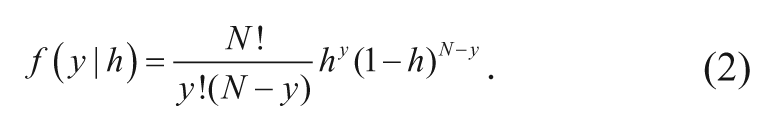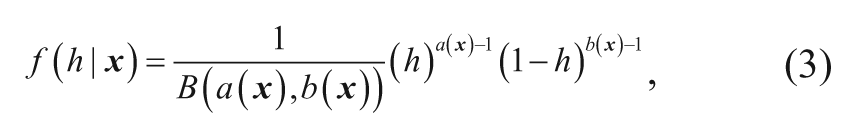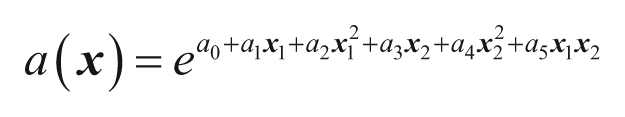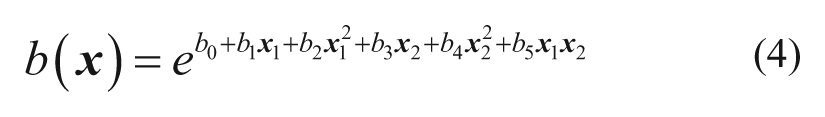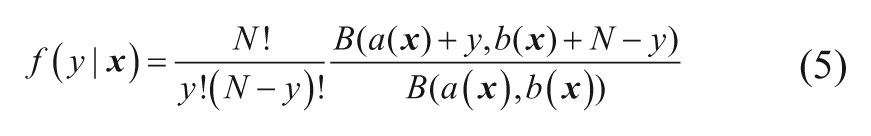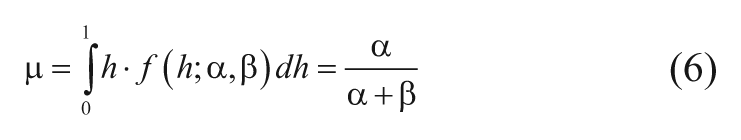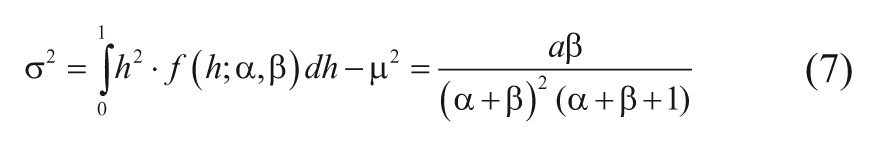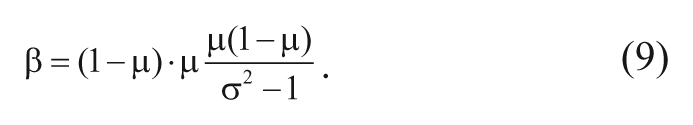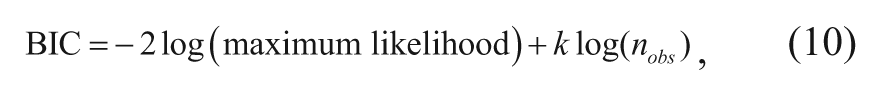Abstract
High-throughput screening (HTS) is widely used in the pharmaceutical industry to identify novel chemical starting points for drug discovery projects. The current study focuses on the relationship between molecular hit rate in recent in-house HTS and four common molecular descriptors: lipophilicity (ClogP), size (heavy atom count, HEV), fraction of sp3-hybridized carbons (Fsp3), and fraction of molecular framework (fMF). The molecular hit rate is defined as the fraction of times the molecule has been assigned as active in the HTS campaigns where it has been screened. Beta-binomial statistical models were built to model the molecular hit rate as a function of these descriptors. The advantage of the beta-binomial statistical models is that the correlation between the descriptors is taken into account. Higher degree polynomial terms of the descriptors were also added into the beta-binomial statistic model to improve the model quality. The relative influence of different molecular descriptors on molecular hit rate has been estimated, taking into account that the descriptors are correlated to each other through applying beta-binomial statistical modeling. The results show that ClogP has the largest influence on the molecular hit rate, followed by Fsp3 and HEV. fMF has only a minor influence besides its correlation with the other molecular descriptors.
Introduction
High-throughput screening (HTS) has become a standard lead generation approach in the pharmaceutical industry. 1 In the quest to improve the success rate of HTS-based drug discovery, it is important to understand the relationship between the likelihood of a small molecule being biologically active in an arbitrary HTS assay and its physicochemical and structural properties. Such knowledge can provide guidance, for instance, in the prioritization of compounds for HTS when selecting compounds for the more resource-consuming hit confirmation and validation steps or for enhancement and exploitation of screening compound collections.
A compound’s probability to be active in an arbitrary HTS is closely related to its ability to interact with different target proteins. This is often referred to as molecular promiscuity. 2 Large pharmaceutical companies have accumulated abundant in-house screening databases that encompass the biological activity of millions of compounds across hundreds of different types of HTS assays. Analysis of such data can provide valuable information for identifying influencing factors on molecular promiscuity by estimating the promiscuity with the molecular hit rate (MHR), that is, the fraction of times a compound has been assigned as active in HTS. Published computational models that investigate the relationship between promiscuity and adverse effects are based on data sets that contain compounds screened against panels of protein targets.3–5
Several recent studies have shown that lipophilicity (ClogP) is a principal determinant of molecular promiscuity. Other molecular properties such as ionization state, basicity, and presence of certain functional groups have also been found to influence promiscuity.6–10 Hopkins et al. 6 reported that compound promiscuity decreases with higher molecular weight (MW) based on HTS data. Diller and Hobbs 11 published a statistical analysis of a company’s historical HTS data, in which they examined the relationship between physicochemical properties, substructures, and likelihood of displaying biological activity, irrespective of target class.
Recently, investigations of molecular topology–related descriptors and their relation to compound quality have been published. Aromatic ring count, 12 fraction of sp3-hybridized carbon atoms (Fsp3), 13 the difference between aromatic and sp3 atom count, 14 and fraction of molecular framework (fMF) 15 are examples of topology descriptors that have been studied in addition to the commonly investigated descriptors, ClogP and MW. Another study of an in-house HTS data set indicated a positive correlation for MHR with ClogP and with MW and a negative correlation for MHR with Fsp3. 16 However, to our knowledge, the correlation between the descriptors has never been addressed in these analyses. For instance, ClogP and Fsp3 might be correlated in an HTS data set, making it questionable if Fsp3 has any additional influence on the MHR besides being correlated with ClogP. It is therefore important to revisit the correlation between molecular descriptors and MHR with rigorous statistical methods that are capable of estimating quantitatively the relative influence on the MHR of the different descriptors. Further investigations are also needed to establish the correlation between MHR and size, since earlier investigations have shown conflicting results. 16 In addition, topological descriptors have so far not been thoroughly investigated in relation to MHR.
Here, the relationships between MHR, based on in-house HTS screening data, and the molecular topology descriptors Fsp3 and fMF in addition to ClogP and heavy atom count (HEV) are investigated. To quantify the independent contributions of the molecular descriptors on the MHR, beta-binomial statistical models were built to model the probability distribution of hit rate as a function of the descriptors. The importance of each descriptor’s influence on the MHR was judged by how well the beta-binomial statistical models could estimate the observed MHR.
Materials and Methods
Data Set
The initial data set used in this study contains single-point (SP, screened at single concentration) data from 260 assays, which have been carried out at AstraZeneca since 2008. The data were heterogeneous and therefore had to be cleaned before the statistical analysis. In the first step in this cleaning procedure, assays that did not contain any active compounds were excluded. For the compounds tested at several concentrations in the same assay, only data points for the most occurring screening concentration were kept. Activity labels other than active or inactive were discarded. In cases where compounds had been tested multiple times in an assay, the activity flag was set to the majority label, while for equal numbers of active and inactive data points, the compound was labeled as inactive. Only assays with more than 100,000 unique compounds were retained. Furthermore, only compounds with an HEV between 6 and 45, ClogP between −2.0 and 6.0, and compounds assigned as neutral, acidic, or basic ionization state were included. Compounds that had been tested in less than 15 HTS assays were discarded. The cleaned data set used in the analysis consisted of 1.3 million compounds tested across 102 assays. The cleaning procedure was done by running an in-house python script. 17
Molecular Hit Rate and Properties Calculations
The MHR for each compound was calculated as the ratio of the number of times a compound was active and the number of times it has been tested (equation (1)).
Molecular lipophilicity (ClogP) was estimated with a commercial program. 18 HEV, Fsp3, and ƒMF were calculated with an in-house C++ program based on Openeye toolkits. 19
Fitting MHR with Beta-Binomial Distribution
Beta-binomial model for MHR
The influence of the different descriptors on MHR was quantitatively measured by empirical hit rate probability models, as follows. The hit rate, h, corresponds to the probability of a compound being active in a random assay. In a fixed number of assays and for a fixed compound, it might be reasonable to assume that the outcomes are independent and that h is constant, so that the number of times the compound would be registered as active has a discrete probability distribution called the binomial distribution. With these assumptions, the probability of a compound being active y times in a fixed number of assays, N, is given by the binomial distribution with probability mass function according to equation (2).
However, the hit rate for compounds with the same descriptor value varies, and the hit rate is actually unknown. An approximate way to model this is to assume that for a randomly chosen compound, the hit rate will vary accordingly to a beta distribution with a probability density function (pdf) depending on the molecular descriptor values according to equation (3).
where a(x) > 0, b(x) > 0, and 0 < h < 1.
The variable
In the current study, the functional form of the exponents is modeled using polynomial terms of the descriptors. Equation (4) shows an example of these functions when using squared terms for two descriptors, including cross terms.
Now, equations (2) and (3) describe the full probability distribution of y and h together, for any given
This is therefore the probability distribution describing the observations, since the unknown h has now been removed. Equation (5) is obtained by multiplying equations (2) and (3) together and integrating out the unknown hit rate variable (applying the usual rules of probability). This probability distribution is used to build the hit rate models.
Parameter estimation of the beta-binomial model
The coefficients a0, a1, a2, . . ., b0, b1, b2, . . . (corresponding to the number of descriptors used in the model) need to be estimated to obtain the a(
The difficulty for such procedures is to find good initial values, so that the iterations can converge quickly to a good solution of the likelihood equations. To achieve this, the process is broken down into a sequence of steps (
First, the descriptor ranges were divided into small bins so that the functions a(
So for each small bin, there was a simpler problem with just two constants to estimate, and a statistical method called method of moments 20 was used to get initial guesses of α i and β i . It seemed reasonable at this stage to try relatively crude values as a quick solution, so the sample estimates of MHR (namely, y/N) were used to estimate the mean (m) and variance (σ2) of the hit rate distribution (equation (3), with constant a and b at this stage), given in equations (6) and (7).
where f(h;α,β) is the beta-probability density function (equation (5), with constant parameters). Clearly, this yields the expressions in equations (8) and (9).
Substituting the MHR estimates of the mean and variance into equations (8) and (9), initial estimates for α and β were obtained for each bin, which were used in the optimization procedure to obtain the local optimal maximum likelihood estimates of α, β. As stated above, ordinary least squares polynomial fitting was done to estimate the global coefficients (a0, a1, b0, b1, . . .) of the functions a(
Estimating the polynomial degrees
To choose the best “model dimension” (i.e., number of descriptors and/or polynomial orders), the Bayesian information criterion (BIC) (equation (10)) was used. 21
where k is the number of estimated parameters and nobs the number of data points. The first term in this expression is a measure of the “lack of fit” of the model, which will decrease every time a parameter is added to the model. The second term is a penalty value, which increases with the number of independent variables added to the model. So, broadly speaking, it is not allowing the possibility of continually improving the fit by just adding more and more terms and thus balances the improvement in fit with the increase of model complexity. A smaller BIC value represents a better fitted model, so the aim is to find the model with the smallest BIC.
An R 22 script was written to do the parameter estimation for beta-binomial distribution models and BIC score calculation. The numerical optimization was done by using the BFGS algorithm 23 implemented in the “Optim” module of R.
Results
Relationship between the MHR and Descriptors for the HTS Data
Before applying the beta-bionomial statistical model, the influence of each descriptor on MHR was investigated individually. The relationship between MHR and lipophilicity (ClogP), size (HEV), Fsp3, and fMF is shown in Figure 1 . For each property plot, compounds were binned according to their property value and plotted against the average hit rate of each bin. Bins containing less than 1% of the whole compound set were removed.
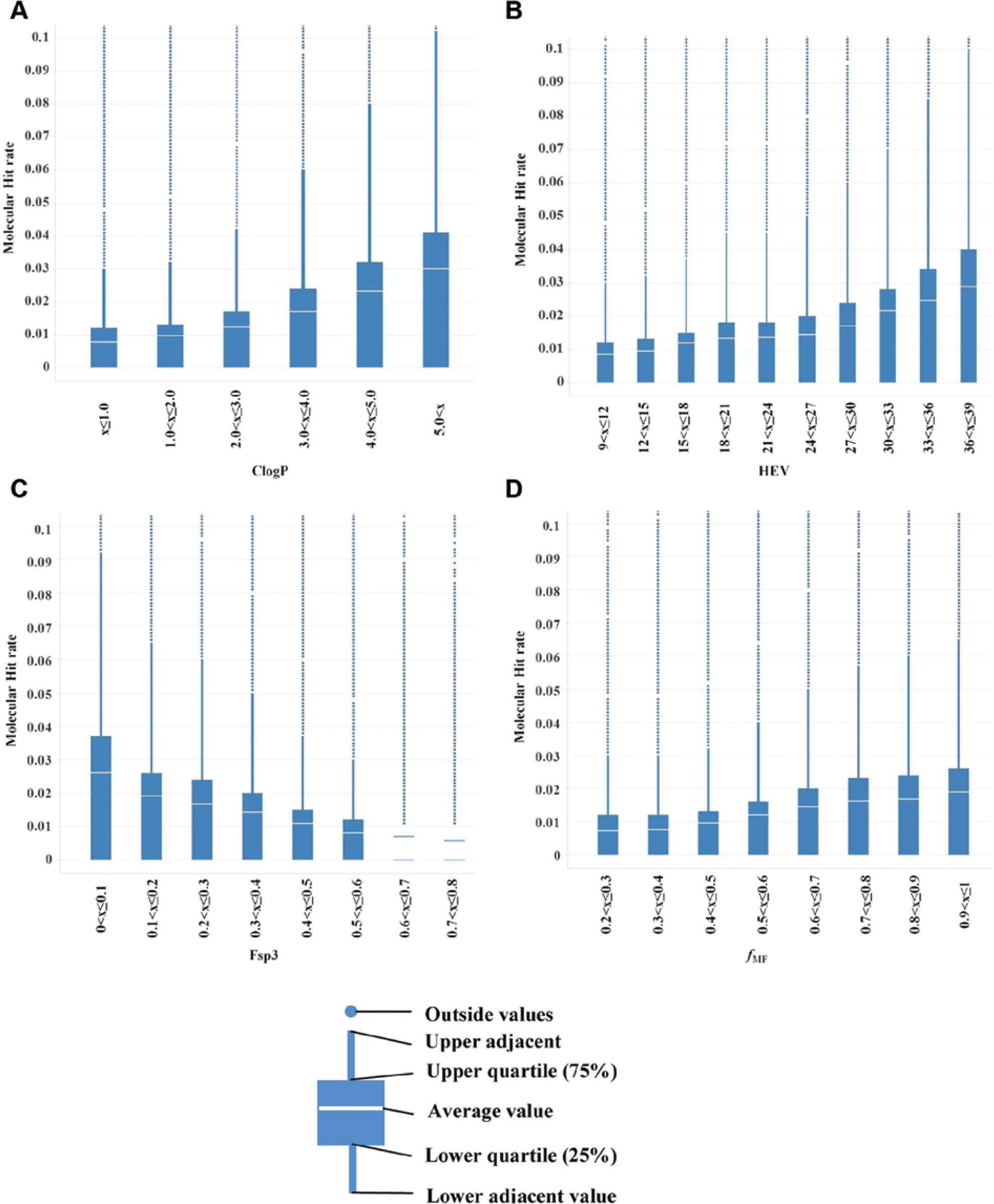
The relationship between molecular hit rate (MHR) and (
ClogP, HEV, and fMF are here positively correlated with MHR ( Fig. 1 ). This is consistent with previous studies.15,16,24 It was previously found that compounds with fMF values above 0.65 are considerably more promiscuous than those with lower fMF values. However, a more linear relationship between MHR and fMF was found in this study. This difference might be due to the difference in data sets used in the two studies. On the other hand, Fsp3 shows a negative correlation with MHR (shown in Fig. 1c ), which is in line with earlier investigations.16,25 The correlations correspond to a shift in the location of the MHR distribution. What is also striking from these graphs is the suggestion of a long upper tail in the distribution of MHR, which appears in all bins. These are features that can be captured by the beta distribution. Generally, it is not surprising to observe compounds with high MHR values in HTS data collections, considering the conserved binding pocket for kinases and the historical popularity of the kinase family in drug discovery, as well as other types of frequent hitters, such as compounds that interfere with the HTS signal readout technology. Actives from each primary assay were identified in their follow-up screening cascades to filter out false positives by using the corresponding concentration-response (CR) tests to calculate the MHR. Even though the hit rates decreased at CR compared with SP data, the same trends in the correlations between MHR and the four different properties were observed (data not shown).
Descriptor Intercorrelation Analysis
The intercorrelations among the descriptors are shown in Figure 2 . For each plot, the descriptor on the x-axis is binned in the same way as in Figure 1 and plotted against another descriptor. The box plots visualize the compound distribution of the descriptor on the y-axis in each bin of the descriptor on the x-axis. ClogP and HEV show a positive correlation ( Fig. 2a ), while ClogP is negatively correlated with Fsp3 ( Fig. 2b ). Figure 2f shows that Fsp3 and fMF are negatively correlated. No monotonic relationships could be found for ClogP versus fMF ( Fig. 2c ), HEV versus Fsp3 ( Fig. 2d ), and HEV versus fMF ( Fig. 2e ). Thus, it is clear that in an analysis of the MHR versus the descriptors, the cross-correlation between the molecular descriptors must be taken into account. This was accomplished by modeling the MHR as a function of the descriptors with a beta-binomial statistical model. These graphs illustrate that complex relationships exist between the descriptors, with a large variation in the values throughout the ranges. Thus, to help determine the dependence of hit rate distribution on the descriptors in light of these complex relationships, we needed to resort to a complex probability model for that distribution.
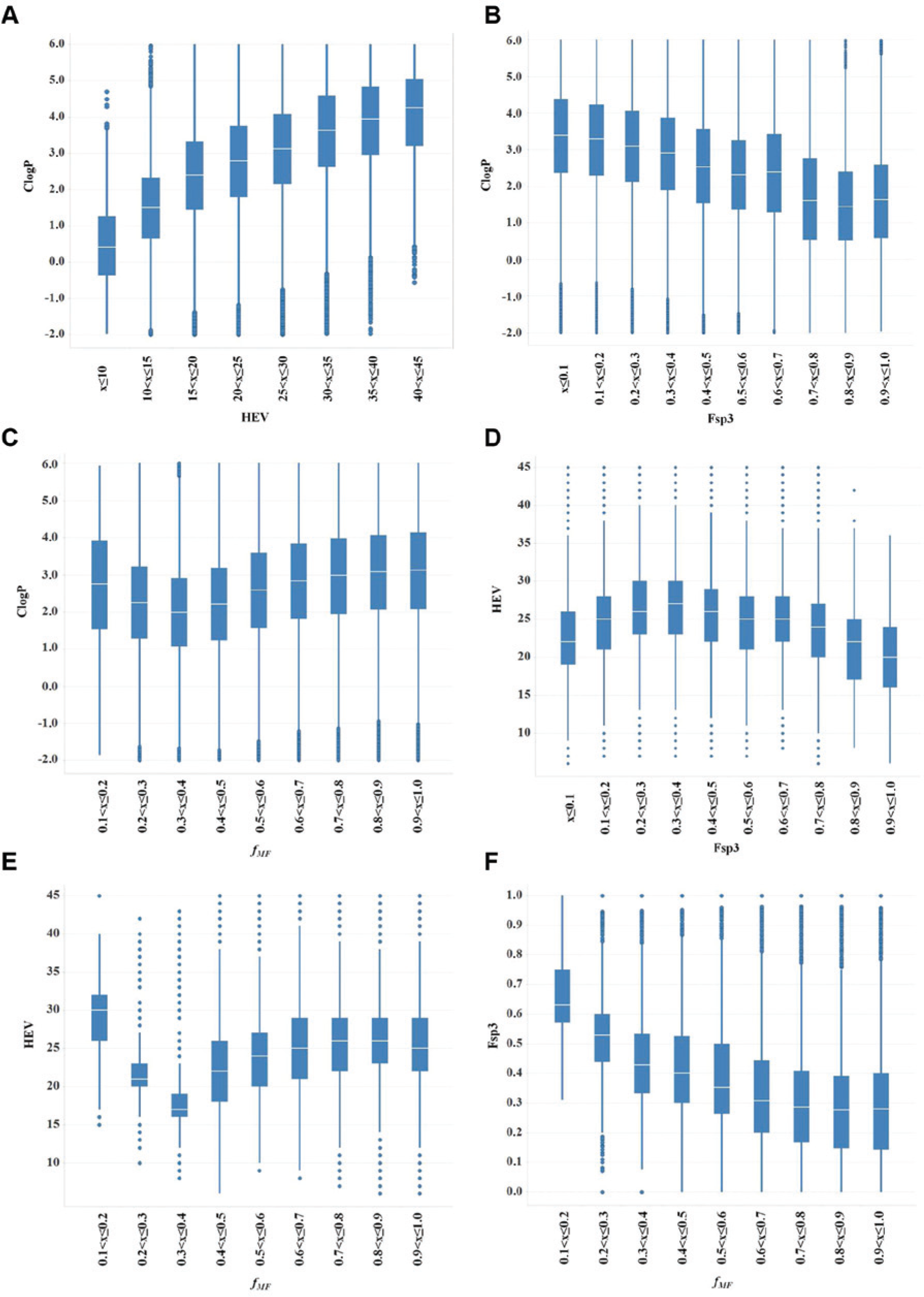
Relationship between (
Fitting the MHR with a Beta-Binomial Statistical Model
Statistical models were built to quantitatively assess the influence of different descriptors on the distribution of MHR. The influence of the descriptors on the distribution of MHR could then be measured by examining the difference between the observed and modeled hit rate distributions.
ClogP, HEV, Fsp3, and fMF were used as independent variables to build the hit rate beta-binomial models, and as a first step, only linear terms of the descriptors were employed for model building. As explained above, a systematic procedure of using BIC was employed to select the most appropriate model for the MHR distribution. The models were built in a stepwise manner. First, only one descriptor was added into the model, and the best one-descriptor model was identified. Then, each descriptor was added, one at a time, into the best model from the previous cycle. This procedure was repeated until no improvement of adding a new variable could be found. The BIC score value was used to measure the model quality at each cycle, and Table 1 demonstrates how the BIC score varied during this process. For the single-descriptor models, the lowest BIC value was found for ClogP. ClogP is thus regarded by the procedure as the most influential of the four descriptors on the MHR distribution. The best two-descriptor model was accordingly the one combining Fsp3 and ClogP. Furthermore, the best three-descriptor model was obtained when HEV was added. In the end, upon adding fMF, the model exhibited only a small additional decrease of the BIC value, indicating that fMF does not have as great an independent influence on the hit rate as the other descriptors.
Bayesian Information Criterion (BIC) Values in the Four Rounds of the Model Building Procedure with the Descriptors Added One at a Time to the Model.
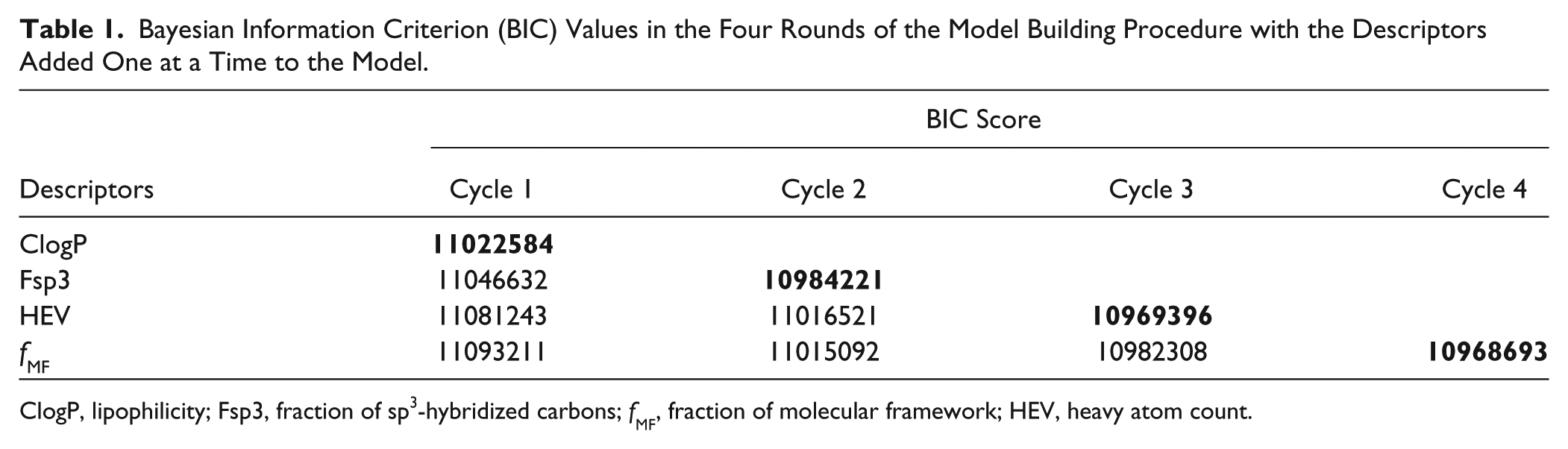
ClogP, lipophilicity; Fsp3, fraction of sp3-hybridized carbons; fMF, fraction of molecular framework; HEV, heavy atom count.
When looking at the decrease of the BIC values for the best model in each round of the model-building procedure, it can be seen that each time a descriptor was added, the BIC decreased, and especially when adding Fsp3, a large improvement was obtained. When HEV was added, the decrease was smaller than that of Fsp3, and adding fMF gave the smallest decrease. This suggests that the contributions of the four descriptors to modeling the MHR distribution are ordered as ClogP > Fsp3 > HEV > fMF. The extra contribution from Fsp3 on top of ClogP was especially large. When fMF was added, the improvement of the model was much smaller.
To further improve the flexibilty of the model for better capturing the experimental hit rate data, higher degree polynomial terms were included as previously discussed. As shown in Figure 3 , higher degree polynomial terms were added to the ClogP, ClogP + HEV, ClogP + Fsp3, ClogP + Fsp3 + HEV, and ClogP + Fsp3 + HEV + fMF models, which demonstrated the improvement in BIC of the ClogP model when Fsp3 and HEV were added to the model, respectively, as well as when polynomial terms were gradually added to the three different models. For the ClogP, ClogP + HEV, and ClogP + Fsp3 models, the BIC scores decreased up to degree five, while in the ClogP + Fsp3 + HEV model, the BIC score decreased until degree four and then slightly increased at degree five. It is also interesting to see that the BIC scores of the ClogP + Fsp3 models were much lower than those of the ClogP + HEV models. This clearly shows that the influence of Fsp3 on MHR was larger than that of size, which is consisent with the conclusion drawn from the previous linear-only term hit rate models. In addition, when quadratic and cubic terms were added, BIC decreased. When higher polynomial terms were included, although the decrease of the BIC value was much smaller, it was still much larger than the nominal level of 10 that is regarded as a statistically significant decrease. 20 The curve for the four-descriptor ClogP + Fsp3 + HEV + fMF model was also generated. The degree of polynomial was increased until three, and the BIC value of the four-descriptor model was very close to that of the three-descriptor ClogP + Fsp3 + HEV model. When the polynomial degree was increased to four, the BIC value increased dramatically to a level well above the BIC range in Figure 3 . Therefore, no further increase of the polynomial degree was done. It is noted that the model with fMF included will not beat the fourth-degree three-descriptor model, as seen in Figure 3 . Since the improvement to the model was relatively small compared with the other three descriptors when fMF was added, for practical reasons, this descriptor was excluded from further consideration in the model building. The polynomial models at degree four were chosen for each descriptor combination for further studies due to the generally low BIC values at the fourth-degree level.
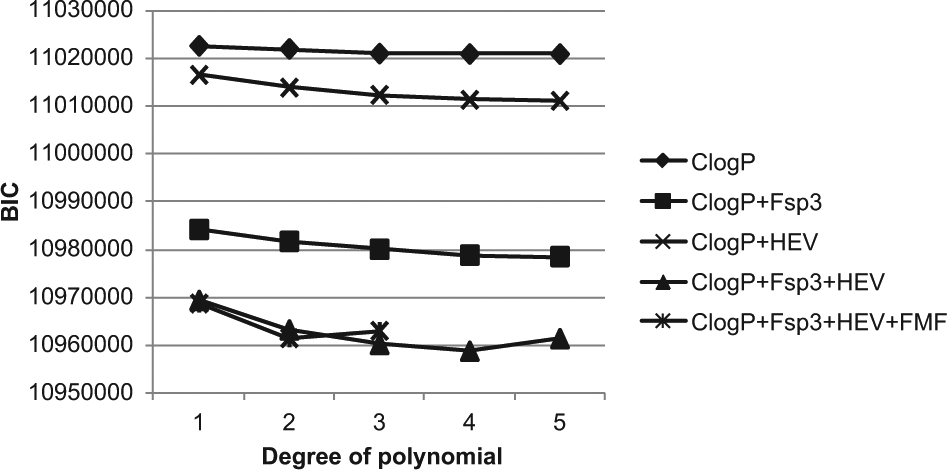
Decrease of the Bayesian information criterion (BIC) value as the degree of polynomial increased for the beta-binomial statistical models. Due to the very large number of parameters for the five-degree three-descriptor and three-degree four-descriptor models, it is difficult for the optimization algorithm to converge to the optimal solution of the likelihood equations. As with all numerical optimization procedures, there is no guarantee of obtaining an optimal solution, especially with very large numbers of variables to optimize over. Given the size of these two models, we felt justified in calling off the search and used the suboptimal solutions in the figure. ClogP, lipophilicity; Fsp3, fraction of sp3-hybridized carbons; fMF, fraction of molecular framework; HEV, heavy atom count.
The cumulative distribution functions (cdf) for the ClogP and ClogP + Fsp3 polynomial models (both with degree four) are shown in Figure 4 . The cdf at any hit rate value is the area under the pdf (the beta distribution, equation (3)) between zero and the specific value. This value indicates the cumulative fraction of compounds whose hit rates are lower than a specific MHR for a given descriptor value. Each plot in Figure 4 corresponds to compounds with a given ClogP value, and four representative ClogP values (ranging from high to low ClogP values) are displayed. The black curves in these plots always refer to the cdfs for the ClogP model. The colored curves in each plot refer to the cdfs of the ClogP + Fsp3 model, and the color scheme corresponds to different Fsp3 values in the model. It can be seen from these cdf curves that for compounds with a fixed ClogP value, an increase in Fsp3 will decrease the probability of having a high MHR, and this is true at all four ClogP values. This means that increasing Fsp3 will tend to decrease the MHR, and the effect is independent of the influence of ClogP. Figure 1c has already shown that Fsp3 is negatively correlated with MHR and, not surprisingly, the effect is reflected in the cdf of the ClogP + Fsp3 model as well, giving us some confidence that the model is valid.
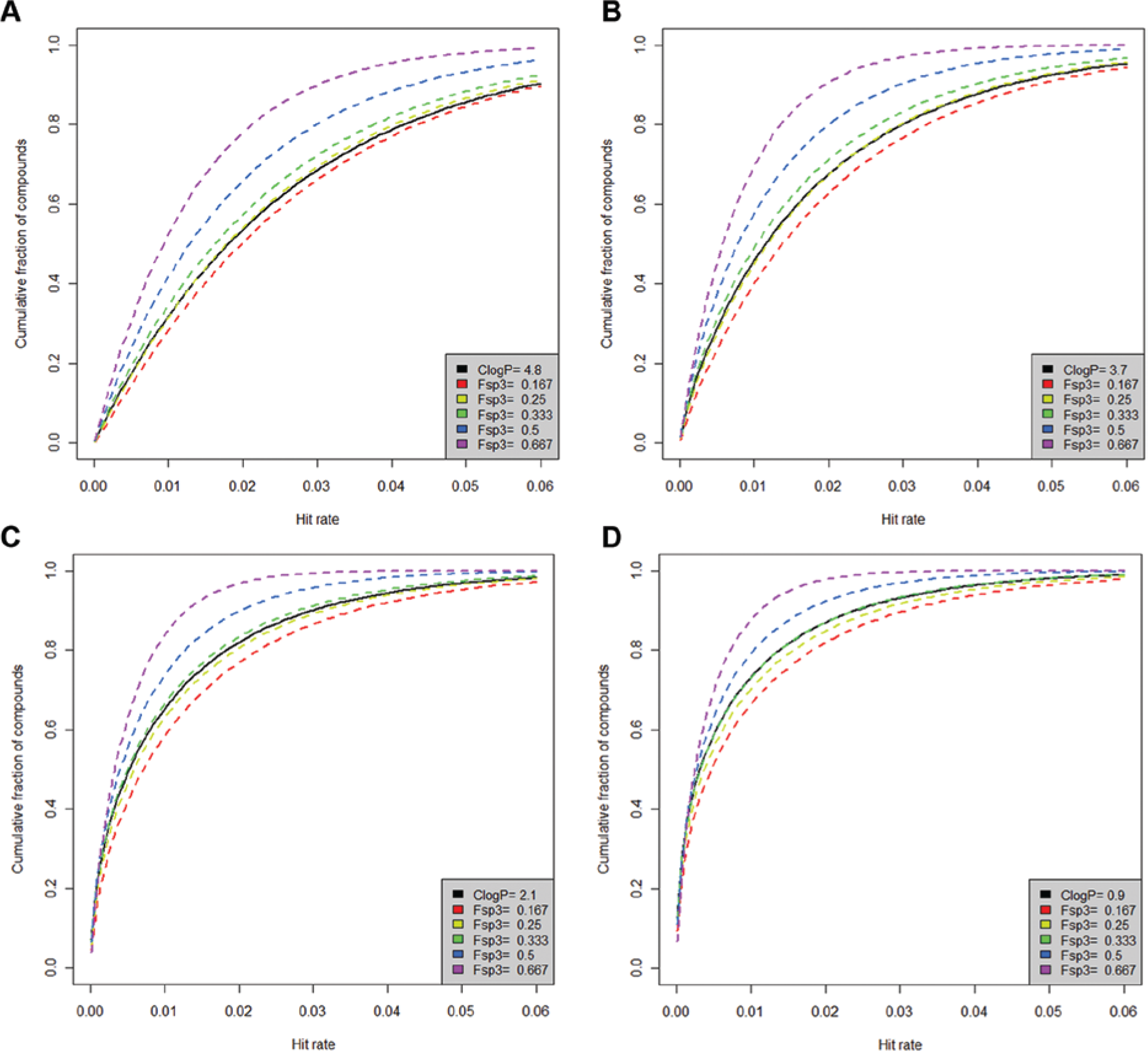
Molecular hit rate (MHR) cumulative distribution function plots for the ClogP model and the ClogP + Fsp3 model. Four plots correspond to compounds with four representative ClogP values: (
It is also of interest to examine the additional influence of molecular size on top of ClogP and Fsp3. The cumulative distributions for compounds in each subgroup that have the same fixed ClogP and Fsp3 values but with different HEV levels are generated and displayed in each plot of
To visually display the difference between the predicted hit rate from the beta-binomial statistical model and experimental hit rate, the surfaces for predicted mean MHR were generated and compared with experimental mean hit rates (
Fig. 5
). The surface in each figure represents the predicted mean hit rate from the model. The colored dot represents the mean experimental hit rate for a given descriptor bin, and the color of the dots corresponds to the compound density in each bin, where green represents the lowest density and purple represents the highest.
Figure 5a
shows the comparison between the four-degree ClogP + Fsp3 polynomial model fitted mean hit rate and mean experimental hit rate data. It can be seen in the figure that the model (blue surface) in general aligns well with the dots, which indicates that the model has a good fit with the experimental mean hit rate. In
Figure 5b
, another surface (red) is added for comparison, which is generated from a four-degree ClogP polynomial model. The red surface has a larger deviation to the green dots, which have high Fsp3 in general, while the surface from the ClogP + Fsp3 model can match them nicely. Similar surface plots are made for the ClogP + HEV model (
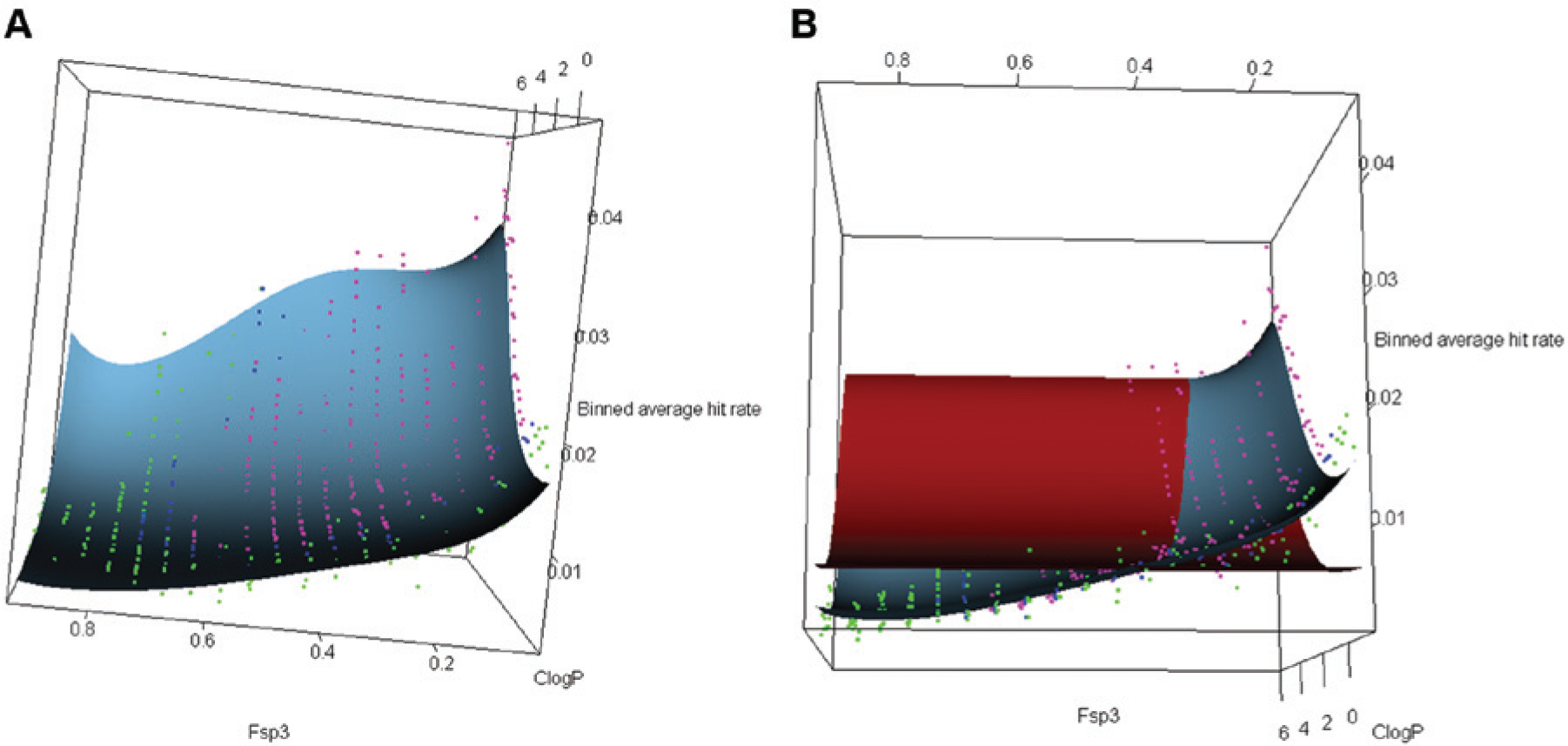
Surface plots comparing (
Discussion
The aim of this study was to investigate the relationship between MHR and the molecular descriptors ClogP, HEV, Fsp3, and fMF. The initial results show the following trends: (1) ClogP and molecular size (HEV) are positively correlated with the MHR data, (2) Fsp3 is negatively correlated with the MHR, and (3) fMF is positively correlated with the MHR, but its effect is smaller than those of ClogP, Fsp3, and HEV. An intercorrelation study of the descriptors has also been carried out, and it was found that ClogP is correlated with Fsp3 and HEV. To find out if the influence of Fsp3 and HEV had additional contributions to the MHR besides being correlated with ClogP and also to quantitatively measure the influence of each descriptor, various beta-binomial statistical models were built by fitting experiment hit rate data to a beta-binomial distribution. ClogP, Fsp3, HEV, and fMF descriptors were used as independent variables, and a BIC score function was proposed to assess the model quality. Different combinations of the four descriptors were used to build beta-binomial statistical models, and their BIC scores were compared. It was found that the descriptor impact on MHR is in decreasing order: ClogP > Fsp3 > HEV > fMF. The analysis of the cdf curves of the models demonstrates that both Fsp3 and HEV have an influence on MHR, which is independent of ClogP, the most influential descriptor. To further improve the model quality, higher degree polynomial terms were added, showing that higher degree polynomial models fit better with the experimental data than the linear term models.
According to this study, the most important property to be considered in finding an active compound is the lipophilicity, followed by Fsp3 and heavy atom count. To our knowledge, for the first time it has been possible to rank the independent influence of the descriptors on the molecular HTS hit rate, taking intercorrelation between the descriptors into account. It is intuitive and has been shown previously that compounds with high lipophilicity and a flat shape are more promiscuous, meaning that they bind more frequently to different targets than hydrophilic and nonflat compounds. The results from this investigation support that promiscuous compounds are more likely to have high lipophilicity and high aromaticity, with an additional smaller contribution from molecular size and fMF. The identified results can be used as guidance in, for instance, the prioritization of compounds for HTS campaigns when selecting compounds for the more time- and resource-consuming hit confirmation steps as well as for general enhancement and exploitation of screening compound collections.
Footnotes
Acknowledgements
The authors are grateful to their AstraZeneca colleagues in the TS department for generating HTS data over the years.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research leading to these results has received support from the Innovative Medicines Initiative Joint Undertaking under grant 115191 (Open PHACTS), resources of which were composed of financial contribution from the European Union’s Seventh Framework Programme (FP7/2007-2013) and EFPIA Companies’ in kind contribution.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.