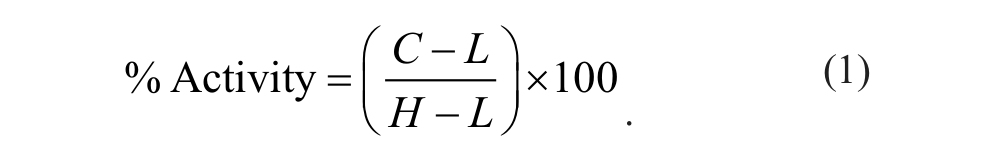Abstract
The synthesis of ribosomes is an essential process, which is aided by a variety of trans-acting factors in bacteria. Among these is a group of GTPases essential for bacterial viability and emerging as promising targets for new antibacterial agents. Herein, we describe a robust high-throughput screening process for inhibitors of one such GTPase, the Escherichia coli EngA protein. The primary screen employed an assay of phosphate production in a 384-well density. Reaction conditions were chosen to maximize sensitivity for the discovery of competitive inhibitors while maintaining a strong signal amplitude and low noise. In a pilot screen of 31,800 chemical compounds, 44 active compounds were identified. Furthermore, we describe the elimination of nonspecific inhibitors that were detergent sensitive or reactive as well as those that interfered with the high-throughput phosphate assay. Four inhibitors survived these common counterscreens for nonspecificity, but these chemicals were also inhibitors of the unrelated enzyme dihydrofolate reductase, suggesting that they too were promiscuously active. The high-throughput screen of the EngA protein described here provides a meticulous pilot study in the search for specific inhibitors of GTPases involved in ribosome biogenesis.
Introduction
In prokaryotes, the 70S ribosome is composed of a 30S small subunit and a 50S large subunit. Bacterial ribosome biogenesis involves the coordinated production, folding, and assembly of 54 proteins and 3 ribosomal RNAs (rRNAs). 1 Ribosomal subunit assembly is aided by several classes of ribosome biogenesis factors, including RNases, helicases, methyltransferases, pseudouridylases, folding chaperones, and GTPases. 1 Deletion of the genes coding for ribosome biogenesis factors often leads to promising phenotypes from a drug discovery perspective, including strains that are nonviable or exhibit slow-growth phenotypes. Nevertheless, our understanding of the pathway for assembly of ribosomal subunits in vivo is in its infancy, and there are no specific inhibitors of the process. Indeed, a chemical inhibitor of ribosome biogenesis would have exciting potential as an antibacterial lead with a novel mechanism of action, and it would also be a useful probe of this complex process. Where bacterial ribosomes are assembled on a time scale of minutes, a chemical probe would be an elegant alternative to perturbing the system by genetic means.
Four GTPases—EngA, CgtA, YihA, and Era—are among the most important ribosome biogenesis factors. Each of these factors was found to be essential for bacterial viability. 2 These factors were also demonstrated to bind to ribosomal subunits, and depletion of each protein led to the accumulation of 30S and 50S subunits as well as unprocessed rRNA.2,3 Where the precise roles of these proteins in ribosome biogenesis remain obscure and the indispensable phenotypes present an obstacle to genetic perturbation, the GTPases are a particularly exciting area of focus for new probe discovery. Likewise, the essential nature of these genes makes them ideal targets for drug discovery. Nevertheless, there are challenges to effective inhibitor screens of these factors that include slow GTPase activity and relatively high affinities for GTP. 2 In the work reported here, we present a high-throughput screen and follow-up assays toward the identification of specific inhibitors of GTPase function where the prototype is the ribosome biogenesis factor EngA.
EngA is a broadly conserved bacterial GTPase that lacks a human orthologue and has been shown to be indispensable to a variety of Gram-positive and Gram-negative organisms.4,5 EngA appears to be important for maturation of the 50S subunit. Polysome profiles of EngA-depleted cells revealed a decrease in the level of 70S ribosomes and an accumulation of 30S and 50S ribosomal subunits compared with wild-type cells.4–6 Depletion of EngA also led to accumulation of the unprocessed rRNA precursors pre-23S and pre-16S. 6 When lysates of wild-type Escherichia coli were fractionated on sucrose gradients, EngA cofractionated with the 50S subunit.4,6 In Bacillus subtilis, the 50S subunit from cells that were depleted of YphC (the EngA orthologue) migrated slower than mature 50S on a sucrose cushion. 5 These slower migrating subunits had reduced levels of the ribosomal proteins L16, L27, and L36. 5 Although EngA is important for producing mature 50S subunits, its precise role in the process is not known.
The EngA protein contains tandem guanosine triphosphate (GTP)–binding domains. We previously characterized extreme cooperativity in these domains where mutations targeting either G-domain abolished the in vitro GTPase activity of EngA and led to an inviable phenotype. 4 Thus, an inhibitor of either G-domain will inevitably inactivate the entire protein. Here, we describe the optimization of a 384-well density high-throughput assay for GTP hydrolysis, designed for favorable sensitivity, signal, and noise characteristics. A pilot screen of 31,800 compounds validated this primary screening assay. Also presented is a thorough collection of secondary assays aimed at the elimination of false positives and nonspecific inhibitors. This screen of EngA may represent a prototypic approach for discovery of specific inhibitors of GTPases involved in ribosome biogenesis.
Materials and Methods
Materials
We screened a collection of 31,800 diverse chemical compounds derived from the Custom Library of 16,000 compounds (Maybridge, Cornwall, UK), the DIVERSet of 9989 compounds (ChemBridge, San Diego, CA), the Prestwick Chemical Library of 1120 compounds (Prestwick, Washington, DC), the Natural Products Library of 361 compounds (Enzo Life Sciences, Farmindale, NY), the LOPAC1280 (International Version) library of 885 compounds (Sigma-Aldrich, St Louis, MO), the Spectrum Collection of 1214 compounds (MicroSource, Gaylordsville, CT), a synthetic library of 1200 compounds (in house), and a targeted Kinase Library of 1000 compounds (Chemical Diversity Labs, San Diego, CA). GTP sodium salt, TrisHCl, MgCl2, KCl, malachite green oxalate, and ammonium molybdate tetrahydrate were from obtained from Sigma-Aldrich (Oakville, Ontario, Canada). High-purity OmniSolv DMSO, 98% sulfuric acid, and KH2PO4 were from EMD Biosciences (Gibbstown, NJ).
Phosphate-Based GTPase Assay
All enzymatic assays were carried out with recombinant untagged EngA that was purified by Q-Sepharose Fast Flow anion exchange chromatography (Amersham Biosciences, Baie D’Urfe, Quebec, Canada). 4 One micromolar EngA was incubated with 300 µM GTP in assay buffer (100 mM TrisHCl, 20 mM MgCl2 and 400 mM KCl, pH 7.5) containing 5% vol/vol DMSO in a final reaction volume of 50 µL in 384-well microplates. Reactions were mixed well and incubated for 25 min at ambient temperature before addition of 20 µL malachite reagent containing 1.65 M sulfuric acid, 0.99% wt/vol ammonium molybdate tetrahydrate, 0.066% wt/vol malachite green oxalate, and 0.1% Tween-20 (detection method adapted from Baykov et al. 7 ). This quenched assay was incubated for 25 min at ambient temperature for color development. Optical density at 600 nm was read in an EnVision multilabel plate reader (PerkinElmer Life Sciences, Waltham, MA). The OD600 was related to the amount of phosphate produced using the equation of the line of a phosphate standard curve. The standard curve contained a 2-fold serial dilution of 5 to 80 µM KH2PO4.
Primary Screen
EngA was screened in high throughput using the GTPase assay described above. We tested 31,800 compounds in duplicate in 384-well plates. Reactions were set up on a SAMI EX Assay Workstation (Beckman Coulter, Fullerton, CA). Compounds dissolved in DMSO were added to empty wells to final concentrations of 20 µM compound and 2% vol/vol DMSO. An equivalent volume of DMSO was added to the controls. Next, assay buffer containing GTP was added to each well. Reactions were initiated with 10 µL of an enzyme stock that was kept on an integrated static Peltier cooling block. Each 384-well plate contained 32 high-activity controls (uninhibited reaction) and 32 low-activity controls (lacking enzyme) in the first two and last two columns with high and low controls alternating from top to bottom.
The quality of the screen was evaluated by calculating the Z′ factor of the high- and low-activity controls as described by Zhang et al. 8 The percent activity of each well was calculated from the absorbance data using equation (1), where C represents the compound value and H and L represent the averages of the high and low controls, respectively, on the same plate as the compound.
A hit was defined as a compound with a residual activity of less than three standard deviations below the average of the compound data in either replicate.
Secondary Screen
The 44 compounds that were selected from the primary screen were retested at 50 µM using the GTPase assay described above in the presence or absence of 0.01% Triton X-100 or 2 mM dithiothreitol (DTT). Susceptibility to Triton X-100 or DTT was defined as restoration of the GTPase activity of EngA to >70% compared with uninhibited controls. The compounds that were not sensitive to Triton X-100 or DTT were checked for interference with the malachite green assay by measuring the signal produced by 30 µM phosphate (KH2PO4) in the presence of 50 µM compound. Compounds that reduced the signal of the phosphate standard by >30% were eliminated.
High-Performance Liquid Chromatography Analysis of GTPase Activity
The effect of the four candidate inhibitors on the enzymatic conversion of α–32P-GTP to α–32P-GDP was monitored by paired ion chromatography (PIC) on a Waters 600 HPLC (Milford, MA). Reactions were quenched with two volumes of 8 M urea and loaded onto an Inertsil ODS-3 column (4 × 150 mm, 5 µm) (GL Sciences, Torrance, CA). Resolution and elution of GTP and guanosine diphosphate (GDP) were achieved with a 2-min linear gradient from Pic A (15 mM dibasic potassium phosphate and 10 mM tetrabutylammonium hydrogen sulfate, pH 7.0) to Pic B (Pic A containing 30% vol/vol acetonitrile), followed by 5 min of Pic B. Analytes were visualized by in-line scintillation counting and quantified by integration of GTP and GDP peaks using the Waters Millennium software.
Dose-Response Assays
Various concentrations of the four active molecules were tested in the GTPase assay containing 1 µM EngA and either 100 µM or 1 mM GTP. To obtain dose-response curves, the data were fit to a four-parameter logistic model using SigmaPlot (Systat Software, San Jose, CA).
The Cheng-Prusoff equation 9 was used to calculate the expected change in IC50 at low and high substrate concentrations. The Ki of a competitive inhibitor is constant and does not depend on the substrate concentration. Since the KM of EngA for GTP is 150 µM, the IC50 of a competitive inhibitor should be 4.6-fold lower at 100 µM GTP than at 1 mM GTP.
Dihydrofolate Reductase Assay
The counterscreen assay was carried out with the dihydrofolate reductase (DHFR) assay kit (Sigma-Aldrich, St. Louis, MO) in a 96-well microplate. Each reaction contained 0.001 U DHFR, 150 µM NADPH, and 30 µM dihydrofolate in 50 mM Tris (pH 7.5) in a final reaction volume of 200 µL. Compound (or an equivalent volume of DMSO) was added to a final concentration of 100 µM. The absorbance at 340 nm was monitored continuously for 5 min in a SpectraMax spectrophotometer (Molecular Devices, Sunnyvale, CA). The rate of each reaction was expressed as a percentage of the rate of the uninhibited reaction.
Results and Discussion
Assay Optimization
Although whole-cell active inhibitors of protein translation are commonplace, a specific inhibitor of ribosome biogenesis has never been reported. We present a pilot chemical screening study of the bacterial ribosome biogenesis factor EngA from E. coli. To measure the rate of production of phosphate by EngA, variations of the ammonium molybdate/malachite green assay and commercial kits were compared. The protocol published by Baykov et al. 7 produced the greatest amplitude between the signals from the high- and low-activity controls, which afforded a more substantial screening window. Linear detection of phosphate was observed from 5 to 80 µM ( Fig. 1A ).
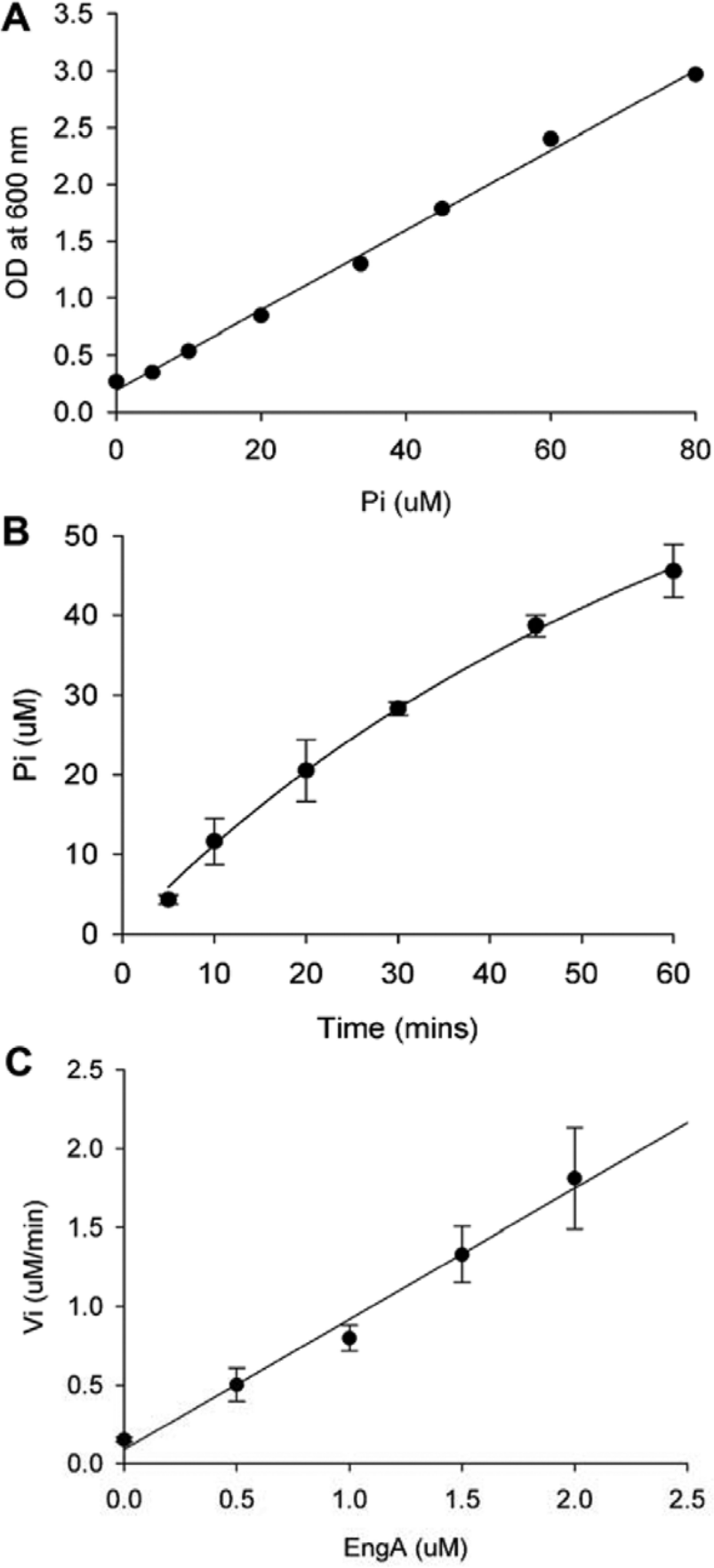
Defining conditions for a GTPase assay of EngA that is sensitive to inhibition. (
Previous kinetic characterization of EngA revealed a Michaelis-Menten constant (KM) of 150 µM. 4 Reactions are more sensitive to competitive inhibition at lower substrate concentrations where the reaction rate varies linearly with substrate concentration. 10 The screen would ideally be carried out at a substrate concentration that is less than or equal to KM; however, due to the relatively high affinity of EngA for GTP, a concentration of 2 × KM (300 µM) of GTP was used. This substrate concentration afforded a reasonable compromise between sensitivity to competitive inhibitors and sufficiently high signal amplitude in the enzyme assay.
We chose conditions where the reaction was linear with both time and enzyme concentration to ensure that the rate of the reaction was first order and, thus, sensitive to inhibition. In a reaction progress curve containing 300 µM GTP, the amount of phosphate produced was fairly linear up to 25 min, which corresponded to 8% depletion of substrate ( Fig. 1B ). Due to the relatively slow catalytic rate of EngA (kcat of 70 h−1), 4 1 µM enzyme was necessary to achieve this amount of substrate conversion. Comfortable that compounds screened at 20 µM would be far in excess of this enzyme concentration, we verified that EngA at 1 µM was within the linear range of enzyme concentrations for product formation ( Fig. 1C ). In all, these assay development efforts afforded primary screening conditions that were sensitive to inhibition while remaining practical for high-throughput chemical screening.
Primary Screen of EngA
EngA was screened against 31,800 diverse chemical compounds that were a compilation of synthetic molecules, known natural products, and a kinase-directed collection that was chosen because EngA is a nucleotide-binding protein. The high-activity controls contained uninhibited GTPase reactions, whereas the low-activity controls lacked enzyme ( Fig. 2A ). The Z′ factor, which accounts for the separation between the high and low controls and the noise of the assay, was 0.68. A Z′ factor of 0.5 to 1.0 describes a robust primary screening assay with good signal amplitude and relatively low noise. 8
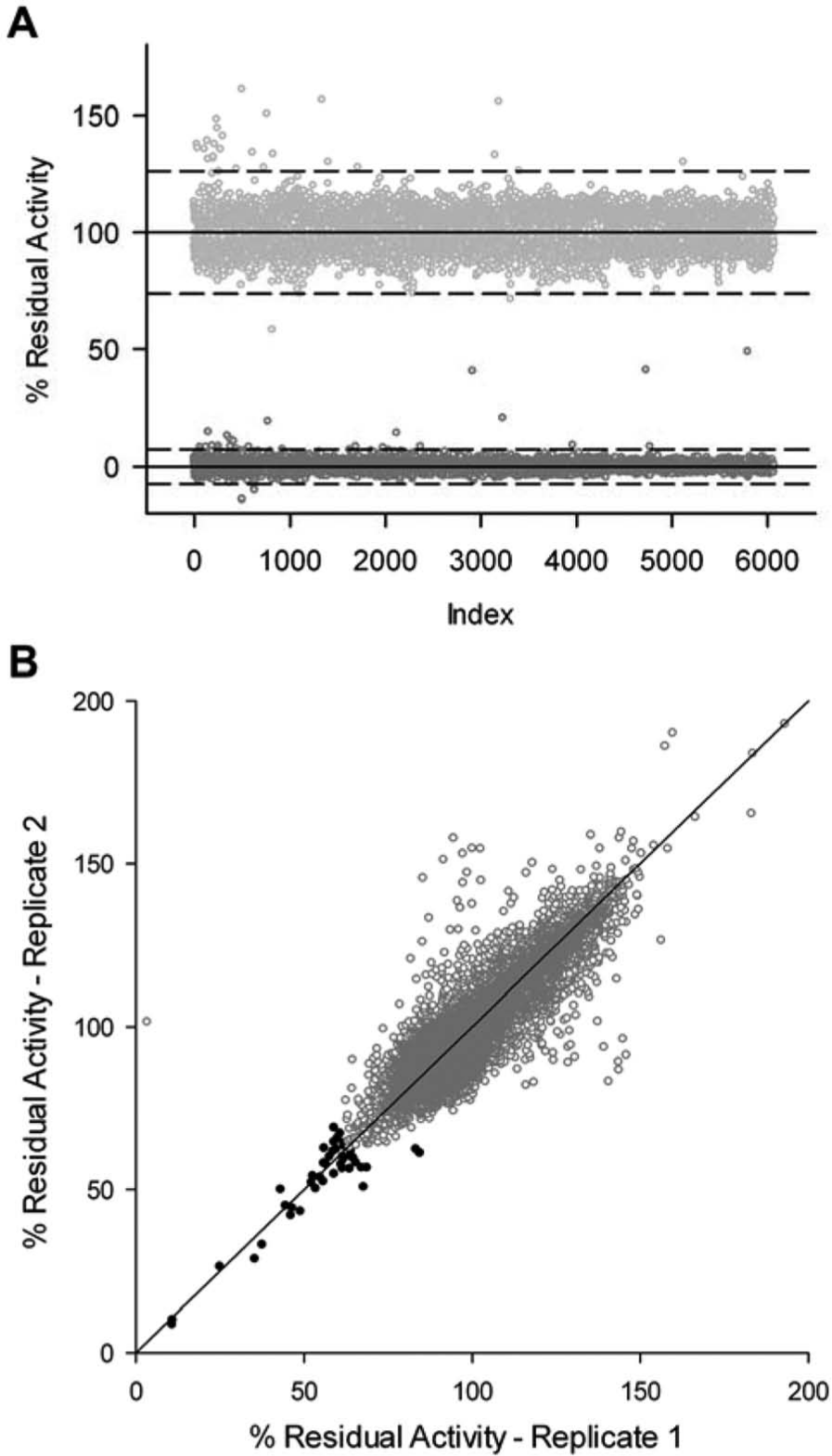
The signal-to-noise ratio and reproducibility of a pilot screen of EngA. (
Each compound was screened in duplicate at 20 µM. A replicate plot of the primary screen showed strong reproducibility between the two replicates, yielding an R2 value of 0.77 ( Fig. 2B ). A hit was defined as a molecule with residual activity that was more than three standard deviations below the mean of all compounds, which corresponded to 63% residual activity. There were 44 hits, representing an overall hit rate of 0.15%. For compounds that were sourced from Chembridge, BioMol, Prestwick, or the Kinase Library, each collection had a hit rate around 0.2%, whereas compounds from Maybridge had a lower rate (0.03%), and those sourced from Microsource had a comparatively higher rate (1%). The Microsource collection comprises previously approved drugs, molecules with known biological activity, and natural products.
Thus, this pilot screen of 31,800 compounds had a robust Z′ factor and showed good correspondence between the two replicates of the screening data. Similar methodology might be extended to other bacterial ribosome biogenesis GTPases to find specific inhibitors of ribosome biogenesis. There are no reports of ribosome stimulation of the GTPase activity of EngA, but for ribosome-stimulated GTPases, this biochemical trait may be exploited in primary or secondary screens to identify inhibitors of ribosome-stimulated GTPase activity.
Secondary Screen of EngA to Eliminate False Positives
The structures and retested activities of the 44 hits from the primary screen and the reasons for elimination are provided (see
Chemical Structures and Dose-Response Curves of the Four Inhibitors of EngA
Of the four remaining compounds, MAC-0174809 and MAC-0080023 are synthetic molecules that are structural analogues, whereas MAC-0182099 and MAC-0182344 are natural products with more complex structures ( Fig. 3 ). The minimum inhibitory concentrations yielding 50% inhibition (IC50) were 13.3 µM for MAC-0174809, 12.5 µM for MAC-0080023, 14.3 µM for MAC-0182099, and 4.6 µM for MAC-0182344 ( Fig. 3 ). All four compounds produced steep slopes in the dose-response curves with Hill values close to 4. Reversible and specific inhibitors that bind stoichiometrically to enzymes reduce activity from 90% to 10% over an 81-fold change in inhibitor concentration. This yields a slope in the dose-response curve that is assigned a Hill slope value of 1. In some cases, high Hill slopes may indicate that the inhibitor is acting by a nonspecific mechanism such as colloidal aggregation, enzyme denaturation, or micelle formation.12,14
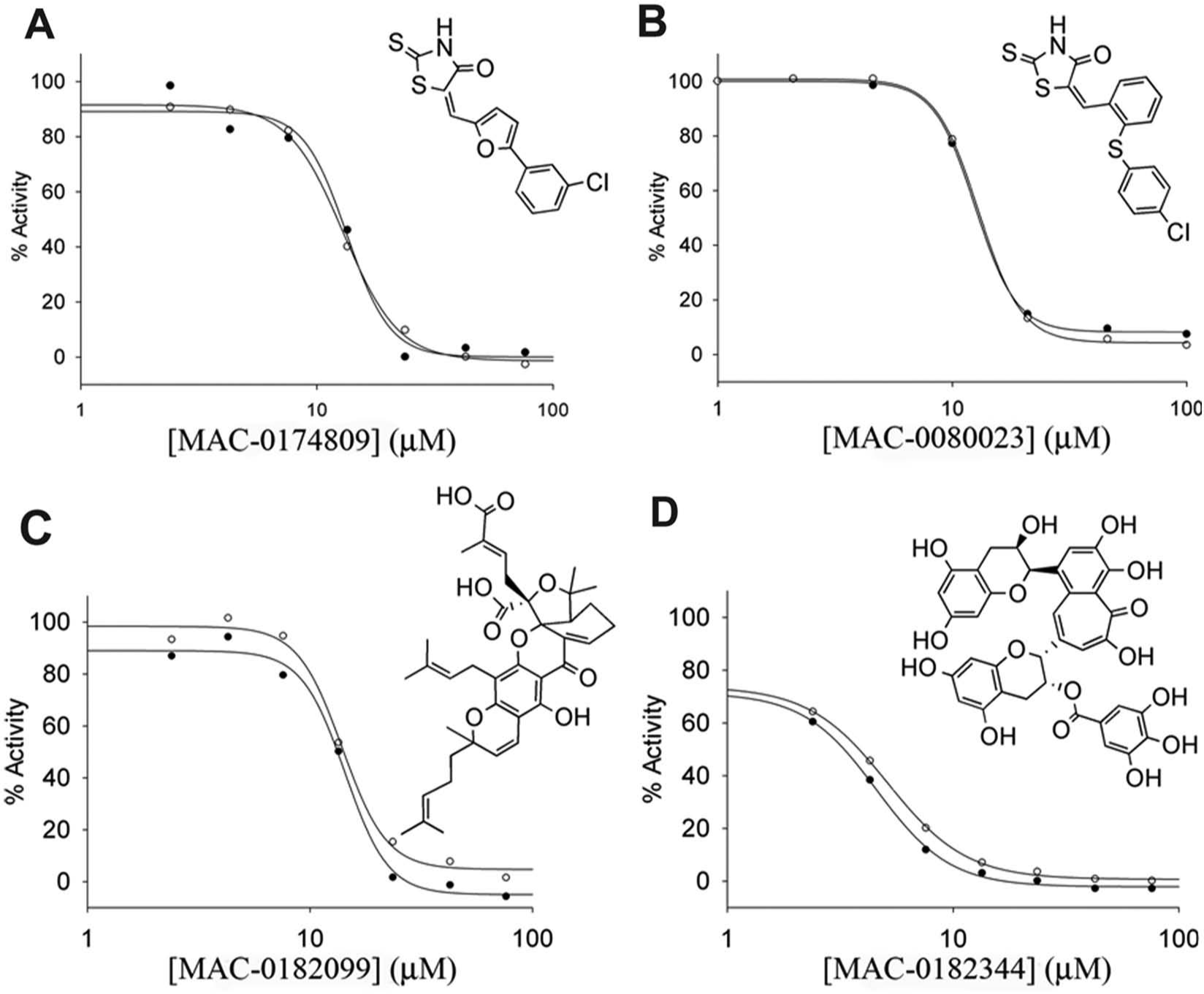
Chemical structures and dose-response curves of the four inhibitors of EngA. The effect of various concentrations of inhibitor on the GTPase activity of 1 µM EngA was tested at 100 µM guanosine triphosphate (GTP; open circles) and 1 mM GTP (closed circles). GDP and GTP were resolved by paired ion chromatography high-performance liquid chromatography (HPLC). SigmaPlot software (Systat Software, San Jose, CA) was used to fit the data to a four-parameter logistic model. (
To test if each inhibitor was competitive with GTP, we obtained the dose response of the inhibitor at two concentrations of substrate. For competitive inhibitors, an increase in substrate concentration from 100 µM GTP to 1 mM GTP should cause a 4.6-fold increase in the IC50. For these four compounds, no increase in the IC50 was observed at the higher substrate concentration, indicating that they are not competitive with GTP ( Fig. 3 ).
Specificity of Inhibition
Although high Hill values and lack of competition with substrate are common traits of promiscuous inhibitors, specific inhibitors may also display these traits. 14 We obtained more definitive evidence of the lack of specificity of these four compounds using a counterscreen against DHFR. DHFR reduces dihydrofolate to tetrahydrofolate using the cofactor nicotinamide adenine dinucleotide phosphate (NADPH), where the consumption of NADPH can be monitored by the decrease in absorbance at 340 nm. At 100 µM, all of the compounds reduced the activity of DHFR to 20% to 40% of the activity of uninhibited controls (data not shown). The counterscreen against DHFR represented a low stringency test of specificity because this enzyme acts on an unrelated pathway. Since all four molecules inhibited DHFR, we did not perform higher stringency tests of specificity against GTPases.
The four most promising inhibitors were not sensitive to detergent or DTT, but due to the lack of competition with substrate, steep IC50 curves, and coincident inhibition of DHFR, we have concluded that these molecules are not specific inhibitors of EngA. MAC-0174809 and MAC-0080023 both satisfy Lipinski’s rule of 5, but MAC-0182099 has a high molecular weight and a high calculated partition coefficient, whereas MAC-0182344 has a high molecular weight and a high number of hydrogen bond donors and acceptors. 15 In future screening campaigns, it would be useful to include 0.01% Triton X-100 and 2 mM DTT to reduce the selection of promiscuous actives. We also recommend prioritizing the test for assay interference in secondary screening.
The GTPase activity of EngA was shown to be an amenable target for a chemical screen using a convenient low-cost assay that was practical for high-throughput experiments. The assay was also optimized for low noise and good separation between the high-activity controls and low-activity controls to maximize the screening window. The robust Z′ and high replication suggest that this high-throughput assay can be used with larger compound libraries to continue the search for an inhibitor of ribosome biogenesis.
Footnotes
Acknowledgements
We thank Jenny Wang from the McMaster High Throughput Screening Laboratory for providing technical assistance with the screen.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: E.D.B. was supported for this work by an operating grant from the Canadian Institutes of Health Research (grant number MOP-64292).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.