Abstract
The insulin-degrading enzyme (IDE) cleaves numerous small peptides, including biologically active hormones and disease-related peptides. The propensity of IDE to degrade neurotoxic Aβ peptides marks IDE as a potential therapeutic target for Alzheimer disease. Using a synthetic reporter based on the yeast
Introduction
According to the Aβ hypothesis, Aβ peptides contribute to Alzheimer disease (AD) by serving as a source of neurotoxicity. It remains unclear whether neurotoxicity stems from accumulation of the monomeric, oligomeric (ADDLs), and/or fibrillar forms of Aβ. 1 Independent of the neurotoxic species, strategies aimed at reducing Aβ levels hold promise for the treatment of AD. Because biosynthetic and clearance activities regulate steady-state Aβ levels, either or both activities could be manipulated to reduce Aβ. Although most research on Aβ has centered on its production (i.e., the α, β, and γ secretases), several mechanisms are known to clear Aβ. 2
One prominent Aβ clearance mechanism involves proteolytic cleavage of the Aβ monomer, which eliminates its ability to form higher order structures. The insulin-degrading enzyme (IDE; EC 3.4.24.56) is one of several Aβ-cleaving enzymes; others include neprilysin, plasmin, matrix metalloprotease-9, angiotensin-converting enzyme, and endothelin-converting enzyme. 2 In animal models, IDE deficiency correlates with increased Aβ levels and increased risk of AD, whereas IDE overexpression appears to protect against AD.3,4 Genetic linkage and biochemical analyses also strongly support a connection between IDE and AD.5,6 IDE also cleaves other small molecules (e.g., insulin, glucagon, and amylin) that can adopt β secondary structure and form amyloid fibrils. 7
IDE belongs to the M16A family of zinc-dependent metalloproteases, which are evolutionarily widespread and highly conserved in sequence and structure.8,9 An inverted zinc-binding motif is characteristic of this family. The recent elucidation of several M16A enzyme structures, including that of human IDE, pitrilysin (Escherichia coli), and the related M16C enzyme PreP1 (Arabidopsis thaliana), reveals a structure resembling that of a clamshell in which substrates form β-sheet contacts with β-strands of IDE.8,9 Members of the M16A family can recognize each others’ substrates as evident by the ability of mammalian IDE and pitrilysin to substitute for the yeast M16A enzymes Axl1p and Ste23p in production of the yeast
The proposed physiological role of IDE in Aβ clearance has led to the hypothesis that its hyperactivation could be therapeutically beneficial. Studies of IDE involving under- and overexpression in transgenic mice support this hypothesis.3,4 Small-molecule activators of IDE can thus be considered potential therapeutic agents, and several such molecules have been reported.14,15 Nucleotide triphosphates (i.e., adenosine triphosphate [ATP]) activate IDE to cleave certain small substrates other than Aβ. This effect occurs at ATP concentrations (i.e., 0.1–1 mM) much higher than that thought to exist physiologically in the extracellular environment where Aβ is supposedly proteolyzed by IDE (i.e., 5–50 nM) but within the range expected intracellularly (4–5 mM).16,17 ATP-dependent enhancement of IDE activity does not involve ATP hydrolysis; the effect is also buffer dependent. 14 High-throughput screening (HTS) has identified two additional small-molecule activators of IDE. 15 Curiously, these molecules enhance IDE-mediated cleavage of an Aβ reporter only when the synthetic non-Aβ HTS substrate is also present. The synthetic compound suramin has also been indicated as an IDE activator. 18 To date, data pertaining to its effectiveness as an IDE activator or AD therapeutic have not been released.
In this study, we report seven chemical activators of IDE that were identified by HTS from a pharmacophore-rich small-molecule library provided by the National Institutes of Health (NIH) Developmental Therapeutics Program. We demonstrate the ability of identified compounds to enhance the activity of rat IDE toward both synthetic and natural peptides based on the yeast
Materials and Methods
Peptides
Two synthetic substrates were used to monitor the in vitro activities of the IDE orthologs evaluated in this study. One was an internally quenched, fluorogenic peptide Abz-SEKKDNYIIKGV-NitroY-OH (AnaSpec, Inc., San Jose, CA; CHI Scientific, Inc., Maynard, MA), where Abz is aminobenzoic acid and NitroY is 3-nitro-tyrosine. The peptide is based on the sequence flanking the M16A cleavage site found in the yeast
The substrates used to monitor activities of yeast Rce1p and yeast Ste24p were internally quenched, fluorogenic, farnesylated peptide substrates that are based on the K-Ras C-terminus.19,20 Product formation for these substrates was measured using a 320/420-nm excitation/emission filter set.
Recombinant Enzymes and Other Reagents
Plasmids encoding rat and yeast Ste23p have been described.13,21 The plasmid encoding human IDE was constructed by amplifying the human IDE open reading frame from plasmid IDE-pSRα using oligos designed to encode a 6X-His tag at the N-terminus and restriction sites for subcloning the PCR product into the XbaI and NotI sites of pET30b(+) (Novagen, Madison, WI). 22 The plasmid encoding Caenorhabditis elegans IDE was constructed by amplifying the cDNA sequence of F44E7.4 from the RB1 cDNA library by PCR and subcloning into the XbaI and NotI sites of pET30b(+). 23 Recombinant rat, human, C. elegans, and yeast IDE were inducibly expressed in BL21 (DE3) E. coli and recovered by immobilized nickel affinity chromatography essentially as previously described.13,21 Purified IDE was stored at –80 °C as 1 mg/mL aliquots in storage buffer (50 mM HEPES, 140 mM NaCl, 20% glycerol [pH 7.4] or 25 mM KPi, 200 mM NaCl, 20% glycerol [pH 7.6]). Membranes containing yeast Rce1p or yeast Ste24p activity were prepared as previously described. 20 Bovine trypsin, pronase E, proteinase K, ATP, bovine serum albumin (BSA), and human recombinant insulin were all from Sigma-Aldrich (St. Louis, MO). Ia1 and Ia2 were from Key Organics (London, UK).
Chemical Library
The Diversity Set compound library was obtained through the NIH Developmental Therapeutics Program (DTP). 24 This compound library contains 1981 compounds with unique pharmacophore characteristics that were reduced from a parent set of more than 70 000 compounds with the use of the 3D structure generation program Chem-X (Chemical Design Ltd., Mahwah, NJ).
High-Throughput Screen
Assays were conducted in the wells of a black, clear-bottom, 96-well microplate (Costar, Corning, Inc., New York, NY) in 0.1 M potassium phosphate buffer (KPi), pH 7.6. The fully assembled reactions contained rat IDE (10 µg/mL), compound (100 µM), and
Chemical Analyses
The structures of the identified IDE activators were rendered in ChemBioDraw Ultra (v. 11.0; CambridgeSoft, Cambridge, MA). Molecular weight, Log P, and other quantitative structure-activity relationship (QSAR) values were calculated using the Analysis and Chemical Properties tools of ChemBioDraw. In addition, molecular weights of the compounds were confirmed by electrospray ionization (ESI) analysis (Proteomic and Mass Spectrometry Core Facility, University of Georgia) using an API I Plus, PE Sciex mass spectrometer (Perkin Elmer, Waltham, MA) in positive or negative ion mode depending on the biophysical properties of the compounds. All samples were dissolved in methanol for ESI analysis.
Target Specificity Analyses
The effects of compounds on the activities of other enzymes were assessed using methods similar to that outlined above for HTS. Reaction rates were determined in triplicate after a 10-min preincubation with 100 µM compound. Single-dose effects on rat IDE (10 µg/mL), Ste23p (10 µg/mL), trypsin (0.5 µg/mL), and pronase E (0.5 µg/mL) were determined using the
Dose-Response, Kinetic, and Other Assays
Values for dose-response curves were determined according to the conditions outlined above (i.e., KPi buffer) using a range of activator concentrations (up to 1 mM) and trypsin/pronase normalization. Half-maximal activating concentration values (AC50) were derived from best-fit curves obtained by plotting the data points in Prism 4.0 (GraphPad Software, Inc., La Jolla, CA). For compounds lacking sigmoidal dose response-curves, the concentration resulting in half-maximal activation ([Max]50) was used.
Data for kinetic analyses were determined over a range of substrate concentrations (0–750 µM) in the absence and presence of activator (100 µM). Data collected were subjected to nonlinear regression analysis in Prism 4.0 to extract kinetic parameters. Curves were fitted using the equation Y = Vmax * X/[Km + X]. Because internal quenching effects were observed with the
The effects of BSA, ATP, insulin, and buffer composition were determined in the context of 50 µM
The activating properties of two previously reported IDE activators (Ia1 and Ia2) were also evaluated for comparison to the hits obtained in this study. 15 They were evaluated as for the single-dose target specificity studies described above using previously reported concentrations of Ia1 (50 µM) and Ia2 (10 µM).
Enzyme-Chemical Interaction Studies
For determining thermal melt profiles, a solution of recombinant rat IDE (0.5 µM) and SYPRO Orange (5× working solution diluted from a 5000× stock solution; Sigma-Aldrich) was prepared in 0.1 M KPi (pH 7.6) and dispensed as 100-µL aliquots into a 96-well plate (HSP-9601; MJ Research, Waltham, MA) containing DMSO or a candidate activator (100 µM). The wells were overlaid with 10 µL mineral oil and incubated in the dark at room temperature for 60 min. Samples were heated from 28 to 70 °C (Δt increment = 1–3 °C) and fluorescence measured (465 nm excitation, 590 nm emission) at appropriate intervals using a FluoDia T70 high-temperature fluorescence plate reader (Photon Technology International, Birmingham, NJ). Thermal melt mid-points (Tm) were determined from a fluorescence versus temperature plot that was fit to a four-parameter sigmoidal dose-response (variable slope) equation using Prism software. Similar results were obtained when IDE was evaluated in 0.1 M HEPES buffer (pH 7.5) containing 150 mM NaCl.
For native polyacrylamide gel electrophoresis (PAGE) profiles, rat IDE (0.5 µg) was preincubated with compound (100 µM) for 60 min at 37 °C, mixed with equal volume of 2× native gel sample buffer (0.1% bromophenol blue, 20% glycerol, 0.6 M Tris-HCl, pH 6.8), and analyzed by 10% native PAGE at 4 °C. Gels were stained with silver and scanned using a flatbed scanner.
Yeast Strains and Plasmids
The yeast strains used in this study were IH1783 (MAT
Yeast Toxicity Assay
To determine whether compounds could exert their activating effects in vivo, it was first critical to determine whether any were cytotoxic, which would complicate in vivo analyses. The effect of compounds on yeast growth was assessed by determining the density of compound-treated cultures after incubation for 44 h at 30 °C. Briefly, a saturated starter culture of yeast y272 cotransformed with pWS192 and pWS496 was diluted (1:2000) into SC-UW and split into equal-volume aliquots (1 mL), and aliquots were treated with DMSO or compound (100 µM). Each condition was evaluated in triplicate, and the average absorbance observed for each condition after 44 h was calculated as a percentage of that observed for the DMSO-treated condition, which was set at 100%. To obtain accurate measurements of culture densities, saturated cultures were typically diluted 10-fold, and A600 values obtained were expressed as percent of untreated. Where toxicity was observed, a lower dose allowing for saturated growth after 44 h of incubation at 30 °C was empirically determined. For compounds that retained toxicity at these lower concentrations, the yeast cultures were grown for 72 h to allow saturated growth.
a-Factor Production Assay
A bioassay was used to assess the impact of compounds on the ability of IDE to promote
Results
An a-Factor–Based Internally Quenched Fluorogenic Peptide Is Cleaved by IDE
The yeast mating pheromone
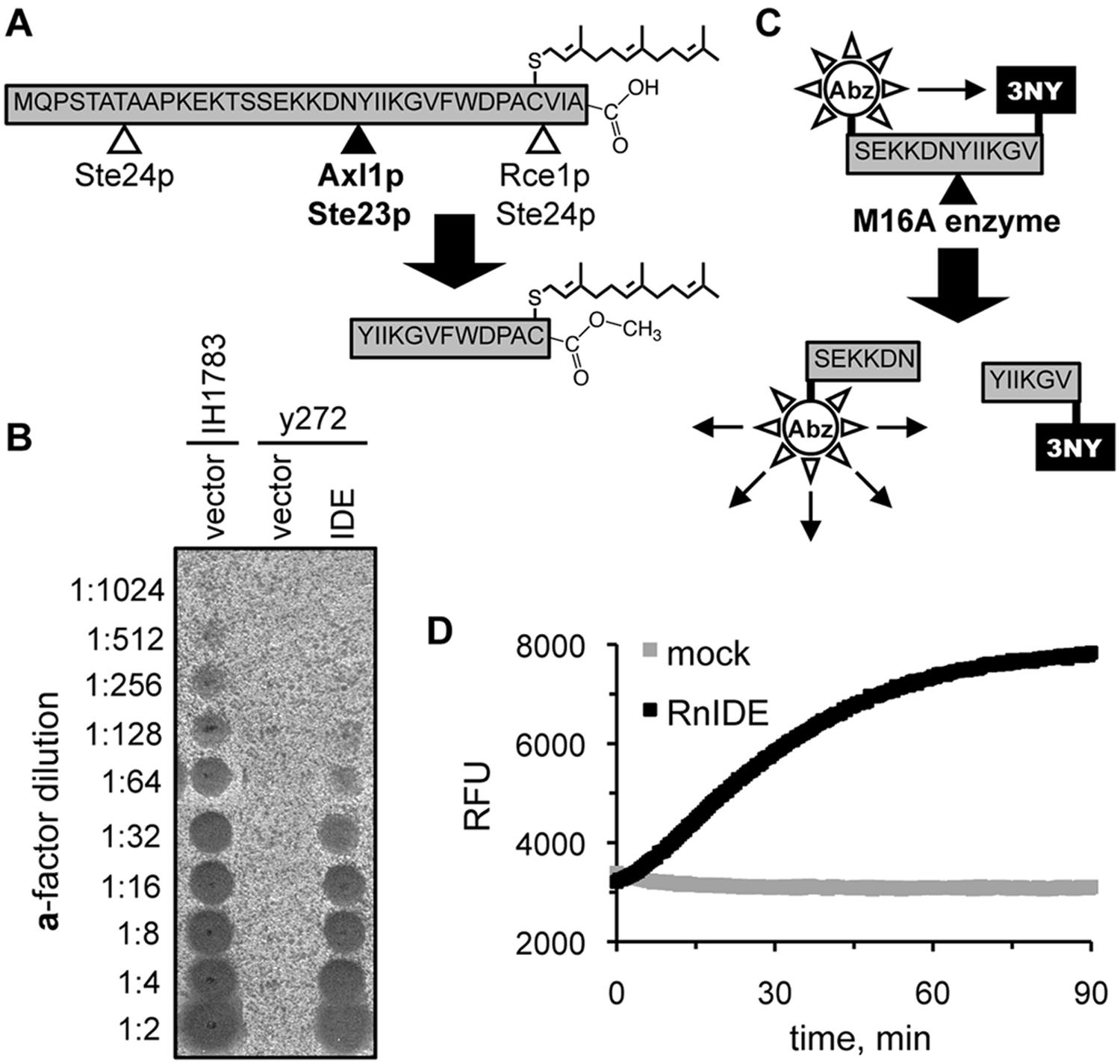
Reporters of M16A enzyme activity. (
Identification of IDE Activators by HTS
The assay described above was optimized for a 96-well format and used to screen the NIH DTP Diversity Set library for modulators of IDE activity.
24
Under the conditions of the screen, the assay had a calculated Z′ factor of 0.86 and a signal-to-noise ratio of 7.0. The compounds of the library had a variety of effects on IDE activity. Of the 1981 compounds within the DTP library, 451 partly inhibited IDE (<50% activity relative to untreated), and 1495 had a minor impact on activity (50%–150% activity observed). Two compounds could not be assessed due to strong quenching effects exerted on the substrate. Of specific interest were 33 compounds (1.7% hit rate) that enhanced the rate of fluorescence output ≥150% relative to that observed for DMSO-treated IDE; DMSO represents the solvent for the library collection and was used as a control. The set of hits was further reduced to eight compounds after elimination of autofluorescing and fluorescence-enhancing compounds. An additional compound (
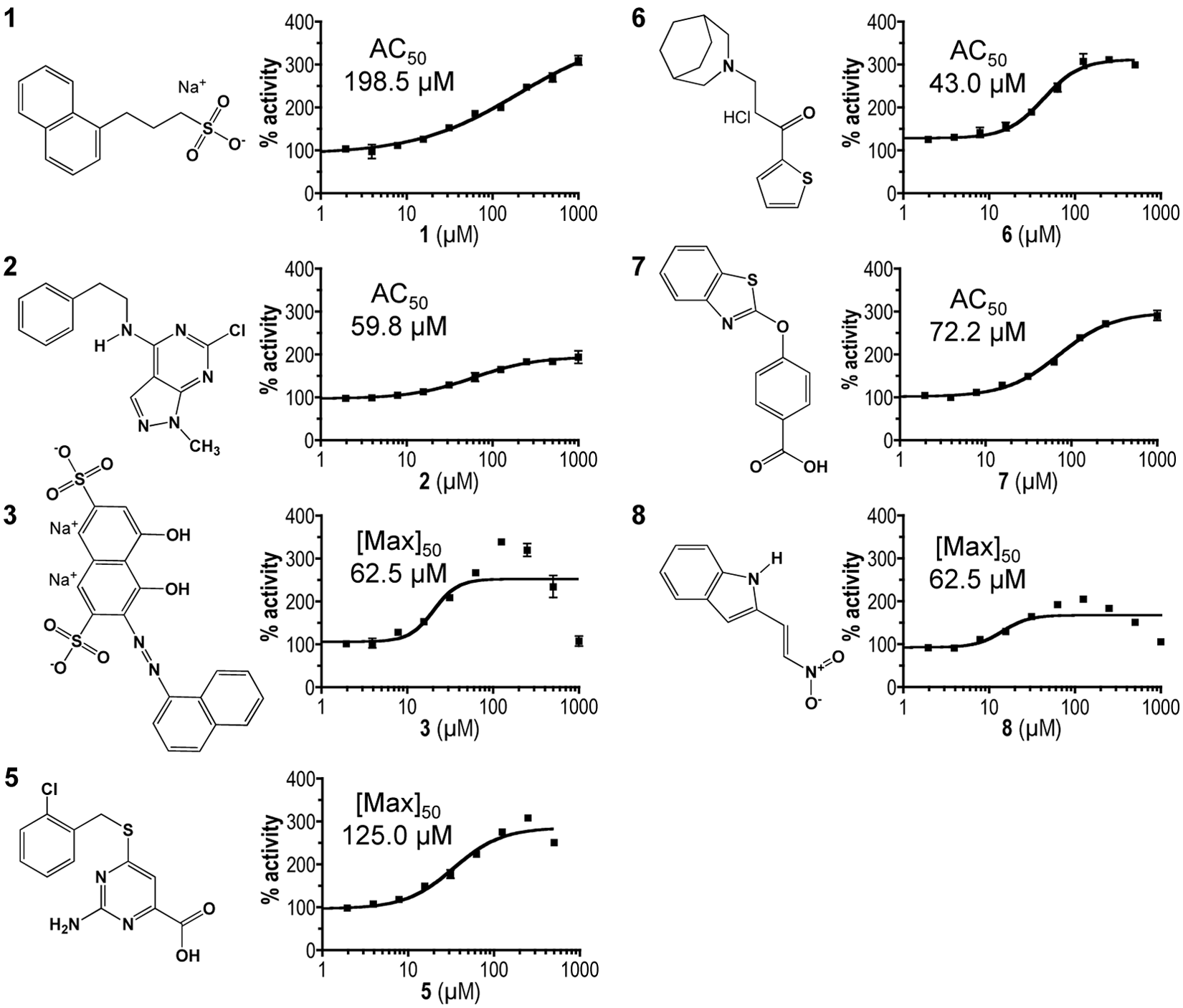
Chemical structures and dose-response profiles of insulin-degrading enzyme (IDE) activators. Compounds were identified by their ability to enhance rat IDE-mediated in vitro cleavage of the peptide reporter depicted in
Figure 1C
. Structures were downloaded from the Developmental Therapeutics Program (DTP) structure database (http://dtp.nci.nih.gov/branches/dscb/diversity_explanation.html). Compound
The compounds all adhere to Lipinski’s rule of 5 and have other drug-like characteristics.
32
The compounds range in mass from 188 to 472 Da (associated counterions excluded) (
Activators Are IDE Specific
The impact of compounds (100 µM) on the activities of IDE and a select group of other proteases was examined (
Table 1
). The maximum activation of IDE observed was 356% (compound
Activities of Various Proteases in the Presence of Rat Insulin-Degrading Enzyme (IDE) Activators
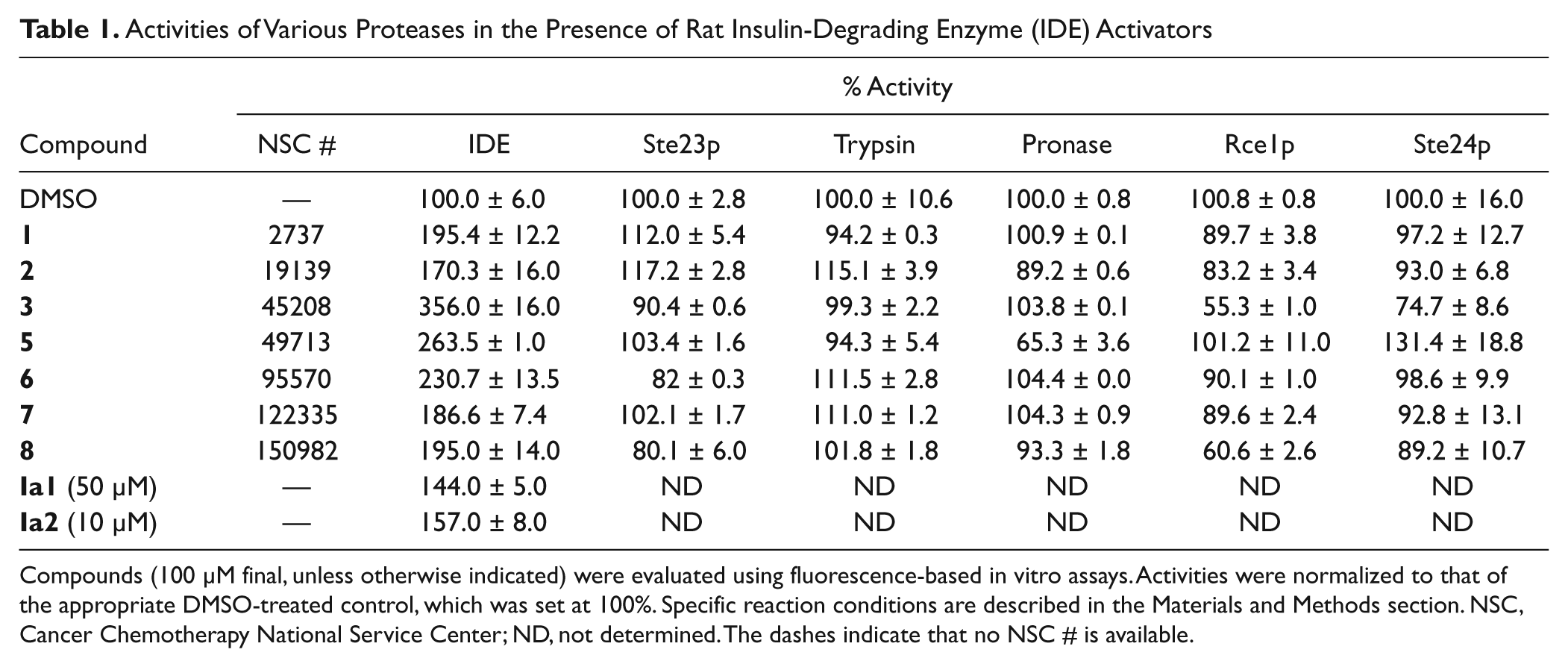
Compounds (100 µM final, unless otherwise indicated) were evaluated using fluorescence-based in vitro assays. Activities were normalized to that of the appropriate DMSO-treated control, which was set at 100%. Specific reaction conditions are described in the Materials and Methods section. NSC, Cancer Chemotherapy National Service Center; ND, not determined. The dashes indicate that no NSC # is available.
Dose-Response and Kinetic Studies
The HTS screen for IDE activators used a fixed concentration of compound. To determine the optimal activating concentration of compounds, dose-response assays were performed. Compounds
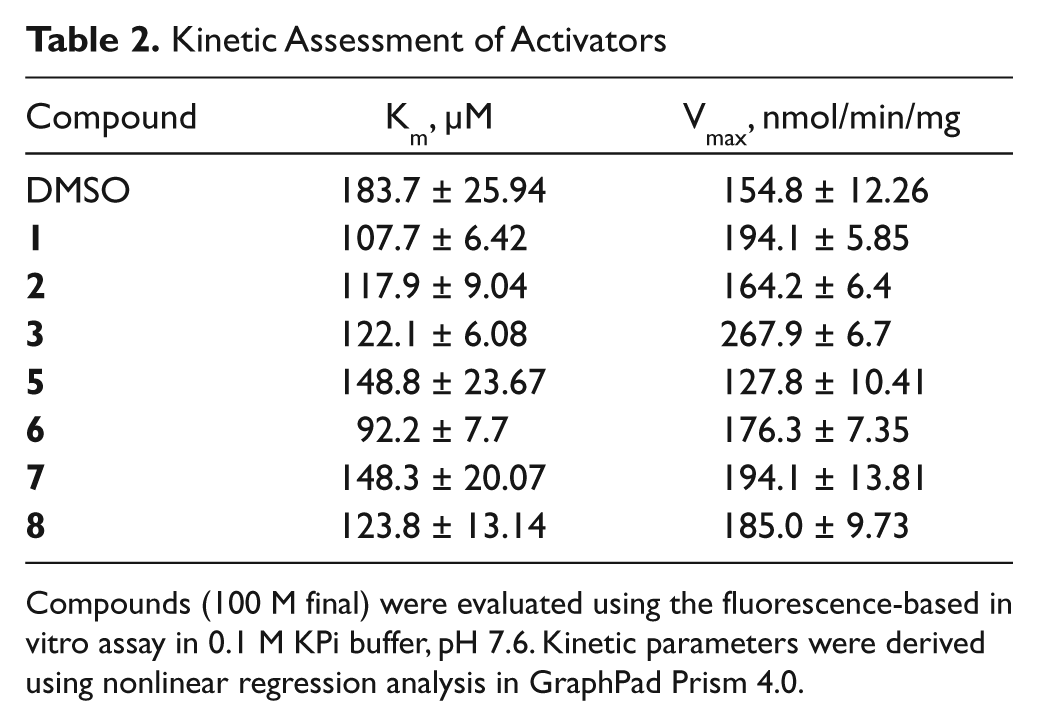
Enzyme kinetic studies were performed in the presence of 100 µM compound. All compounds reduced the Km of IDE, and all but compound
Kinetic Assessment of Activators
Compounds 3, 6, and 8 Physically Interact with IDE
To determine whether the activators interact with IDE to exert their effects, we performed two types of studies. First, we hypothesized that ligand binding would alter the biophysical properties of IDE in some manner. Thermal melt analyses revealed a substantial shift in the melting profile of IDE in the context of compound
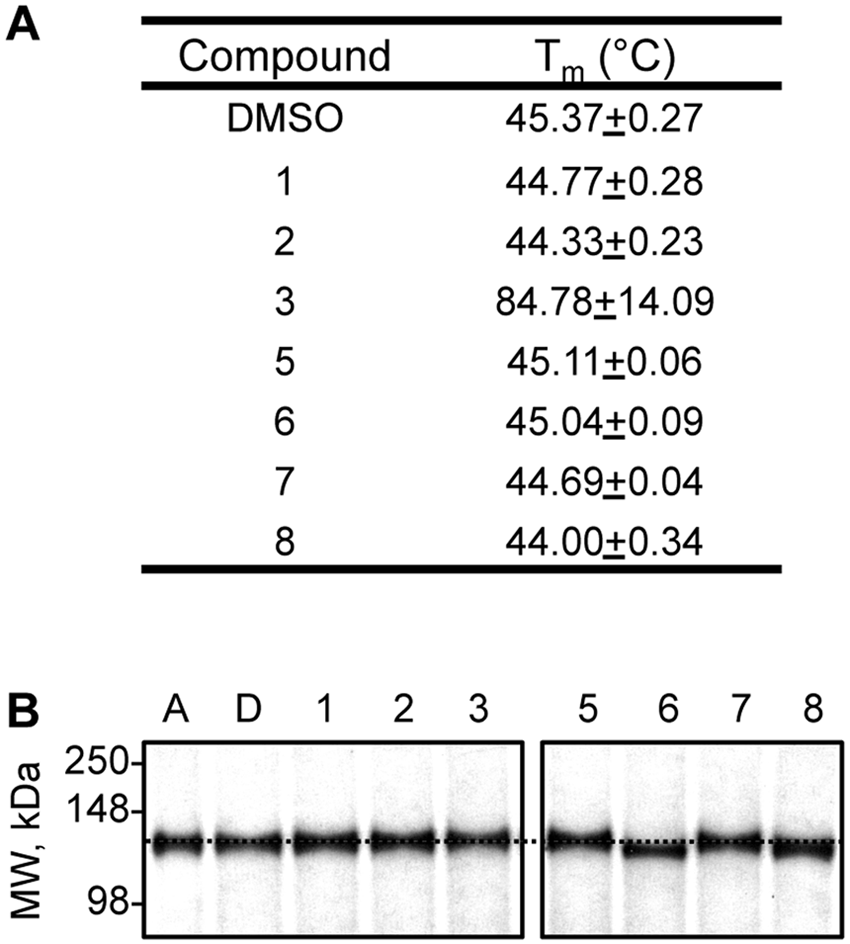
Effect of compounds on the biophysical properties of rat insulin-degrading enzyme (IDE). (
Effect of Assay Conditions on Activator Potency
Promiscuous activation has been reported for certain enzyme/activator combinations. 25 Often, the activating effects of such compounds are mitigated in the presence of BSA or low concentrations of detergent. In the presence of low amounts of BSA (≤0.02%), a modest increase in IDE activity was observed ( Fig. 4A ). The reason for this effect is unknown and could reflect stabilization of a more active form of IDE or better substrate/enzyme availability (i.e., less nonspecific adsorption by plasticware). Importantly, all of the compounds retained their ability to activate IDE in the presence of an optimal dose of BSA ( Fig. 4B ). The relative amount of activation observed was less than that observed in the absence of BSA ( Table 1 ), which is due to the higher baseline activity of the BSA-treated control. Similarly, detergent treatment (0.1% CHAPS) also reduced but did not completely neutralize compound-induced activation of IDE (S. P. Manandhar and W. K. Schmidt, unpublished observation).
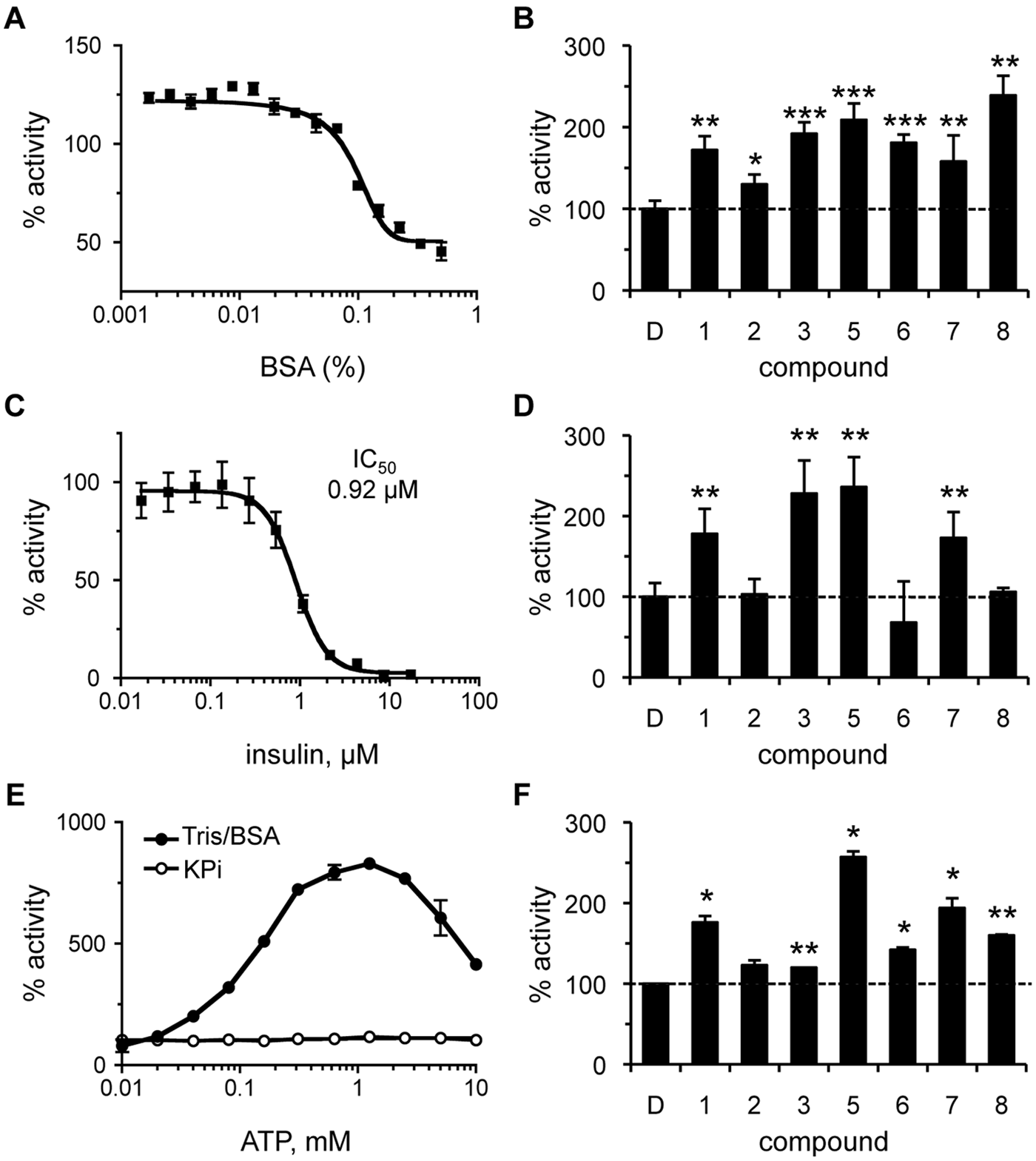
Effect of assay conditions on the properties of rat insulin-degrading enzyme (IDE) activators. The effect of bovine serum albumin (BSA), insulin, and adenosine triphosphate (ATP) on the activity of IDE was determined both in the absence and presence of compounds. Rat IDE was used (87.7 nM) in the assay described in
Figure 1
. Values are reported as percentages relative to a water or DMSO-treated control as appropriate (n = 4); *p < 0.05, **p < 0.01, and ***p < 0.001 relative to the mock-treated control. For dose-response curves, data points were plotted, and a best-fit nonlinear dose-response curve was determined using GraphPad Prism 4.0 as described in
Figure 2
. (
On the basis of the fact that IDE can cleave multiple substrates, we investigated the ability of our compounds to hyperactivate IDE under mixed substrate conditions. The compounds were evaluated in the presence of insulin, which disproportionately inhibits IDE-mediated Aβ degradation.
21
The effect of insulin on degradation of the
Last, we examined whether our compounds were activating IDE in an ATP-like manner.
14
We expected this to be unlikely since ATP-dependent activation was not observed under the assay conditions used for HTS. To resolve whether the lack of ATP-dependent activation was due to our substrate or assay conditions, we evaluated the effect of ATP under assay conditions where effects had been observed.14,15 The switch from a KPi buffering system to one containing Tris/BSA resulted in ATP-dependent activation of IDE, indicating that using KPi buffer indeed neutralizes ATP-dependent activation of IDE (
Fig. 4E
). The maximum activating concentration in the Tris/BSA buffering system was approximately 1 mM. Evaluation of the activators under these conditions revealed that all the compounds, with the exception of compound
Effect of Activators on Yeast Growth and IDE-Mediated Production of a-Factor
The ability of compounds to enhance IDE-mediated cleavage of an
Using conditions guided by the toxicity data, we next addressed whether any of the compounds could stimulate IDE-mediated
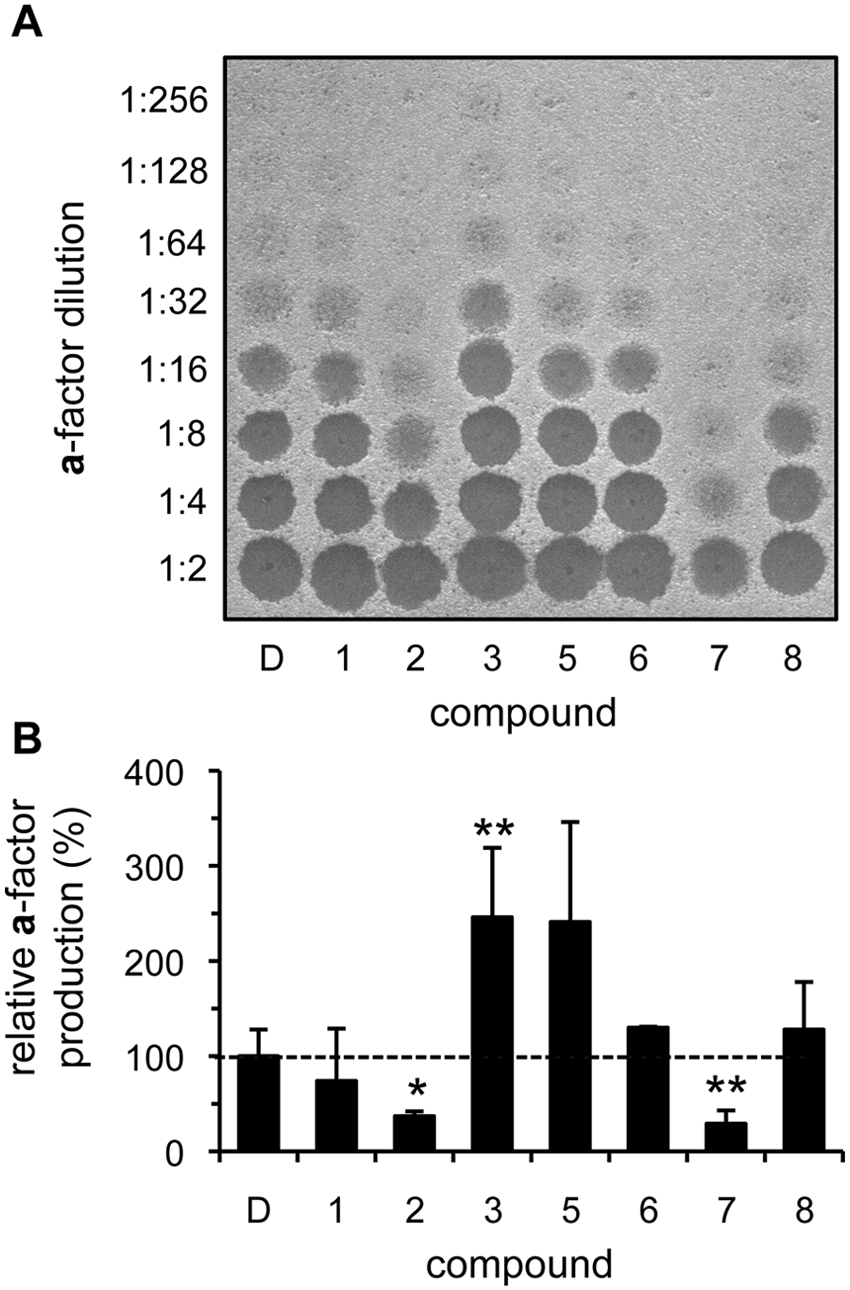
Select compounds enhance in vivo insulin-degrading enzyme (IDE)–dependent production of yeast
Where changes in net
Substrate and Species Specificity of IDE Activators
Although the synthetic and in vivo
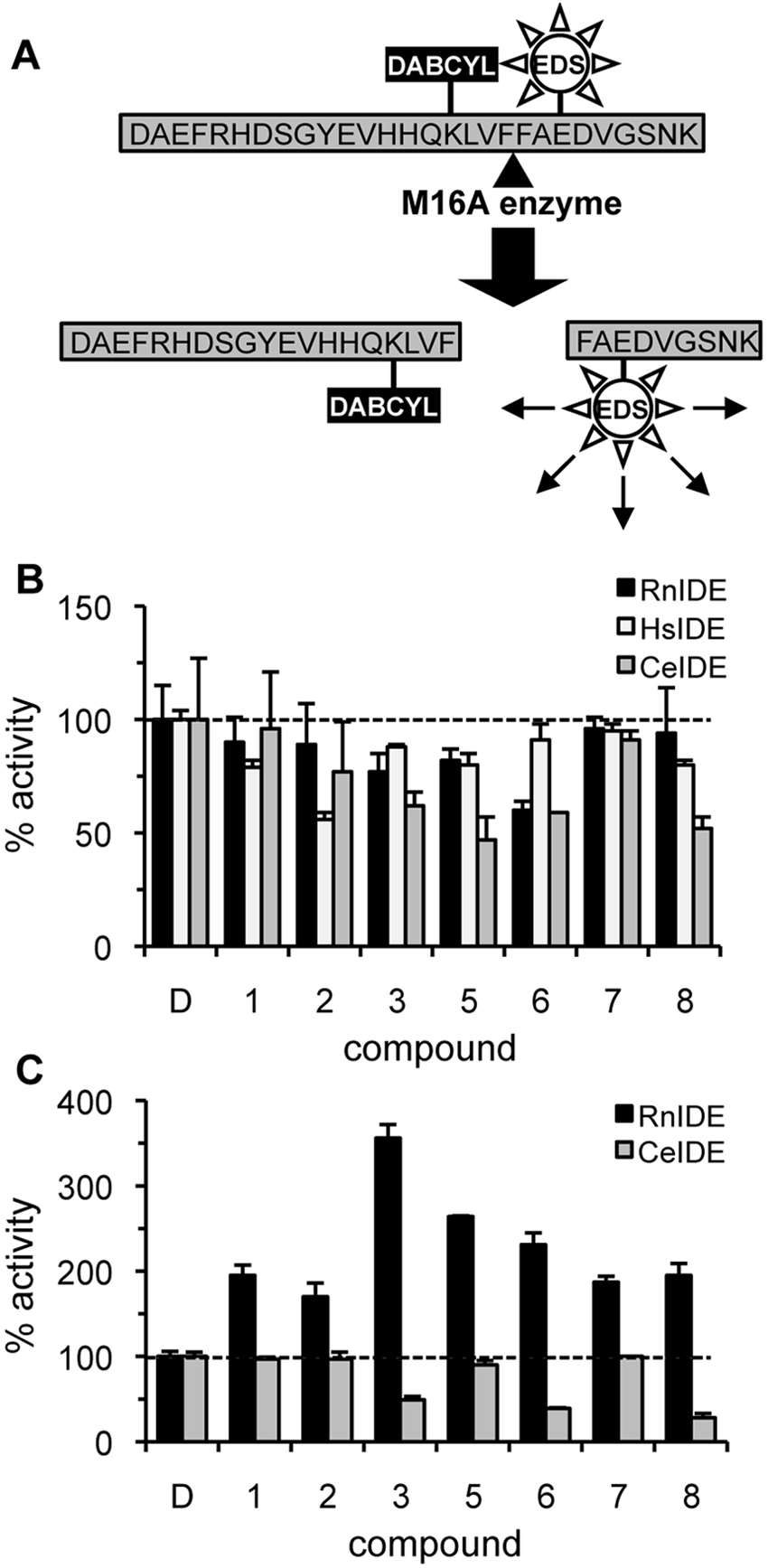
Rat insulin-degrading enzyme (IDE) activators are substrate and species specific. (
Discussion
IDE cleaves small amyloidogenic peptides. 35 Thus, the activation of IDE has been viewed as therapeutic for the clearance of Aβ. 36 Indeed, IDE overexpression reduces amyloid deposits in the brain of amyloid precursor protein (APP) transgenic mice. 4 IDE can also be rendered hyperactive through mutations that expose the active site for a longer period of time. 9 As documented by our findings and related studies, chemical activation of IDE is also feasible.15,18 We have identified seven small-molecule activators of rat IDE that lower the Km of IDE, retain activation in the presence of a competing substrate (insulin), and are effective in an in vivo assay. The activators are structurally distinct from previously reported IDE activators, consistent with the possibility that there may be multiple modes of IDE activation.
An important and critical outcome of our study revolves around the use of nonnative IDE substrates as reporters (i.e.,
We expect our current findings to provide guidance for future studies aimed at identifying IDE activators. Foremost, the observation that the compounds identified in this study do not generally activate IDE orthologs emphasizes the importance of using the appropriate target enzyme (i.e., human IDE) in future high-throughput screens aimed at generating therapeutic agents. Our study also provides evidence that a hormetic in vitro dose-response profile (e.g., compounds
Perhaps the most important impact of our study will be on the design of future IDE reporters, which our study suggests should resemble Aβ as closely as possible. A key step toward this goal is our development of a FRET-based Aβ reporter. Our reporter is based on Aβ1–28 with a strategically positioned quencher (DABCYL) and fluorophore (EDANS) on K16 and E22, respectively. Choice of the 1–28 peptide versus a longer Aβ species was driven by the technical issues frequently reported for syntheses and storage of longer forms of Aβ. 39 Moreover, residues beyond E23 lack ordered density in the IDE/Aβ1–40 co-crystal structure (PDB 2g47), suggesting that they do not affect substrate binding. 9 K16 and E22 were chosen for side-chain modification because our analysis of the IDE/Aβ1–40 co-crystal structure suggested that these residues are in reasonable proximity and could be modified without impairing interactions between Aβ and IDE. Importantly, the Aβ1–28 reporter has a free N-terminus and thus retains an important exosite binding capability that is not found on other IDE HTS-compatible reporters (e.g., FAβB). 40 An Aβ1–28 reporter having an N-terminal fluorophore has been reported as a relatively poor IDE substrate, but specific data were not reported, and direct comparison between the reporters is thus not possible. 40 We contend that N-terminal modifications may lead to binding interference at the IDE exosite and complicate the determination of kinetic parameters. We also acknowledge that the failure of our activators to enhance cleavage of the FRET-based Aβ1–28 reporter could be indicative of interference by the EDANS and/or DABCYL moieties leading to inappropriate folding of the peptide. It is unknown whether alternative fluorophores and quenchers will yield similar results.
There are many examples of enzymes that are activated by biological small molecules (e.g., PKA by cAMP). There are, however, relatively few examples of enzymes that can be activated through synthetic small molecules. This list of enzymes is growing and includes notable targets such as glucokinase, AMP-activated protein kinase (AMPK), and the sirtuins. 41 IDE activators can now be added to the limited set of small-molecule enzyme activators, and it is expected that our development of a FRET-based Aβ1–28 reporter will help drive the identification of species and Aβ-specific activators of IDE.
Footnotes
Acknowledgements
We thank the Developmental Therapeutics Program (National Institutes of Health) for providing the Diversity Set compound library and the following (all at the University of Georgia): Dr. Z. A. Wood for help with IDE purification and design of the Aβ1–28 reporter, Dr. J. P. Rose for assistance with thermal shift studies, the Bioexpression and Fermentation Facility for help with HPLC analyses, Dr. T. M. Dore, and members of the Schmidt laboratory for critical discussions and technical assistance.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Alzheimer’s Drug Discovery Foundation/Institute for the Study of Aging (grant 271223 to WKS) and the National Institutes of Health (5R01GM067092 to WKS).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
