Abstract
Activation of the antioxidant response element (ARE) upregulates enzymes involved in detoxification of electrophiles and reactive oxygen species. The induction of ARE genes is regulated by the interaction between redox sensor protein Keap1 and the transcription factor Nrf2. Fluorescently labeled Nrf2 peptides containing the ETGE motif were synthesized and optimized as tracers in the development of a fluorescence polarization (FP) assay to identify small-molecule inhibitors of the Keap1-Nrf2 interaction. The tracers were optimized to increase the dynamic range of the assay and their binding affinities to the Keap1 Kelch domain. The binding affinities of Nrf2 peptide inhibitors obtained in our FP assay using FITC-9mer Nrf2 peptide amide as the probe were in good agreement with those obtained previously by a surface plasmon resonance assay. The FP assay exhibits considerable tolerance toward DMSO and produced a Z′ factor greater than 0.6 in a 384-well format. Further optimization of the probe led to cyanine-labeled 9mer Nrf2 peptide amide, which can be used along with the FITC-9mer Nrf2 peptide amide in a high-throughput screening assay to discover small-molecule inhibitors of Keap1-Nrf2 interaction.
Introduction
The upregulation of oxidative stress response enzymes by natural products from fruits and vegetables represents the focus of current chemoprevention research.
1
The expression of genes that encode these enzymes are regulated through a
By comparing the amino acid sequences of Nrf2 across different species, six conserved homologous domains (Neh1 to Neh6 domains) of Nrf2 have been identified.
10
Of these domains, the
The
The DLG and the ETGE motifs within the Neh2 domain of Nrf2 are two evolutionary conserved motifs of the CNC family of proteins. The DLG motif has been reported to be involved in ubiquitin-dependent degradation of Nrf2, whereas the ETGE motif has been reported to be essential for the Keap1 regulation of Nrf2 activity.24,25 Although both motifs bind to the same site in Keap1, the ETGE motif was reported to be approximately 100-fold stronger in affinity to Keap1 than the DLG motif.
14
Several different ETGE-containing Nrf2 peptides have been reported to displace Nrf2 from Keap1.
26
The longer ETGE-containing 14mer and 16mer Nrf2 peptides (H-LQLDEETGEFLPIQ-OH and H-AFFAQLQLDEETGEFL-OH) were able to effectively displace the Nrf2 protein from the Keap1:Nrf2 complex; the binding affinity (
We have recently reported the development of surface plasmon resonance (SPR)–based assays to examine the interactions between Keap1 Kelch domain and Nrf2, to measure the inhibitory activity of Nrf2 peptides against the Keap1-Nrf2 interaction and to determine the minimal Nrf2 peptide sequence required for Keap1 binding. 27 Despite their advantages of label-free detection, full automation, and real-time monitoring of binding interactions, the SPR assays require separate injections for each test sample, are of relatively low throughput, and do not meet the throughput requirement for high-throughput screening (HTS) applications. To facilitate the discovery of small-molecule inhibitors of Keap1-Nrf2 interaction, we report the development of a homogenous fluorescence-based competition assay that can be adapted to the high-throughput screening of chemical libraries. This assay uses a peptide tracer selected from a series of Nrf2 peptides of varying length based on the high-affinity ETGE motif in the Neh2 domain of the Nrf2 protein. We demonstrated that the assay can distinguish inhibitory Nrf2 peptides that have varying affinities. The assay is robust and can be adapted to a high-throughput format for the discovery of small-molecule inhibitors of the Keap1:Nrf2 complex. Molecules that interfere with the Keap1:Nrf2 complex formation can be used to induce the expression of ARE genes and could potentially be developed into chemopreventive and therapeutic agents.28,29
Materials and Methods
Peptide Synthesis
All peptides were synthesized and purified as described previously. 27 Briefly, standard Fmoc chemistry was used for the synthesis of Nrf2 peptides on either Wang or 2-chlorotrityl chloride (2CTC) resin. After cleavage, the peptides were purified on a Gilson preparative high-performance liquid chromatography (HPLC) system using a reversed-phase BDS-Hypersil C18 column (150 × 20 mm). The peptides were obtained at >95% purity and their identities confirmed by LC-MS and high-resolution mass spectrometry.
Peptide Modification and Labeling
Fluorescein-labeled peptides were prepared by modification of purified peptides with fluorescein isothiocyanate (FITC) according to the manufacturer’s standard protocol. Briefly, the peptide (1 mg) was dissolved in 0.5 mL of 100 mM sodium carbonate buffer at pH 8.5, and FITC (2 eq.) was added to the solution. The reaction was stirred at room temperature for 12 h while protected from light. The progress of the labeling reaction was monitored by the disappearance of the starting peptide using LC-MS. A 1 N solution of sodium bisulfate (NaHSO4) was added to reduce the pH of the reaction to 2, and the precipitated crude modified peptide was collected by centrifugation. The product was purified by reversed-phase HPLC. For labeling of the 9mer Nrf2 peptide amide (H-LDEETGEFL-NH2) with
The
The
Peptide Concentration Determination
The concentrations of unlabeled Nrf2 peptide solutions were determined by amino acid analysis through acid hydrolysis and
Protein Expression and Purification of Human Keap1 Kelch Domain
The recombinant human Keap1 Kelch domain was expressed and purified as previously described.31,32 Briefly, the Kelch domain (residues 321–609) was cloned into a Novagen pET15b vector (EMD Chemicals, San Diego, CA) between an
Anisotropy Measurements
All fluorescence polarization (FP) assays were performed on a Wallac Victor 3V multilabel counter/plate reader (PerkinElmer, Shelton, CT) using the excitation and emission filters appropriate for each fluorophore used in the binding experiment. The plates used for the FP measurements were the black nonbinding surface Corning 3650 96-well or Corning 3575 384-well plates, loaded with 80 or 40 µL of assay solution per well. For fluorescein and BODIPY, 485 nm excitation and 535 nm emission filters were used. For Cy3B, excitation and emission filters of 560 nm and 650 nm were used, respectively. For Cy5, 579 nm excitation and 670 nm emission filters were used. The FP assays were performed in 10 mM HEPES buffer, pH 7.4, containing 50 mM EDTA, 150 mM NaCl, and 0.005% Tween-20. All aqueous solutions were prepared using deionized water collected from a Millipore water purification system.
FP was determined by measuring the parallel and perpendicular fluorescence intensity (F║ and F┴) with respect to the linearly polarized excitation light. The fluorescence polarization (

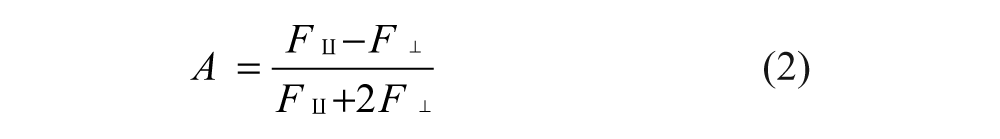

We elected to use anisotropy in our quantitative analysis because anisotropy values can be combined additively. In addition, total fluorescence (F║ + 2F┴) was calculated for each well to ensure that the fluorescence intensity was consistent with the amount of fluorescent probes used across all assay wells.
Measurement of Binding of Fluorescent Peptide Probes to the Keap1 Kelch Domain
The binding of the fluorescently labeled Nrf2 peptides to the Keap1 Kelch domain was determined by dose titration of the fluorescent peptide probes with varying concentrations of Keap1 Kelch domain protein. The maximum anisotropy achieved at a high concentration of Keap1 Kelch domain protein defines the upper end of the dynamic range for the peptide probe, and the total fluorescence (F║ + 2F┴) at the high concentration of Keap1 Kelch domain protein can be used to calculate the
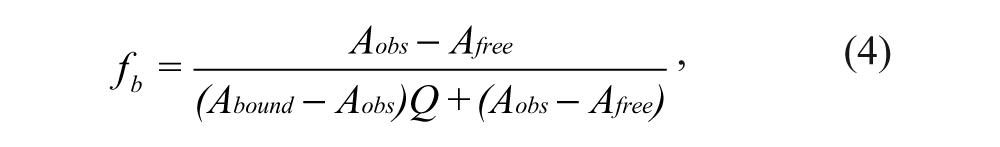
where
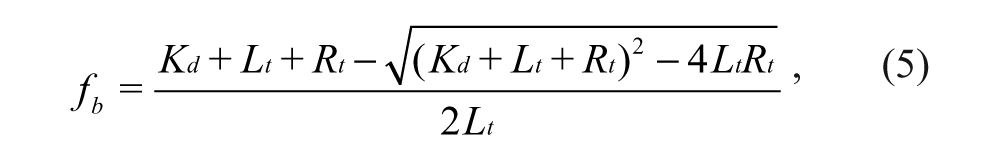
where
FP Competition Assays to Determine the Inhibitory Potency of the Keap1-Nrf2 Interaction
A competition assay using the conditions described above was established and miniaturized to a 384-well plate format to determine the potency of inhibitors of the Keap1-Nrf2 interaction. A Packard Bioscience (now PerkinElmer) Multiprobe II HT EXpanded liquid-handling robot was used to deliver small volumes of solutions to each of the 384 wells. Each well had a final volume of 40 µL that consisted of 10 µL of 40 nM FITC-9mer Nrf2 peptide amide and 10 µL of 400 nM Keap1 Kelch domain protein, 10 µL of HEPES buffer, and 10 µL of an inhibitor sample of varying concentrations. The binding experiments were performed in triplicates, with initial concentration of the inhibitor typically set between 10 µM and 100 µM depending on the inhibitor potency and serially diluted twofold to give a concentration range down to 0.5 nM or 5 nM. The plate was centrifuged at 370
The percentage inhibition of the competitor at each concentration point was determined by using equation 6, and the IC50 of an inhibitor was determined from the plot of %inhibition against inhibitor concentration using equation 7.
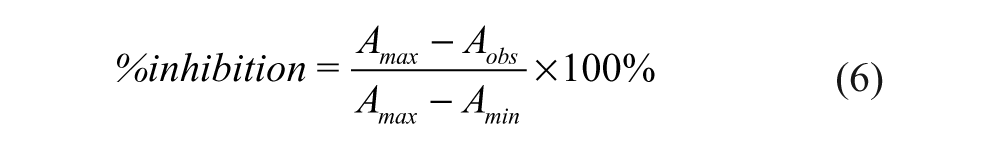
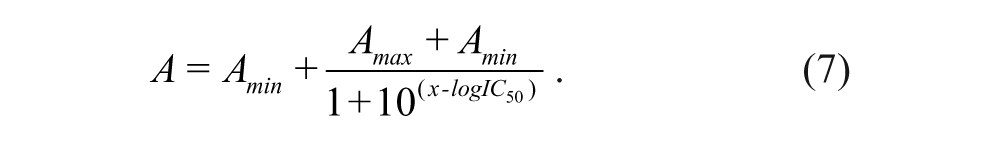
IC50 is the concentration of an inhibitor needed to inhibit 50% binding of the fluorescently labeled peptide probe, and
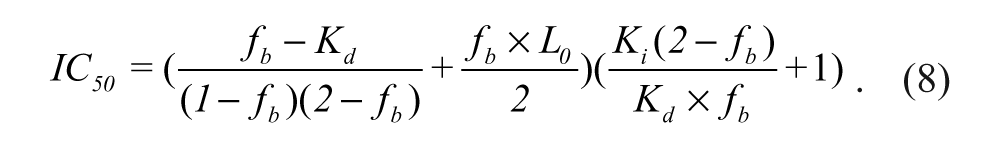
For the measurement of Z′ factor, a 384-well plate was prepared with 10 nM FITC-9mer Nrf2 amide and 100 nM Keap1 Kelch domain in 10 mM HEPES buffer in the presence or absence of 1 µM of
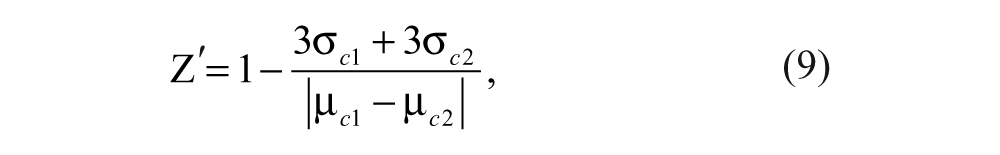
where µ
The Effect of DMSO on Anisotropy Measurement
The effect of DMSO on our FP assay was determined by measuring the anisotropy change of the Keap1-bound and free fluorescent probe in the presence of DMSO in the assay solutions at concentrations up to 20%. Briefly, 20 µL of the assay buffer containing varying concentrations of DMSO and 10 µL of 40 nM FITC-9mer Nrf2 peptide amide was added to a Corning 3575 384-well plate. Then, either 10 µL of 400 nM Keap1 Kelch domain protein or 10 µL of buffer was added to each well, and the plate was centrifuged for 2 min at 370
SPR Binding Assay
The SPR competition binding assay was carried out on a Biacore 3000 biosensor (GE Healthcare) using the immobilized
Pilot Screen of the NCI Diversity Set II and Clinical Collection
Test compounds were dissolved in DMSO to give 10 mM stock solutions like most library compounds are. The pilot screen was performed in duplicates at 100 µM of test compound, 100 nM Keap1 Kelch domain, and 10 nM FITC-9mer Nrf2-NH2 in 10 mM HEPES buffer, pH 7.4, containing 2% DMSO with a total volume of 40 µL in 384-well Corning 3650 plates.
Results and Discussion
Selection of the FP Assay Conditions
Nrf2 peptides of different length (8mer to 16mer) based on the ETGE motif were synthesized using Fmoc chemistry. The ETGE motif is known to bind to Keap1 with 100-fold higher affinity than the DLG motif
14
; thus, we hypothesized that fluorescently labeled peptides based on the ETGE motif could be used as the probe with high binding affinity to the Keap1 Kelch domain. The length of the Nrf2 peptides also affects Keap1 binding with the ETGE-containing 16mer Nrf2 peptide (H-AFFAQLQLDEETGEFL-OH) reported to have a
To determine the optimal conditions for our FP assay, we used the FITC-labeled 16mer Nrf2 peptide as the initial fluorescent probe to explore different assay conditions. We measured the anisotropy of the fluorescently labeled 16mer Nrf2 peptide as a function of its concentration to select the optimal concentration of the probe for use in our FP binding assays. Anisotropy is a size-dependent, not concentration-dependent, property, and the observed anisotropy in the presence of protein is related to the amount of the protein-bound probe relative to the free probe. However, too low of a probe concentration would lead to increased errors in anisotropy measurements. It is thus necessary to select a probe concentration that will provide reliable anisotropy signal. As shown in Figure 1A , 10 nM of the FITC-16mer Nrf2 peptide gave a reliable anisotropy measurement that is within the fluorescence linear range. Concentrations higher than 10 nM did not significantly improve the error of anisotropy measurement, while at lower concentrations, the anisotropy signals fluctuate as the errors become more prominent due to background noise in the fluorescence measurements.
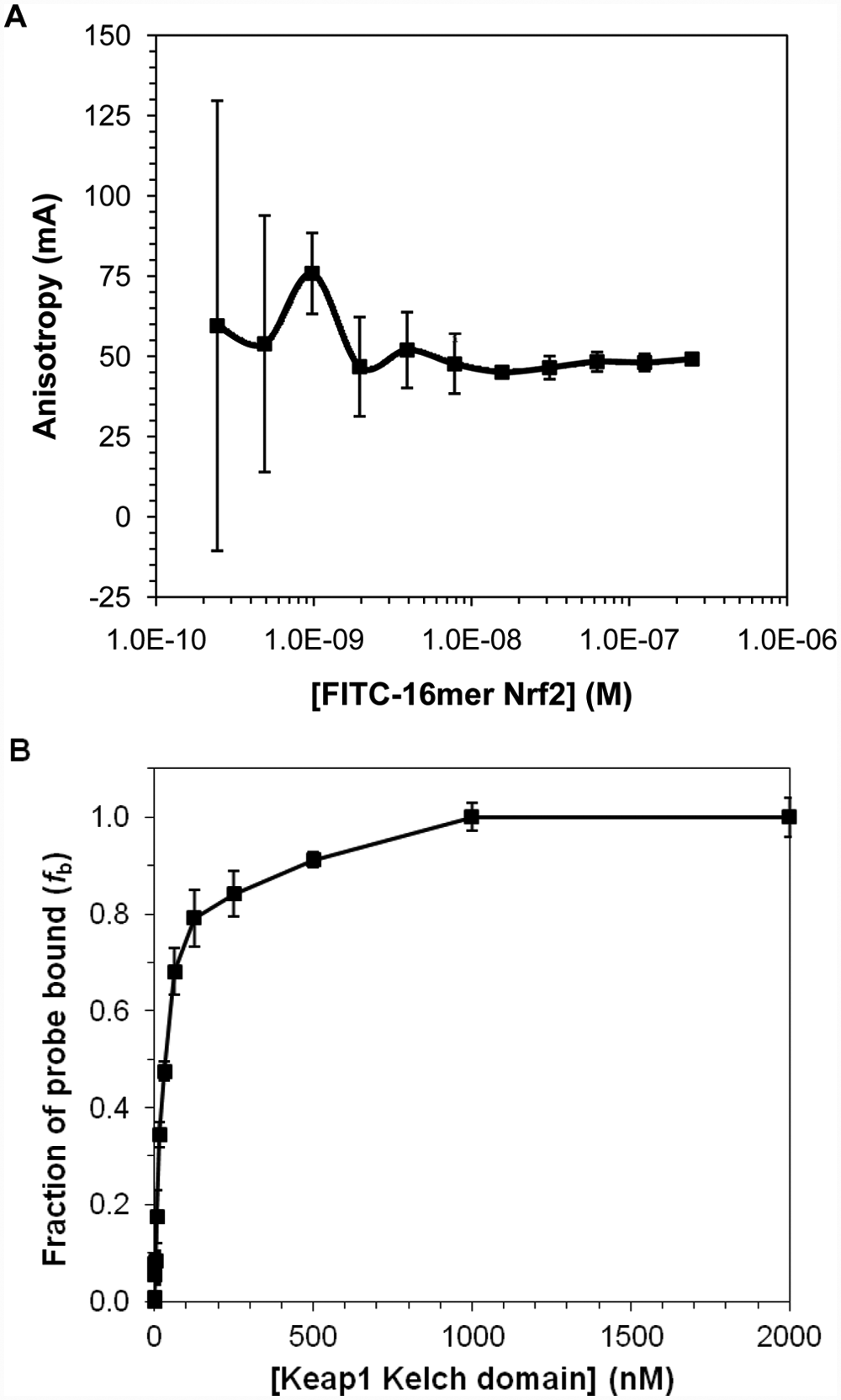
(A) The anisotropy of FITC-labeled 16mer Nrf2 peptide as a function of peptide probe concentration and (B) the plot of fraction bound of FITC-16mer Nrf2 peptide to Keap1 Kelch domain against Keap1 concentration. For the fluorescence polarization assay shown in (B), a fixed concentration (10 nM) of FITC-16mer Nrf2 peptide was incubated with a range of concentrations of Keap1 Kelch domain for 30 min at room temperature. The assay plate was centrifuged at 370
The equilibration time required for the assay was determined by following the increase in anisotropy signal every 10 min over a 3 h incubation period after mixing the human Keap1 Kelch domain protein with the fluorescent probe. The time it took for the anisotropy signal to plateau was used as the binding period for all of the assays. The equilibration was complete within 30 min at room temperature, which was selected as the incubation time for our FP assay.
Based on the FP binding assay shown in
Figure 1B
, the FITC-labeled 16mer Nrf2 peptide has a
Optimization of Fluorescently Labeled Nrf2 Peptide Probes for the FP Assay
The X-ray crystal structure of 16mer Nrf2 peptide-Keap1 Kelch domain complex has shown that the 16mer Nrf2 peptide is much longer than the DxETGE motif that is needed for binding to the Keap1 Kelch domain.
26
Several
As shown in
Table 1
and
Figure 2A
, the binding of the FITC-labeled 8mer Nrf2 peptide (FITC-DEETGEFL-OH) to the Keap1 Kelch domain was too weak for regression analysis (estimated
The Binding Affinities and Dynamic Range for FITC-Labeled Nrf2 Peptides a
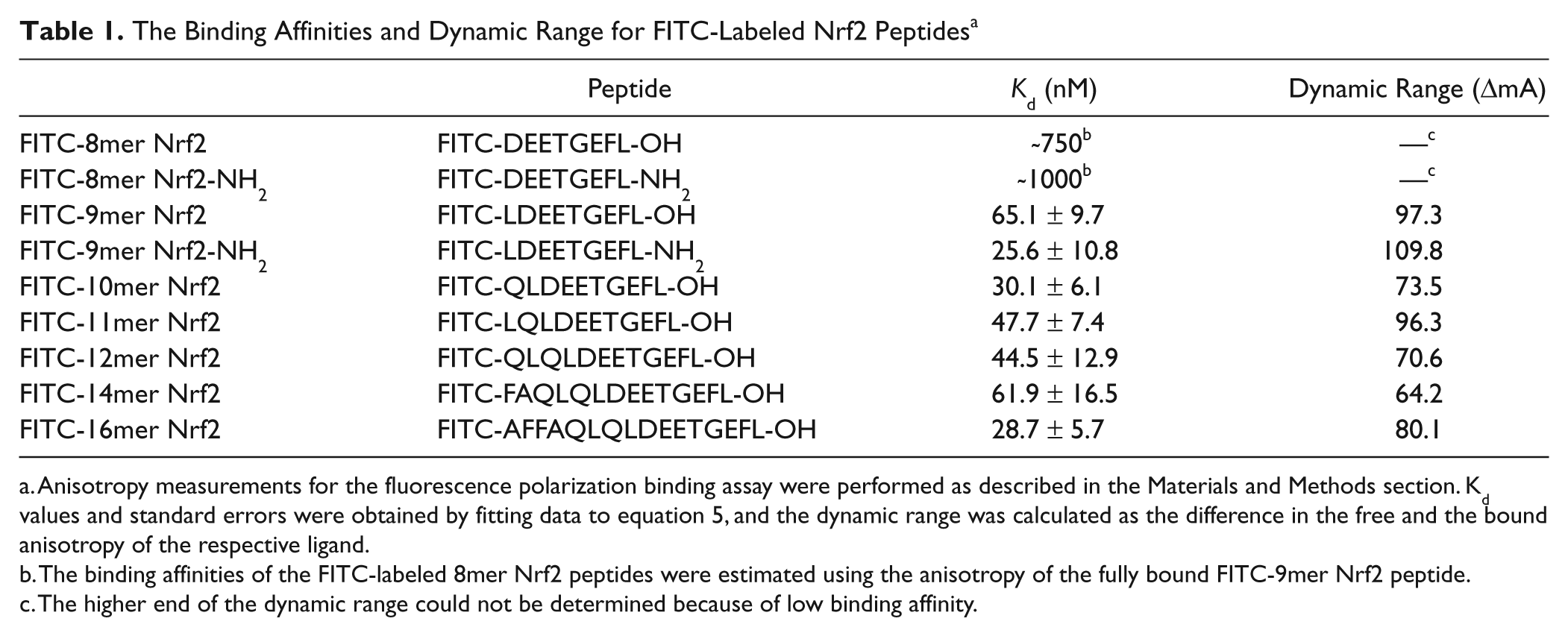
Anisotropy measurements for the fluorescence polarization binding assay were performed as described in the Materials and Methods section. Kd values and standard errors were obtained by fitting data to equation 5, and the dynamic range was calculated as the difference in the free and the bound anisotropy of the respective ligand.
The binding affinities of the FITC-labeled 8mer Nrf2 peptides were estimated using the anisotropy of the fully bound FITC-9mer Nrf2 peptide.
The higher end of the dynamic range could not be determined because of low binding affinity.
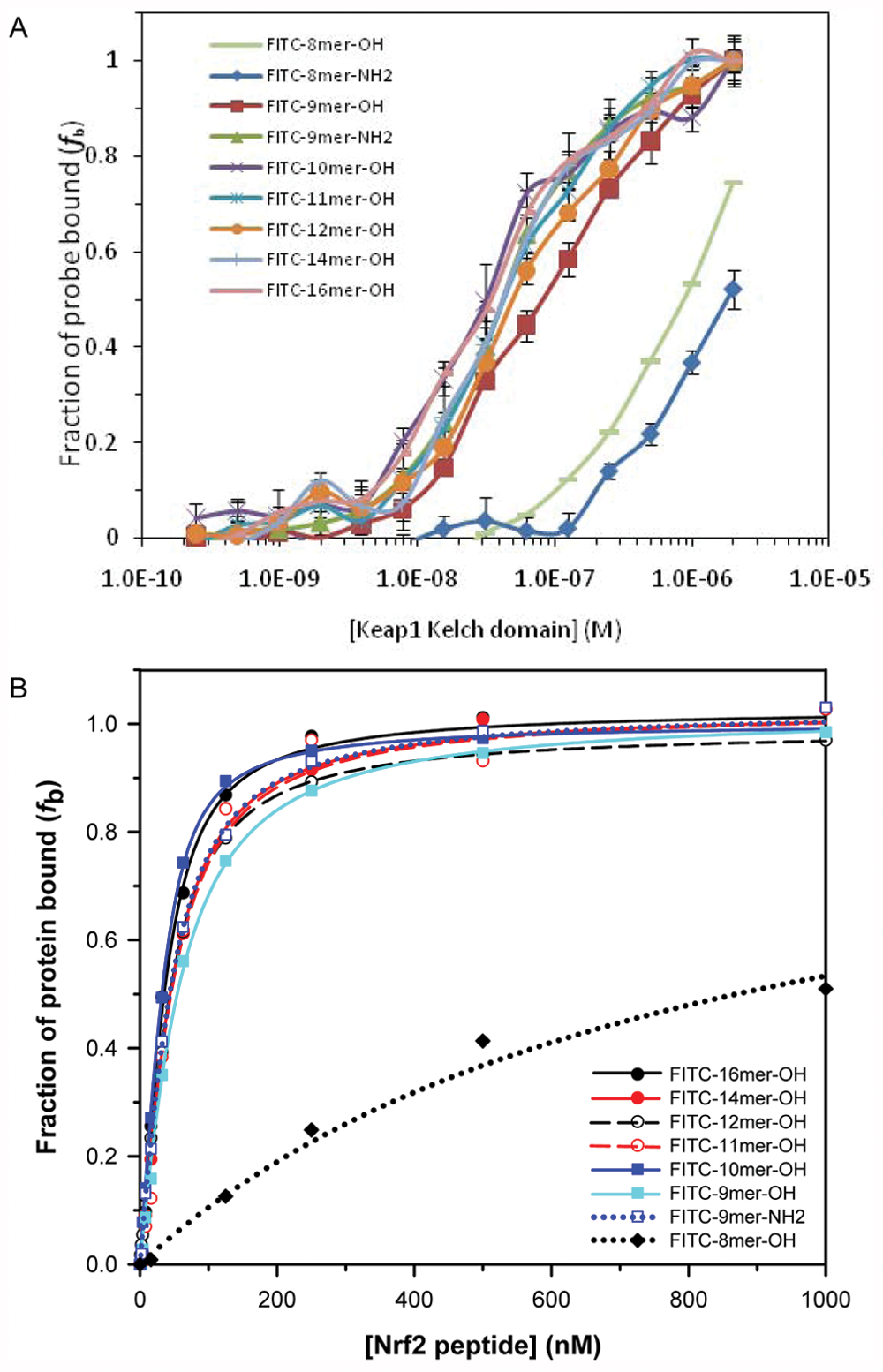
(A) The fraction bound of various FITC-labeled Nrf2 peptides to the Keap1 Kelch domain against the Keap1 Kelch domain concentration based on fluorescence polarization. Each of the fluorescent probes (10 nM) was incubated with varying concentrations of Keap1 Kelch domain (0.2 nM to 2 µM) for 30 min at room temperature before anisotropy measurement. The anisotropy was adjusted with the G-value determined from the literature value of free fluorescein, and the fraction bound (
We have demonstrated in our recent study of the binding of Nrf2 peptides to the Keap1 Kelch domain using an SPR solution competition assay that the binding affinity of the 9mer Nrf2 peptide to the Keap1 Kelch domain can be increased by the
To test the effect of
As shown in
Table 1
, the dynamic range of the FITC-9mer Nrf2 peptide amide (FITC-LDEETGEFL-NH2) was the highest among FITC-labeled Nrf2 peptides tested with 107.8 mA increase in anisotropy upon binding to the Keap1 Kelch domain. The FITC-14-mer Nrf2 peptide (FITC-FAQLQLDEETGEFL-OH) was shown to have the smallest dynamic range of 64.2 mA increase in anisotropy upon binding with a
The relatively high dynamic range of the FITC-11mer Nrf2 peptide (ΔmA = 96.3 mA) as compared with some of the shorter peptides was unexpected because of the additional amino acids outside of the DxETGE sequence. The dynamic range for the longer peptides was expected to be lower because the longer peptides contain several residues that extend out of the binding site of the DxETGE motif of Nrf2 and are exposed to solvent. Therefore, the expected decrease in dynamic ranges of the assay for the longer peptides would have been due to the propeller effect, in which the fluorophore extends out of the binding pocket and the flexibility of the fluorophore depolarizes the bound probe. The data shown in
Table 1
do indicate an overall trend that the dynamic range decreases with increasing length of the peptide. From these studies, it was clear that the FITC-9mer Nrf2 peptide amide has the strongest binding affinity to the Keap1 Kelch domain, with a
SPR Binding Assay of FITC-Labeled Nrf2 Peptides to the Keap1 Kelch Domain
The binding affinities of FITC-labeled Nrf2 peptides were confirmed by an SPR binding assay. As shown in Figure 2B and Table 2 , the general trend for the binding affinities of the Nrf2 peptides was similar to the results obtained using our FP assay. Although the binding affinities of the 10mer and 12mer Nrf2 peptides were better than expected, the general trend shows that the binding affinity increases with increasing peptide length. The SPR data also support that most of the binding interaction between the Keap1 Kelch domain and FITC-Nrf2 peptides is within FITC-9mer Nrf2 peptide, as the binding affinities of 9mer to 16mer Nrf2 peptides to the Keap1 Kelch domain are quite similar. Consistent with the FP assay, the 8mer Nrf2 peptide binds weakest to the Keap1 Kelch domain, and the binding affinity is more than 20-fold weaker than that of the FITC-9mer Nrf2 peptide.
The Binding Affinities of FITC-Labeled Nrf2 Peptides Determined by Surface Plasmon Resonance a
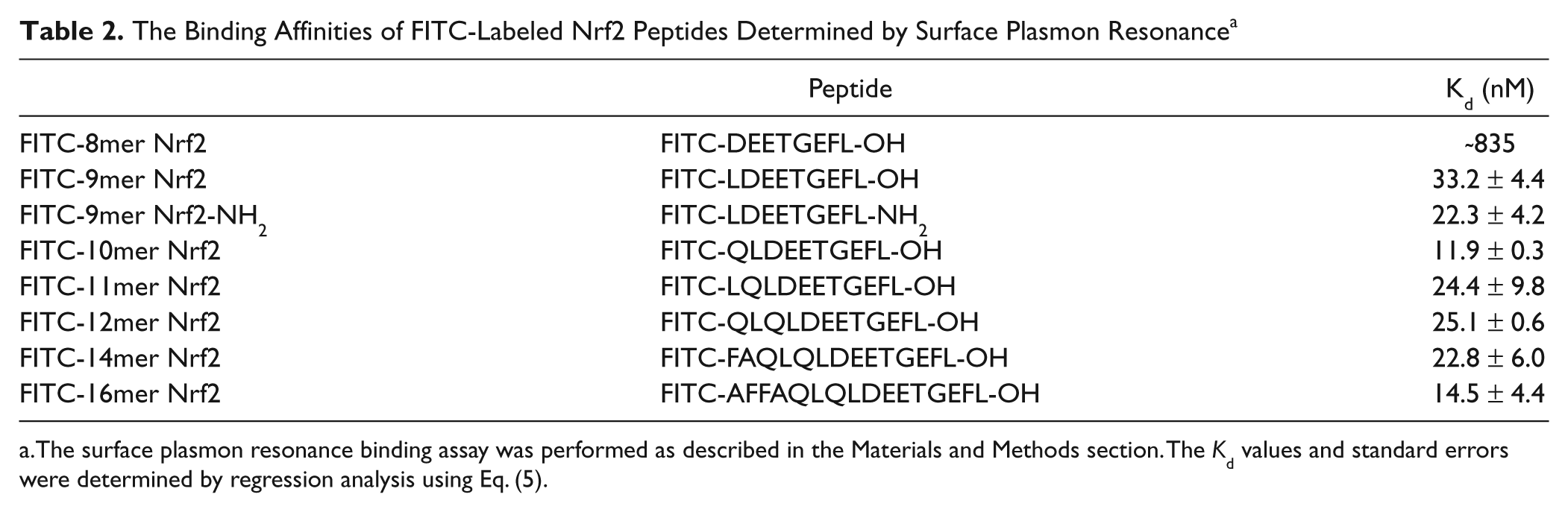
The surface plasmon resonance binding assay was performed as described in the Materials and Methods section. The
FP-Based Competition Binding Assay
Our FP assay using FITC-9mer Nrf2 peptide amide as the probe was used to differentiate a series of Nrf2 peptide as inhibitors of the Keap1-Nrf2 interaction in a competition binding assay. The concentration of the Keap1 Kelch domain was fixed at 100 nM to give an
The Inhibition of the Keap1 Kelch Domain-Nrf2 Interaction by Different Nrf2-Peptides as Determined Using the FP-Based Competition Assay Developed a
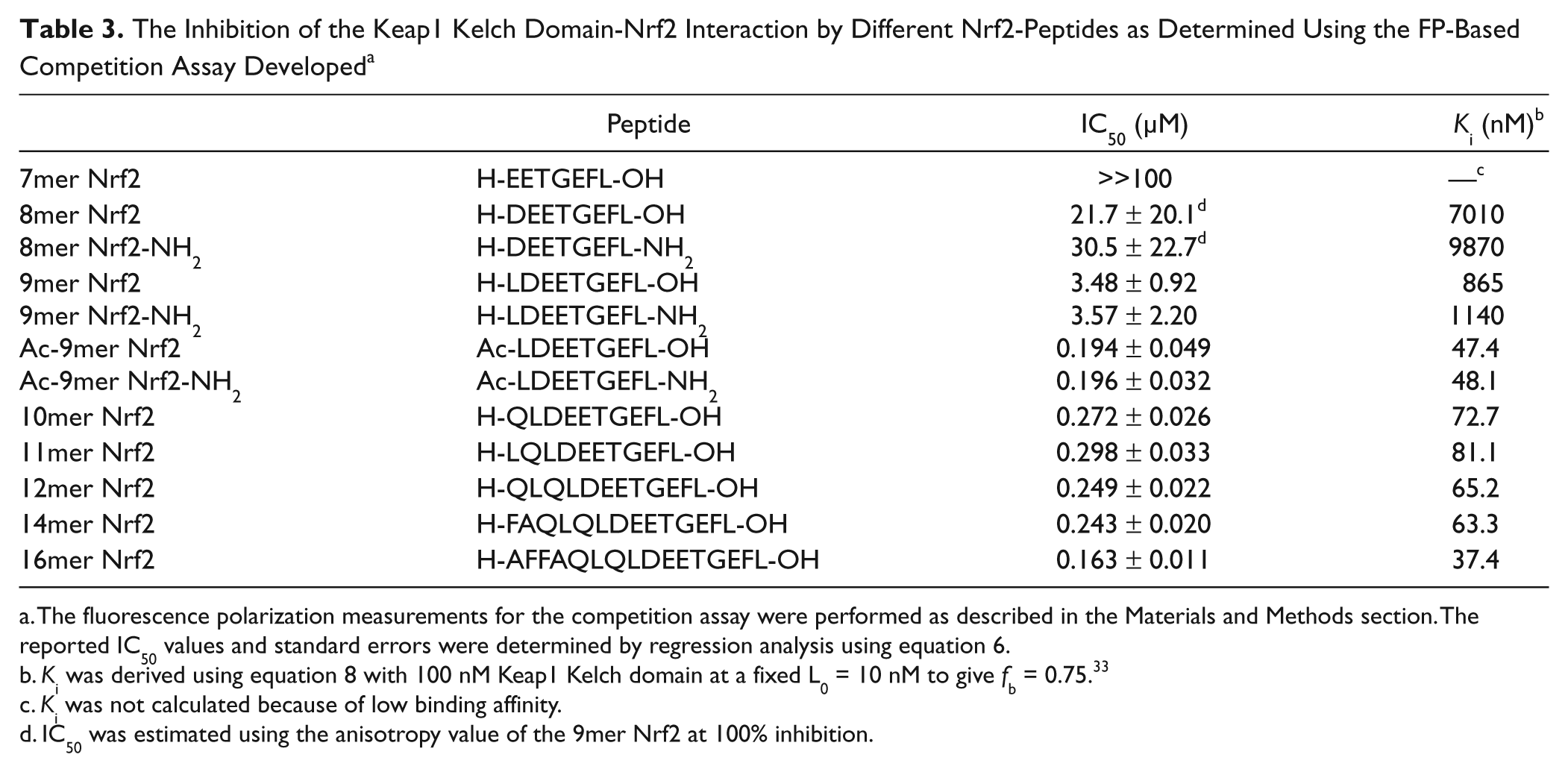
The fluorescence polarization measurements for the competition assay were performed as described in the Materials and Methods section. The reported IC50 values and standard errors were determined by regression analysis using equation 6.
IC50 was estimated using the anisotropy value of the 9mer Nrf2 at 100% inhibition.
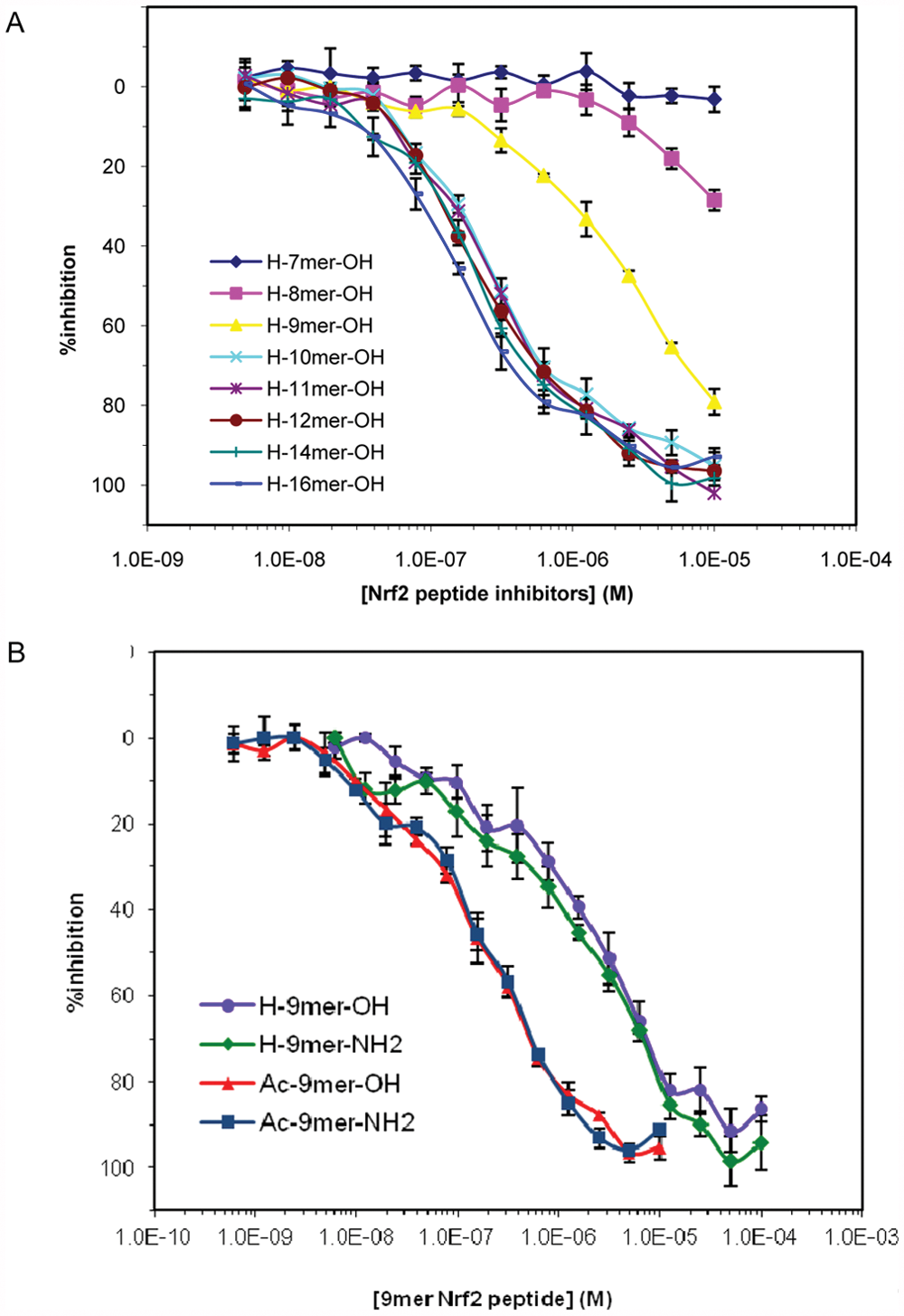
The inhibition of binding of FITC-9mer Nrf2 amide to the Keap1 Kelch domain by (A) Nrf2 peptides and (B) a series of 9mer Nrf2 peptides as determined in the fluorescence polarization–based competition assay. The assay was performed at room temperature using 10 nM FITC-9mer Nrf2 peptide amide as the probe and 100 nM Keap1 Kelch domain in the presence of varying concentrations of the inhibitory Nrf2 peptides. Error bars represent the standard deviations of triplicate measurements.
Our FP data also indicate that the binding affinity of the 16mer Nrf2 peptide is stronger than the shorter Nrf2 peptide and is in good agreement with the previously reported value.26,27 The binding of the 16mer Nrf2 peptide (AFFAQLQLDEETGEFL) was the strongest in our assay, with an IC50 of 163 nM and a calculated Ki of 37.4 nM. The resolution of inhibitor potencies by an FP assay under nonstoichiometric conditions may be limited by the
The inhibitory activities of the
Comparison of Fluorophores for the FP Assay
To select the best fluorophore for our FP assay, the 9mer Nrf2 peptide amide was also labeled with BODIPY-FL, Cy3B, and Cy5. The BODIPY-FL dye is a known replacement for fluorescein with a higher extinction coefficient (>80 000 M−1cm−1), high quantum yield (1.0 in water), insensitivity to environment such as pH,
37
and sharp excitation and emission bands. The cyanine dyes, Cy3B and Cy5, are also known for their excellent fluorescence properties and high water solubility, which may improve the solubility of the labeled compound in aqueous assay buffers.
38
The Cy3B dye is a “brighter” derivative of Cy3 and could be better suited for FP application due to its increased quantum yield and fluorescence lifetime. The ranking of the fluorophores for FP assay application was determined by determining the
As shown in Table 4 and Figure 4 , BODIPY-labeled 9mer Nrf2 peptide amide gave the lowest dynamic range of the four fluorescently-labeled peptides with an assay window of 66.5 mA. The FITC-labeled 9mer Nrf2 amide gave the third largest assay window, with an anisotropy change of 109.8 mA, whereas the Cy3B and Cy5-labeled peptides gave assay windows of 167.7 and 143.2 mA, respectively. Interestingly, the cyanine-labeled peptides had slightly higher binding affinity to the Keap1 Kelch domain, whereas the BODIPY-9mer Nrf2 peptide amide showed similar binding affinity to the FITC-labeled peptide.
The Dynamic Ranges, Assay Quality, and Binding Affinities of the 9mer Nrf2 Peptide Amide Probes with Different Fluorophores under Fluorescence Polarization Assay Conditions Developed a
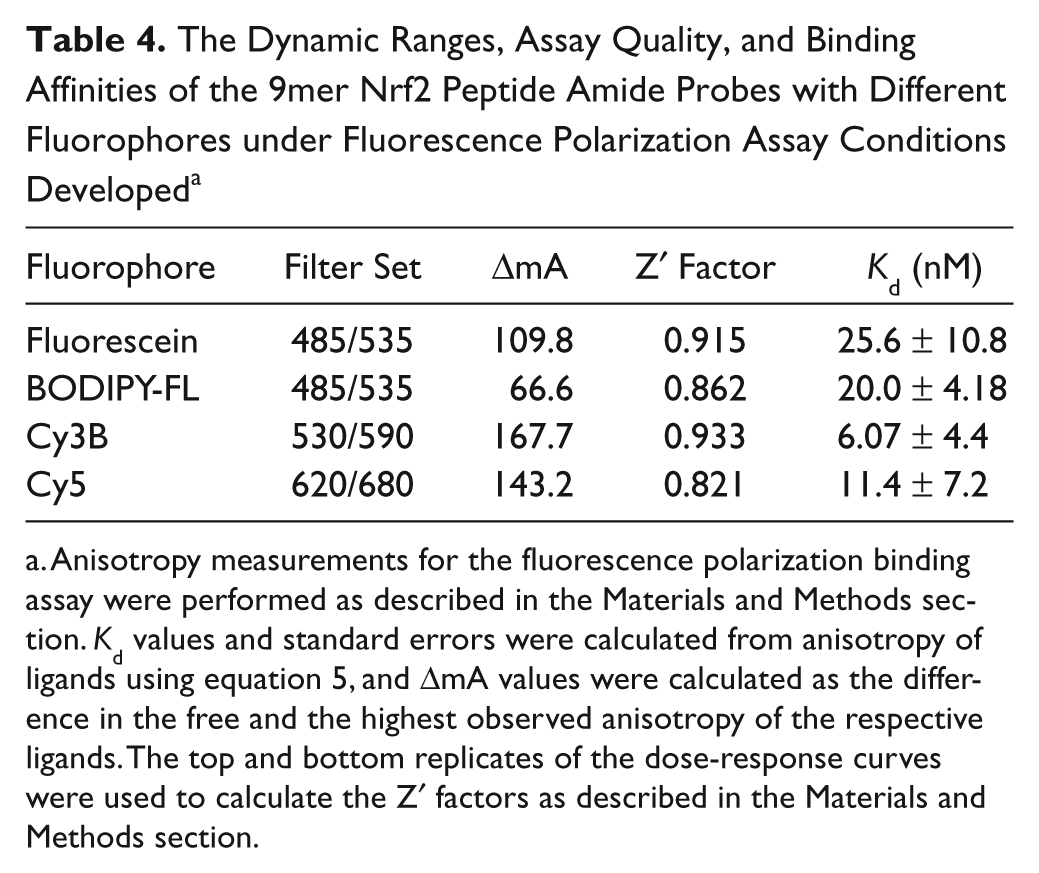
Anisotropy measurements for the fluorescence polarization binding assay were performed as described in the Materials and Methods section.
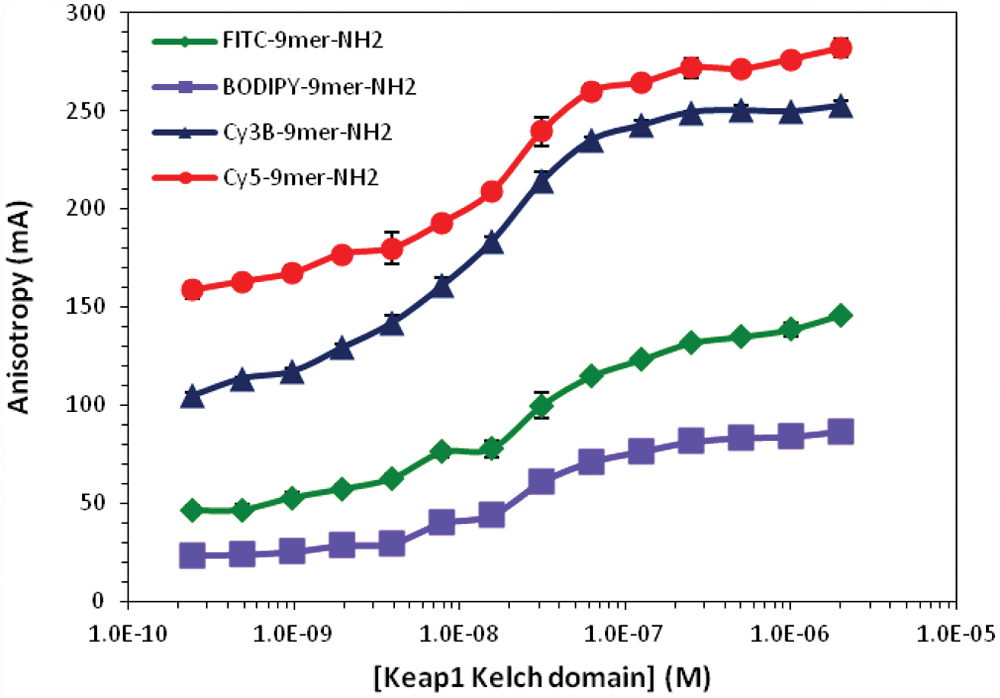
The anisotropy of four 9mer Nrf2 peptide probes plotted as a function of Keap1 Kelch domain concentrations. Each fluorescent probe at a fixed concentration (10 nM) was incubated with varying concentrations of Keap1 Kelch domain (0.24 nM to 2 µM) for 30 min at room temperature. The anisotropy values of each fluorophore were not adjusted by the G-value.
To compare the performance of fluorophores for our assay, the Z′ factor was calculated for each probe. 39 The Z′ factor is a measure of assay performance that incorporates the assay window and the random error present in the FP measurement. According to the Z′ factor model, values greater than 0.5 indicate good assays that may be readily adapted to high-throughput applications. As shown in Table 4 , all of the fluorophores perform well under our assay conditions with the Z′ factors greater than 0.8. Considering the cost of Cy5 and Cy3B label reagents, FITC-labeled peptide is still a viable option for HTS application, as evidenced by the relatively high Z′ factor.
Effect of DMSO on the FP Assay
Compounds in chemical libraries used in HTS are often dissolved in DMSO. Typically, the DMSO concentration for biochemical HTS assays range from 0.5% to 5%, and for FP-based assays, increasing the viscosity of the solution is not desirable, as the polarization of the fluorescent molecules may increase due to slower rotation in viscous solution. The increase in concentration of DMSO does not significantly affect the anisotropy of the bound state of the ligand, whereas the anisotropy of the free ligand slowly increases with increasing concentration of DMSO (
Pilot Screen of NCI Diversity Set II and Clinical Collection Using the FP Assay
We applied the FP assay in a 384-well format in a pilot screen of the NCI Diversity Set II of 1364 compounds and the NIH Clinical Collection (NCC) of 446 compounds. Initial screening of the 1810 compounds in duplicate at 100 µM using %inhibition >20% (mean + 3SD) for hit calling identified 31 primary hits (hit rate 1.7%) after excluding fluorescent compounds based on total fluorescence. The Z′ factors for this pilot screen (26 plates) averaged 0.60 ± 0.05; even though the retest of the 31 primary hits at 10 µM in duplicates did not result in any confirmed hits from this small library of 1810 compounds, this pilot screen did demonstrate that the FP assay is robust and suitable for HTS.
In summary, we optimized the fluorescent probe based on the DxETGE motif of Nrf2 and successfully developed a Keap1 binding FP assay. Our design of the fluorescently labeled probe is based on the 16mer Nrf2 peptide reported, and the probe for the assay was selected based on the binding affinity (
The competition binding assay using FITC-9mer Nrf2 amide as the probe was used to rank the binding affinities of 7mer to 16mer Nrf2 peptide inhibitors. The binding affinity of the 16mer Nrf2 peptide (H-AFFAQLQLDEETGEFL-OH,
Footnotes
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors disclose receipt of the following financial support for the research and/or authorship of this article: This work was supported by grants CA133791, CA125868, and MH093197 from the National Institutes of Health.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
