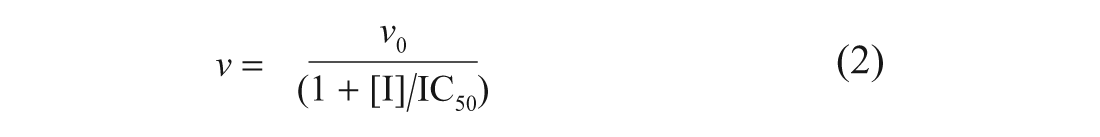Abstract
Histone deacetylase (HDAC) enzymes modify the acetylation state of histones and other important proteins. Aberrant HDAC enzyme function has been implicated in many diseases, and the discovery and development of drugs targeting these enzymes is becoming increasingly important. In this article, the authors report the evaluation of homogeneous, single-addition, bioluminogenic HDAC enzyme activity assays that offer less assay interference by compounds in comparison to fluorescence-based formats. The authors assessed the key operational assay properties including sensitivity, scalability, reproducibility, signal stability, robustness (Z′), DMSO tolerance, and pharmacological response to standard inhibitors against HDAC-1, HDAC-3/NcoR2, HDAC-6, and SIRT-1 enzymes. These assays were successfully miniaturized to a 10 µL assay volume, and their suitability for high-throughput screening was tested in validation experiments using 640 drugs approved by the Food and Drug Administration and the Hypha Discovery MycoDiverse natural products library, which is a collection of 10 049 extracts and fractions from fermentations of higher fungi and contains compounds that are of low molecular weight and wide chemical diversity. Both of these screening campaigns confirmed that the bioluminogenic assay was high-throughput screening compatible and yielded acceptable performance in confirmation, counter, and compound/extract and fraction concentration-response assays.
Introduction
T
The HDAC enzymes are composed of 18 family members classified in four classes depending on sequence identity and domain organization. 6 The 11 so-called classical HDAC enzymes of class I, II, and IV are Zn2+ dependent. 7 The remaining seven class III HDAC enzymes are referred to as sirtuin (SIRT) enzymes and require NAD+ as an essential cofactor. 8 The first-generation pan HDAC enzyme targeted drug approved by the Food and Drug Administration (FDA) was vorinostat, a hydroxamic acid derivative that exhibits broad activity against HDAC class I and II enzymes for the treatment of cutaneous T-cell lymphoma. 9 Subsequent to this, the natural product romidepsin, which targets the Zn2+-dependent HDAC enzymes, was approved by the FDA also for the treatment of cutaneous T-cell lymphoma. 10 Both of these drugs have significant side effect profiles, and thus, a key goal for hit-finding studies, particularly in non-oncology indications, is to identify inhibitors from synthetic small-molecule libraries as well as natural products libraries that are both potent and selective for individual HDAC enzymes and thereby offer the potential to overcome the side effects of the current generation of HDAC-targeted drugs.11–13 This requirement necessitates assays that are both cost-effective and adaptable to high-throughput screening (HTS).
The initial approach for monitoring HDAC enzyme activity required the use of radiolabeled histones as the substrate. 14 However, more recently, HDAC enzyme assays utilized in drug discovery typically rely on artificial substrates, for example, a peptide containing a ε-acetylated lysine that is C-terminally coupled to 4-methyl-coumarin-7-amide. The product of the HDAC enzyme–mediated turnover of this substrate can be monitored after a development step, in a fluorescence intensity format. 15 This type of fluorogenic assay has drawbacks, which if not fully controlled for in counter and confirmation studies, can lead to confounding results, which risk diverting resources from the development of real hits to the pursuit of artifacts.16,17
In this report, we evaluate homogeneous, single-addition, bioluminogenic assays that can be used to monitor the activity of a range of HDAC class I/II and SIRT enzymes. See Figure 1 for a schematic representation of the HDAC-Glo and SIRT-Glo assays. In the case of the HDAC-Glo I/II assay, the proluminogenic substrate contains an acetylated lysine peptide sequence derived from Histone 4 conjugated to aminoluciferin. HDAC enzyme–mediated deacetylation of the lysine residue facilitates luminogenic substrate susceptibility to specific proteolytic cleavage by the enzyme in the developer reagent. The aminoluciferin product of that cleavage can then act as a substrate for luciferase, and the amount of light produced in this reaction is proportional to HDAC enzyme activity. The analogous SIRT-Glo assay makes use of a proluminogenic substrate that has been optimized for class III enzymes. We have investigated the HDAC-Glo Class I/II and SIRT-Glo assays using HDAC-1, HDAC-3/NcoR2, HDAC-6, and SIRT-1 enzymes, and the studies undertaken included determining their sensitivity, scalability, reproducibility, signal stability, robustness (Z′), DMSO tolerance, and pharmacological response to standard inhibitors. Finally, their performance in an automated HTS setting was assessed using 640 FDA-approved drugs and the Hypha Discovery MycoDiverse natural products library, which is a collection of 10 049 extracts and fractions with unusually high chemical novelty. Both of these screening campaigns confirmed that the HDAC-Glo I/II and SIRT-Glo bioluminogenic assays were HTS compatible and yielded acceptable performance in confirmation, counter, and compound/extract and fraction concentration-response assays.
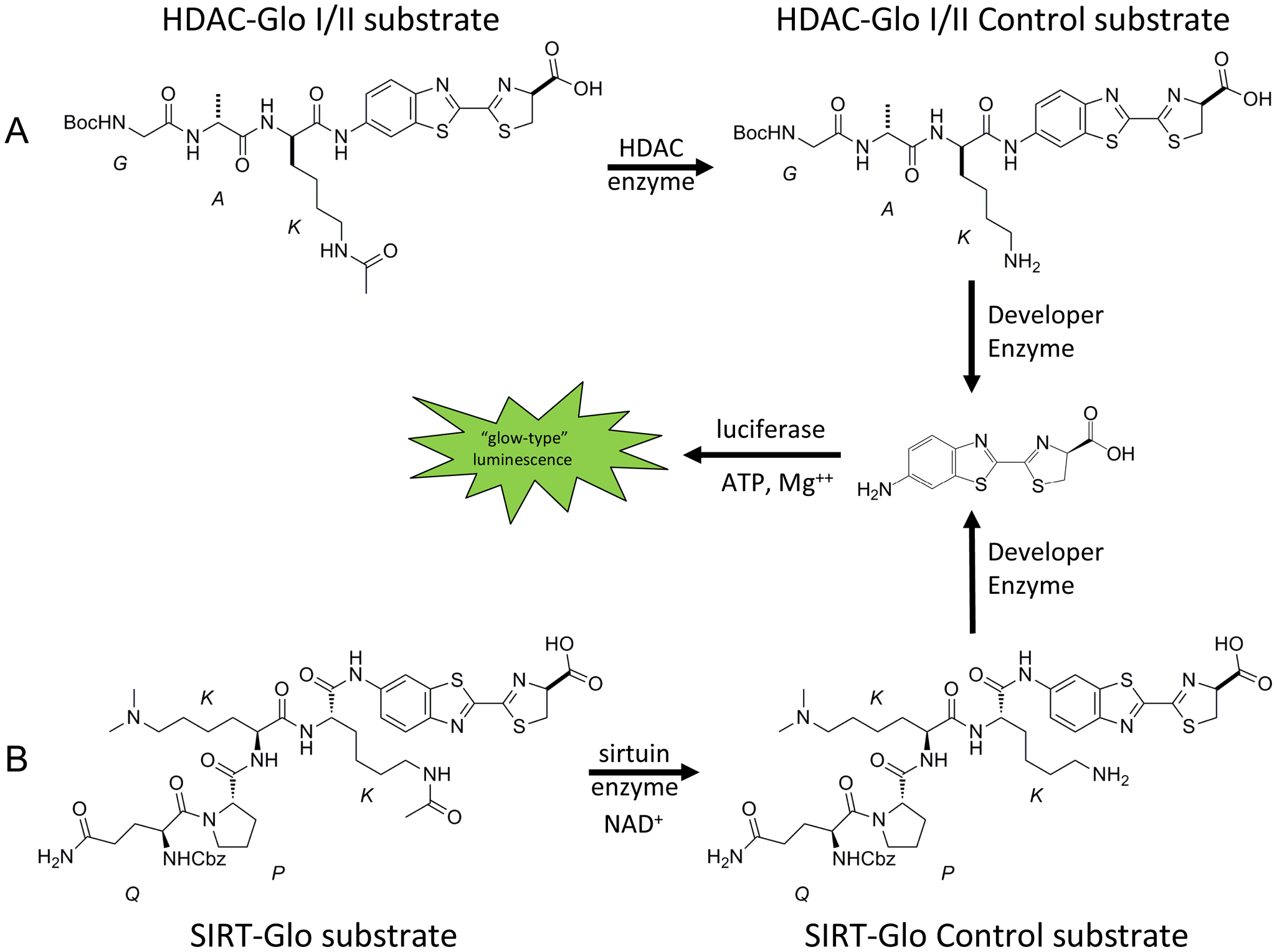
Schematic representation of (
Materials and Methods
Reagents
Human recombinant C-ter-His-FLAG-HDAC1 (50051), HDAC-3/NcoR2 (50003), N-ter-GST-HDAC-6 (50006), and N-ter-GST-SIRT-1 (50012) were purchased from BPS Bioscience (San Diego, CA). HDAC-Glo Class I/II Kits and SIRT-Glo Kits were provided by Promega Corporation (Madison, WI). Porcine trypsin (T-0303), Tergitol NP-9 (NP9), Triton X-100 (X-100), NAD+ (N-1511), and the standard inhibitors valproic acid (P4543), trichostatin A (T8552), sodium butyrate (B5887), and suramin (S2671) were purchased from Sigma-Aldrich (St. Louis, MO). The standard inhibitors were dissolved to yield stock solutions in 100% v/v DMSO purchased from Roth (Karlsruhe, Germany) and stored at −20 °C. The 640 FDA-approved drugs (BML-2841-0100) were purchased from Enzo Life Sciences (Lörrach, Germany) and supplied at a concentration of 2 mg/mL in 100% v/v DMSO. The Hypha Discovery MycoDiverse natural products library was made available by Hypha Discovery as part of a collaboration and is composed of biomass extracts and extracellular fractions from fermentations of higher fungi (mushrooms and toadstools) and contains compounds that are of low molecular weight with wide chemical diversity. Each extract and fraction was dissolved in 100% v/v DMSO. Plate handling was performed using a Cell Explorer HTS platform (Perkin Elmer, Waltham, MA) equipped with an Echo 550 (Labcyte, Sunnyvale, CA) and Multidrop (Thermo, Waltham, MA) liquid-handling systems. Luminescence measurements were made using an EnVision Multilabel Reader (Perkin Elmer) or PolarStar (BMG LABTECH, Ortenberg, Germany). Assays were performed in white 96-well polystyrene, TC-treated, clear, flat-bottom, microtiter plates (3903) purchased from Corning (Lowell, MA) or white 384-well polystyrene LUMITRAC 200 (medium binding), flat-bottom microtiter plates (781075) purchased from Greiner Bio-One GmbH (Frickenhausen, Germany).
Development of HDAC-Glo I/II and SIRT-Glo kits
Luminogenic Substrates
Substrates
The two luminogenic substrates Boc-GAK(Ac)-aminoluciferin (HDAC-Glo I/II substrate) and Z-QPK(Me2)K(Ac)-aminoluciferin (SIRT-Glo substrate) were synthesized in a similar manner to previously described peptidylaminoluciferins. 18 The nonacetylated control substrates Boc- GAK-aminoluciferin (HDAC-Glo I/II control substrate) and Z-QPK(Me2)K-aminoluciferin (SIRT-Glo control substrate) were synthesized in a similar manner as previously described 18 but using orthogonal protecting groups on the lysine rather than an acetyl group, followed by deprotection as the final synthetic step.
HDAC-Glo I/II and SIRT-Glo assay buffers
Assay buffers contained 25 mM Tris buffer, pH 8.0, supplemented with 137 mM NaCl, 2.7 mM KCl, and 1 mM MgCl2. The HDAC-Glo assay buffer also contained 1% v/v Triton X-100, and the SIRT-Glo assay buffer also contained 1% v/v Tergitol NP-9.
The HDAC-Glo I/II and SIRT-Glo assay protocols
The HDAC-Glo I/II assay reagent was prepared by (1) rehydration of lyophilized HDAC-Glo I/II substrate (with an acetylated peptide concentration of 100 µM) in 10 mL HDAC-Glo I/II assay buffer and (2) addition of 10 µL of developer reagent (containing trypsin). The SIRT-Glo assay reagent was prepared similarly but with a final acetylated peptide concentration of 200 µM. The HDAC enzymes diluted as appropriate using the HDAC-Glo I/II assay buffer were dispensed into microtiter plates, followed by an equal volume of HDAC-Glo I/II assay reagent to initiate the reaction. The microtiter plates were mixed briefly by orbital shaking (500–700 rpm) and luminescence measured at steady-state signal:background, which was achieved after 20 min using an Envision or PolarStar. The SIRT-Glo assay protocol was the same as described above except that the SIRT-Glo substrate was used instead with the inclusion of β-nicotinamide adenosine dinucleotide (NAD+) at 5 mM in the assay. During adaptation of the assay to the Cell Explorer HTS platform, it was shown that for each enzyme, no difference in the luminescence signal was obtained when using a low control composed of either no enzyme or an inhibitor at concentrations >>IC50.
Kinetic characterization of HDAC and SIRT enzymes
Enzyme titrations and Km determinations
Enzyme titrations were performed following the HDAC-Glo I/II and SIRT-Glo assay protocol as described above with appropriately diluted HDAC and SIRT enzymes (between 0.1 nM and 10 µM) and steady-state read time points. The Km values for the HDAC and SIRT enzymes for their respective substrates were determined in substrate titration experiments by following the HDAC-Glo I/II and SIRT-Glo assay protocol as described above but by spiking in additional acetylated peptide to a final concentration of 400 µM, then performing twofold serial dilutions of the HDAC-Glo I/II and SIRT-Glo Assay Reagents in acetylated peptide-free luciferin detection reagent (Promega, V859A) rehydrated with assay buffer and developer reagent. The data from these duplicate experiments were fitted using the Michaelis–Menten equation, equation 1, within GraphPad Prism (version 5.02, GraphPad Software, Inc, La Jolla, CA).
IC50 determinations for standard inhibitors
The IC50 values for the standard inhibitors of each enzyme were determined in order to validate the HDAC-Glo I/II and SIRT-Glo assay formats. As the standard inhibitors were dissolved in DMSO, it was necessary to determine the tolerance of the assays to this solvent. The DMSO tolerances of the HDAC-Glo I/II and SIRT-Glo assays were investigated by titration up to 10% v/v DMSO, which was substantially above its concentration proposed in any screening campaign. For standard inhibitor concentration-response experiments, the IC50 values were calculated by fitting the duplicate data to the four-parameter logistic equation, equation 2 using GraphPad Prism (version 5.02, GraphPad Software, Inc).
Adaptation of the HDAC-Glo I/II and SIRT-Glo assays for use in HTS campaigns
The parameters that were determined to ensure that the assays were HTS compatible included their scalability, reproducibility, signal stability, robustness (Z′), and pharmacological response to standard inhibitors. The concentration of enzyme used in these experiments was determined from the enzyme titration experiments in which linear luminescence signal was observed and there was <20% substrate turnover, and these were as follows: for HDAC-1, 0.1 nM; HDAC-3/NcoR2, 0.15 nM; HDAC-6, 0.1 nM; and SIRT-1, 2 nM.
Prior to initiating the screening campaigns, the assays were miniaturized to a volume of 10 µL, and no change in assay performance was observed. They yielded reproducible results in terms of signal stability and robustness (Z′). As the libraries to be screened were stored in 100% v/v DMSO, it was possible to obtain a suitable screening concentration of test compounds/extracts and fractions for the 640 FDA-approved drugs and Hypha Discovery MycoDiverse natural products library with 0.25% and 1% v/v DMSO, respectively. In all screening campaigns, the low control population was based on a no-enzyme control. The quality and robustness of the assay, represented as Z′, were calculated as previously described. 19
In all cases, test compounds/extracts and fractions (25 nL) were added to microtiter plates. Subsequent to this, the HDAC-Glo I/II and SIRT-Glo assay protocols described above were followed. Counter assays were used to determine whether the hits identified from the screening campaigns were inhibiting the luciferase and enzymes in the developer reagent. The counter assay was essentially identical to the HDAC-Glo I/II and SIRT-Glo assay protocols except that (1) the developer reagent was used at a 1:100-fold dilution and (2) the nonacetylated control substrates were used instead at concentrations that produced luminescence comparable to their respective deacetylase assays. All screening data were processed using Activity Base XE (IDBS, UK) and visualized using Vortex (Dotmatics, Herts, UK).
Results
HDAC and SIRT enzyme substrates
The key features of the HDAC and SIRT substrates were that they contained (1) an acetylated lysine residue that would undergo HDAC and SIRT enzyme-mediated deacetylation and (2) recognition features of each enzyme as reported in the literature.20,21 Upon HDAC or SIRT enzyme–mediated deacetylation of the lysine residue within their respective substrate, the product becomes susceptible to protease cleavage, resulting in the formation of aminoluciferin, which subsequently acts as a substrate for the luciferase enzyme, resulting in light production. The concentrations of the luciferase and trypsin enzyme in the developer reagent were at suitably high concentrations so that they were not rate limiting (data not shown). Sequence-matched, nonacetylated, luminogenic control substrates were developed for a counter assay to assess potential compound interference directed at luciferase or enzymes in the developer reagent. The concentrations of the control substrates used in the counter assays were such that they produced similar relative luminescence to that observed in their respective deacetylase assays. Any decrease in light observed in the counter assays is expected to be due to inhibition of the enzymes in the developer reagent or luciferase, indicating that the compound is a false-positive rather than a true inhibitor of the HDAC or SIRT enzymes.
Kinetic characterization of HDAC and SIRT enzymes
Enzyme titrations
The initial kinetic characterization of the HDAC and SIRT enzymes involved titration experiments that allowed an assessment of the sensitivities of the HDAC-Glo and SIRT-Glo assay reagents. These studies also allowed the determination of appropriate enzyme concentrations that should be used in their subsequent kinetic characterization while providing a suitable assay window for their use in HTS campaigns. Each enzyme was titrated at concentrations dictated by their stock concentrations, and the profiles for HDAC-6 and SIRT-1 are shown in
Figures 2A
and
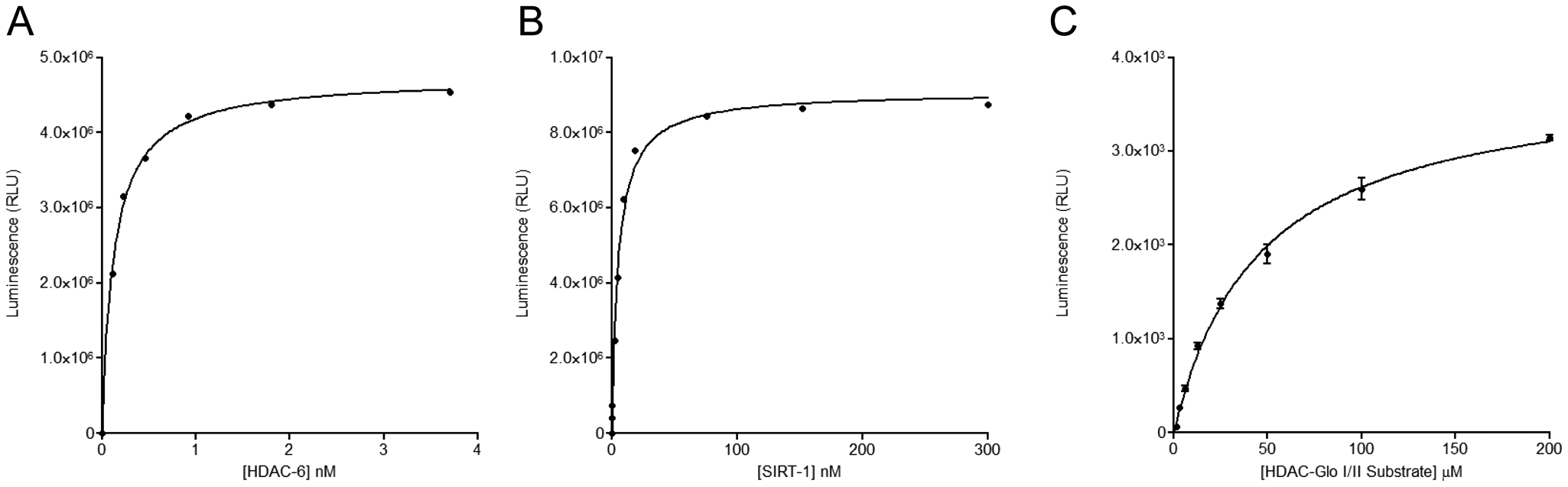
Titration of (
Substrate Km and IC50 determinations for standard inhibitors
The Km for the HDAC-Glo and SIRT-Glo substrates for each enzyme were also determined in this study. The Km determinations were made by titrating the substrates with an appropriate fixed concentration of each enzyme when <20% substrate turnover took place and fitting the data using the Michaelis–Menten equation, equation 1 within GraphPad Prism (version 5.02, GraphPad Software, Inc.). The Km value determined for HDAC-1 was 45 ± 3 µM (see Fig. 2C ). The Km values determined for the other enzymes were as follows: for HDAC-3/NcoR2, 192 ± 15 µM; for HDAC-6, 31 ± 3 µM; and for SIRT-1, 183 ± 16 µM (data not shown). All Km values were comparable to values reported in the literature. 21
The tolerance of the HDAC-Glo I/II and SIRT-Glo assays toward DMSO up to 10% v/v was investigated. The HDAC-6 enzyme was able to tolerate up to 1% v/v DMSO with negligible loss in enzyme activity (see
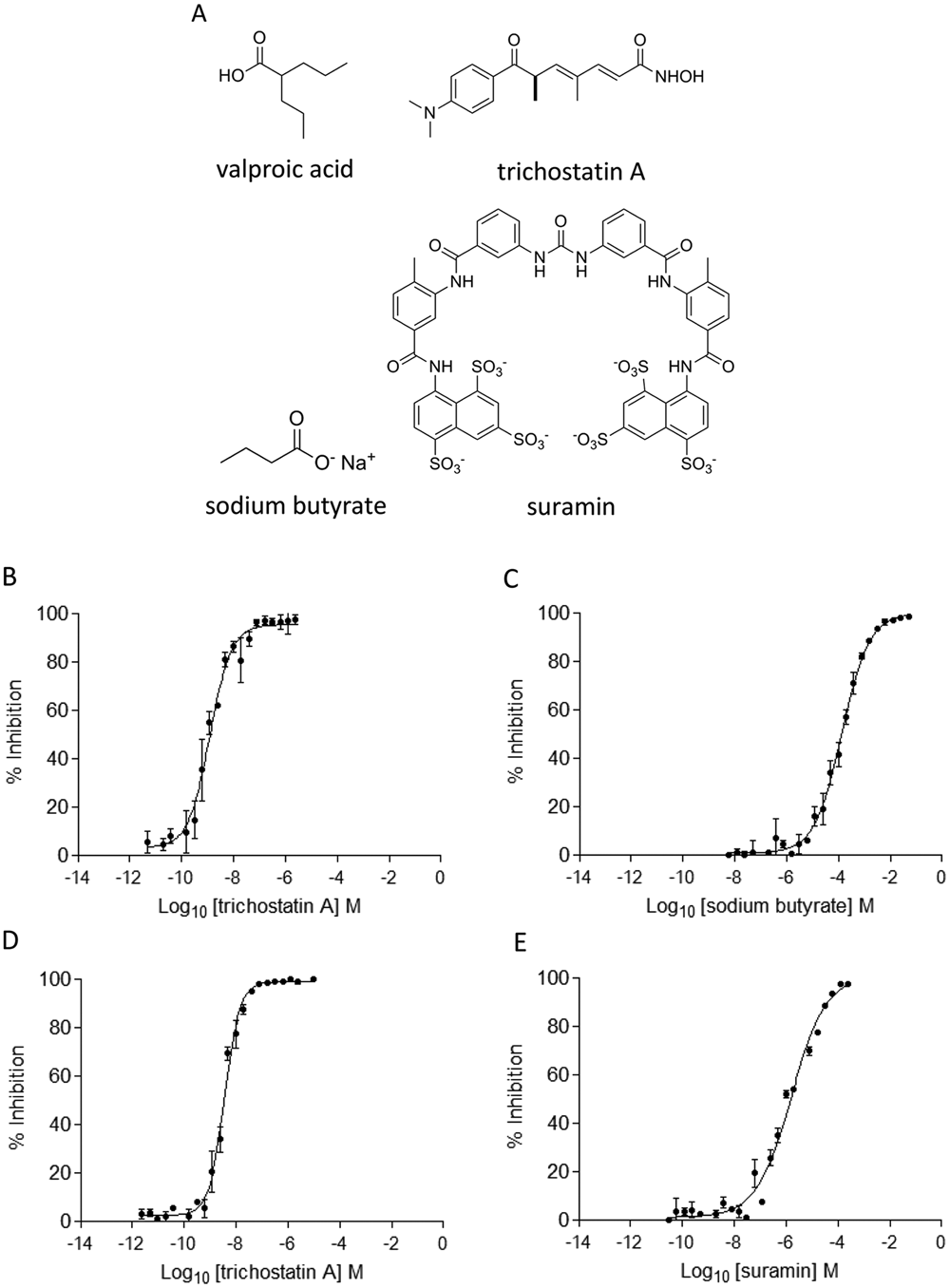
(
Summary of the Results from the Evaluation of Standard Inhibitors (Valproic Acid, Trichostatin A, Sodium Butyrate, and Suramin) against HDAC-1, HDAC-3/NcoR2, HDAC-6, and SIRT-1
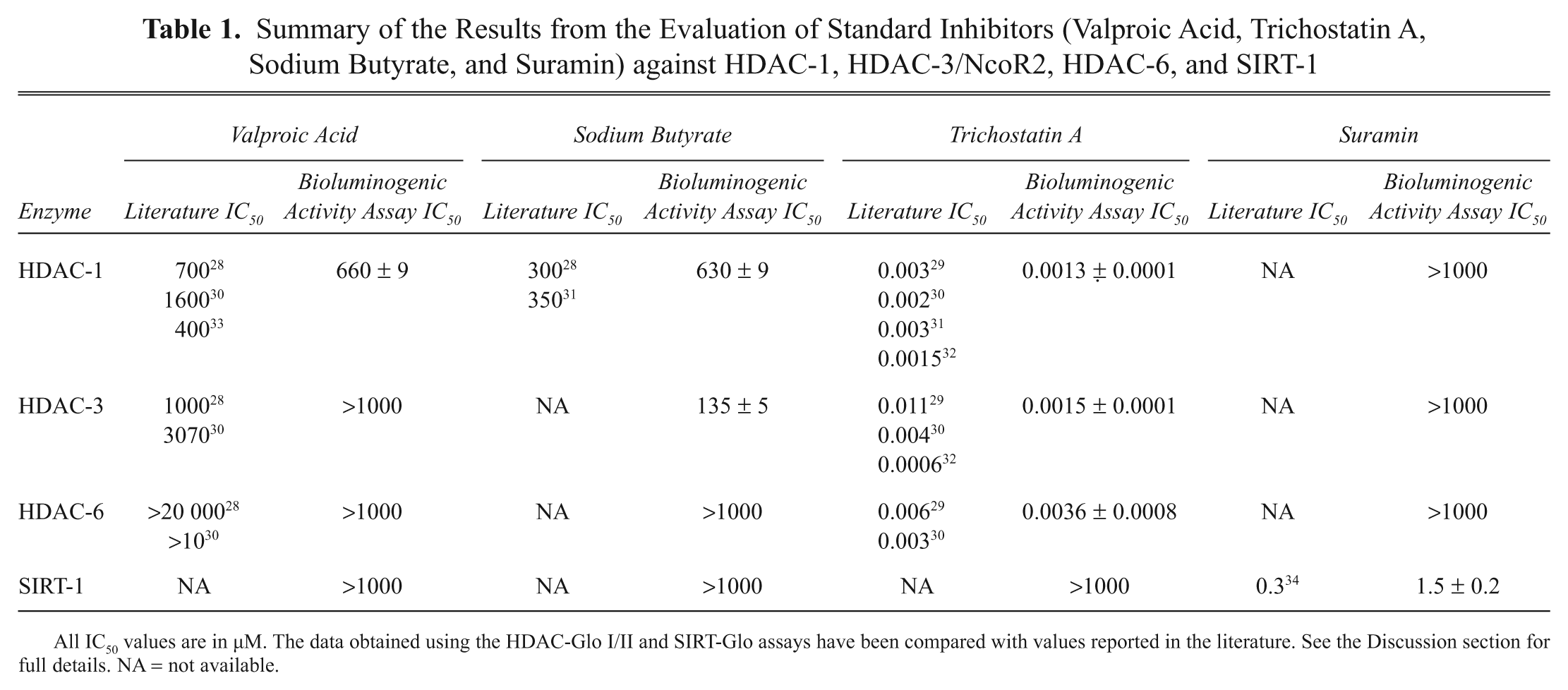
All IC50 values are in µM. The data obtained using the HDAC-Glo I/II and SIRT-Glo assay have been compared with values reported in the literature. See the Discussion section for full details. NA = not available.
Automation of the HDAC-Glo I/II and SIRT-Glo assays and execution of the HTS campaigns against the 640 FDA-approved drugs and Hypha Discovery MycoDiverse natural products library
After the preliminary enzyme characterization experiments as described above were completed, the key parameters necessary for their successful adaptation to HTS were determined. These included their sensitivity, scalability, reproducibility, and robustness (Z′). The concentrations of each enzyme for the screening and compound/extract and fraction concentration-response experiments were fixed as follows: for HDAC-1, 0.1 nM; HDAC-3/NcoR2, 0.15 nM; HDAC-6, 0.1 nM; and SIRT-1, 2 nM, and reactions were allowed to proceed that yielded <20% substrate turnover. The assay volume was also reduced from 30 µL to 10 µL with minimal effort while retaining assay robustness and pharmacology with respect toward the standard inhibitors. The read time per microtiter plate was approximately 4 min, and this level of throughput would enable >50 000 assays to be performed per day on an automated screening platform. The overall quality and robustness of the assays were determined in 384-well microtiter plates containing 352 test compounds (located in columns 1–22), the high control (DMSO alone, located in column 23), and the low control (no enzyme, located in column 24). Each assay well contained either 0.25% or 1% v/v DMSO depending on the library screened.
The output from the 640 FDA-approved drugs screening campaign
HDAC-1, HDAC-3/NcoR2, HDAC-6, and SIRT-1 enzymes were screened in duplicate against the 640 FDA-approved drugs. The Z′ values were >0.7 for each of the 16 microtiter plates (see
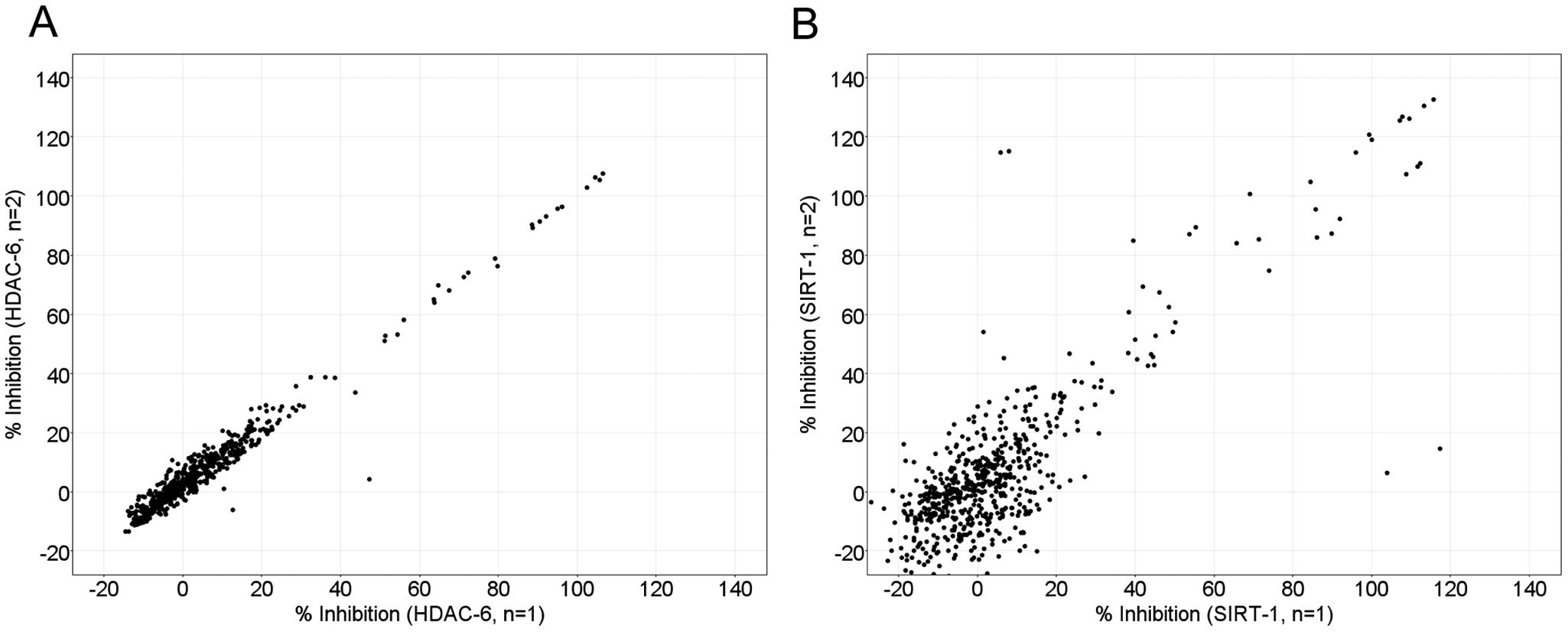
Correlation plots for the screening of 640 Food and Drug Administration–approved drugs against (
The output from the Hypha Discovery MycoDiverse natural products library screening campaign
HDAC-6 and SIRT-1 enzymes were screened against the Hypha Discovery MycoDiverse natural products library using the same strategy as for the 640 FDA-approved drugs screening campaign but in singlicate. The Z′ values were >0.65 and >0.85 for the 34 microtiter plates screened against HDAC-6 (see
Discussion
The validation of HDAC and SIRT enzymes as targets for therapeutic intervention by small molecules has now been established by the FDA approval of a number of vorinostat and romidepsin.9,10 These drugs have significant side-effect profiles, and there is a need to identify novel and selective HDAC and SIRT enzyme inhibitors, in particular in non-oncology indications. One method that is used to identify chemical starting points for drug discovery makes use of HTS against small-molecule libraries containing >500 000 compounds, and these are often screened in standard microtiter plates in 384- and 1536-well format. Although fluorescence-based assay formats have been used in drug discovery for determining in vitro HDAC activity,20,21 they have drawbacks, and it needs to be shown that any hits that are identified from such screening campaigns are not artifacts arising from interference mechanisms due to the intrinsic fluorescence properties of compounds. 22 The potential impact of interfering compounds arising from fluorescence intensity–based assays was brought into focus by the controversy involving compounds identified as activators of SIRT-1 using fluorescence-based assays. 23 Subsequent independent side-by-side comparisons using TAMRA-p53 and native-p53 peptide substrates, high-performance liquid chromatography separation, and ultraviolet detection of the substrates and deacetylated products have shown deacetylase activity only with the TAMRA-p53 peptide substrate. Further biophysical studies using nuclear magnetic resonance, surface plasmon resonance, and isothermal calorimetry showed that one of the compounds interacted directly with TAMRA peptides in a dose-dependent manner; however, the native peptides did not.24,25 This episode demonstrates the need for additional assays to validate the hits identified from fluorescence–based assays.
In this article, we report the evaluation of homogeneous, single-addition, bioluminogenic HDAC enzyme assays with respect to their sensitivity, scalability, reproducibility, signal stability, robustness (Z′), DMSO tolerance, and pharmacological relevance toward standard inhibitors. Is it noteworthy that these assays met all criteria required for them to be compatible for screening purposes. The assays required relatively low concentrations of enzyme (nM range), thus allowing the identification of potent inhibitors and were successfully miniaturized to 10 µL assay volumes and yielded IC50 values for the standard inhibitors that were comparable to those reported in the literature (see
The screening of targets against known drugs or compounds that might have failed to demonstrate efficacy against the primary disease in clinical trials offer the potential for drug repositioning with reduced cost and time scale for evaluation against the alternative diseases.
26
The 640 FDA-approved drugs were used for this purpose and screened against HDAC-1, HDAC-3/NcoR2, HDAC-6, and SIRT-1 enzymes. These activities made use of 16 microtiter plates in total, and they all yielded a Z′ >0.7 (see
In addition to synthetic small-molecule libraries, natural products are also valuable sources for identifying starting points in drug discovery, and this is exemplified by the FDA approval of the natural product romidepsin as an HDAC inhibitor.
10
On this basis, the Hypha Discovery MycoDiverse natural products library was screened in singlicate against HDAC-6 and SIRT-1 enzymes. These activities made use of 34 microtiter plates for each enzyme. For the HDAC-6 screening campaign, all microtiter plates yielded a Z′ >0.65 (see
Footnotes
Acknowledgements
We thank Hypha Discovery for providing the MycoDiverse natural products library for use in this study; Björn Windshügel for helpful discussions; Katarina Bohm, Tom Livelli, Eric Muhr, and Terry Riss for support; and Paul Steckelberg and Lisa Cherradi for technical assistance.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.