Abstract
The pregnane xenobiotic receptor (PXR) is a key transcriptional regulator of cytochrome P450 (CYP) 3A, a crucial enzyme in the metabolism and detoxification of xenobiotics and endobiotics. PXR is activated by a wide variety of chemicals and serves as a master regulator of detoxification in mammals. Here, we report a fast evaluation method for PXR-drug interactions using differential scanning fluorometry (DSF). DSF analysis revealed that PXR associates with a fluorescence dye in the native state as well as in the unfolded state, which prevented precise evaluation of any shift in the transition midpoint (ΔTm) due to association with a drug. Hence, we defined a new parameter, (dF/dT)50, where F is fluorescence intensity and T is temperature, to describe the ligand concentration. (dF/dT)50 exhibited better correlation with EC50 (r2 = 0.84) than with ΔTm (r2 = 0.71). The correlation of ΔTm measured using differential scanning calorimetry (DSC) with EC50 (r2 = 0.86) was similar to the above (dF/dT)50 correlation. Therefore, the use of (dF/dT)50 enables DSF to be used for the rapid evaluation of PXR-drug interactions and could provide prescreening to narrow down the collection of candidate ligands that most likely result in transcriptional activation of CYP3A4.
Keywords
Introduction
Cytochrome P450 (CYP) 3A plays a crucial role in the metabolism and detoxification of xenobiotics and endobiotics. Normally, when a cell is exposed to a xenobiotic, genes encoding chemoprotective proteins are induced. This adaptive response provides a mechanism for protecting the organism against foreign chemicals, including pharmaceuticals, pesticides, and steroids. The pregnane X receptor (PXR) is a nuclear receptor for transcription factors that regulate CYP3A transcription and is activated by most CYP3A inducers, despite their structural diversity.1–7 PXR consists of an N-terminal DNA binding domain and a C-terminal ligand binding domain (LBD), which is a characteristic feature of other nuclear receptors such as constitutive androstane receptor, estrogen receptor, and peroxisome proliferator-activated receptor.7,8 Upon having its LBD associate with a ligand, PXR is released from heat shock protein 90 and is activated by forming a complex with the retinoid X receptor and another member of the nuclear receptor superfamily such as steroid receptor coactivator 1 (SRC1). The activated form of PXR then binds to a xenobiotic response element located in the promoter region of the target CYP3A gene, thereby regulating transcription (
Reporter gene assay is a common method for evaluating CYP3A4 induction and has been applied to the high-throughput screening of the drug candidates.9–11 However, its use is limited, as the cytotoxicity of a drug candidate often causes the death of human-derived cell lines. Furthermore, the assay monitors only the final result of the gene enhancement and does not directly detect binding between the drug candidate and PXR, thereby sometimes resulting in false positives.
Because ligand association is generally accompanied by an increase in the transition midpoint (Tm) of protein unfolding, differential scanning calorimetry (DSC) could provide a convenient and alternative tool for detecting protein-ligand interactions 12 and protein stability. 13 Differential scanning fluorometry (DSF) is another method that can rapidly evaluate the protein association of low-molecular-weight ligands. The temperature at which a protein unfolds is measured by an increase in the fluorescence of a dye, for example, SYPRO orange (SYPRO orange protein gel stain, Sigma-Aldrich, St. Louis, MO), which has a higher affinity for the hydrophobic parts of a protein that are exposed upon unfolding. DSF allows for observation of the Tm of more than 96 samples within an hour and exhibits significantly higher throughput than DSC. However, in principle, DSF cannot be used for proteins that possess an exposed hydrophobic pocket, such as PXR, because association with the fluorescent dye leads to a fluorescence intensity that decreases with the unfolding transition. This property prevents precise determination of the Tm shift (ΔTm) that occurs with drug association.
Here, we employed a new parameter, (dF/dT)50, in which F is fluorescence intensity and T is temperature, which describes the ligand concentration at half the initial dF/dT value. The (dF/dT)50 value exhibited good and better correlation (r2 = 0.84) with the EC50 value from reporter gene assays than did the ΔTm in DSF (r2 = 0.71). The present method should therefore markedly extend the usefulness of DSF for drug discovery.
Materials and Method
Construction of the Expression Plasmid
The expression plasmid for the PXR-SRC1 fusion protein was constructed as follows. The fragment encoding the human PXR (130-434) domain was amplified using the oligonucleotides PXR-f (5′-CCTCTAGACATATGAGTGAA-CGGACAGGGACT-CAGCCACTGGG-3′) and PXR-r (5′-AGAGGTACCGCTACCTGTGATACCGAACAAC-3′) and digested using NdeI and KpnI. The fragment encoding the linker (GSSGSSSG) and human SRC1 (678-710) was amplified using the oligonucleotides SRC-f (5′-CGGGGTACCGGCTCGAGTGGTAGCTCTA-GCGGGTCTTCTCA-TAGCTCATTGA-CAGAACG-3′) and SRC-r (5′-CACTTTGTCTGT-CGAGCCTGATTAATCT-CGAGCGG-3′) and digested using Kpn1 and xhoI. The two PCR products were linked by Ligation high version 2 (TOYOBO, Osaka, Japan). After purification, the ligation product was amplified by PCR using the oligonucleotides PXR-f and SRC1-r, digested using NdeI and XhoI, and cloned into pGBHPS14,15 using the same restriction enzyme cutting sites.
Protein Expression and Purification
The PXR-SRC1 protein was expressed in Escherichia coli at 25 °C as a GB1-fusion protein. GB1 and hexahistidine tags and an HRV3C protease cleavage site were fused to the N-terminus of PXR. Protein expression was induced by the addition of isopropyl-1-thio-β-galactopyranoside to a final concentration of 1 mM at 16 °C overnight. The GB1- and hexahistidine-tag–fused PXR-SRC1 was purified using an Ni-NTA resin (Qiagen, Germantown, MD), had it tags removed by HRV3C protease digestion, and was then further purified using a Superdex 75 gel filtration column (GE Healthcare). The total yield of PXR-SRC1 was 10 mg/L.
Isothermal Titration Calorimetry Measurements
Isothermal titration calorimetry (ITC) measurements were performed using an Auto-iTC200 calorimeter (GE Healthcare). All scans were obtained at 0.05 mM PXR-SRC1 and 0.5 mM ligand and were acquired in phosphate-buffered saline (PBS) buffer (pH 7.4) containing 5% glycerol and 5% DMSO. The samples were degassed under vacuum prior to the titrations, and the reference cell was filled with Milli-Q water. Titration was done 20 times using 4 µL of ligand, with injections occurring at 4 min intervals. A 40 µL syringe with 1000 rpm constant rotation was used for all experiments. The temperature of the titration cell was set at 25 °C. All ITC data were analyzed using Origin 7.0 software (OriginLab, Northampton, MA).
DSC Measurements
DSC measurements were carried out with a VP-DSC microcalorimeter (GE Healthcare) at a scanning rate of 1 °C/min from 20 °C to 80 °C. All scans were obtained at 0.03 mM PXR-SRC1 and 0.5 mM ligand and acquired in PBS buffer (pH 7.4) containing 5% glycerol and 5% DMSO. All DSC data were analyzed using Origin 7.0 software (OriginLab).
DSF Measurements
Thermal shift assays were carried out with StepOnePlus (Applied Biosystems, Foster City, CA). All scans were obtained at a protein concentration of 1 µM and ligand concentrations of 0, 0.3, 1, 3, 10, 30, 100, 200, 400, and 1000 (if needed) µM, and were acquired in PBS buffer (pH 7.4) containing 5% glycerol and 5% DMSO plus Protein Thermal shift Dye (Applied Biosystems) diluted 1000 times. Scans were measured from 25 °C to 95 °C at a scanning rate of 1.75 °C/min. All DSF data were analyzed using StepOnePlus software, thermal shift assay software (Applied Biosystems), and Microsoft Office Excel 2007 (Microsoft, Redmond, WA).
Results
Design, Expression, and Purification of Fused PXR-SRC1
The isolated PXR-LBD domain is relatively unstable and insoluble because it possesses a large, highly hydrophobic, and expandable ligand binding pocket.16–18 This property enables PXR to bind to various type of compounds. SRC1 contains an LxxLL motif that binds to the surface of PXR-LBD. This binding reduces the conformational fluctuation of PXR-LBD. SR12813 ( Figure 1a ) is a novel cholesterol-lowering drug 19 and efficacious activator of human and rabbit PXR. 20 In the absence of SRC1, SR12813 binds at multiple orientations to PXR, 16 whereas in the presence of SRC1, the binding occurs in a fixed orientation. 21 Recently, a tethered PXR-SRC1 construct was reported to be stable and to represent an excellent model system for evaluating the interaction between ligands and PXR.22,23 Hence, we constructed a fusion protein that included PXR-LBD(130-434), a linker (10 amino acids), and SRC1(678-710). The fusion protein was expressed in E. coli as an N-terminus GB1- and hexahistidine-tag fusion protein and purified from the supernatant of the lysate using an Ni-NTA resin (Qiagen). The GB1 and hexahistidine tags were then removed by HRV3C protease digestion. The fusion protein was further purified by gel filtration using Superdex 75 (GE Healthcare) to a single band in sodium dodecyl sulfate polyacrylamide gel electrophoresis analysis. The total yield of PXR-SRC1 was about 10 mg from 1 L of culture medium, an amount sufficient for the following biophysical analysis, which included ITC, DSC, and DSF.
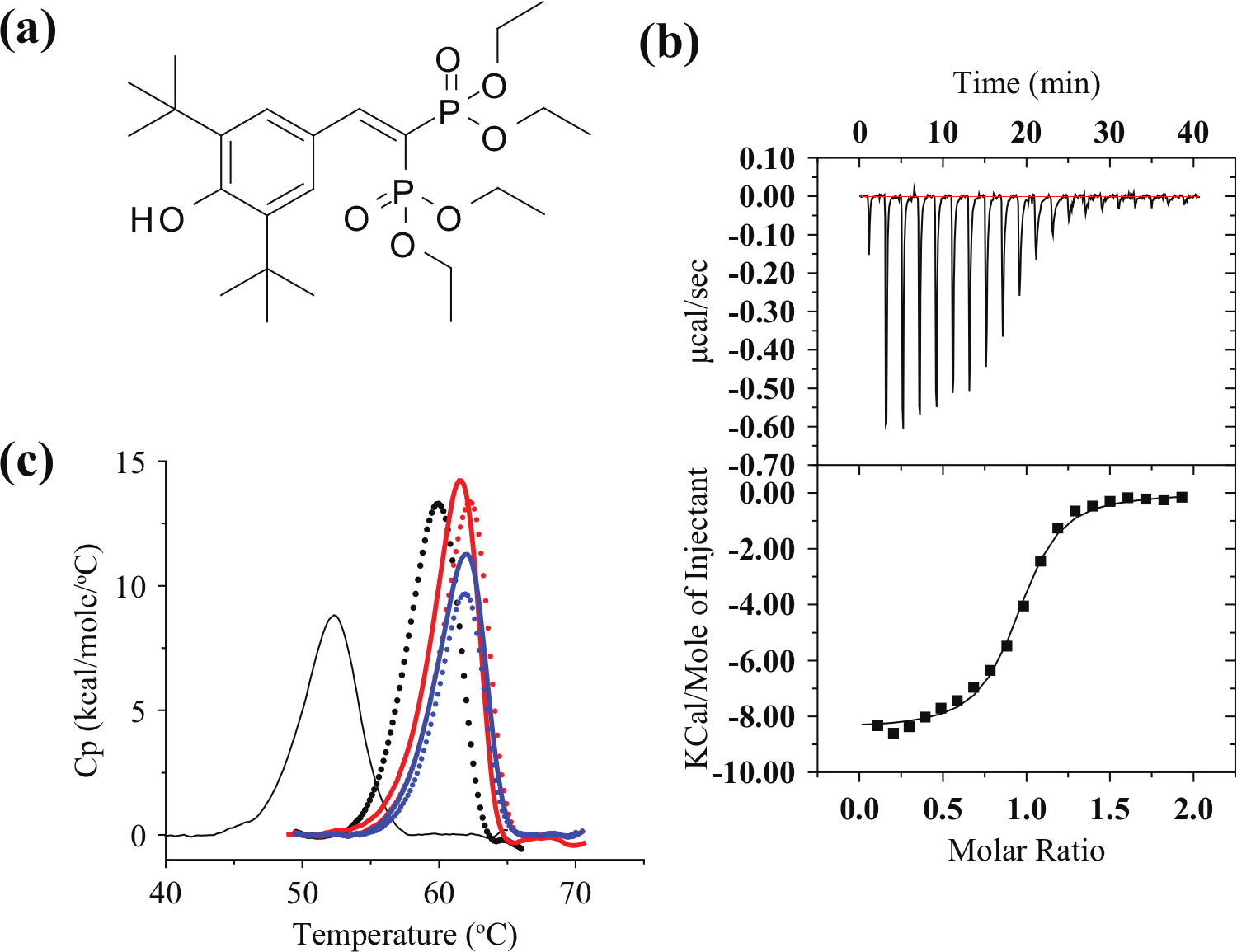
(
Binding Analysis between PXR and SR12813 Using ITC, DSC, and DSF
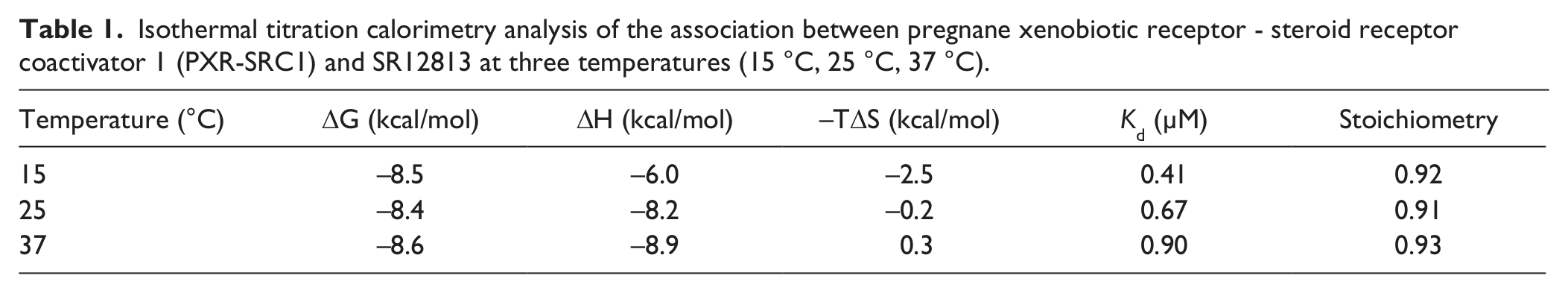
ITC measurements were performed for PXR-SRC1 and SR12813 ( Fig. 1a ) to confirm the activity of PXR-SRC1. Stoichiometry of the association of PXR-SRC1 to SR12813 was estimated to be about 0.9, which indicates that one molecule of PXR-SRC1 associates with one molecule of SR12813 ( Fig. 1b ). The dissociation constant (Kd) of PXR-SRC1 from SR12813 was estimated to be 0.41, 0.67, and 0.90 µM at 15 °C, 25 °C, and 37 °C, respectively (ΔCp = −0.13 cal/mol). Because the association of PXR-SRC1 to SR12813 was exothermal (ΔH < 0), it is reasonable that Kd increases with decreasing temperature. Furthermore, the small ΔCp suggests that a large conformational exchange did not occur when SR12813 interacted with PXR-SRC1. These results allowed us to conclude that a properly folded form of PXR-SRC1 was successfully prepared for the unfolding experiments done using DSF and DSC.
Next, we performed DSC measurements of PXR-SRC1 at different SR12813 concentrations ( Fig. 1c ). The DSC curve of PXR-SRC1 (30 µM) in the ligand-free state exhibited a single peak with a Tm of 52.6 °C that shifted to higher temperatures with increasing concentrations of SR12813.
We also performed DSF measurements of PXR-SRC1 at different SR12813 concentrations (
Fig. 2
). In the absence of SR12813, fluorescence intensity began at a peak and decreased as protein unfolding proceeded. The initial fluorescence intensity decreased as the ligand concentration increased. At higher ligand concentrations, the nature of the DSF curve changed, with the fluorescence intensity starting low and increasing with protein unfolding. These results suggest that Protein Thermal shift Dye could bind to PXR-SRC1 by directly competing for its hydrophobic pocket with SR12813. We therefore performed further DSC measurements in the presence of Protein Thermal shift Dye diluted 1000 and 300 times, finding that PXR-SRC1 at 30 µM exhibited a single peak with a Tm of 53.1 °C and 53.4 °C, respectively (
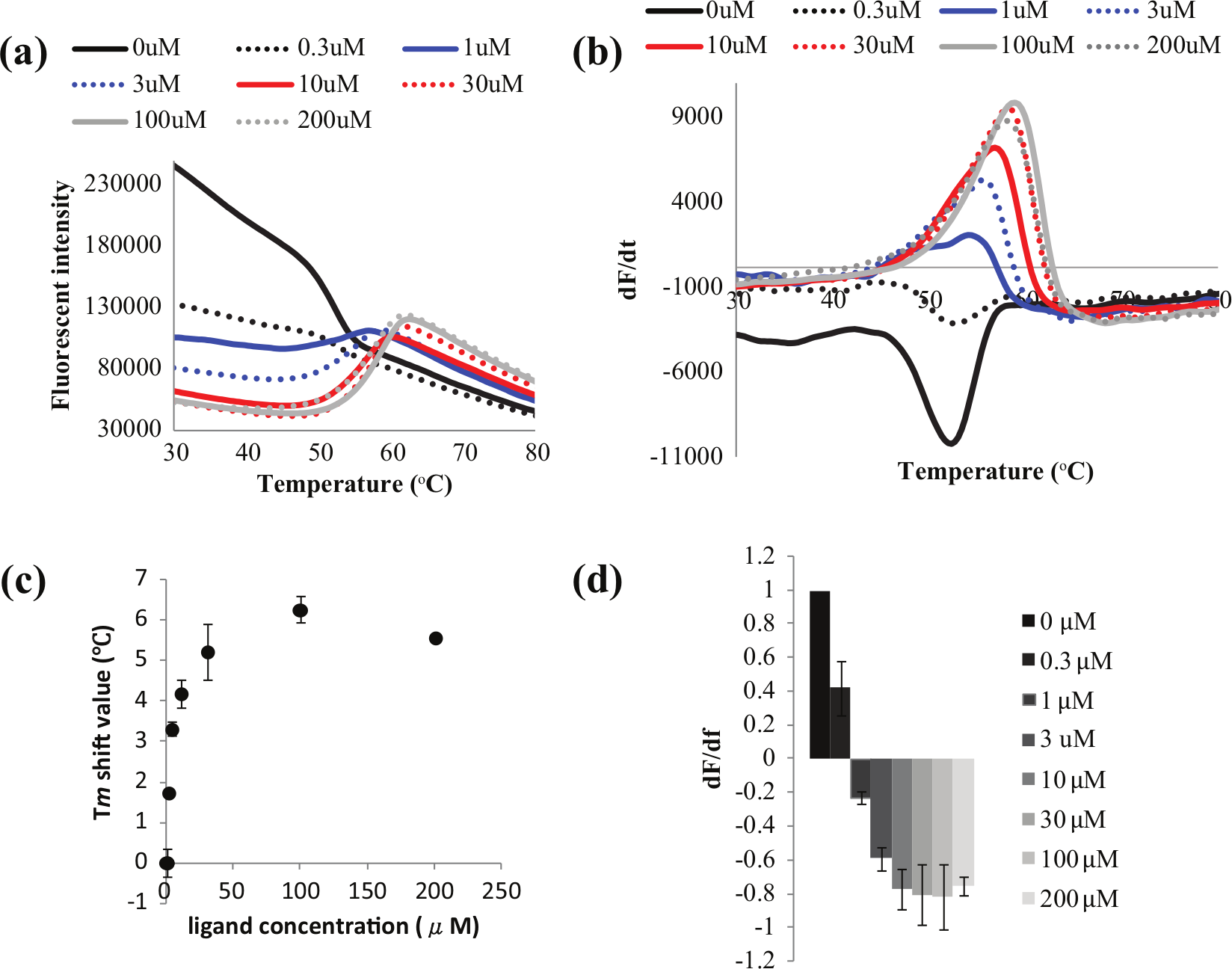
Differential scanning fluorometry at 1 µM pregnane xenobiotic receptor– steroid receptor coactivator 1 and different concentrations of SR12813: 0 µM (black straight), 0.3 µM (black dot), 1 µM (blue straight), 3 µM (blue dot), 10 µM (red straight), 30 µM (red dot), 100 µM (gray straight), and 200 µM (gray dot). Fluorescent intensities (
Evaluation of the Association of 29 Compounds to PXR-SRC1 by DSC and DSF
We also conducted DSC and DSF measurements for PXR-SRC1 in the absence and presence of 29 different ligands. Upon the addition of rifampicin, the Tm of DSC measurements shifted from 52.6 °C to 58.2 °C, indicating thermodynamic stabilization of PXR-SRC1. In the presence of phenytoin, Tm shifted to 54.3 °C, which indicates that phenytoin binds to PXR-SRC1 with slightly less affinity than SR12813 or rifampicin. No apparent Tm shift was observed in the presence of nanodol or caffeine, indicating that these compounds do not bind to PXR-SRC1 (
In DSF, we estimated Tm shift values and peak dF/dT values for different ligand concentrations ( Fig. 3 ). In contrast to the DSC measurements, we could not detect a Tm shift in DSF measurements for weak affinity compounds. On the other hand, the dF/dT values of all 29 compounds could be measured. For example, upon the addition of nifedipine (0, 0.3, 1, 3, 10, 30, 100, and 200 µM), the maximum value of dF/dT decreased (1, 1.28, 1.63, 1.15, 0.85, 0.44, 0.26, and 0.15, respectively) in proportion to nifedipine concentration, and (dF/dT)50 was estimated to be 27 µM. The (dF/dT)50 values for nicardipine, ticlopidine, phenytoin, and pregnenolone-16 alpha-carbonitrile were estimated to be 0.92, 46, 240, and 290.3 µM, respectively. Table 2 shows results for all 29 compounds.
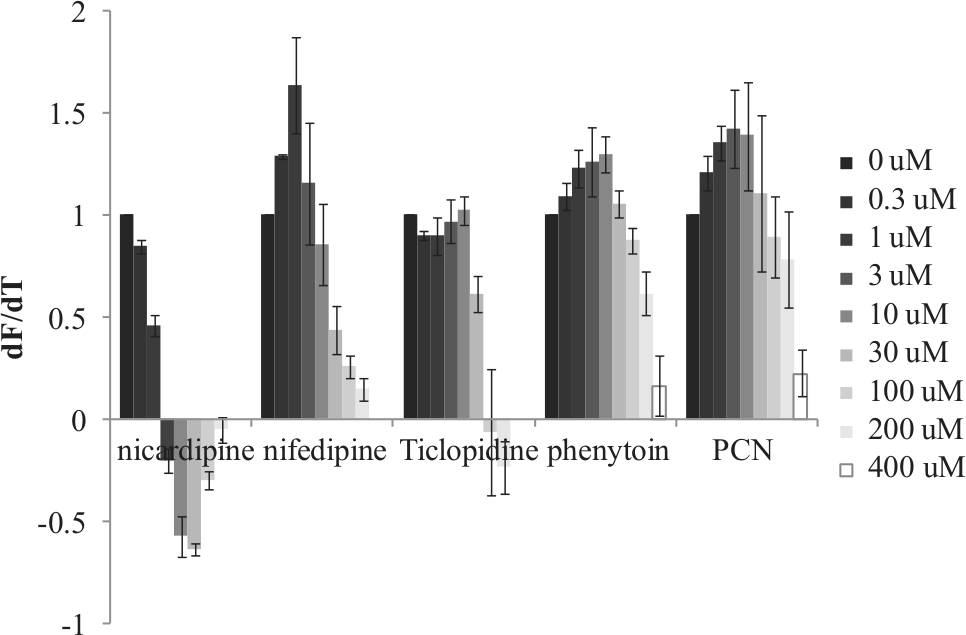
dF/dT values obtained by differential scanning fluorometry are plotted against different ligand concentrations.
Isothermal titration calorimetry analysis of the association between pregnane xenobiotic receptor - steroid receptor coactivator 1 (PXR-SRC1) and SR12813 at three temperatures (15 °C, 25 °C, 37 °C).
Summary of 29 compounds.
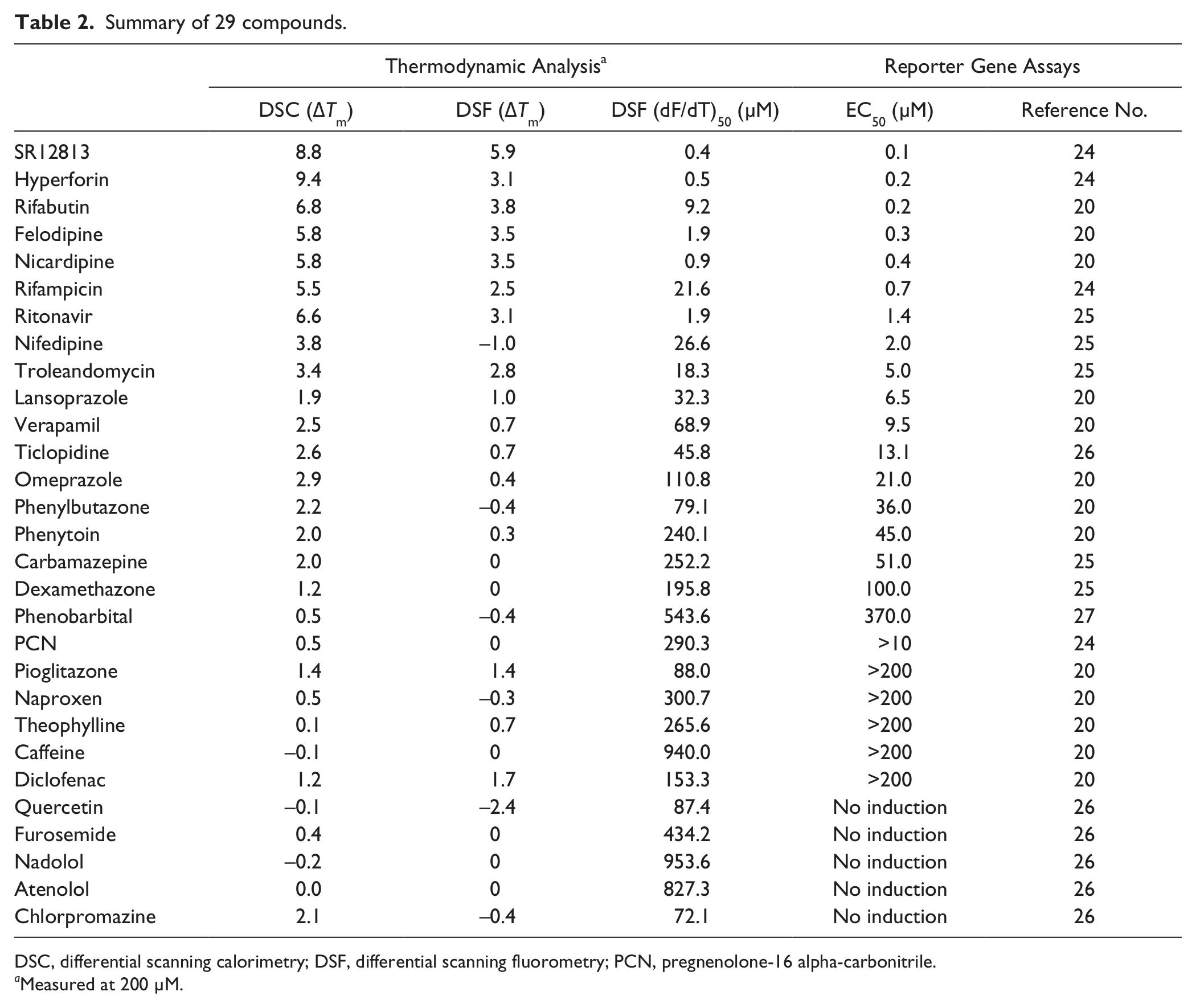
DSC, differential scanning calorimetry; DSF, differential scanning fluorometry; PCN, pregnenolone-16 alpha-carbonitrile.
Measured at 200 µM.
Discussion
Cytochrome P450 (CYP) 3A is widely recognized as a key enzyme for the metabolism of clinically used drugs. The PXR is a regulator of CYP3A transcription and is activated by most CYP3A inducers, despite their structural diversity. Reporter gene assay is widely used clinically to evaluate the PXR induction of CYP3A. However, this assay exhibits several problems regarding the cytotoxicity, membrane permeability, and solubility of the drug. Furthermore, it cannot directly observe the association of a drug to PXR, which risks false positives. Moreover, the concentration of DMSO is required to be less than 0.1%, because higher concentrations induce reporter signals, even though most drug candidates have low solubility for water, meaning organic solvents such as DMSO are necessary, often at concentrations greater than 0.1%. These disadvantages therefore indicate the need for other methods, which led us to construct a fast evaluation method for PXR-drug interactions using DSF. DSF offers an alternative method for evaluation, especially for compounds with high cytotoxicities, low membrane permeabilities, or low solubilities.
In the present study, we tested 29 compounds by DSC analysis and compared their Tm with their EC50 values from reporter gene assays in the literature. As shown in Figure 4a , a good correlation (r2 = 0.86) was observed between the ΔTm value of PXR-SRC1 and its EC50, although some compounds could not be assessed because of low induction. Although DSC is a powerful tool for measuring the thermal stability change of proteins upon ligand association, it requires significantly large amounts of protein (about 200–600 µg for a single measurement in the 10–30 µM range) and is time-consuming (about 2 h/single measurement), limiting its throughput capability for drug screening. DSF, a fluorescence-based assay, on the other hand, is easy to set up and compatible with multiwell plate devices (such as 96-well plates).13,28 Using DSF, we measured the fluorescence intensity of an extrinsic probe that bound to the hydrophobic environment on the protein surface. Normally, the fluorescence intensity increases with the thermal unfolding that occurs with exposure of the hydrophobic interior of the protein.
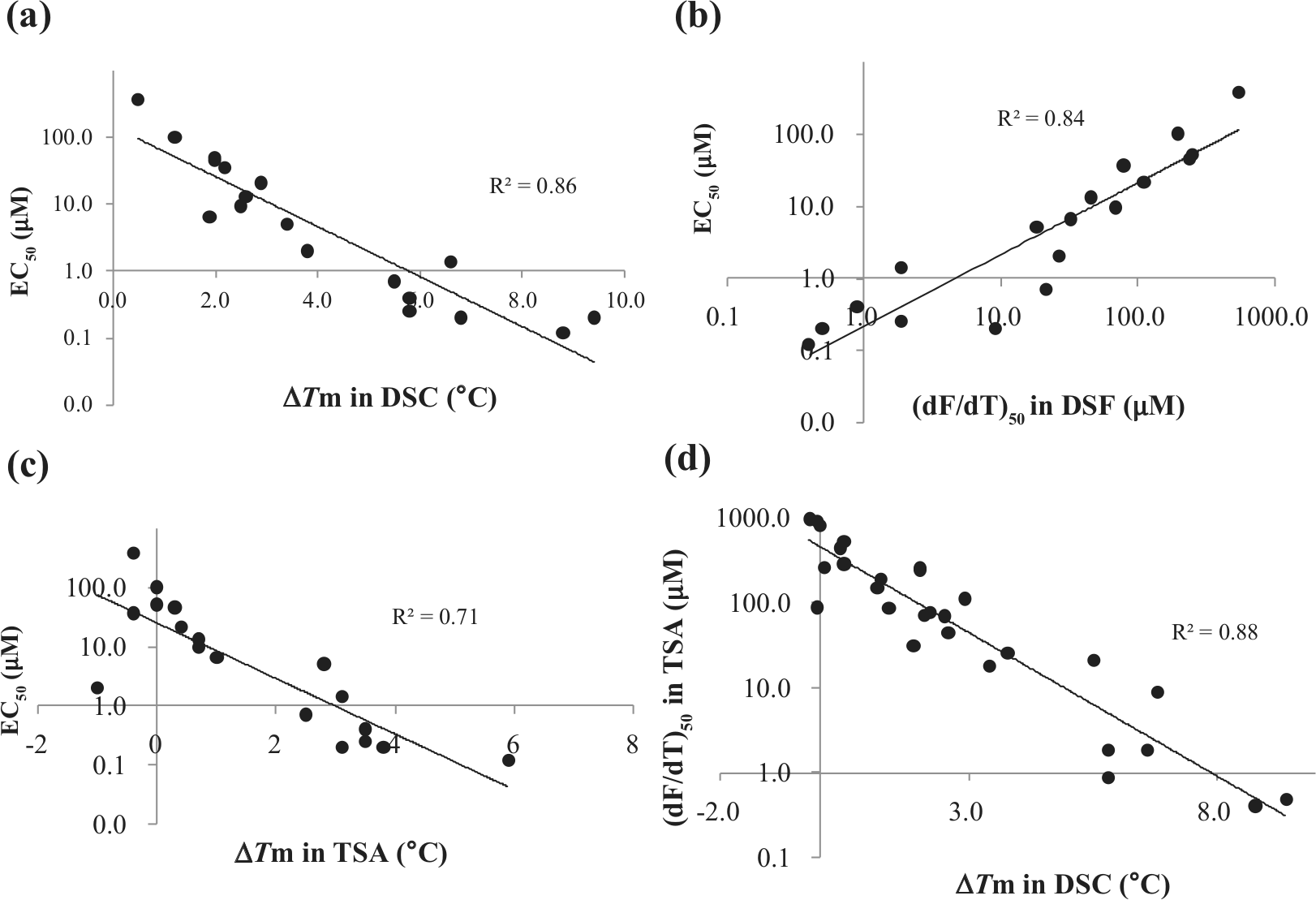
(
Ligand association raises the Tm of a protein and can therefore be used for drug discovery.29–31 In the case of PXR, a Tm shift was not observed for weak to medium affinity compounds, because Protein Thermal shift Dye competitively competed for the binding pocket. To overcome this problem, we measured (dF/dT)50, finding it had good correlation (r2 = 0.84) with EC50 ( Fig. 4b ). It should be noted that (dF/dT)50 also exhibited better correlation to EC50 than did the Tm shift value in DSF (r2 = 0.71; Fig. 4c ). Furthermore, (dF/dT)50 exhibited good correlation (r2 = 0.88) with the Tm shift value in DSC ( Fig. 4d ). In vitro PXR-based assays using temperature-dependent circular dichroism (TdCD) and the automated ligand identification system (ALIS) have been recently described. 22 DSF showed a correlation with the reporter gene assay that was comparable to correlations with TdCD and ALIS. These previous and our present results suggest that the reporter gene assay closely correlates with binding to PXR. Further, DSF exhibited markedly higher throughput than TdCD or ALIS. Thus, our DSF-based method is useful for prescreening to narrow down the collection of candidate ligands that most likely result in transcriptional activation of CYP3A4. Finally, (dF/dT)50 should be applicable to other proteins that possess a large flexible hydrophobic pocket.
Abbreviations
ALIS, automated ligand identification system; CYP, cytochrome P450; DSC, differential scanning calorimeter; DSF, differential scanning fluorometry; ITC, isothermal titration calorimeter; LBD, ligand binding domain; PXR, pregnane X receptor; SRC1, steroid receptor coactivator 1; TdCD, temperature-dependent circular dichroism.
Footnotes
Acknowledgements
We thank Mr. Koichi Shiozuka for his support of our experiments.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partly supported by the Targeted Proteins Research Program, the Funding Program for World-Leading Innovative R&D on Science and Technology, and a Grant-in-Aid for Scientific Research on Innovative Areas from the Ministry of Education, Science and Culture, Japan.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
