Abstract
β-Site amyloid precursor protein cleaving enzyme–1 (BACE-1) is a transmembrane aspartic protease that mediates the initial cleavage of the amyloid precursor protein (APP), leading to the generation of amyloid-β (Aβ) peptides that are thought to be causative of Alzheimer’s disease (AD). Consequently, inhibition of BACE-1 is an attractive therapeutic approach for the treatment of AD. In general, in vitro biochemical assays to monitor BACE-1 activity have used the extracellular domain of the protein that contains the catalytic active site. This form of BACE-1 is catalytically active at acidic pH and cleaves APP-based peptide substrates at the β-site. However, this form of BACE-1 does not mimic the natural physiology of BACE-1 and shows minimal activity at pH 6.0, which is more representative of the pH within the intracellular compartments where BACE-1 resides. Moreover, high-throughput screens with recombinant BACE-1 at pH 4.5 have failed to identify tractable leads for drug discovery, and hence, BACE-1 inhibitor development has adopted a rational drug design approach. Here we describe the development and validation of a novel membrane assay comprising full-length BACE-1 with measurable activity at pH 6.0, which could be used for the identification of novel inhibitors of BACE-1.
Introduction
Alzheimer’s disease (AD) is a devastating neurodegenerative disorder characterized pathologically by the presence of extracellular senile plaques and intracellular neurofibrillary tangles in the brain. 1 The core constituents of senile plaques are insoluble aggregates of the 4-kDa amyloid-β (Aβ) peptides, Aβ40 and Aβ42, that are generated following sequential proteolytic processing of the amyloid precursor protein (APP) by β-site APP cleaving enzyme 1 (BACE-1) and γ-secretase. 1
BACE-1 is a type I transmembrane aspartic protease that comprises a large extracellular domain containing the catalytic active site, a single transmembrane domain, and a short cytoplasmic tail.2–5 In cells, BACE-1 co-localizes with its substrate APP within the Golgi and endocytic compartments,2,3,6 where it cleaves APP at the amino-terminus of the Aβ domain. Overexpression of BACE-1 in cells augments Aβ production, whereas knockdown of BACE-1 expression precludes Aβ generation.3–5 Moreover, genetic ablation of BACE-1 abolishes Aβ production 7 and can reverse the cognitive deficits observed in APP transgenic mice. 8 In addition, BACE-1 knockout mice are viable and fertile, suggesting inhibition of BACE-1 is unlikely to be associated with mechanism-based toxicities. 7 Due to its pivotal role in Aβ generation, BACE-1 is an attractive therapeutic target for AD, and inhibition of BACE-1 has been the focus of extensive drug discovery efforts for many years. 9
The development of in vitro biochemical assays to monitor BACE-1 activity is pivotal in drug discovery for the screening and identification of BACE-1 inhibitors. Since the catalytic active site of BACE-1 is localized within the extracellular region of the protein, soluble forms of BACE-1 comprising the ectodomain have been generated and used to develop sensitive in vitro assays to monitor enzyme activity.10–14 Using a range of assay formats, including fluorescence resonance energy transfer (FRET)11–13 and enzyme complementation, 14 the purified soluble ectodomain of BACE-1 was shown to be catalytically active, cleaving APP-based substrates at the expected site with optimal activity at pH 4.5. Very little or no activity was evident at higher pH.3,11
In cells, BACE-1 has been shown to reside and co-localize with APP within the Golgi and endocytic compartments.3,6 Intriguingly, the pH of these cellular compartments is higher (~ pH 6.0)15,16 than the pH at which recombinant BACE-1 is optimally active in vitro. This disparity in pH for recombinant BACE-1 activity versus intracellular BACE-1 activity is noteworthy. In addition, it has been reported that full-length BACE-1 but not the soluble ectodomain of BACE-1 can form homodimers and that this requires the attachment of BACE-1 to the membrane.17,18 Interestingly, the BACE-1 homodimers appeared to be catalytically more active than monomeric soluble BACE-1. 17 The potential influence of the transmembrane domain and cytoplasmic tail in the regulation of BACE-1 activity is also evident from reports that several proteins that interact with these regions of BACE-1 can regulate its activity.19–21 Hence, despite the obvious advantages of assays using the soluble ectodomain of BACE-1, these assays do not mimic the natural physiology of BACE-1.
In general, high-throughput screens with recombinant BACE-1 failed to identify tractable hits for BACE-1 inhibitor development. Consequently, BACE-1 inhibitors were initially developed using a rational drug design/transition-state mimetic approach. 9 Although this approach was successful in generating potent BACE-1 inhibitors, these compounds exhibited poor pharmacokinetic properties, including low brain penetration and cytochrome P450 liabilities, which hindered their subsequent development. 22 More recently, fragment-based drug discovery has been applied for the identification of BACE-1 inhibitors.23,24 Because of the limited success with BACE-1 inhibitor development, the availability of alternative screening assays for BACE-1 could be advantageous.
The aim of this study was to develop and characterize an in vitro membrane assay for full-length BACE-1 at pH 6.0 that would permit the screening of BACE-1 inhibitors under more physiologically relevant conditions.
Materials and Methods
Recombinant BACE-1 Activity Assays
Recombinant BACE-1 was obtained from Proteros (Martinsried, Germany). The BACE-1 FRET assay (β-secretase) kit and the LanthaScreen BACE-1 assay kit were from Invitrogen (Paisley, UK). The HitHunter β-secretase kit was from DiscoveRx (Fremont, CA). All assays were conducted at pH 4.5 as described by the manufacturer. Assays at pH 5.2 and 6.0 were conducted using sodium acetate buffer pH 5.2 or 50 mM PIPES pH 6.0 in place of the assay buffer provided with the kits.
Generation of HEK Cell Line Overexpressing BACE-1
Human embryonic kidney (HEK) cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Invitrogen, Carlsbad, CA) containing 10% fetal bovine serum (FBS) (Lonza, Basel, Switzerland). Cells were transfected with human BACE-1/pcDNA3.1 using Fugene HD transfection reagent (Roche Applied Science, Indianapolis, IN), and stable clones were selected using medium containing 200 µg/mL hygromycin (Invitrogen).
Gel Electrophoresis and Western Blot Analysis
HEK cells stably overexpressing BACE-1 (HEK–BACE-1) were lysed in 50 mM Tris HCl (pH 7.4), 150 mM NaCl, 1% Triton X-100, 0.2% sodium dodecyl sulfate (SDS), and 0.5% Nonidet P-40 containing a Complete protease inhibitor cocktail (Roche Applied Science). Cell lysates were resolved on a 4% to 12% Bis-Tris NuPAGE gel (Invitrogen) under reducing conditions and transferred to a nitrocellulose membrane for Western blot analysis for BACE-1. Membranes were incubated with Odyssey blocking buffer (diluted 1:2 in phosphate-buffered saline [PBS]) (LI-COR Biosciences, Lincoln, NE) and then probed overnight at 4 °C with anti-BACE-1 antibody (1:500 dilution in blocking buffer) (Abcam, Cambridge, UK). Following washes with PBS and 0.2% Tween-20, membranes were incubated with a goat anti-rabbit IRDye 800CW secondary antibody (1:10 000 dilution in PBS) (LI-COR Biosciences) for 1 h at room temperature. After several washes, the membrane was visualized using the Odyssey infrared imaging system (LI-COR Biosciences).
Membrane Preparation
Membranes from HEK–BACE-1 cells were prepared as follows. Cells from 10 × T225 flasks or cell factories (6300 cm2, Nunc or Corning) were harvested in PBS containing 1 mM EDTA and centrifuged at 1500 rpm for 10 min. The cell pellet was resuspended in 40 mL 20 mM HEPES (pH 7.3) and 10 mM KCl and incubated for 8 min at 4 °C with gentle mixing prior to centrifugation at 2300 rpm for 10 min at 4 °C. The cell pellet was resuspended in 40 mL 20 mM HEPES (pH 7.3) and 90 mM KCl and homogenized using a Dounce homogenizer with a Teflon pestle. The cell extract was centrifuged at 2300 rpm for 10 min to remove cell debris and nuclei. The supernatant was collected and subject to ultracentrifugation at 100 000 g for 30 min at 4 °C to pellet the membranes. The membranes were resuspended in PBS containing 5% glycerol and stored at −80 °C for subsequent use. Prior to solubilization, membranes were collected by ultracentrifugation and resuspended in 2 mL 50 mM sodium acetate (pH 4.5) and 150 mM NaCl. The membranes were either incubated at 4 °C for 30 min or alternatively homogenized using the Dounce homogenizer prior to ultracentrifugation. The membranes were resuspended in 2 mL 20 mM 2-(N-morpholino)ethanesulfonic acid (MES) (pH 6.0) containing a 2.5× Complete protease inhibitor cocktail that does not inhibit BACE-1 activity (data not shown). Triton X-100 (Sigma-Aldrich, Saint Louis, MO) was added to give a 1% final concentration and the membranes triturated using a 25-gauge needle. Alternatively, membranes were resuspended in 50 mM sodium acetate (pH 4.5) and 150 mM NaCl containing 2.5× Complete protease inhibitor and 1% Triton X-100 prior to homogenization with a Dounce homogenizer. Following centrifugation at 15 000 g for 10 min at 4 °C, the supernatant was collected and the protein concentration determined using a BCA assay (Thermo Scientific, Rockford, IL). The solubilized membranes were aliquoted and stored at −80 °C for subsequent use. For the detergent screen, membranes were solubilized in CHAPS (Interchim, Montluçon, France), octyl thioglucoside (Sigma-Aldrich), Tween-20 (Sigma-Aldrich), saponin (Sigma-Aldrich), or Triton X-100 to give the required final concentration.
BACE-1 Membrane Activity Assay and Compound Screening
During assay development and optimization, the enzymatic activity of BACE-1 in solubilized membranes was determined as follows. All assays were conducted in 96-well plates in a final volume of 200 µL/well. Solubilized membranes (equivalent to 0.4 µg/well membrane protein) were incubated in the absence or presence of 1 µM BACE-1 inhibitor, BSI IV (Calbiochem, Darmstadt, Germany), in assay buffer (either 20 mM sodium acetate [pH 4.5], 75 mM NaCl or 20 mM sodium acetate [pH 6.0], 75 mM NaCl) containing 2.5× Complete protease inhibitor cocktail and 50 µM pepstatin A (Calbiochem) for 30 min at room temperature. Then, 10 µM fluorogenic BACE-1 substrate IV [H-RE(EDANS)EVNLDAEFK-(DABCYL)R-OH] (Calbiochem) was added and the reaction incubated for 90 min at room temperature. Fluorescence was measured on a Fluoroskan Ascent FL microplate reader (Thermo Scientific). During the development of a high-throughput BACE-1 membrane assay, BACE-1 activity in the solubilized membranes was determined using a BACE-1 FRET assay (β-secretase) kit (Invitrogen) in 384-well plates as described below. The solubilized membranes were diluted 20-fold in assay buffer prior to use. A Multidrop Combi (Thermo Scientific) was used for reagent dispensing, and plates were analyzed using a Tecan Ultra (Tecan, Männedorf, Switzerland). For the high-throughput screening (HTS) and compound profiling, BACE-1 activity in the solubilized membranes was determined using a BACE-1 FRET assay (β-secretase) kit, and plates were analyzed using a 2101 EnVision plate reader (PerkinElmer, Waltham, MA). The assay was performed according to the following procedure in either a 384-well (compound profiling) or a 1536-well (HTS) format. The volumes given in parentheses are those used in the 1536-well HTS assay. Solubilized membranes, 5 µL (2 µL), were incubated with 2.25 µL (0.9 µL) compound in plates (Corning, New York, NY) for 20 min at room temperature. The FRET-substrate, 5 µL (2 µL), was added and the plate(s) centrifuged at 1200 rpm for 50 s. Fluorescence was measured at 550/595 nm prior to a 50-min incubation at room temperature. Fluorescence was measured at 550/595 nm at the end of the assay. All reagents were kept cool at 4 °C. The compounds were applied from a predilution in 20 mM HEPES (pH 7.2) and 5.44% DMSO. To minimize the effects of autofluorescent or quenching compounds during the HTS, the difference between the two fluorescence measurements was calculated as the readout. For compound characterization, only the second fluorescence measurement was used as the readout. The final reagent concentrations in the assay were 40 mM PIPES (pH 6.0) or 50 mM sodium acetate (pH 4.5) for the assays run at the respective pH, 1% DMSO, 250 nM substrate, 15 µg/mL solubilized membranes, 0.013% lipid-free BSA (Roche Applied Science), 2.5× Complete protease inhibitor cocktail, and 10 µM compound (or vehicle). The HTS was performed on a Beckman Coulter (Brea, CA) screening platform integrating a dual bridge Biomek FX liquid handler, incubators, and robotic plate transport and applying SAMI assay design and scheduling software.
Synthesis of Compounds
BACE-1 inhibitors BSI III and BSI IV were purchased from Calbiochem. BACE-1 inhibitors A to D were synthesized as described in the patent literature.25–29
HEK-APPwt Cell Assay and Aβ Analysis
For profiling of compounds in a 384-well format, HEK cells overexpressing human APP wild-type variant (HEK-APPwt) were seeded at 20 000 cells per well in 40 µL DMEM containing 10% fetal calf serum (FCS), 1 mM sodium pyruvate, 10 mM HEPES, and 1% penicillin/streptomycin. Compounds (2.4 µL in 20 mM HEPES [pH 7.2], 8.3% DMSO) were added and plates incubated at 37 °C for 20 to 26 h. Supernatant (4 µL) was transferred into 384-well AlphaPlates (PerkinElmer) and analyzed for Aβ using the AlphaLISA human amyloid β 1-42 immunoassay kit as described by the manufacturer (PerkinElmer).
For profiling of compounds in 96-well plates, HEK-APPwt cells were cultured in DMEM containing 10% FBS and treated with inhibitor or 0.5% DMSO for 16 h. Medium was collected and assayed for total Aβ using a sensitive immunoassay. Medium was incubated in the presence of a ruthenylated anti-Aβ antibody, 6E10 (Covance, Princeton, NJ), in avidin plates (Meso Scale Discovery, Gaithersburg, MD) coated with a biotinylated midregion Aβ antibody, 4G8 (Covance). Plates were washed with PBS, 0.1% Tween-20, and 1× Read Buffer T (Meso Scale Discovery) added to the plates. Plates were analyzed on a SECTOR Imager 6000 (Meso Scale Discovery). The effect of compounds on cell viability was assessed using alamarBlue (Invitrogen).
Results
Development of a Recombinant BACE-1 Enzyme Assay at Physiological pH
Before embarking on the development of a BACE-1 membrane assay, we initially investigated whether it would be feasible to develop an assay for recombinant BACE-1 enzyme at pH 6.0 to resemble the pH of the Golgi and endocytic compartments where BACE-1 resides in cells. Several commercially available assay formats were explored, including a LanthaScreen BACE-1 assay kit, a FRET-based BACE-1 (β-secretase) assay kit, and a HitHunter β-secretase assay kit in the absence or presence of a transition-state mimetic BACE-1 inhibitor (BSI) to determine the assay window. Recombinant BACE-1 was catalytically active at pH 4.5 and, to a slightly lesser extent, at pH 5.2 in all three assays, with signal windows of approximately 5 and 3, respectively, in the FRET-based BACE-1 assay ( Fig. 1 ). However, no specific BACE-1 activity was evident at pH 6.0 ( Fig. 1 ). The signal obtained at pH 6.0 was significantly lower than that observed at pH 4.5 (8.6-fold, p < 0.0001) and pH 5.2 (7-fold, p < 0.001). In addition, there was no difference in signal at pH 6.0 in the absence or presence of the BSI, suggesting the activity observed was nonspecific ( Fig. 1 ). A small assay window was evident in the HitHunter assay format at pH 6.0, but this required a lengthy reaction time of 16 h (data not shown). Attempts were made to increase BACE-1 activity at pH 6.0 by applying excipients to the enzymatic reaction mix. Sixty different additives were tested, among which were salts (e.g., MgCl2, NaCl), organics (dithiothreitol [DTT], glycerol, urea, guanidine), buffers (e.g., MES, CAPS, MOPS), sugars (e.g., glucose, saccharose, ficoll), and detergents (e.g., CHAPS, Tween-20, Triton X-100, Zwittergent, dodecylmaltoside). All excipients were tested at two concentrations with two different recombinant BACE-1 protein sources. However, no reasonable assay window could be established by applying any of the additives (data not shown). On the basis of these findings, we abandoned this approach and embarked on the development of a BACE-1 membrane assay.
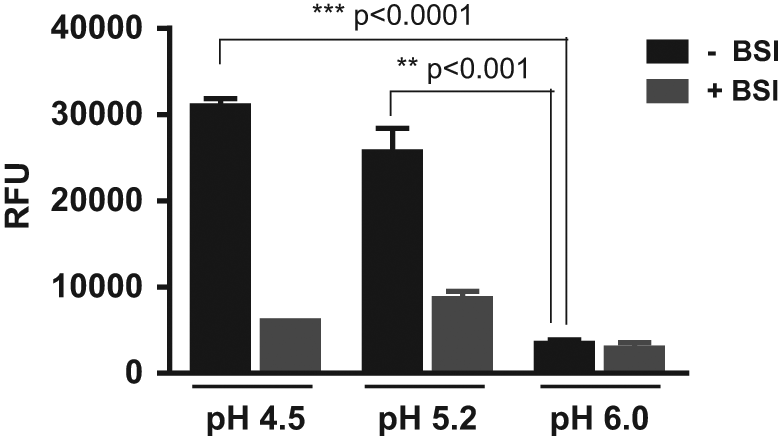
pH profile of recombinant β-site amyloid precursor protein cleaving enzyme–1 (BACE-1). The enzymatic activity of recombinant BACE-1 was determined at various pH using a LanthaScreen BACE-1 assay, a fluorescence resonance energy transfer (FRET)–based BACE-1 (β-secretase) assay, and a HitHunter β-secretase assay in the absence or presence of a BACE-1 inhibitor (BSI). Representative data for FRET-based BACE-1 assay are shown. Data are means ± SEM. Statistical analysis was conducted using a one-way analysis of variance with Bonferroni’s multiple comparison test. RFU, relative fluorescence units.
Development and Optimisation of a BACE-1 Membrane Assay
To establish an in vitro assay for full-length BACE-1, solubilized membrane preparations from HEK–BACE-1 cells ( Fig. 2a ) were generated. Evaluation of these membranes by Blue Native polyacrylamide gel electrophoresis (PAGE) demonstrated that the majority of BACE-1 in these membranes was monomeric (data not shown). To monitor BACE-1 activity, solubilized membranes were incubated with a fluorogenic APP-based peptide substrate containing the β-cleavage site of Swedish mutant APP. Assays were conducted at either pH 4.5 or pH 6.0 and in the absence or presence of a BACE-1 inhibitor (BSI) to determine the assay window. Initially, a detergent screen was conducted to identify the optimal detergent for solubilization of BACE-1–containing membranes ( Fig. 2b ). Solubilization of the membranes with 1% Triton X-100 or 0.5% CHAPS was found to efficiently solubilize BACE-1, resulting in the highest signal-to-background (S/B) window in assays conducted at pH 4.5 (14- and 11-fold, respectively) and pH 6.0 (5- and 6-fold, respectively). In contrast, solubilization with other detergents resulted in smaller assay windows at pH 4.5 and, importantly, did not result in any appreciable signal window at pH 6.0. In addition, solublization with a lower concentration of Triton X-100 (0.05%) did not efficiently solubilize BACE-1, resulting in a small assay window (data not shown). Based on these results, solubilization with 1% Triton X-100 was selected for future studies. In addition, the inclusion of a protease inhibitor cocktail containing ETDA in both the MES buffer used for resuspension of the membranes prior to their solublization with detergent and in the buffer used for the BACE-1 activity assay resulted in a larger S/B window at pH 6.0, but this had no marked effect on the assay window at pH 4.5 (data not shown). Finally, resuspension of the membranes in a small volume of MES buffer (0.2 mL per T225 flask) prior to solubilization with detergent resulted in a lower background signal compared with the use of higher volumes of buffer (1 mL per T225 flask) (data not shown). Although the latter method was more efficient at extracting proteins from the membranes and led to a greater total protein yield, it was found to be detrimental as it was associated with a higher background signal in the assay (data not shown).
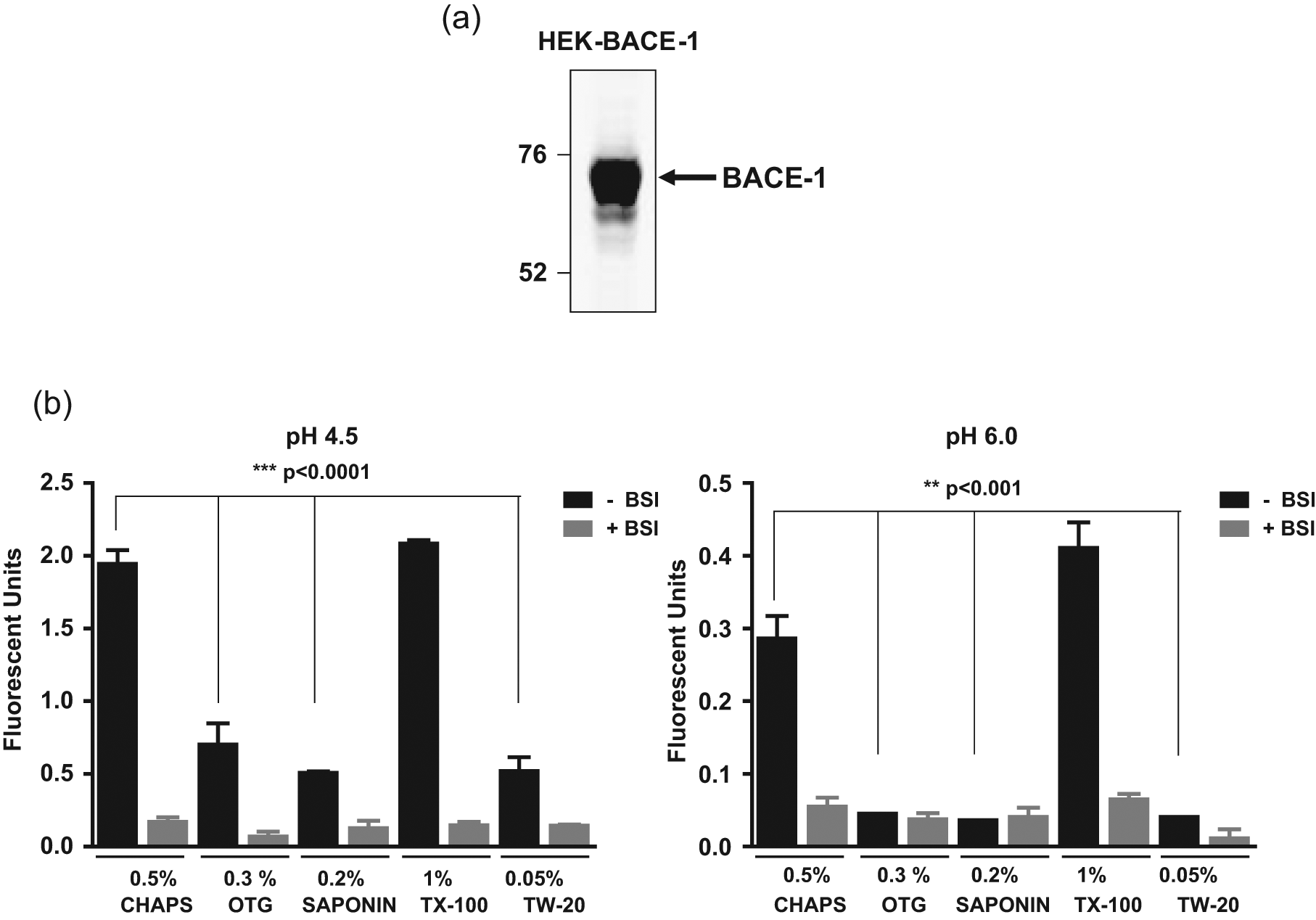
β-Site amyloid precursor protein cleaving enzyme–1 (BACE-1) expression and solubilization. (
Development of a High-Throughput BACE-1 Membrane Assay
The optimized assay parameters described above subsequently formed the basis for the development of a high-throughput BACE-1 membrane assay in a 384- and 1536-well format. Initially, time course studies with solubilized BACE-1 membranes (equivalent to 7.5 or 15 µg/mL total protein at pH 4.5 and pH 6.0, respectively) and increasing substrate concentrations (62.5–2000 nM) were performed using a BACE-1 FRET assay (β-secretase) kit. At pH 4.5, the reaction was linear with a substrate concentration-dependent increase in signal during the 30-min time period ( Fig. 3a ). A similar profile was observed at pH 6.0, although the reaction proceeded at a much slower rate, despite the presence of twice as much membrane proteins ( Fig. 3b ). In a longer time course study, we confirmed that the reaction at pH 6.0 was linear over a 50-min period over the same substrate concentration range (data not shown). One of the major considerations for a high-throughput assay is cost-effectiveness. Consequently, to minimize costs, the assay conditions were set to 15 µg/mL solubilized BACE-1 membranes (~ 2.5 nM BACE-1 enzyme) and 250 nM substrate. This gave an assay window of approximately 6 and a Z′ value >0.6—parameters that were sufficient for a robust HTS. The final assay format chosen for the HTS and compound characterization used the BACE-1 FRET assay (β-secretase) kit (Invitrogen) since it was very comparable to the LanthaScreen BACE-1 assay (data not shown) and was more cost-effective.
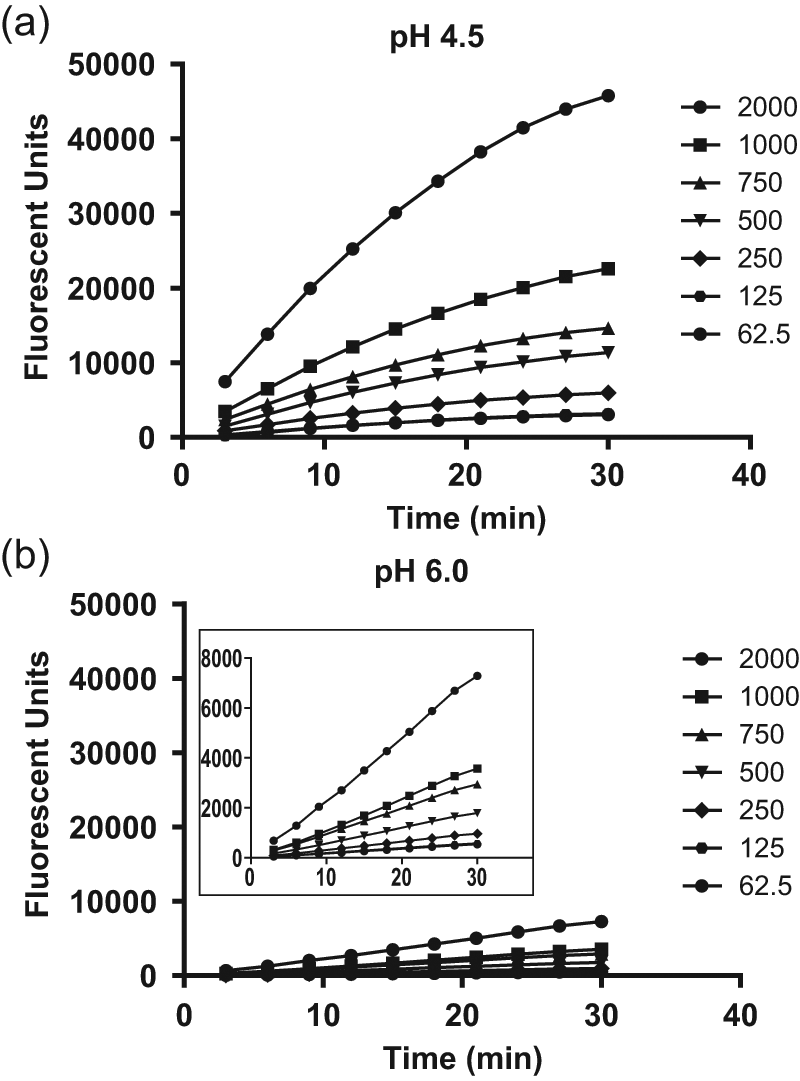
Time course studies at pH 4.5 and pH 6.0. A time course study was conducted using β-site amyloid precursor protein cleaving enzyme–1 (BACE-1) solubilized membranes in a 384-well format in the presence of varying concentrations of substrate at (
Inhibitor Profiling and HTS in BACE-1 Membrane Assay
Prior to running an HTS, several BACE-1 inhibitors were selected for evaluation in the BACE-1 membrane assay ( Fig. 4a ). These included a statine inhibitor (BSI III), a hydroxyethylamine transition-state analog inhibitor (BSI IV), and four novel small-molecule inhibitors of BACE-1 (compounds A–D).25–29 The potency of each compound was determined in the membrane assay at both pH 4.5 and pH 6.0 ( Table 1 ). With the exception of the statine BSI III, which appeared to be approximately 15-fold less potent at pH 6.0 compared with pH 4.5, the BACE-1 inhibitors exhibited similar potencies at both pH. A compound library of 550 000 compounds was subsequently tested at a screening concentration of 10 µM in the 1536-well assay format. Applying a cutoff value of −40% inhibition of BACE-1, a hit rate of 0.11% was obtained. The active 617 hits were picked for confirmation, and out of these, 35% gave an IC50 value of <10 µM in two independent IC50 runs, which resulted in a final 0.04% total hit rate. The mean S/B on the HTS plates was ~6, and the mean Z′ was 0.66. To determine if screening at pH 6.0 rather than at pH 4.5 was advantageous with regard to the identification of chemical leads, the activity of the confirmed hits was subsequently determined at pH 4.5. Interestingly, the majority of the hits were found to be markedly less active at pH 4.5 compared with pH 6.0 ( Fig. 5 ).
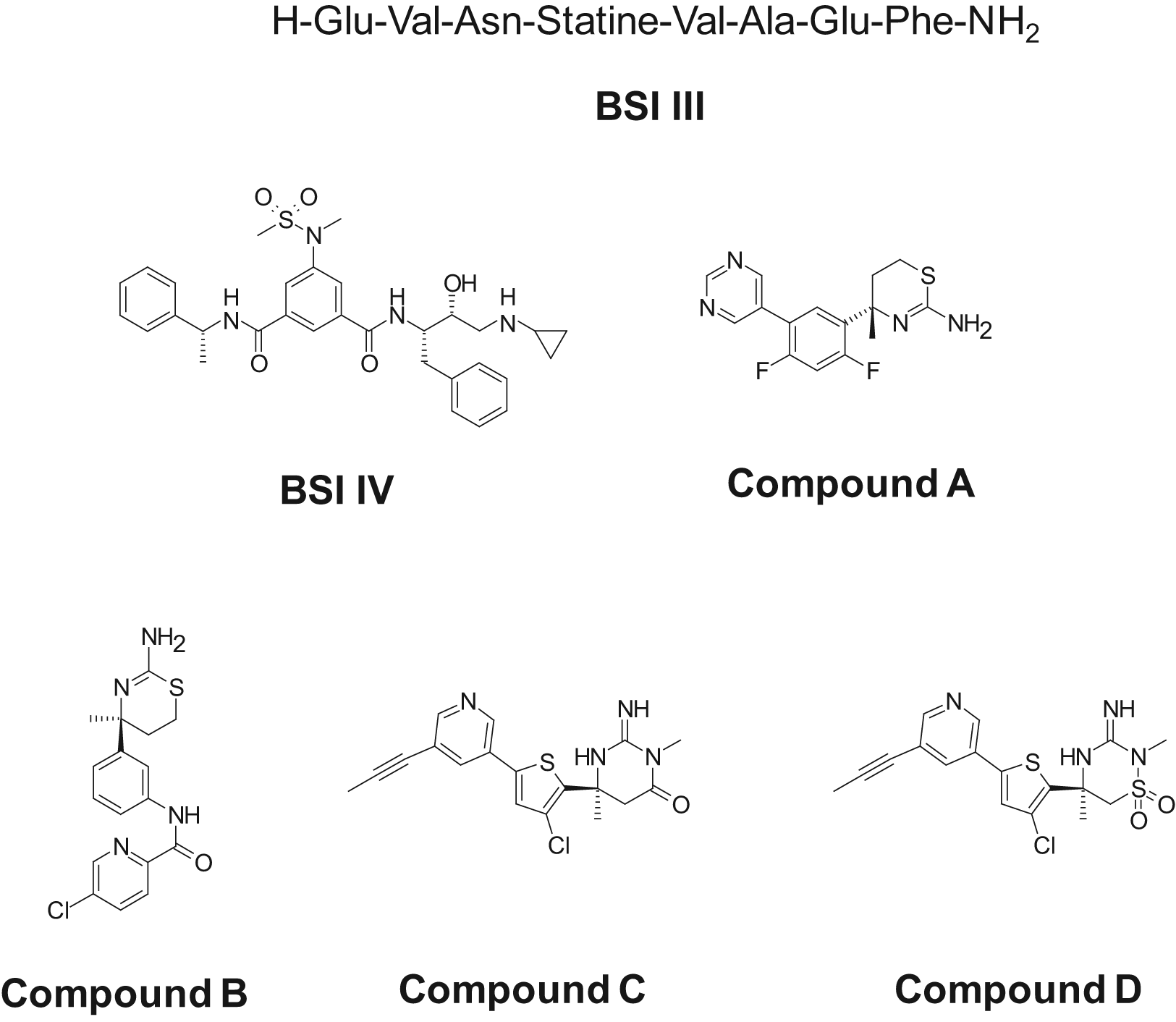
Chemical structures of β-site amyloid precursor protein cleaving enzyme–1 (BACE-1) inhibitors.
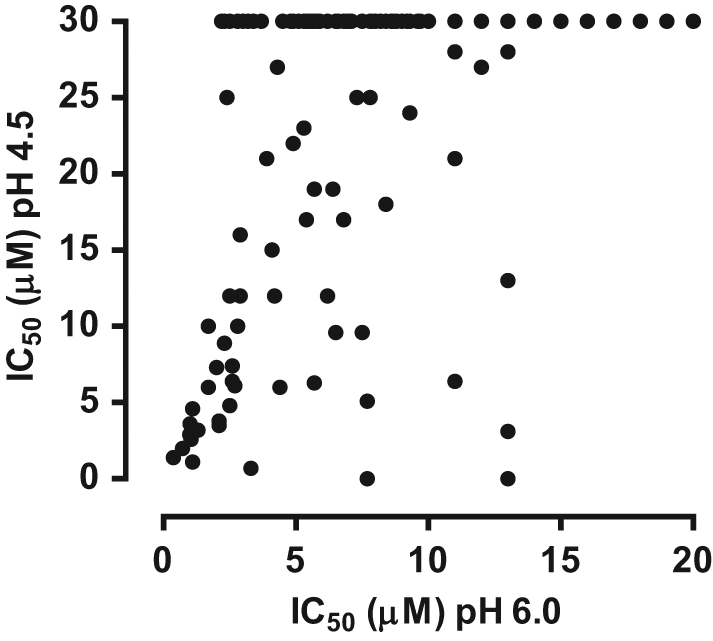
Profiling of high-throughput screening (HTS) hits at pH 4.5 and pH 6.0. The activity of the confirmed HTS hits was evaluated in the β-site amyloid precursor protein cleaving enzyme–1 (BACE-1) membrane assay at pH 4.5 and pH 6.0. Only data for compounds with an IC50 of ≤20 µM at pH 6.0 are shown.
BACE-1 Inhibitor Profiling in the Membrane Assay at pH 4.5 and pH 6.0
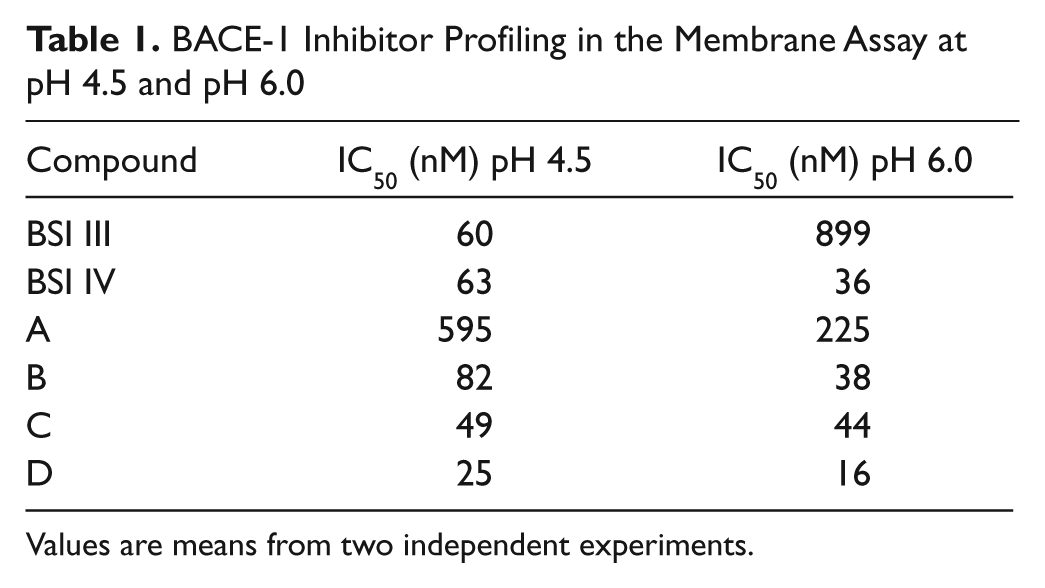
Values are means from two independent experiments.
Inhibitor Profiling in HEK-APPwt Cells
To evaluate the potencies of BACE-1 inhibitors in cells, HEK cells overexpressing APP wild type (HEK-APPwt) were incubated with compound and the effect on the level of Aβ42 in the medium determined. The small-molecule BACE-1 inhibitors caused a concentration-dependent decrease in Aβ42 levels with IC50 values in the nM range ( Table 2 ). BSI IV was also active in cells, whereas BSI III was only weakly active ( Table 2 ). The hits from the HTS were also evaluated in this cellular assay. Initial screening identified 19 hits, which reduced Aβ42 levels in cells with IC50s <14 µM. Upon further screening and evaluation of the hits in a cell viability assay, only two hits were confirmed as cell active BACE-1 inhibitors, decreasing total Aβ levels with IC50s of approximately 2 to 3 µM.
BACE-1 Inhibitor Profiling in HEK-APPwt Cells
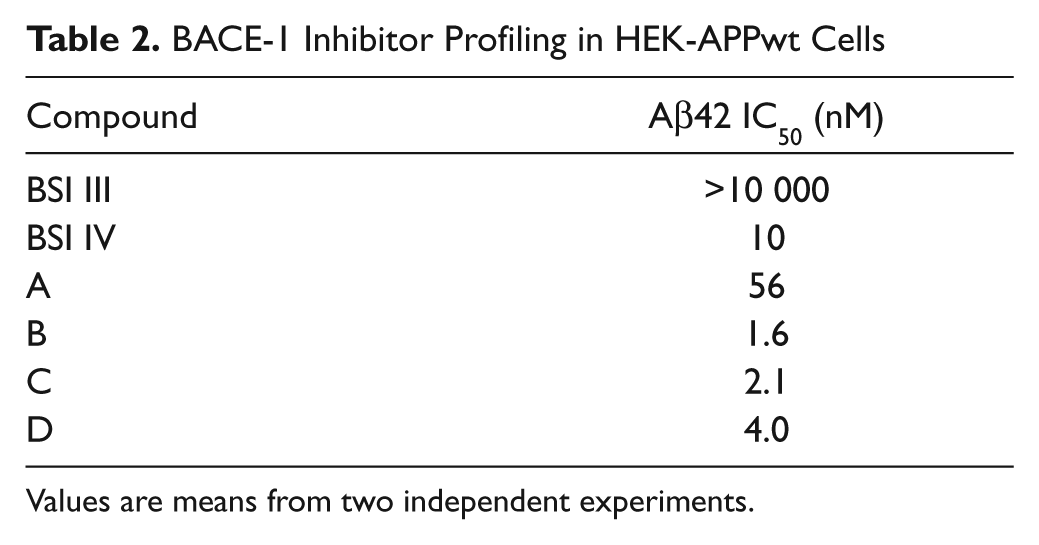
Values are means from two independent experiments.
Discussion
Here we describe the development and characterization of a novel membrane assay for full-length BACE-1. Compared with conventional biochemical assays for BACE-1 that have relied predominantly on the use of the soluble ectodomain of BACE-1 that contains the catalytic active site, this novel membrane assay offers several advantages. First, it incorporates full-length BACE-1 in its native membrane environment rather than a truncated form of the enzyme. This form of BACE-1 can be posttranslationally modified and could potentially interact with other proteins via its transmembrane domain and cytoplasmic tail. Second, full-length BACE-1 in the membranes exhibits measurable activity at pH 6.0, which more closely resembles the pH of the Golgi and endocytic compartments in cells where BACE-1 resides and cleaves APP.3,6 This contrasts with our observations and published findings with recombinant BACE-1, which exhibits minimal activity at pH 6.0.3,11 Hence, the BACE-1 membrane assay is physiologically more relevant and could favor the identification of novel BACE-1 inhibitors, including active site and non–active site binders. Indeed, a noncompetitive BACE-1 inhibitor, TAK-070, that binds to full-length BACE-1 but not to truncated BACE-1 lacking the transmembrane domain was previously identified from a cell-based assay, 30 thereby highlighting the potential value of alternative, novel BACE-1 screening assays.
In initial experiments, we determined the optimal protocol for preparation of the solubilized membranes from HEK–BACE-1 cells. A number of parameters were found to be critical to deliver a robust membrane assay for BACE-1 with measurable activity at pH 6.0. These included solubilization with 1% Triton X-100 and the use of an EDTA-containing protease inhibitor cocktail during membrane preparation and in the activity assay. The latter suggested the presence of an EDTA-sensitive metalloprotease in the membranes, which contributed to the background signal at pH 6.0, which was successfully reduced in the presence of EDTA. Interestingly, BACE-1 activity in the membranes was higher at pH 4.5 compared with pH 6.0, consistent with the pH profile described for the soluble ectodomain of BACE-1.3,11 These findings suggested solubilized BACE-1 membranes could be used to run screens to identify novel BACE-1 inhibitors at either pH 4.5 or pH 6.0. However, we favored the latter as these assay conditions would more closely mimic the cellular environment where BACE-1 is active.
The assay was subsequently modified to deliver a high-throughput 1536-well BACE-1 membrane assay that was HTS compatible and could be used for the characterization of BACE-1 inhibitors. An important consideration for the HTS was to deliver a cost-effective assay. This was successfully achieved by identifying the minimum amount of solubilized BACE-1 membranes and BACE-1 substrate required to give a robust assay window of approximately 6 and a Z′ value of >0.6 using the cost-effective BACE-1 FRET assay format.
Profiling of BACE-1 inhibitors in the membrane assay at pH 4.5 and pH 6.0 demonstrated that the small-molecule BACE-1 inhibitors were equipotent in both assays and were active in the cellular HEK-APPwt assay. In contrast, the peptidic statine inhibitor (BSI III) was markedly less potent at pH 6.0 and only weakly active in the cellular assay. The lack of activity in the cellular assay most likely reflects the poor cell permeability of this large peptidic compound. Overall, these findings suggested that an HTS at pH 6.0 would offer the additional advantage that it would disfavor the identification of peptidic scaffolds that were not tractable starting points for BACE-1 inhibitor development. An HTS was subsequently run at pH 6.0. The confirmed hit rate was relatively low, which is typical for this target class. However, a number of hits were identified, including the known small-molecule BACE-1 inhibitors present in our compound library. Interestingly, the majority of these hits were less potent in assays run at pH 4.5, thereby supporting the rationale for screening at higher pH.
In summary, we have developed a novel BACE-1 membrane assay that mimics the natural physiology of BACE-1. The data presented here validate the utility of this assay for the identification and characterization of small-molecule inhibitors of BACE-1 for the treatment of AD.
Footnotes
Acknowledgements
We thank Aurelie Baguet for the generation of the BACE-1 cDNA; Birgit Hartmann, Karlheinz Hergesell, Michael Kiaser, and Helga Drosdat for technical support of the HTS; and Heinz Thoma for technical support of the cell assay.
Declaration of Conflicting Interests
All authors are full time employees of Merck Serono S.A.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by Merck Serono S.A.
