Abstract
Recent industry–academic partnerships involve collaboration among disciplines, locations, and organizations using publicly funded “open-access” and proprietary commercial data sources. These require the effective integration of chemical and biological information from diverse data sources, which presents key informatics, personnel, and organizational challenges. The BioAssay Research Database (BARD) was conceived to address these challenges and serve as a community-wide resource and intuitive web portal for public-sector chemical-biology data. Its initial focus is to enable scientists to more effectively use the National Institutes of Health Roadmap Molecular Libraries Program (MLP) data generated from the 3-year pilot and 6-year production phases of the Molecular Libraries Probe Production Centers Network (MLPCN), which is currently in its final year. BARD evolves the current data standards through structured assay and result annotations that leverage BioAssay Ontology and other industry-standard ontologies, and a core hierarchy of assay definition terms and data standards defined specifically for small-molecule assay data. We initially focused on migrating the highest-value MLP data into BARD and bringing it up to this new standard. We review the technical and organizational challenges overcome by the interdisciplinary BARD team, veterans of public- and private-sector data-integration projects, who are collaborating to describe (functional specifications), design (technical specifications), and implement this next-generation software solution.
Introduction
The recent introduction of disruptive models of “open innovation” in drug discovery and development fueled by government and philanthropic organizations in collaboration with industry requires “open access” to chemical and biological data from public data sources. Pharmaceutical companies have released selected data sets as part of their open-innovation collaborations.1–5 The genesis of the majority of public biological and chemical data repositories, however, has been in academic, government-sponsored, or biotechnology environments. 6 These repositories of compound structure and activity data enable structure–activity data mining and hypothesis generation in advancement of discovery efforts. They also enable and support scientific collaborations between commercial and academic institutions. The largest public repository of these data is PubChem. 7 Within PubChem, approximately 90% of the data on active substances, a large part of the current chemical-biology data in the public domain, have been generated by the nine centers comprising the Molecular Libraries Probe Production Centers Network (MLPCN) ( Table 1 ). Funding for this National Institutes of Health (NIH) Roadmap Molecular Libraries Program (MLP) is scheduled to end in May of 2014. In an effort to ensure the integrity, accessibility, use, and impact of this valuable data set, the MLP leadership funded an administrative supplement in 2012 to create the BioAssay Research Database (BARD) and migrate existing Molecular Libraries Screening Centers Network (MLSCN) and MLPCN PubChem data into it. The processes and standards developed and optimized for data extraction, transformation, curation, and migration during this phase of BARD will be used on a continuing basis to migrate relevant new data from PubChem and from additional data sources into BARD. PubChem will continue in its current mission.
Summary of Data and Results in PubChem Generated by the Molecular Libraries Screening Centers Network (MLSCN) and Molecular Libraries Probe Production Centers Network (MLPCN).
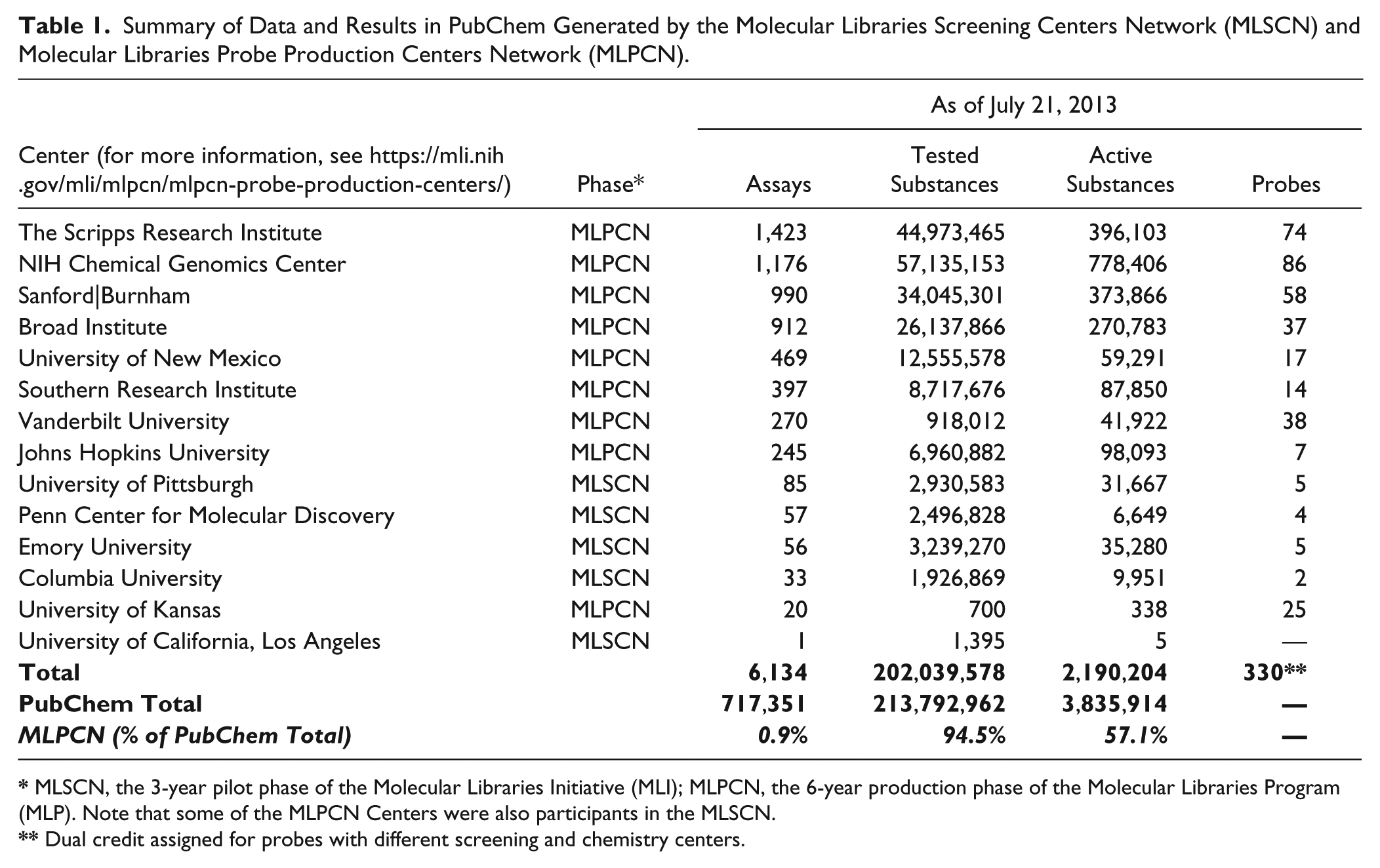
MLSCN, the 3-year pilot phase of the Molecular Libraries Initiative (MLI); MLPCN, the 6-year production phase of the Molecular Libraries Program (MLP). Note that some of the MLPCN Centers were also participants in the MLSCN.
Dual credit assigned for probes with different screening and chemistry centers.
PubChem was originally designed as a data archive with many of its current data-analysis functions released in the years following the start of data deposition. It was chiefly designed to be a flexible archive of public data, able to hold types of data that had not yet been anticipated. But, when reviewing how MLPCN data were put into PubChem, the BARD team determined that better structuring the data would improve its visibility and usability. This structure required the development and adoption of a data model and a standard set of terms that could form the basis of the vocabulary for assay and result descriptions. The data dictionary in BARD, which leverages the BioAssay Ontology (BAO), 8 provides a foundation that is well controlled, suitably extensible, and consistent in its usage of terms and definitions. Analysis tools built on top of this system are expected to be intuitive, user-friendly, and graphically compelling to facilitate the information analysis that today’s research scientists need. BARD also has an extensible “plug-in” architecture to facilitate the deployment of community-developed computational algorithms. Because BARD users are expected to be either casual infrequent users or more advanced users, the query tools should support both customer types. The diverse user base should be able to load data, execute their queries, and find the data they need without extensive training or reading of user manuals.
The BARD project has intentionally focused its work on bioassay descriptions, their experimental conditions, and the corresponding results in the context of the specific projects. Importantly, BARD does not present itself as a general-purpose laboratory information-management system (LIMS). Instead of focusing on the normalization of plate-based data, for example, BARD focuses on abstracting and structuring screening results to allow comparisons among data sets and support their long-term storage and use. Although we expect some overlap with existing systems, BARD has evolved the current standard for research data management for both academia and industry, and it is increasing the quality of structured annotations in the appropriate technical architecture.
The bio-pharmaceutical industry is also embracing limited open-innovation and open-access projects through Linked Data, a foundation for pharmaceutical R&D data sharing, at Eli Lilly, Johnson & Johnson, and UCB Pharma. 9 Linked Data was created by the Linked Open Drug Data (LODD) task force of the World Wide Web Consortium (W3C) Health Care and Life Science Interest Group (HCLS IG), which aims to address the challenges of data representation, connection, and access by harnessing the power of new web technologies, then uncovering what questions can be answered once the data sets are connected in Linked Data. Endeavors such as Bio2RDF 10 and Chem2Bio2RDF 11 represent additional efforts to formalize and integrate biology and chemical-biology data using semantic-web technologies. Eli Lilly, a visible proponent of open innovation, offers a set of Open Innovation Drug Discovery computational tools on its website. 12 Furthermore, their Open Innovation Drug Discovery screening panel, 13 which is available to authorized users, includes both phenotypic and target-based assay modules, with information on the collected and curated best practices for the enablement, statistical validation, operations, and quality control of various in vitro assay formats. The website also provide links to the program background and its approaches to drug discovery, a list of available “open” targets, assay-validation protocols, testing flow schemes, and descriptions and standards for in vitro, phenotypic, and tuberculosis assays. To continue to be relevant and align with the NIH’s “public-good” and open-innovation mission for MLPCN, BARD was developed with a commitment to open-source software. All of the source code and vocabularies developed for BARD will be made public along with the necessary documentation required for the community to reuse and extend the software. BARD leverages open-source libraries and tools extensively, using commercial components only in limited cases when an open-source solution did not meet our needs. For example, structure rendering is provided by ChemAxon’s JChem toolkit. These commercial components will not be distributed with the BARD source code but are required to run the software.
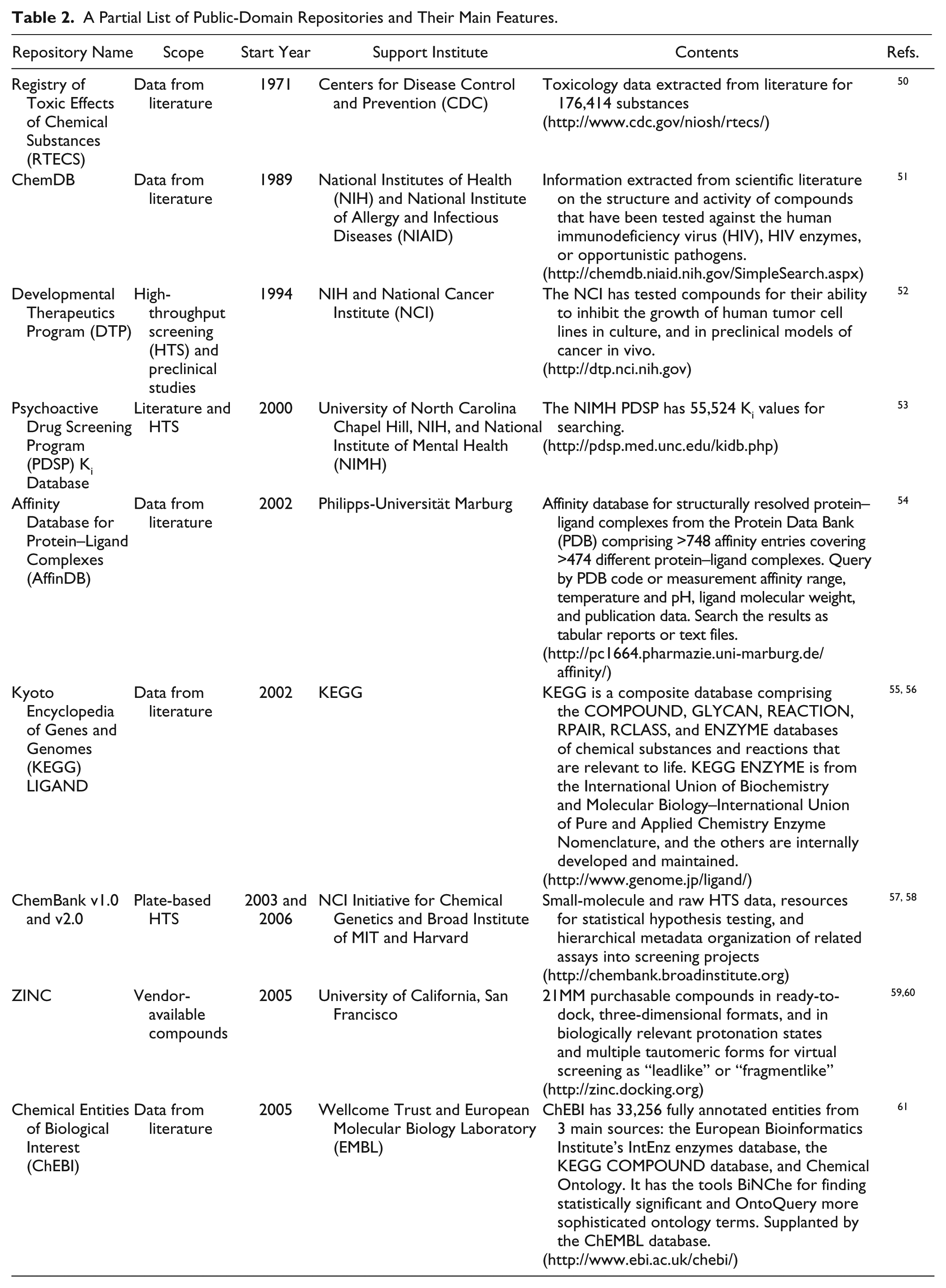
These data-sharing efforts and the proliferation of multiple public data repositories ( Table 2 ) have resulted in an informatics maze that is difficult to navigate and decipher because each repository uses a different dialect, which causes confusion regarding common terms. The language barrier is compounded by complex interfaces, which prevent the efficient and effective use of these data to advance understanding of the underlying biology and chemical structure–activity relationships. These challenges prevent scientists from understanding the data and using them to generate novel hypotheses. It should be noted that BARD does not aim to play the role of the “mother of all databases” by integrating various public repositories. Instead, when required and relevant, BARD makes extensive use of external life-science data sources such as Gene Ontology (GO), 14 UniProt, 15 BAO, 8 and others, but it uses them by reference rather than by maintaining local copies of these sources.
A Partial List of Public-Domain Repositories and Their Main Features.
Pharmaceutical Industry Best Practices
As the first step of the project, the BARD project team undertook a comprehensive review of the pharmaceutical and biotechnology industry’s best practices, as that industry has significant experience with merging and integrating different commercial and public data sets. With significant investments, many have implemented effective solutions. But the challenges are manifold: multiple legacy systems, different research cultures, independently developed processes, incompatible or competing technology choices, legal restrictions, lack of commonly accepted and understood terminology, intellectual property concerns, and (above all) plenty of “opinion” (i.e., scientists with strongly held beliefs and little willingness to compromise for the common good). Although worthy endeavors have been reported throughout the years (e.g., Cousin–Upjohn, 16 VERDI–Vertex, 17 ArQiologist–ArQule, 18 Avalon–Novartis,19–21 ABCD–Johnson & Johnson (J&J), 22 Mobius–Lilly, 23 OSIRIS–Actelion, 24 and Chemistry Connect–AstraZeneca 25 ), the proprietary nature of the data has not always incentivized disseminating best practices. Many have not built from scratch but integrated commercial components with some custom development. The latter approach includes inherent challenges such as resource constraints, the limited scalability of the tools, and restricted access to data because some vendor solutions lack the appropriate application programming interfaces (APIs). Perhaps the biggest challenge is that the complexity of the biological research data requires people with expertise in data integration who also understand the complex research domain. This combination of domain and technical expertise also impedes software vendors, who are not embedded within research organizations, lack exposure to real data, and rely on more formal (and less efficient) mechanisms for getting user feedback and suggestions for improvements. The BARD team has a great deal of scientific and technical experience from participating in or leading previous efforts in the private sector, and this experience allowed the team to hit the ground running at the start of the project by quickly converging on a data-management philosophy and architecture, as well as data structures, to work with.
Research Data Management
One distinguishing feature of BARD is the use of research data-management (RDM) principles 26 to organize and manage the data lifecycle. RDM defines the policies, standards, and infrastructure that are needed to ensure the integrity, accessibility, and stewardship of research data. RDM affects several features of the BARD interface, including assay definition workflows, assay query tools, and result displays. A key principle of RDM is the use of standard terms and structures for the description of both experimental contexts and the corresponding result data. This standardization enables a common understanding among groups with different internal terminology and allows machine mining of structured terms that can be rigorously defined for use in computational modeling. We contend that, as with other deposition standards, 27 a public bioassay database needs to mandate that all data being deposited be annotated using a common language with a well-defined standard. The system of voluntary annotation with “controlled comments” that is used by PubChem 28 is not sufficient to enable robust cross-assay data mining.
In defining such standardized terms, a significant challenge was reaching consensus about how the experimental data generated from bioassays should be described. Even within the MLP network centers, different terminology or concepts were considered the norm based on scientists’ previous industry experience, academic training, and/or level of focus on different specialized assay types. To achieve consensus, we convened these scientists and agreed on a concept hierarchy that describes how an experiment was run, its purpose, how it relates to other experiments, and what the data values signify. These concepts are captured in a hierarchy of terms with the corresponding top levels of Assay (definitional protocols), Biology (descriptions of the biological system intended to be studied by the protocol), Project (aggregations of different instances of execution of a protocol for a larger interpretive purpose, as well as per-run details such as compound collection or test concentration), and Result (standard terms for values calculated as measurements).
As a foundation for several parts of this hierarchy, we relied on previous annotation of assay experimental data by BAO.8,29,30 During our review of industry best practices, it was clear that there was wide interest in leveraging BAO, but no one had yet deployed a production system based on the ontology. We used a subset of BAO terms that focuses on the formal description of experimental protocols, and from that foundation we added terms or concepts with input from domain experts. Notably, BAO is a formal ontology constructed as a subsumption hierarchy that allows formal inference modeling, whereas the BARD data dictionary is intended to be a collection of terms that allows users to rapidly locate concepts, select a standard vocabulary term for their intended purpose, and understand the term’s usage in relationship to other terms. Although this loosened relationship of terms does not allow for formal machine modeling of concepts (e.g., automatically indicating that a bioluminescence assay has luciferase as a component), we have maintained a consistent mapping of all BARD terms to BAO to allow for future conversion of assay protocols into BAO and their subsequent use by that system. At the same time, we have included some basic concepts that will assist the user with defining assays in a logical manner, such that the system can offer suggestions of workflows or flag possible inconsistencies within a protocol.
Data-Structure and Data-Migration Challenges in BARD Development
BARD was created to deliver a well-curated set of results that can be mined by either an end-user biologist or chemist, or a more computationally savvy scientist. Beyond data-dictionary standards, BARD implements business rules and data-stewardship procedures to resolve some of the current challenges with cross-center assay comparisons. The BARD dictionary is focused on the minimum vocabulary required to effectively and efficiently compare compounds, assays, and results. This vocabulary includes key assay parameters (e.g., target, cell line, assay stage, and assay type), experimental variables (e.g., concentration, number of points in the curve, and route of administration), and results [e.g., half maximal inhibitory concentration (IC50), activity score, and half-life]. An assay-protocol catalog also stores the relationships among the results (dependent variables) and assay parameters and experimental choices (independent variables). The result hierarchy is also captured to specify the calculation precedence [e.g., percentage IC50 (pIC50) is derived from IC50, which is derived from multiple percent-inhibition values at a range of concentrations].
Key business rules facilitate data extraction, normalization, and aggregation across the centers, including the computation of statistics and conversion of IC50 to pIC50, unit conversion, display rules, and the handling of custom data types (structures, images, result modifiers, scientific notation, and curves). We implemented data-stewardship and -governance policies by which new items, some classes of which are expected to be frequently supplemented as more data are deposited (e.g., result types, assay categories, and instrumentation), are rationalized with existing terms to ensure a logical description of new data without corrupting existing information. To meet the challenge of maintaining a standard set of terms among a diverse research community, the dictionary requires ongoing curation as more data are added to the system; these are ultimately reconciled with component ontologies such as BAO. From the initial term set, we have added synonyms or new terms as data have been migrated from PubChem. A curation team will continue to provide support for the system as it is released to data depositors outside of the MLP that will have their own understanding of terms and experimental concepts. The ultimate goal is to allow the system to adapt to how scientists use language in their research by recognizing similar concepts that may be described differently but ultimately have a common scientific meaning.
BARD’s engineering approach included a focus on usability and functionality to enable scientists to use MLP data prospectively to generate new hypotheses. The team leveraged the research described in the preceding sections to focus on how BARD could go beyond existing public resources to facilitate (1) the capture of higher-quality data, (2) assay data capture with relevant context, (3) data querying and mining, and (4) a modern technical architecture that includes the segregation of transactional and analytical layers with the relevant APIs. BARD does not provide functionality for compound registration and instead leverages PubChem to provide this function. Examples of questions that BARD will enable include finding summary information for any MLP probe or all results for a given list of compounds. BARD will be able to find a biological profile (among many assays) for a given compound, or it will find a summary of all assays or experiments in which the compound was tested. BARD will also be able to provide new information about a list of compounds resulting from a query, for example displaying the list as a structure–activity relationship (SAR) table or performing promiscuity analysis on the set. Finally, BARD aims to connect compounds and compound sets to target and pathway information.
Our experience and industry survey indicated that data generators and data consumers have different needs. Specifically, the role of data generator must be acknowledged and respected. We have designed a data-registration system on the assumption that the person designing and performing the assay (i.e., the data generator) has the necessary background and scientific expertise to properly process the data to generate high-quality bioactivity results. This assumption accords with the key data-integrity principle recommended by the National Academy of Sciences that “researchers themselves are ultimately responsible for ensuring the integrity of research data.” 26
Our approach has been to encourage and enforce common quality-control standards through the use of a hierarchical dictionary of terms that integrates with publicly available ontologies. A common ontology or dictionary is essential for effective navigation, search, and comparison of assays and results among sites and institutions. To ensure that this common vocabulary is properly used, BARD provides a new assay-registration tool, which is by definition transactional in nature, for use by MLPCN Centers and other interested groups. This transactional subsystem, the Catalog of Assay Protocols (CAP), is a foundational component of BARD that interacts with other layers and technological components of BARD to provide functions that enable sophisticated hypothesis generation from the data by both novice (or “casual”) and expert users ( Fig. 1 ).
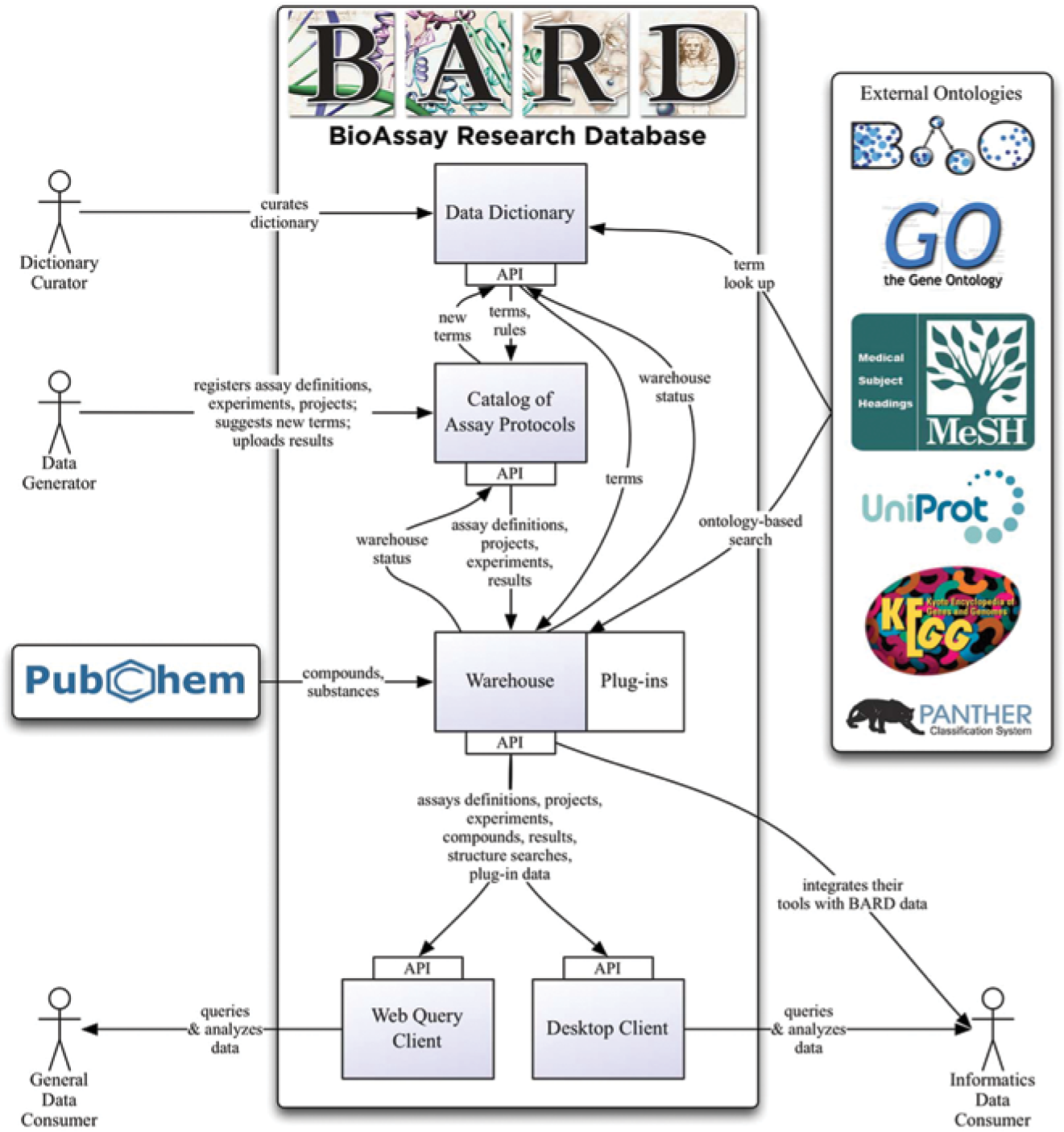
Overview of BioAssay Research Database (BARD) technology components and associated user-interaction points.
Organizational and Social Challenges in BARD Development
The organizational challenges of a geographically dispersed team with different institutional cultures have required flexibility and determination to make the project succeed as a whole—a trait perhaps shared by many successful open-source initiatives. BARD is a cross-functional national program that is being co-led by the Broad Institute and the National Center for Advancing Translational Sciences (NCATS) with extensive participation from regional experts and teams from Cambridge, MA (Broad Institute); Jupiter, FL (Scripps); Miami, FL (University of Miami); Rockville, MD (NCATS); San Diego, CA (Sanford|Burnham); Nashville, TN (Vanderbilt University); and Albuquerque, NM [University of New Mexico (UNM)]. Large face-to-face meetings were limited and usually conducted only for major milestone reviews. The team, however, found great benefit in smaller, focused, face-to-face meetings and opportunities to work in one place, as they discovered each other’s working styles, strengths, and skills. The net impact of the geographical separation and multiple subcultures meant that relationship development was critical to the creation of a high-performance team.
Most software development was managed by two primary centers. The Broad Institute is an experienced user of Agile (http://www.agilemanifesto.org) methodologies 31 and follows the Scrum techniques 32 for managing and executing projects. NCATS uses a rapid-prototyping approach. The diverse subcultures in the organizations and their differing approaches to the relative importance of various roles and skill sets in the project team highlighted that good communication between the centers facilitates the development process and creates a working rapport. To mitigate the increased risks associated with these challenges, the team often attended daily video meetings (not just telephone communication and shared documents); divided work into major streams that could operate independently for stretches of time, thereby bringing the immediate team size down to 5–7 people; and organized a series of more formal requirement-gathering and usability sessions with scientists who would become users of the system when it was released. Many of these sessions were recorded to share them among the teams. Much of the daily contact was accomplished with one or two experienced developers taking the lead role in core discussions and working with a dedicated project manager. The project team also agreed to general interaction guidelines to ensure that all stakeholders committed sufficient time and resources to BARD. Each center agreed to be accountable for ensuring that their staff followed the RDM standards.
Each of the six main work streams was led by one of the centers involved: CAP and data migration was led by Broad, data warehouse and API by NCATS, desktop clients by NCATS, web query clients by Broad, API plug-ins by the UNM, and outreach, training, and support by Vanderbilt University. An external scientific panel (ESP) that includes industry and academic participation reviews program progress toward project milestones and evaluates overall direction and relevance. The ESP has been selected as an expert panel that understands the data and information needs (both functional and technical requirements) with respect to high-throughput screening, hit-to-lead, and lead optimization, including in vitro and in vivo ADME and pharmacology. The panel consists of Lawrence Barak, Duke University; Tom Ferrin, University of California, San Francisco; Sam Gerritz, Bristol-Myers Squibb; Kip Guy, St. Jude Children’s Research Hospital; Adam Margolin, Sage Bionetworks; and Art Morales, Novartis. In addition, a subset of the NIH MLP Program Team actively participated in biweekly conference calls to review and shepherd the BARD development progress.
Data Migration, Deposition, and Curation
Data Migration
As anticipated at the outset of BARD, the most challenging aspect of the project was migrating existing data into new data structures and raising the level of annotation to the minimum standard recommended by RDM. The source data from PubChem have little or no controlled vocabulary—they are nearly all free-text entries—and a simple computer transfer of data was not feasible due to inconsistent usage of terms and many exceptional cases. To address this issue, the BARD team engaged scientists at the MLP Centers who were familiar with the experiments to manually annotate their data using BARD’s controlled vocabulary. Because the availability of these experienced scientists was limited, we further augmented the manual-annotation effort by training a team of junior biologists in the RDM standard, and we then had them focus on providing an initial attempt at manual annotation by reading the descriptions in PubChem. The experienced scientists then worked with the annotation team to review and approve the annotations.
The volume of MLP result data in PubChem is fairly high (~1.5–5 billion data points), whereas the volume of assay-definition and experiment data is reasonably low [~6000 PubChem assay identifiers (AIDs)], so we used a different strategy to handle results. We divided the problem into three stages: mapping PubChem column headers [i.e., target identification (TID) labels] to BARD result types and standard units of measure, specifying the relationships among these result types for each experiment, and loading the results into BARD. The first stage was handled programmatically by developing heuristics to map synonymous terms (e.g., PI and PercInhib both map to percent inhibition). The annotation team, using a software interface, performed the second stage manually. The third stage was again handled programmatically, combining the data from the first two stages with raw PubChem result files. To get the maximum value from the resources available, we prioritized migration, annotation, and quality review of the most recent probe projects, working backward to the earlier probe projects and then focusing on projects that did not generate probes.
As data migration progresses, each center has taken on the task of verifying that the migration is accurate and that the data loaded into BARD are meaningful. The effort required for this data migration should not be underestimated—it has been and is a huge task, requiring a significant portion of the project’s resources. One of PubChem’s drawbacks has been that there was no definition of an assay protocol at a more generic level than a run or experiment. This structure means that each AID in PubChem maps to an experiment in BARD, and then BARD must deduplicate identical assay protocols to create a higher-level “assay definition.” Although this deduplication has added to the complexity of the data migration, the benefit to scientific analysis is enormous—particularly for such things as promiscuity analysis, in which the hit count against a given target can then be more accurately assessed.
Separation of Transactional and Analytical Logic
Databases intended for online analytical processing (OLAP) are fundamentally different from those intended for online transactional processing (OLTP). OLTP systems (also known as operational or transactional systems) are optimized for storing data. They apply database-normalization principles so that there is very little or no data redundancy, and they apply the ACID (atomicity, consistency, isolation, and durability) database standard to ensure that data can be safely written using discrete transactions. 33 A consequence of storing data in normalized form is that OLTP databases do not perform well when dealing with queries that combine many different tables of information. These are the types of queries most commonly used for analyzing BioAssay screening data. Also, OLTP databases always contain the most recent version of the data, so it is difficult to perform queries that look at how data have changed throughout time.
OLAP databases are optimized for querying data across multiple dimensions of information. To achieve this optimization, the data are highly denormalized, which means that storage of data is highly redundant. This structure reduces the number of memory-intensive table joins for queries. The data in an OLAP database is more static than in an OLTP database, so different indexing strategies can be applied to optimize query performance. OLAP databases can store historical information to enable queries that look at how data have changed throughout time. 34 The different requirements for OLTP and OLAP systems make it difficult to design a single database that can do both, and past attempts to do so have resulted in complex, high-maintenance, poor-performing systems. BARD’s architecture contains both OLTP and OLAP subsystems, using the former to meet the needs of data generators and the latter to meet the needs of data consumers. We chose to use a relational database-focused architecture because it is a well-proven technology and is widely used in the pharmaceutical industry to solve similar problems of data capture and data query.
A team of dictionary curators, scientists who are skilled in RDM, use the Data Dictionary component to maintain the BARD dictionary, which includes the standardized language for describing the minimum information about an assay as well as business rules that enforce what information is optional and what is required. The Data Dictionary component exposes this information via a representational state transfer (REST) API.
Scientists use the CAP component to define assays and upload results in BARD. The academic community, unlike industry, has historically preferred to annotate and curate their assay and result data after experiments have been completed and through the use of additional resources. Industry requires biologists to learn the corporation’s controlled vocabulary and define their assays in compliance with established data standards. The BARD initiative seeks to educate and help academic scientists understand the value of using a controlled vocabulary for target naming, result types, units of measure, experimental methods, and so on, with reusable assay definitions that can be leveraged for multiple comparable experiments. BARD encourages assay annotation and curation to occur before the results are registered to the system by the scientist. The BARD CAP tool enables scientists to provide canonical classification of assay protocols and associated results using predefined data dictionaries ( Fig. 2A ).

(
CAP enables scientists to describe the sequence of experiments conducted throughout the course of a project and to specify the stage for each experiment ( Fig. 2B ). The resulting graph will help other scientists understand the decisions made to arrive at a probe compound. CAP also enables scientists to load results from suitably formatted files. To handle the wide variety of existing bioassay data types and to support future data types, we have designed a flexible file format that supports both flat and hierarchical data ( Fig. 2C ). During the load process, CAP applies validation rules to the incoming data based on the annotations defined for the experiment and alerts the scientist to any problems.
Data Design
The structure of the data in the CAP is a Codd’s third-normal form database,35–37 and it uses name–value pairs to record many of the assay-definition items. This structure provides users with the flexibility to define new assay parameters and experimental choices without additional engineering support. The use of controlled vocabularies in the dictionary enforces consistency and validation with business rules for the assay definition to be implemented. The disadvantage of this approach is that the software code becomes more abstract and is harder to design to the required level of robustness. The assay metadata itself can become part of the coding, and the division of responsibilities between the database and the application code requires careful consideration.
Overall, the BARD data-migration process is adding a significant amount of useful metadata to each assay definition that did not exist in PubChem, except as sentences in free-text protocols and descriptions.
Warehousing the Data
During the past decade, most bio-pharmaceutical companies have segregated their data-capture subsystems into a transactional layer that is separate from a warehouse layer and has sophisticated querying, reporting, and visualization tools. 22
In line with this practice, BARD was required to address two related but distinct data-storage requirements. The data-deposition component of BARD represents the transactional system in which data are strictly normalized. Importantly, the data-deposition system is not required to store external information (such as metadata or derived data) and, in general, is a short-term store. The second storage component of BARD is the data warehouse. This store is a permanent store of the data received from the data-deposition system, and its primary function is to serve data in response to arbitrary, user-defined queries. A key feature of the data that are stored in the warehouse is that certain data are transformed into a form that is more useful for general queries. In addition to such transformations, data imported into the warehouse from the deposition system may be enriched with external data or linked to preexisting data.
The BARD data warehouse is currently based on MySQL 5.1 (Structured Query Language) and implements a number of optimizations to ensure high query performance. Chief among these optimizations is the use of solid-state drives (SSDs) to host database files. Although appropriate indexing coupled with large amounts of random-access memory allow the database to avoid spending significant time reading from disk, the use of SSDs allows the system to minimize the time spent reading from disk in the case of cache misses. We have also implemented multiple levels of caching to ensure that queries are not repeated. Although MySQL implements its own caching, the BARD warehouse implements a layer of caching so that if a query is repeated, the results are instantaneously available. Finally, we have implemented a form of cluster computing, such that we have multiple instances of the database running on a cluster of computers. Incoming requests are distributed among these instances. It is important to note that the warehouse is essentially database agnostic, allowing one to replace MySQL with other open-source or proprietary database software.
Although the BARD warehouse supports rapid queries, such queries must still be formally defined (usually via SQL). In contrast, most modern search systems provide users with the ability to enter arbitrary text searches. To enable such capabilities, BARD uses the Lucene full-text search engine to extract and analyze descriptions, annotations, and even numeric and categorical fields. 38 As a result, BARD supports arbitrary queries as well as filters that allow a user to progressively drill down to increasingly specific results. Finally, the BARD warehouse supports extremely rapid chemical-structure searches (similarity, substructure, and superstructure) using custom indexing technology. 39
The BARD warehouse component periodically queries the data-deposition components (i.e., the Data Dictionary and CAP) for new data. When such data are available, the warehouse performs the extraction, transformation, loading, and indexing procedures to make data available for query.
Data Access and Visualization
Chemical biology data are inherently multidimensional as they connect chemical-structure to biological responses as well as to higher-level biological entities such as pathways and interactions. As a result, the intuitive summary and visualization of such data are nontrivial tasks. A variety of commercial and open-source tools have been developed that attempt to tackle this problem. The ABCD platform from J&J 22 is an exemplar of an integrated informatics system that was designed to support multiple visualization schemes ranging from enhanced SAR maps 40 to single R-group polymorphisms. 41 Cytoscape 42 is an open-source tool that has been designed with a focus on network visualizations but has extensive support for data integration and the visualization of new data types (such as chemical structures).
Both examples highlight the need for a flexible platform on top of which multiple different interfaces and visualization paradigms can be built. Furthermore, the BARD platform represents a next-generation toolset and resource for public chemical-biology data and must be capable of supporting a diverse community that includes less (computationally) experienced biologists and chemists, informatics-savvy power users, and computational chemists and biologists. From a scientist’s perspective, the ability to pose relevant questions, retrieve results quickly, and present information in a way that offers insights into the underlying biological processes is more important than how the data are stored.
At the lowest level of access to BARD is the availability of a REST API, which is primarily intended for programmatic interactions and has emerged as a predominant web API design model. 43 The API provides access to individual entities (assays, projects, and experiment results) at varying levels of detail, allowing for the development of responsive and diverse clients (e.g., desktop, tablet, and mobile clients). Currently, the API supports JavaScript object notation (JSON) as the only response format, although chemical structures can be accessed in simplified molecular-input line-entry specification (SMILES) or structure-definition file (SDF) formats. Importantly, the API abstracts the details of the underlying warehouse to allow for robust client development in the face of changes to data structures and storage details.
The REST API is not generally intended for consumption by practicing bench chemists or biologists, but rather by informatics-savvy consumers who can integrate their own tools with BARD data (see Fig. 1 ). In this way, the API serves as a foundation for more traditional graphical-user-interface development, allowing future development of this sort to be explored by the wider community. Building on the success of prior successful projects by team members in either industry or academia, the BARD team has designed two specific query tools, a Web Query Client and a Desktop Client, that address a broad range of data-analysis and -visualization needs for chemical-biology scientists, such as SAR tables ( Fig. 3A ), histograms ( Fig. 3B ), and sunburst visualizations ( Fig. 3C ). 44 These tools allow users to interrogate the data without knowing SQL or the underlying data structures. Scientists can also retrieve the results in a variety of tabular or other formats.

(
A key aspect of the BARD platform is that it does not claim to solve every challenge in visualization and analysis. To this end, the BARD architecture supports the concept of plug-ins—user-provided software packages that can be deployed within BARD. On deployment, the plug-in appears as a new resource within the REST hierarchy. The plug-in architecture has nearly unlimited flexibility in the nature of services that can be provided, from a simple plain-text interface to a rich HTML interface, or from simple computations on single compounds or assays to complex analyses of large data sets with rich HTML output. Thus, a plug-in could conceivably present a novel user interface to the BARD data warehouse, with the advantage over a client application of the plug-in having direct access to the underlying database.
To validate the plug-in architecture and help drive its development, the UNM team adapted their BADAPPLE promiscuity tool45,46 to work as a plug-in, and they extended their methodology to take advantage of the structure and annotation provided by BARD. Additional plug-ins are under development and will be deployed when ready to further enhance the use of BARD.
Outreach and Community Adoption
An outreach work stream, led by Vanderbilt University and with participation from all of the institutions collaborating on BARD, was charged with connecting with a wide variety of stakeholder groups in the chemical-biology community. In the early stages of the project, we focused on recruiting interested scientists to participate in the development of RDM standards, persona-research interviews, 47 requirements definitions, and user–interface mockup feedback sessions. As the development of BARD progressed, we conducted multiple rounds of usability studies to gather additional feedback. The formal review performed by the ESP provided further feedback that guided the development of BARD. More recent outreach activities have involved engaging scientists knowledgeable about MLP data to help verify their quality and presenting BARD at conferences and user-group meetings to generate interest within the community.
Future Directions
One of the most important advances of BARD is the creation and adoption of an internal standard for exchanging assay definitions and their associated results among different MLPCN Centers. The addition of a well-curated dictionary connected to relevant external ontologies would, if adopted by the community, greatly enhance the community’s ability to share information and develop interinstitutional science in the future. The broad range of collaborating partners in BARD along with the extensive outreach efforts to engage industry and academia have enabled major progress toward a standard that builds on current best practices in RDM. This early engagement of the scientific community will also increase the likelihood of widespread adoption of BARD as a bioassay-results repository.
The next step in this process will be to develop a formalized assay definition standard (ADS) based on the logical model, data dictionary, and business rules developed by the BARD team. The physical implementation of this standard will be the assay definition format (ADF). The ADF will be used within BARD to simplify and make consistent the transfer of data from the transactional (input) system to the warehouse (storage) and the analytical tools (output). The ADF can also be used to load large result sets into CAP and BARD. The ADF will include sections for defining the assay protocol, the experiment, and then the resulting data. Each of these sections could be used separately, if desired. Also, the standard allows for the transport of new vocabulary terms from one system to another under the management of the RDM processes and guidance.
As a practical usable standard, the BARD team anticipates that with the appropriate outreach and support of industry groups, such as the Pistoia Alliance (http://www.pistoiaalliance.org) and OpenPHACTS,48,49 this standard could facilitate assay data exchange between industry and academia. For example, OpenPHACTS has provided a useful and intuitive web portal through its Explorer. They have also provided a framework for the “critical pharmacological questions” that is useful for defining the kinds of research questions that are impactful and relevant to compounds (Cluster I) and pathways (Cluster II). 49 BARD has a significant focus on structured data, as a large proportion of the BARD project concerns structured-data capture and value-added data migration, which is quite different from OpenPHACTS. 49
Conclusions
This case study reviews the motivations and context for BARD, and the organizational challenges surmounted by the multicenter BARD team, with a view to evolving the existing best practices for chemical-biology data. The BARD team is partnering with peers in industry and academia to formalize and adopt an assay-definition standard. Current BARD development activities include creating a technical toolbox for innovation that includes a Catalog of Assay Protocols, Web Query, and Desktop Client tools, with a plug-in infrastructure that enables both novice and expert users to query, mine, and interpret chemical-biology data, specifically the MLP data. As the BARD project team looks toward the future, team members and other early adopters must continue to serve as agents of change that encourage the use of the developed research data-management standards.
Footnotes
Acknowledgements
The authors wish to thank the principal investigators of their home MLPCN Centers—Prof. Stuart Schreiber (Broad), Dr. Larry Sklar (UNM), Dr. Chris Austin (NCATS), Prof. Hugh Rosen (Scripps), Dr. Craig Lindsley (Vanderbilt), and Dr. John C. Reed and Dr. Michael R. Jackson (Sanford|Burnham)—for their support of the respective teams. We especially thank Prof. Schreiber for his leadership in catalyzing the discussion within the MLP that led to BARD; Dr. Jackson for inviting key pharmaceutical information technology representatives to demonstrate their solutions to the MLPCN Steering Committee, which catalyzed additional discussions; and Dr. Chris Austin for vigorously publicizing and supporting the BARD initiative. We also wish to thank the NIH MLP Program Team (PT) biweekly BARD teleconference participants: Drs. Ajay Pillai (NHGRI), Carson Loomis (NHGRI), Ronald Margolis (NIDDK), Yong Yao (NIMH), Jamie Driscoll (NIMH), Linda Brady (NIMH), Ingrid Li (NIMH), Christine Colvis (NIDA) and Enrique Michelotti (NIMH). In addition, we thank the subset of the NIH MLP PT that is active in usability studies to review and shepherd BARD’s development progress. Additionally, we wish to thank Dr. Pillai for his participation in biweekly engineering teleconferences and his valuable feedback and suggestions. We also wish to thank Mr. Michael North (NIMH) for his coordination and succinct minutes of BARD biweekly and ESP meetings.
Abbreviations
BARD, BioAssay Research Database; MLP, Molecular Libraries Program; BAO, BioAssay Ontology; CAP, Catalog of Assay Protocols; RDM, Research Data Management; OLAP, online analytical processing; OLTP, online transactional processing; REST API, representational state transfer application programming interface.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding for BARD came directly from the Common Fund through the Molecular Libraries Probe Production Center Network (RFA-RM-08-005) as an administrative supplement directly to the centers following receipt and review of center proposals in response to an request for proposal crafted by Dr. Ajay Pillai (NHGRI).
