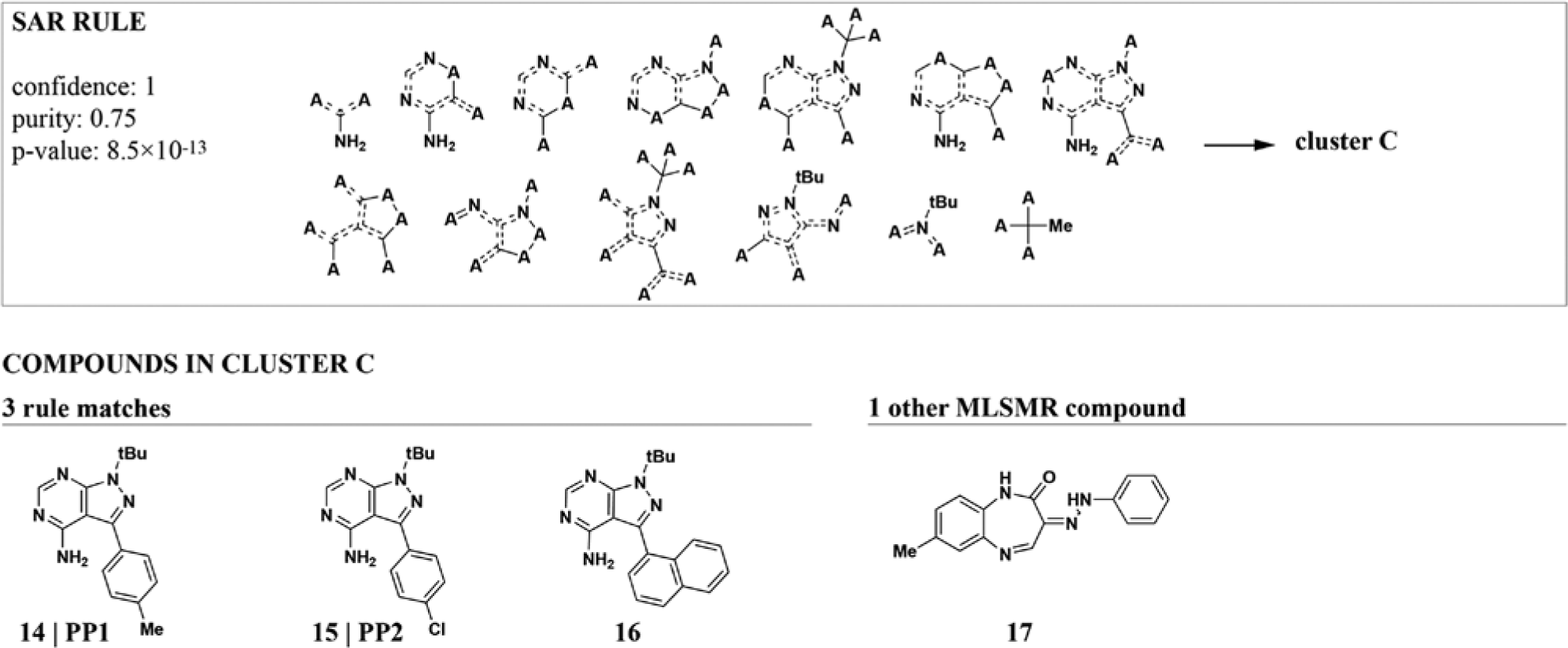
Extended-connectivity fingerprint (ECFP4) features map arbitrary parts of molecules with overlapping substructures. The highest-scoring ECFP4-based rule that involves known bioactive compounds describes a set of structurally and functionally related kinase inhibitors. Although ECFP4 fragments correctly identify their overlapping substructures, feature redundancy and abstractness make them harder to interpret than synthesis-oriented substructures like diversity-oriented synthesis fingerprints (DOS-FPs) (
Direct Mining of Attributes Identifies Biological Signatures for Sets of Related Compounds
Clustering biological profiles captures global similarities and differences between the biological effects of compounds. Information on individual features (e.g., genes from a GE profile) is, however, lost in this process. Because ARM is designed to operate on high-dimensional data, it can be used to mine biological features without intermediate aggregation. Useful applications include mining compound-target profiles or GE signatures (i.e., sets of genes that change expression levels in response to compound treatment).
We generated a signature of up- and downregulated genes for each compound by applying a
The second-ranked rule is highly specific for a small set of compounds with a signature of 13 upregulated genes (
Fig. 5
). A functional analysis strongly related the 13-member gene set to sterol and cholesterol synthesis (
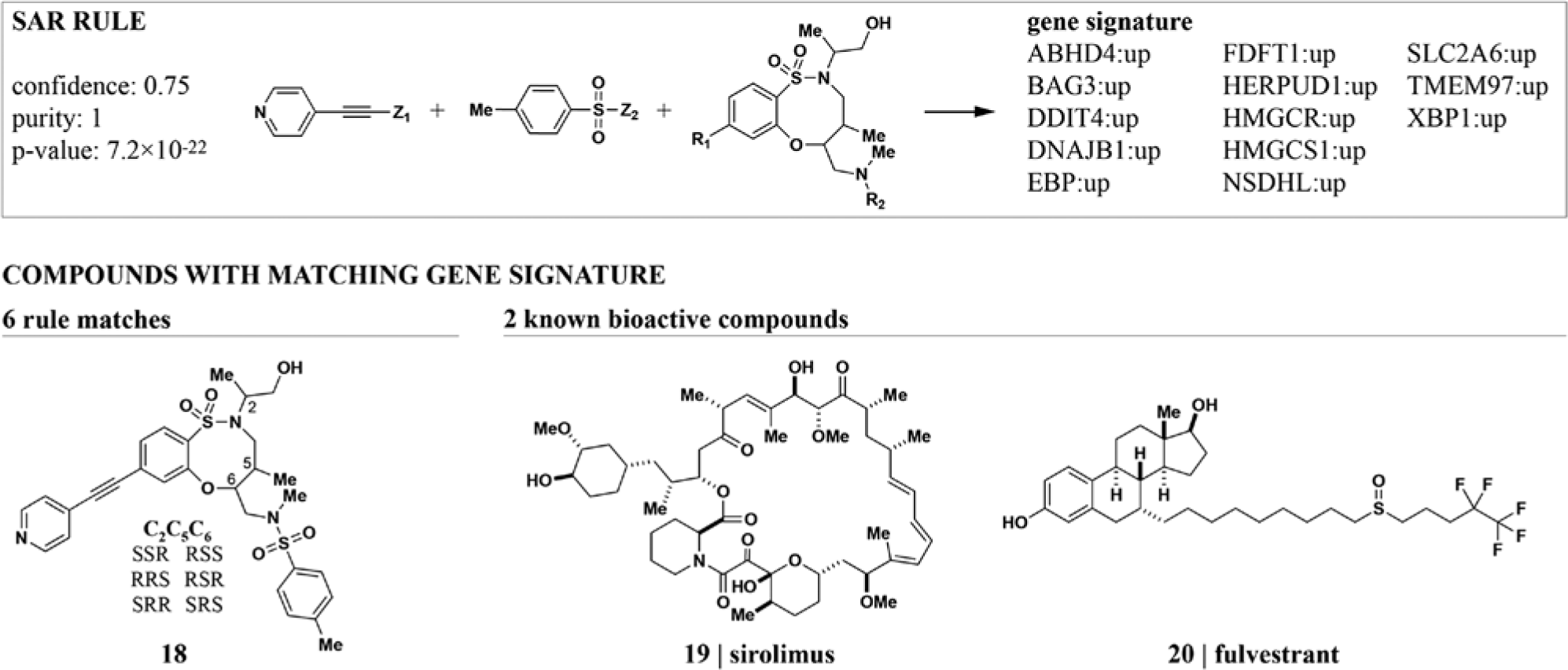
ARM can identify biological signatures for groups of structurally related compounds. A set of DOS stereoisomers was found to regulate 13 genes enriched for functions in cholesterol metabolism. The same genes are regulated as part of the signatures of sirolimus and fulvestrant, two mechanistically unrelated compounds that have both been linked to cholesterol metabolism.31,32
Discussion
Small-molecule profiling data sets are promising resources for the elucidation of SARs with high granularity. We have illustrated in several examples that association-rule mining can identify interpretable SAR rules from large sets of high-dimensional profiling data.
Our method is designed to support compound selection and synthesis decisions in early-stage drug- and probe-discovery projects. The data in such projects often consist of high-throughput measurements for large compound libraries. Such data are expected to be noisy and contain only small groups of related compounds with similar biological effects. Acknowledging these limitations, we chose a data-mining approach to analyzing such data. Rather than attempting automated classification, our method supports hypothesis generation and human decision making.
These goals further led us to choose interpretable structural fragments as chemical features to obtain rules that can directly inform synthesis decisions. Our method is not, however, limited to structural descriptors. ARM can be used with any descriptor that can be expressed through binary attributes. Categorical, ordinal, or continuous attributes like physicochemical properties or calculated descriptors can usually be represented by binning and proper encoding without too much loss of information. In any case, a balance between generality and interpretability of chemical attributes needs to be found. Custom synthesis-oriented representations like DOS-FP are highly interpretable but not applicable to all libraries. By contrast, generic fingerprints like ECFP4 rely on more abstract features that usually do not directly relate to synthetically accessible building blocks.
Similar considerations apply to the biological descriptors. First, different experimental methods will likely capture and emphasize different biological effects. We found that, even though several top-scoring rules were identified by both GE and MC, the particular composition of compound groups often differed. A comprehensive comparison between such methods has not yet been performed, however. Second, the computational representation of biological profiles and measures for their comparison will influence the quality of the resulting rules. Global similarity measures like correlation used to cluster profiles before SAR mining can be more suitable for compounds that elicit subtle effects across a wide range of features. By contrast, if a phenotype results from strong effects on only a few features, signature-based approaches (i.e., methods that use lists of the most strongly altered features for a perturbation) 33 are likely to perform better than whole-profile similarity methods. The generality of the ARM approach allows using arbitrary combinations of chemical and biological features, and comparing different feature choices can greatly increase the confidence in individual results.
Finally, the scoring function can be tailored toward individual needs. A number of objective measures of interestingness have been suggested that can be used with any attributes. 16 More domain-specific measures like intracluster distances, for example, can easily be integrated into the score as addends. Weighting individual features gives control over their relative importance to each other and allows different rule types to be prioritized. For example, putting high weight on the purity of a rule would lead to preferential selection of rare bioactivity patterns over common ones. We believe that these features make ARM an efficient and versatile approach for automated SAR mining of biological profiling data.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Cheminformatics and data-mining work was supported by the National Institute of General Medical Sciences (P50-GM069721, awarded to S.L.S.), as part of the Center of Excellence for Chemical Methodology and Library Development. Profiling measurements were supported as part of the National Institutes of Health (NIH) RoadMap Molecular Libraries Initiative (U54-HG005032, awarded to S.L.S.). Associated data can be accessed at ![]() . D.M.F. and S.J.H. were supported through funding from the NIH (R01DA028301) and the Stanley Medical Research Institute. S.L.S. is an investigator at the Howard Hughes Medical Institute.
. D.M.F. and S.J.H. were supported through funding from the NIH (R01DA028301) and the Stanley Medical Research Institute. S.L.S. is an investigator at the Howard Hughes Medical Institute.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
