Abstract
Identification of tick-protective antigens remains the limiting step in vaccine development. The authors have generated several B cell epitope candidates by fingerprinting Rhipicephalus (Boophilus) microplus proteins that were characterized through bioselection of random peptide phage display libraries against polyclonal antibodies antitick proteins. From 280 clones selected and sequenced, 107 distinct reactive clones were validated by dot-blot assays. Eight consensus motifs were generated, and the most frequent ones were PxxKxH, NxxKxxL, and HTS (68.2%, 65%, and 42%, respectively). The consensus sequences identified potential vaccine targets by alignment with the protein database of R. microplus, which may have putative roles in the host response. Sequences that did not align with known proteins but shared extensive homology among each other were classified as conformational epitopes. Sequence alignments also recognized multiple targets, and the most predominant proteins were identified. Finally, immunized mice sera recognized tick proteins, demonstrating that functional epitope profiles can be identified through selection of phage-displayed peptide libraries with hyperimmune sera and revealing that the epitope-displaying phages can be used as potential vaccine immunogens.
Introduction
T
Methods that employ affinity selection of random peptides fused to the phage capsid (phage display, PD) provide an unbiased approach for epitope mapping, allowing detection and characterization of protein–protein interactions.8,9 We have previously demonstrated that PD can be used to characterize target epitopes of polyclonal antibodies, 10 and more recently, we have also applied this technology to monoclonal antibodies. 11 In addition, we have also identified R. microplus mimetic antigen epitopes (mimotopes) with immunogenic potential for eliciting specific host immune response after mice and cattle immunizations. 12 These mimotopes may represent linear B cell epitopes, conformational epitopes, and conformational discontinuous epitopes of tick proteins, as shown elsewhere.10,13
Another important and complementary approach for antigen discovery is in silico analysis, which has been used to determine protein features that are important for antigen selection and may also lead to the identification of potential vaccine candidates. 14
In this investigation, we have used the strategy to immunize chickens to generate a very large response to R. microplus proteins, as well as use the hyperimmune sera for a large-scale antigen fingerprinting based on phage display, which generated a reservoir of novel tick antigens with immunogenic potential.
Materials and Methods
Preparation of polyclonal antibodies
Total proteins were obtained from larvae and adult R. microplus as previously described, 15 with some modifications. Samples of whole fed females were washed with phosphate-buffered saline (PBS) at 4 °C to remove bovine blood present in the adult tick stomach, and whole unfed larvae were individually macerated in liquid nitrogen. The extraction buffer (40 mM HEPES [pH 7.4], 10 mM EDTA, 2 mM EGTA, 1 mM dithiothreitol [DTT], 1 mM benzamidine, and 0.5 mM PMSF) was immediately added to the samples and centrifuged for 40 min with 40 000 g at 4 °C. The supernatant was collected and dialyzed in PBS at 4 °C. Pierce BCA Protein Assay Reagent (Pierce Biotechnology, Rockford, IL) was used to determine protein concentrations with bovine serum albumin as a standard. No significant proteolysis occurred during the purification (results not shown). Samples were stored at −20 °C until use.
The chicken anti–R. microplus hyperimmune sera were prepared as previously described. 9 For the immunizations, four White Leghorn chickens (6-week-old) were used: two positive inoculums and two negative controls (with buffer without proteins). The animals were kept at the Animal Facility of the Department of Veterinary Medicine at the Federal University of Uberlândia, supervised by the Committee of the Ethical Use of Animals for Education and Research. The first dose contained 200 µg, and the three subsequent doses contained R. microplus larvae or adult proteins administered by intramuscular route, with a 14-day interval between doses. The first dose contained complete Freund’s adjuvant (Sigma-Aldrich, St. Louis, MO), and the subsequent doses contained incomplete Freund’s adjuvant (Sigma-Aldrich, St. Louis, MO). Immunized animals were bled from the third dose and measured for production of antibodies using enzyme-linked immunosorbent assay (ELISA; results not shown). The chicken serum titers were greater than 1:145 800 and 1:16 200 for larvae and adult tick proteins’ immunizations, respectively. Serum samples from chickens were collected and processed. 9 The chicken serum was submitted to IgG purification through an IgY HP column (GE Healthcare, Uppsala, Sweden). Samples were dialyzed in PBS for 12 h at 4 °C in regenerated cellulose dialysis tubing (Fisherbrand, Pittsburgh, PA) and quantified in a spectrophotometer at 280 nm and stored at −20 °C until use.
The chicken serum was submitted to IgG purification through an IgY HP Column (GE Healthcare, Uppsala, Sweden). Samples were dialyzed in PBS for 12 h at 4 °C in regenerated cellulose-dialysis tubing (Fisherbrand, Pittsburgh, PA) and quantified in a spectrophotometer at 280 nm and stored at −20 °C until use.
Phage library screenings
Selection of specific ligands from random peptide phage libraries was performed as described elsewhere.9,16 Phage libraries Ph.D.12mer and Ph.D.C7C were purchased from New England Biolabs (NEB, Beverly, CA) and phage libraries f3-6mer, f88-15mer, f88-cys3, and f88-cys6 were kindly donated by Dr. George Smith (University of Missouri, Columbia). These libraries covered linear and cyclic random peptides, fused on the pIII or the pVIII phage capsid protein, with a size spanning from 6 to 16 amino acids.
Seven biopanning strategies 17 were used to improve the selection of specific ligands to tick antigen-associated IgG. Prior to selection, phage libraries were preadsorbed onto purified IgG from a negative control chicken to subtract phage clones with nonspecific binding. Subtracted libraries were then selected onto purified IgG from immunized chickens against adult (S1 selection) or larvae (S2–S7 selections) R. microplus total proteins. Phage titer was estimated (plaque-forming units, PFU) for the M13 virus, 9 and the concentration of the transducing unit (TU) was used for fd-tet phages by UV absorbance at a wavelength between 240 and 320 nm and the highest peak of absorption at 269 nm. 9 Viral concentration was based on the following formula: viral particles per mL = (A269 × 6 × 1016/number of nucleotides in the phage genome). Productivity was calculated as output/input PFU or TU per 10 µL, and enrichment was determined for each selection by dividing the productivity obtained on the third cycle by that of the first cycle. The final phage stock was stored at −20 °C.
DNA sequencing and bioinformatics
After the last round of selection, individual clones were isolated and subjected to DNA sequencing. Single-stranded DNA was prepared from purified phages
9
and sequenced with the following primers: M13 (5′-CCCTCATAGTTAGCGTAACG-3′; Invitrogen, Carlsbad, CA) and fUSE 5/6-mer (5′-GGAGTATGTCTT TTAAGT-3′; Invitrogen, Carlsbad, CA) for the pIII-fusion sequences and the primer f88-4/15-mer (5’-AGAAGTCCGAAGA CGATA-3′; Invitrogen, Carlsbad, CA) for the pVIII-fusion sequences. DNA sequences were translated (ExPASy; http://www.expasy.ch/tools/dna.html) and aligned by Clustal W2 multiple-sequence alignment software
18
and by manual annotation to characterize mimotopes and motifs. Peptide sequences that shared at least three residues with known protein sequences and were found in three or more selected clones were classified as a consensus sequence (CS). The BLAST server (http://blast.ncbi.nlm.nih.gov/Blast.cgi) and the RELIC suite were used to search for homologies among consensus and peptide sequences with known proteins from the R. microplus database19,20 (
Immunoreactivity of phage clones to chicken IgG
Dot-blot assays were used to evaluate the binding specificity of selected phage clones against chicken-purified polyclonal IgG.
12
Each phage clone was spotted onto a nitrocellulose membrane, and chicken polyclonal sera were incubated, washed, and detected with anti-chicken antibody conjugated with alkaline phosphatase through a colorimetric reaction. The wild-type M13 phage was used as a negative control, and the R. microplus larvae proteins were used as positive controls (
Recognition of mimotopes by bovine sera naturally infested with ticks
IgG antibody responses of bovines naturally infested with R. microplus against phage-displayed mimotopes were also demonstrated by dot-blot assays. 17 Bovine serum samples naturally infested with R. microplus (Uberlândia, Minas Gerais State, Brazil) were collected from four selected calves that were classified as Holstein cross-breed (Bos taurus and Bos indicus) tick resistant (HR) and susceptible (HS), as well as Nelore (B. indicus) tick resistant (NR) and susceptible (NS). The procedure was the same used in the conventional dot-blot assays, except that the secondary antibody was exchanged by the goat anti-bovine IgG horseradish peroxidase (HRP) conjugated (Sigma-Aldrich, St. Louis, MO), followed by detection with Sigma Fast OPD (Sigma-Aldrich, St. Louis, MO).
Competitive dot-blot assays
Competitive dot-blot assays were used to evaluate the mimicry specificity of selected phages to R. microplus total proteins. Dot-blot experiments were performed by preincubating IgG with R. microplus total proteins before incubation onto membranes containing phage clones blotted and incubated for 1h at 37 °C. Serial dilutions of tick total proteins, ranging from 0 to 200 μg/mL, were used. After washing, phage–IgG binding was detected as previously described for conventional dot-blot assays. Inhibition was determined by the decreased binding of IgG (intensity) as tick protein (competitor) concentration increased.
Mice immunization and immunoassays
Selected phage clones were amplified, purified, and individually used to immunize BALB/c mice to evaluate their immunogenic potential and to check for cross-specificity with each serological response against total protein extracts of ticks as described elsewhere.
12
Briefly, 41 groups of five animals each, at 6 weeks of age and with an average weight of 28 g, were immunized with 36 different clones, wild-type M13KE phage, f88 phage, larvae and adult tick proteins, and Tris-buffered saline (TBS). The animals were inoculated subcutaneously in the neck on days 1, 30, 60, and 90 with complete Freund’s adjuvant (Sigma-Aldrich) in the first inoculation and incomplete Freund’s adjuvant (IFA; Sigma-Aldrich) for the three additional boosters. The phage dosage contained 1 × 1013 viral particles/mL in TBS per sample. Blood was collected from the orbital vein postimmunization at days 1, 60, and 90. Blood sample aliquots were dispensed into 1.5-mL microtubes and stored at −20 °C for subsequent ELISA tests (
Results
Phage peptide selection
Purified IgG from hyperimmune chickens was used in six distinct random peptide phage libraries and screened in seven biopanning experiments to maximize epitope discovery of the R. microplus antigens. The number of phage-displayed peptides bound to IgG gradually increased with successive rounds of selection, with phage enrichment ranging from 5- to 1091-fold, except for the S1 strategy (0.6-fold). In three or four rounds of biopanning selections with increasing stringent conditions, phages bound to IgG were eluted (
Peptide sequence analysis
Randomly selected clones (280) were amplified, sequenced, and translated, revealing 107 distinct peptide sequences (
The selected peptides were aligned by Clustal W software (http://www.ebi.ac.uk/clustalw), and eight consensus motifs, SC1 to SC8 (
Peptide sequences were also aligned with the R. microplus proteins database (http://www.ncbi.nlm.nih.gov/protein) and characterized by in silico analyses through a homology search. As shown in Table 1 , the NxxxKxxL consensus sequence presented five protein hits. The TPDKS motif has not fully matched with any protein in the database, and for alignment purposes, a homology of least three residues was adopted, generating a partial alignment with fifteen proteins. The motifs LHS, LHXXL, HTS, PXFF, and LYGS aligned with 2, 7, 2, 3, and 13 proteins, respectively, and only the PXXKXH motif did not present any alignment with known R. microplus proteins. Motifs and mimotope sequences were further investigated by ranking the 10 proteins with the higher number of hits against those sequences, and most of them are present in many consensus sequences and in most of the mimotopes ( Table 1 ).
Putative R. microplus Protein Targets According to each of the Eight Consensus Sequences Obtained by Alignment and Clustering of Mimotopes Originated from Seven Different Phage Display Selections

Proteins with a higher number of hits of motifs and mimotope sequences were ranked based on their average similarity score (RELIC) and had the following order: GP80 precursor, putative secreted protein, Notch-like protein, putative sodium channel, leucokinin-like receptor, salivary gland metalloprotease, cathepsin-type protein precursor, G-protein-coupled receptor, cell surface glycoprotein BM86, and glucose 6-phosphate dehydrogenase isoform C.
Binding assays: antigenicity and specificity
Dot-blot and competitive dot-blot assays were performed to further validate the specificity of the 107 selected phage clones, which showed that the majority of the phage clones bound specifically to chicken IgG when compared with wild-type phage (M13KE) used as negative control ( Fig. 1 ). The majority of the selected phages showed a stronger hybridization to the specific IgG, and among them, the ones with higher reactivity were phages 9, 11, 17, 22, 25, 26, 27, 28, 40, 41, 47, 50, 67, 69, 70, and 90.
The ability of tick proteins to interfere with the anti–R. microplus IgG binding to the phage clones was evaluated by competitive dot-blot assays ( Fig. 2A , B ). Interestingly, phage clone 50 reacted with anti–R. microplus IgG in a dose-dependent manner ( Fig. 2A ). The pretreatment of chicken anti–R. microplus IgG antibody with R. microplus proteins (200 µg/mL) inhibited or decreased the anti–R. microplus IgG binding to phage 50, adsorbed onto the membrane at concentrations of 109, 1010, and 1011 PFU/spot, using serum dilutions ranging from 1:100 to 1:800 ( Fig. 2A, a ). On the other hand, pretreatment of chicken anti–R. microplus IgG antibody with PBS buffer without R. microplus proteins, in the same conditions, showed no inhibitory or interference effect ( Fig. 2A, b ). Finally, the competitive dot-blot assay was used as a screening test to evaluate the affinity of chicken anti–R. microplus IgG antibody bound to phage clones by displacing them with tick total proteins ( Fig. 2B ). The R. microplus proteins were able to specifically displace several phage-bound IgGs in a dose-dependent manner when compared to the control (lack of inhibition in the absence of R. microplus protein competitors). The pretreated anti–R. microplus chicken IgG with R. microplus total proteins inhibited the binding of phages18, 22, 32, 47, 48, 50, 51, 69, and 70 at concentrations ranging from 0 to 100 µg/mL of tick proteins. Therefore, several selected peptides could indeed bind specifically to the purified IgG and mimic the epitopes on R. microplus.
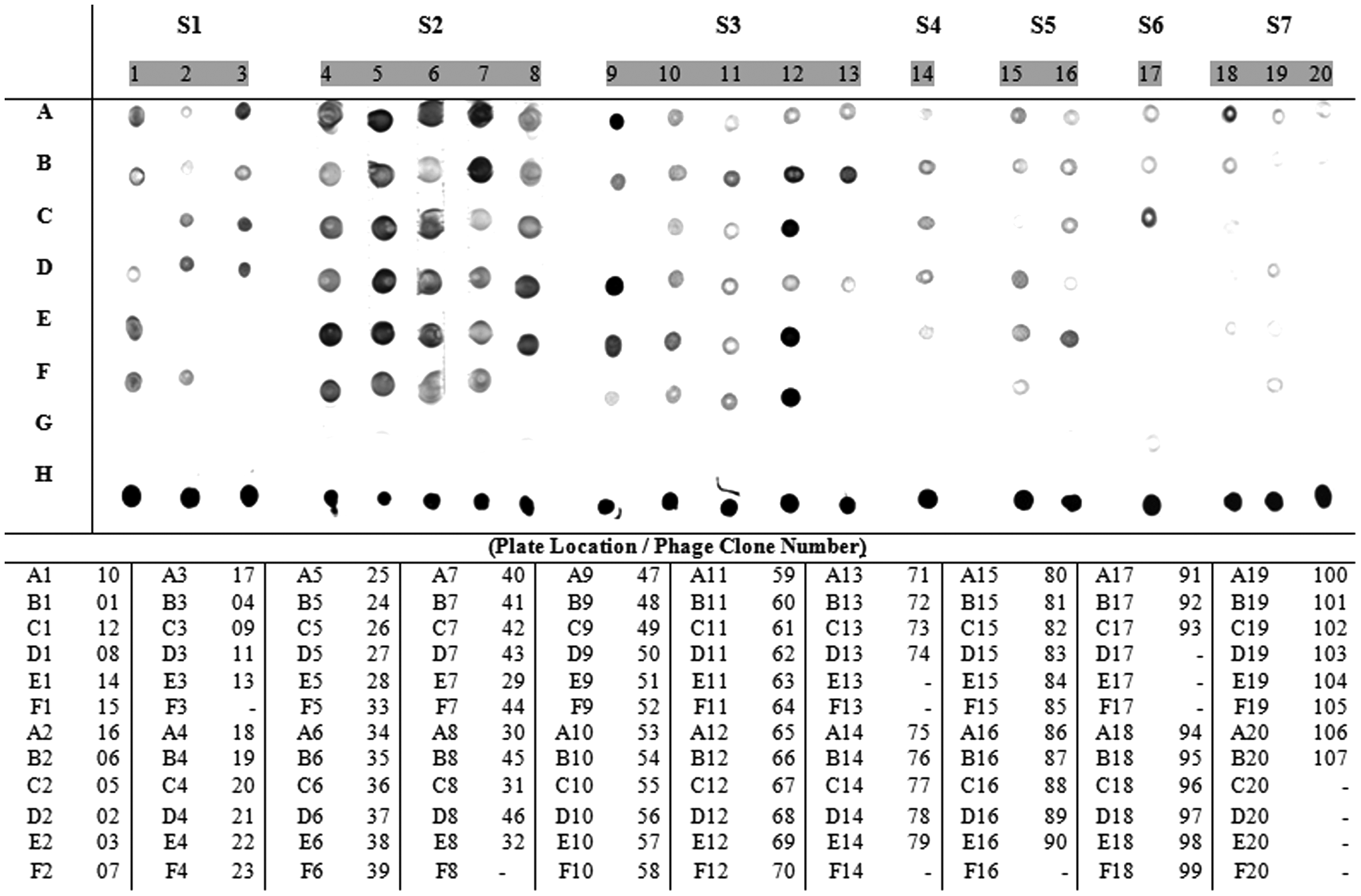
Dot-blot assay to evaluate the specificity of phage clones to chicken antitick polyclonal IgG. The dots are subdivided according to the type of selection (S). The clone location is identified according to its respective lane number (1–20) and column letter (A–F). The negative control (wild-type M13 phage, without peptide) is located in lane G. The positive control (total tick proteins) is located in lane H. The “-” represent blank dots. The clone identification and its location are listed below the assay.
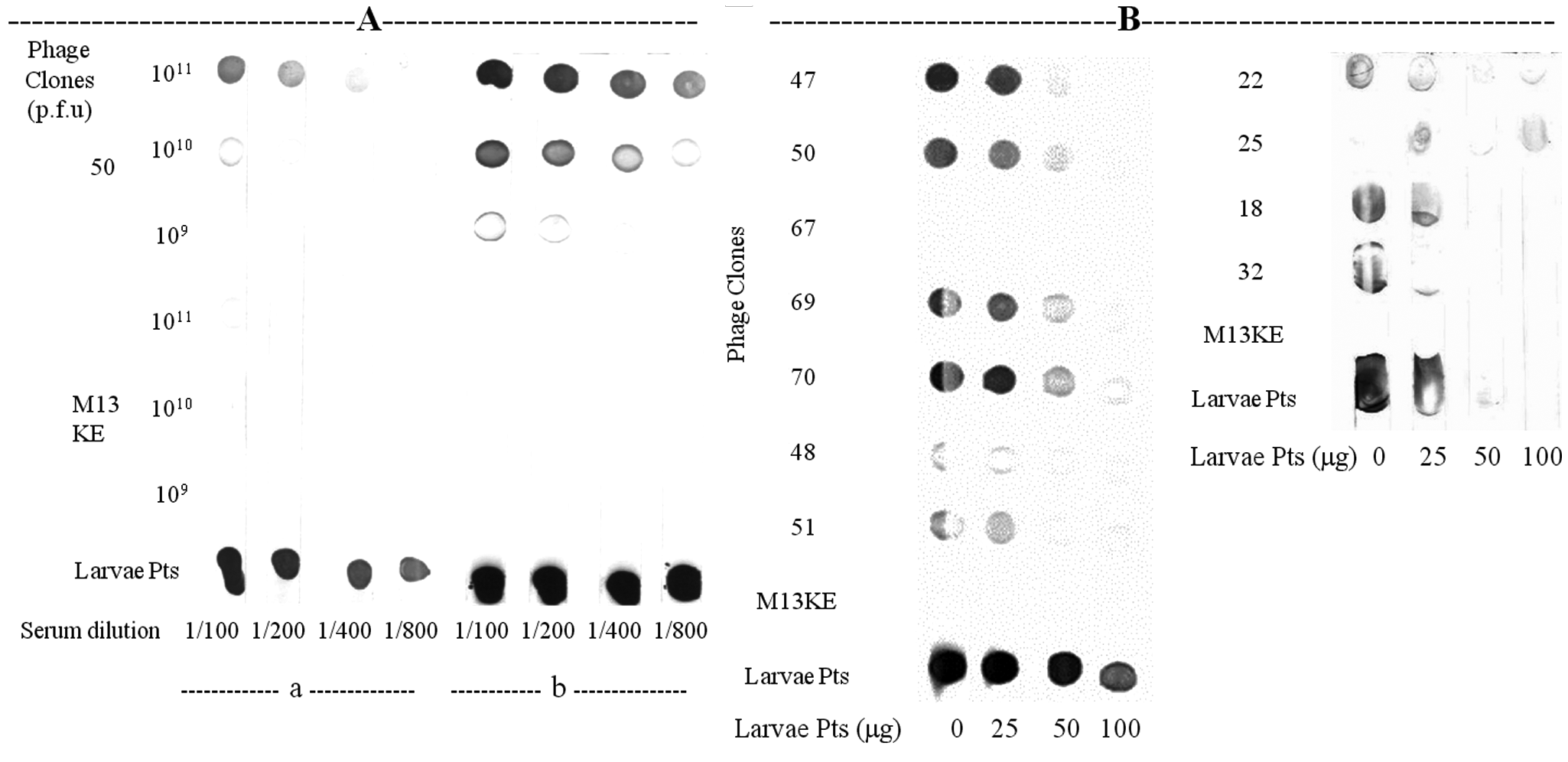
Competitive dot-blot assays between phage clones and tick proteins. (
Furthermore, to verify the IgG recognition of phage-displayed peptides by sera of bovines naturally infested with ticks, the antigen recognition profiles were determined by dot-blot analyses ( Fig. 3 ). The sera were collected after tick counting for two breed animals displaying the lowest and highest infestations, and the means of tick counts for each breed were used to stratify the resistant and susceptible phenotypes. 17 The antigen recognition profiles of 36 phage mimotopes are shown in Figure 3 . Considerable variation was observed between animal sera and the phage-displayed peptides using the wild-type M13 phage as a negative control and tick proteins as a positive control. Among those clones, phage clone 2 was more intensely recognized by the NR animal (animal 163). Phages 12, 15, 75, and 76 were moderately immunoreactive to all sera, except for the HR serum (animal 12). Clones 32, 47, 50, 67, 84, 92, and 94 were well recognized by the four sera tested, but clones 32, 84, 82, and 94 were highly immunoreactive to HS serum (animal 9). Clones 48, 51, and 93 were highly recognized by the HR (animal 12) and HS (animal 9) animals. The results showed that the population of phage clones presented different degrees of immunoreactivity and profiles of binding to tick-infested bovine IgGs, reflecting an antigenic “signature.”
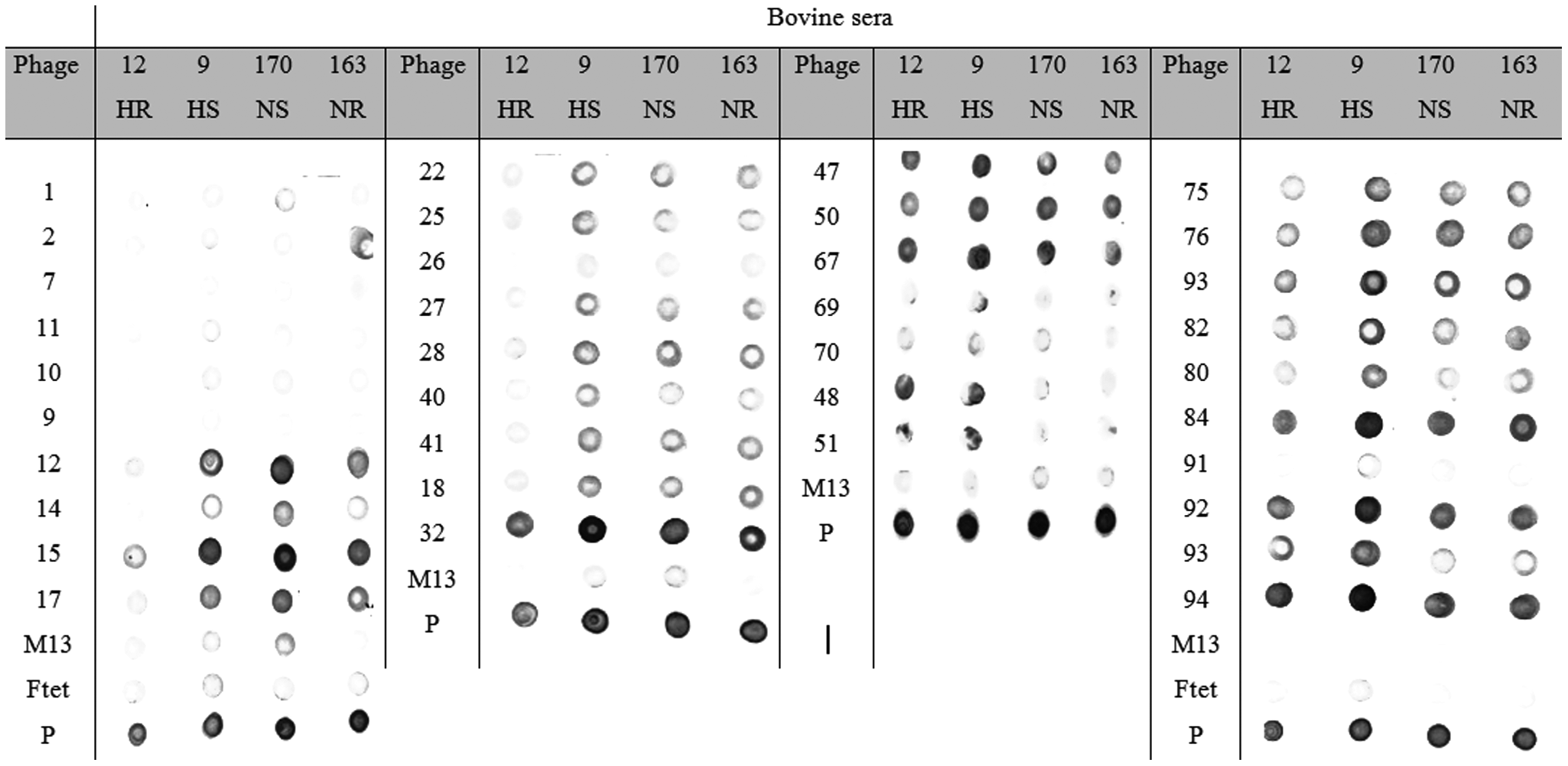
Dot-blot analyses of mimotopes of Rhipicephalus microplus antigens recognized by cattle sera collected under natural tick infestation. Presented are Holstein cross-breed (Bos taurus and Bos indicus) tick resistant (HR-12) and susceptible (HS-9), as well as Nelore (B. indicus) tick resistant (NR-163) and susceptible (NS-170). M13 and Ftet are wild-type phages used as negative controls. Total tick proteins are considered positive controls (P).
Immunogenicity of phage clones
To evaluate the peptide-specific IgG levels in the sera induced by the selected phage clones against R. microplus proteins, 36 phage clones were chosen based on antigenicity and in silico analyses (frequency of multiple copies and similarities) to immunize BALB/c mice ( Fig. 4 ). Most of the mimotopes raised IgG responses against tick proteins with positive ELISA index (EI >1) values. The immune responses due to wild-type phage and TBS were undetectable in the immunized mice. The sera corresponding to immunized groups with phages 32, 26, 75, 25, and 14 presented the highest reactivity indices, respectively. These results clearly demonstrate that mice immunized with phage mimotopes developed IgG antibodies that recognize R. microplus total proteins.
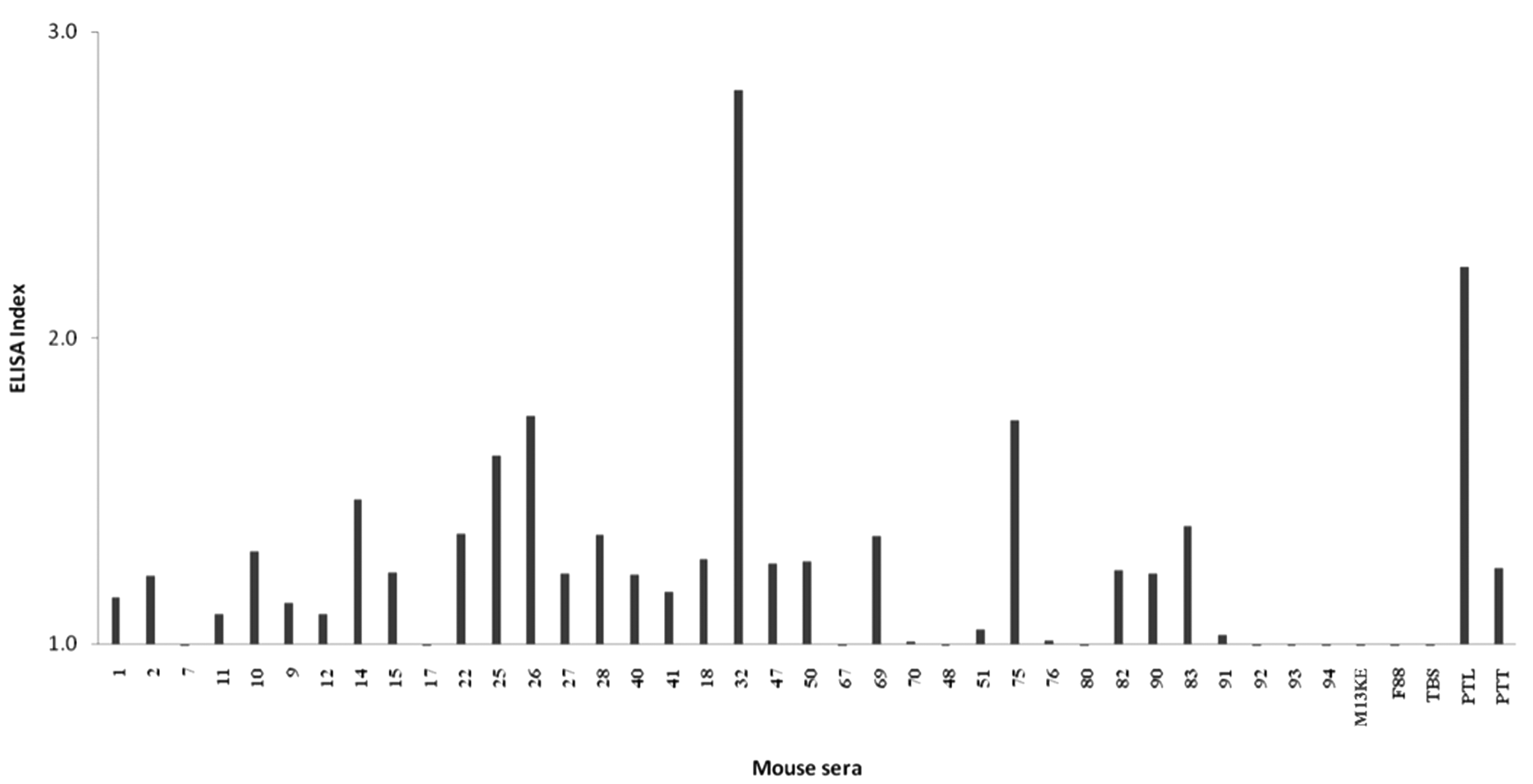
Mouse challenge assays with selected phage clones and immunoreactivity (ELISA index) of mice IgGs against tick total proteins. ELISA, enzyme-linked immunosorbent assay.
Discussion
Vaccines tested against arthropods are based on the generation of host antigen-specific antibodies that results in injury to the arthropod vector during feeding.4,21 Development of improved vaccines against tick infestations aimed at the control of the parasite and the transmission of tick-borne diseases offers a reliable, cost-effective, and environmentally sound control method.2,22 However, new antigens are needed to increase the efficacy of tick vaccines. Although protective antigens against tick infestations have been identified and characterized, discovery of new antigens remains the limiting step for improving the efficacy of tick vaccines.3,7,23–26 Thus, affinity selection of mimotopes against sera of non-host-immunized animals, such as chickens, provides an excellent opportunity to isolate novel ligands without preconceived ideas on which epitopes to target, as a starting point for vaccine development.27,28 The phage display technology is able to detect linear and conformational epitopes, illustrating the whole B cell epitope diversity recognized by polyclonal antibodies. 13 Furthermore, protective immune reactions by mimotope immunizations have been shown for several agents.10,28–32 In the present study, the goal was to perform epitope mapping of R. microplus proteins as the basis for a future tick vaccine design. To our knowledge, this is the most comprehensive study describing the antibody repertoire induced through tick immunizations in a nonnatural host of R. microplus.
Tick proteins were used for chicken immunizations due to the presence of bovine antibodies in a broad range of tick tissues after feeding, 33 which have no neutralization or inhibitory capacity. Furthermore, the interaction of ticks and bovine is a complex relationship. Although the mammalian host tries to avoid the completion of the feeding process, the tick has devised strategies to overcome and evade the bovine immune response, as a result of their coevolution. Consequently, many tick antigens may be masked by repression of signaling pathways and by distorting the host immune response. This diverted immune response may have led to a low reactivity and immunological pressure against tick antigens that mediate protection, as suggested by Waksman’s postulate.34–36 In addition, R. microplus is not a natural chicken ectoparasite, and chickens can raise high-affinity antibodies for highly conserved and low immunogenic antigens than by using bovine native IgGs. 37 A successful selection of highly reactive peptides was obtained probably because of the great diversity of antibodies produced in chickens as a consequence of the large phylogenetic differences from mammalians. However, it is also interesting to note that IgY is exclusively an IgG-isotype antibody, does not activate mammalian complement systems, and does not bind to mammalian Fc receptors. 38 In addition, there are several other technical advantages of chicken IgG as an antibody over mammalian antibodies, such as stability, physicochemical characteristics, productivity, low cross-reactivity, and low cost. 39
Several phage display peptide libraries were screened to obtain highly reactive mimotopes that were validated in a series of in silico, in vitro, and in vivo assays. The identification of relevant epitopes was due to the selection strategy, the use of proper controls, and the greater number of peptides tested in different assays. One hundred seven phage peptides and eight consensus sequences spanning continuous or discontinuous epitopes were identified, which have specifically recognized anti–R. microplus polyclonal IgG from hyperimmune chickens and from R. microplus tick-infested bovines.
The binding of selected peptides to chicken antibodies was dose dependent and showed inhibition or interference on tick proteins in competitive assays. Also, different profiles of immune reactions of bovines naturally infested with ticks were detected, suggesting that mimotopes are representing true epitopes of tick antigens. Interestingly, common mimotopes were not selected in different biopanning strategies, probably reflecting differences among libraries and also the very large repertoire of a polyclonal antibody response.
Mimotopes were also able to generate a specific response against R. microplus proteins in mice and raised IgG antibodies that promptly recognized tick proteins. Sera from mice immunization with phages 32 and 26 presented the highest reactivity indices. Although the immune systems of animal models, such as mouse and chicken, are different from the bovine, we were able to demonstrate antibody cross-recognition among them, and some of the selected peptides may become vaccine candidates in the near future. Our findings are in agreement with the mimotope concept, in which peptides presented on the phage surface resemble the antigenic structure of B cell epitopes of tick proteins.27–29,40 Because the mimotopes may be applicable not only to epitope-specific vaccination, we have also performed analysis of B cell epitopes for target identification through in silico analyses, which provided information on the consensus patterns and similarity among mimotope sequences, and determined new R. microplus epitopes and antigens.
In our in silico analysis, three proteins presented frequent matches against mimotopes, and the GP80 precursor was the protein with highest alignment score, for which a complementary analysis in this study detected putative epitopes that span the SC2, SC5, SC6, and SC8 motifs. The salivary gland metalloprotease spans the SC1, SC2, and SC8 motifs, and the P450 CYP319A1 spans the SC2, SC4, and SC5 motifs. The consensus sequences may reflect major amino acids responsible for the spatial organization of the epitope structure. Interestingly, both GP80 41 and metalloproteases have shown protective effects against ticks. 42 For GP80, it is assumed that critical protective epitopes have been associated with the folding of the protein and/or the oligosaccharides attached to it. 41 It is interesting to note that eight of nine mimotopes (18, 19, 20, 21, 22, 26, 32, and 37) that shared the consensus sequence SC1 were used in a preliminary vaccine trial in bovine, and the cattle immune response induced specific changes in the teleogine physiology, manifested as a hemorrhagic event in the tick gastrointestinal and reproductive tracts. 12 On the other hand, other consensus sequences, such as SC3, which did not match with any known protein but shared extensive homology among peptides, were clustered and classified as conformational epitopes. In this case, it is possible that the displayed peptides were presented to the antibody target as loops and did not show sequence homology to linear sequences of tick proteins.27,28 But it is also possible that the lack of alignment is due to the lower coverage of the R. microplus proteome, which reinforces the use of novel technologies for antigen discovery when the database is limited. Therefore, the selected mimotopes not only revealed antigens with important roles but also provided putative epitope sequences within proteins that may be involved in the host immune response.
Recognition of molecular targets that are relevant to the tick proteome is a key step for the development of vaccines.25,43,44 We have used a proteomic approach in combination with immunizations of a nonnatural host to identify potential epitopes of R. microplus antigens to be explored as vaccine candidates. We hypothesized that parasite proteins with frequent homologous motifs to the selected peptide sequences represent the dominant epitopes of antigens that may play important roles in tick survival and in bovine immune response. The number of times that a specific protein was hit may be a good measure of specificity, especially if the same protein presented contiguous or overlapping sequences in independent clones.20,45
The highest score obtained for the GP80 precursor 14 may be explained by its high antigenicity, 41 its abundance in parasite eggs and larval stages, and the large number of mimotopes that presented high homology to putative epitope regions of the protein. The putative secreted protein has the five-domain Kunitz inhibitor identified in R. microplus, termed penthaplus (UnitProt/TrEMBL entry Q3HYC9). These inhibitors appear to belong to a larger family of homogeneous Kunitz proteins, which have not been well characterized. Finally, the Notch-like protein has a calcium-binding epidermal growth factor (EGF)–like domain, present in a large number of membrane-bound and extracellular (mostly animal) proteins (entry AAN06819.1). Although our selection strategy and the in silico analysis identified important putative antigen targets, none of them were confirmed by purification or immunization.
It is important to emphasize that a different strategy was used in this investigation when compared to another phage display selections recently performed elsewhere. The authors used phage-displayed peptide libraries to detect ligands in eggs and larvae of R. microplus, and although they have attempted to further characterize the targets, data were still preliminary. On the other hand, in our investigation, besides using larvae and adult tick proteins, different types of selections and libraries, and polyclonal sera raised in a nonnatural host, we have further validated mimotopes by using a combination of in vitro, in silico, and in vivo assays that resulted in a large epitope database as potential vaccine candidates for future animal challenges. However, the complete R. microplus proteome is not available, and alignments are only suggestive. The in vitro assays only demonstrate binding efficiency, and the in vivo assays with mice challenges are the first step toward the final validation in cattle immunizations. Importantly, none of the mimotope sequences can be discarded based on isolated or combined assays, but the combination of assays may indicate the most dominant antigens with important roles in the host immune response, and the selection strategy was performed to reduce the amount of antigens for future assays.
In conclusion, we have generated a database of mimotopes through phage display selections against immunized chicken sera with tick total proteins. Several functional antigens were mapped by in silico analysis, and in vitro and in vivo assays were used to identify and validate B cell epitopes. Further experiments are required to confirm if these mimotopes can confer protection against tick infestations. The novel mimotopes selected and prevalidated in all immunoassays are potential vaccine candidates for bovine challenges. The final selection of mimotopes may lead us to develop a recombinant chimeric immunogen as a multiepitope vaccine that may become a novel strategy to control tick infestations and reduce vector capacity to transmit pathogens.
Footnotes
Acknowledgements
We thank Dr. George P. Smith (University of Missouri, Columbia) for providing the phage display vectors and libraries based on the filamentous phage strain fd-tet. This work was supported by FINEP-Brazil, ImunoScan Molecular Engineering Ltda, and Vallée S/A (project 01.05.0303.00). CRP was funded by CAPES-Brazil. The authors thank Dr. Moacir Marchiori Filho and also Dr. Itabajara da Silva Vaz Junior for their valuable comments.
Disclosure: All mimotope sequences presented in this publication are described in the International Patent Application WO/2009/146513.
All authors have equally contributed to this work.
