Abstract
Chronic lung inflammation caused by bacterial pathogenesis through activation of nuclear factor kappa B (NFκB)–responsive proinflammatory genes is a major hurdle in the management of lung disease in cystic fibrosis (CF) patients. The authors generated a disease-relevant cell-based high-content screen to identify novel anti-inflammatory compounds for treating lung inflammation in CF. The human bronchial epithelial cell line KKLEB, harboring the most common form of mutation that causes CF, was modified to express an NFκB-responsive green fluorescent protein (GFP) reporter. After creation, the cell line was tested for its ability to respond to disease-relevant inflammatory stimuli elicited by treatment of cells with filtrates of Pseudomonas aeruginosa isolated from the airways of a CF patient. P. aeruginosa filtrates potently activated NFκB-responsive GFP reporter expression in cells. Subsequently, the assay was optimized for high-throughput screening (HTS) through generation of a Z factor (~0.5) and by testing its tolerance to the commonly used solvents ethanol and DMSO. A pilot library of clinically approved compounds was screened for assay validation. Several compounds with known NFκB inhibitory activity were identified, including several steroidal compounds that have been clinically tested in CF. Thus, the assay can be used in a broader HTS campaign to find anti-inflammatory agents for use in CF.
Introduction
C
Within the airways of CF patients, pathogenic bacterial strains activate different toll-like receptor (TLR) subtypes producing inflammation. However, the extent to which each type of receptor is involved in promoting inflammation upon infection is debated. 6 Bacterial products, including proteoglycans, lipopolysaccharides, lipids, and DNA, have been shown to acutely activate TLRs. 7 Stimulation of the TLR pathway leads to translocation of the transcription factor NFκB into the cell nucleus, 8 which in turn leads to the transcription of a number of NFκB responsive genes, including the proinflammatory cytokines IL-8, IL-6, GM-CSF, and G-CSF and the neutrophil chemoattractant ICAM-1. 8 Concomitantly, anti-inflammatory cytokines such as IL-1 and IL-10 are reduced. 9 An overwhelming number of neutrophils are attracted to the site of infection and release proteases that cause structural damage, which initiates a cyclical chain of events that leads to further recruitment of neutrophils, release of cytokines, cleavage of structural proteins integral to lung function, and persistent bacterial infection. At this juncture, the inflammatory response becomes self-sustaining and eventually leads to loss of lung function and death. 8
Anti-inflammatory agents are used to treat lung inflammation in CF but are somewhat limited in their scope due to adverse effects. 10 Systemic steroids improve lung function but often cause growth retardation, hyperglycemia, and cataracts. 11 The nonsteroidal anti-inflammatory drug (NSAID) ibuprofen at high doses is also effective, 12 but the increased risk of gastrointestinal hemorrhage has limited the use of this agent and other NSAIDs to about 15% of CF patients. Thus, there is a need for compounds that act upon CF airway epithelial cells to inhibit activation of NFκB and the initiation of the inflammatory response without damaging systemic side effects. Although NFκB is ubiquitously expressed, there is considerable evidence suggesting that this pathway in hyperactivated in CF. 13,14 Also, drugs can be specifically delivered to the airways by inhalers, which are commonly used in CF therapy. 15
During the course of this project, an image-based high-content screening (HCS) assay was developed and validated, which can be used to identify novel anti-inflammatory agents for use in CF and other related inflammatory disorders. An immortalized CF airway epithelial cell line (KKLEB) harboring the most common and severe form of mutation seen in CF (ΔF508) was engineered to express a green fluorescent protein (GFP)–based NFκB reporter. 16,17 Filtrates of P. aeruginosa (PAF) originally isolated from a CF patient and containing a mixture of clinically relevant proinflammatory bacterial products were used to stimulate NFκB reporter activity in our assay. 7 Furthermore, a library of clinically approved drugs (NIH Clinical Collection) was used for assay validation and to identify compounds that have anti-inflammatory potential. To the best of our knowledge, the assay reported in this publication is the first example of a disease-relevant high-throughput screening (HTS) assay to identify compounds for use in CF to treat lung inflammation.
Materials And Methods
Creation of KK-NFκB-GFP cells
The KK-NFκB-GFP cell line was created from parental KKLEB cells (gift of S. Randell, University of North Carolina at Chapel Hill), which is an immortalized ΔF508 CF bronchial epithelial-derived cell line. 16 KKLEB cells were trans- duced with the Cignal Lenti-NFκB reporter (GFP) vector (SABiosciences, Frederick, MD) at a multiplicity of infection of 10. Responsive cells were selected by inducing them with 5 ng/mL IL-1β for 24 h and then isolating them using fluorescent-activated cell sorting (DakoCytomation, Glostrup, Denmark) for 2 rounds. The cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM)/F12, with 10% fetal bovine serum (FBS), penicillin, and streptomycin. All cell culture reagents were obtained from Sigma Chemicals (St. Louis, MO) unless otherwise specified.
Screening assay for NFκB activity
P. aeruginosa filtrates were generated by inoculating a clinically isolated strain (CF03114, gifted by Dr. M. C. Wolfgang, University of North Carolina at Chapel Hill) of P. aeruginosa into 50 mL Luria-Bertani (LB) broth and growing for 72 h at 37 °C, shaking. The culture was centrifuged for 20 min at 5000 rpm, and the supernatant was taken and heated at 65 °C for 25 min to inactivate proteases (modified from Wu et al. 7 ).
KK-NFκB-GFP cells were removed from their flasks using a preformulated nonenzymatic solution (Cell Stripper; Mediatech, Manassas, VA) quenched with DMEM/F12 and 10% FBS, centrifuged, and resuspended in the serum-containing media. Cells were counted with a hemacytometer, and ~30,000 cells were transferred to each well of a poly-D-lysine-coated optical bottom 96-well plate. Each plate was incubated at 37 °C for 48 h, at which point the cells were confluent. Then the media were removed and replaced with media containing PAF, lipopolysaccharides (LPS) from PAF (Sigma), LB, or media alone as described for various experiments. Withaferin A (WFA) was used as a positive control for inhibition of NFκB activity during assay development. This compound is a potent inhibitor of NFκB. 17,18
Inhibition of GFP expression stimulated by PAF was performed by pretreatment of the cells with various concentrations of WFA for 2 h. Then PAF was added to each well, and plates were returned to 37 °C for 18 h. On the following day, cells were washed once with Hank’s balanced salt solution (HBSS) and fixed for 10 min at room temperature with 3.7% formaldehyde in HBSS. The fixative was removed and replaced with HBSS with DAPI to stain the cell nucleus, and images were acquired using a Discovery 1 (D1) imager (Molecular Devices, Sunnyvale, CA). Fluorescent images were acquired using a Discovery 1 automated fluorescent microscope (MDS Analytical Devices, Union City, NJ) with filter sets appropriate for FITC (for GFP detection) and DAPI (nuclear stain). Images were analyzed using the integrated MetaExpress software that was provided with the imaging platform. The threshold for positive cells was iteratively set at 300 gray levels above background to ensure that ~10% to 20% of cells in control wells were scored positive. Percent of positive cells per well and integrated intensity for each well were used for calculations. Comparison of the 2 methods used for analysis revealed that both methods yielded directionally similar data sets, but data from integrated intensity calculations added more variability. This is not surprising as an image-guided assay is not a homogeneous method to monitor reporter activity versus other conventional approaches, such as luciferase reporter assays. Therefore, the cell scoring approach was used to calculate most data reported herein. All experiments were repeated at least once to ensure reproducibility.
Z factor determination
The method of Zhang et al. 19 was used to determine the robustness and reproducibility of the assay by calculating a Z factor. Cells were plated in 96-well plates as above and on day 3, half of the wells were pretreated with 3 µM WFA for 2 h, and the other half received fresh media. Then all of the cells were treated with PAF to a final concentration of 5% for 18 h. After the overnight incubation, cells were fixed, stained, and imaged. The average of the percentage of positive cells and the standard deviation within each group was calculated. This experiment was repeated twice more on 2 separate days, and the Z factor was calculated with the following equation:
Solvent sensitivity
The sensitivity of the cells to the presence of ethanol or DMSO was determined by replacing media on cells seeded in a 96-well plate with DMEM/F12/10% FBS containing DMSO, ethanol, or a mix of the two at 1%, 0.5%, or 0.25% of total volume per well. The cells were incubated at 37°C for 2 h. Then PAF was added to a final concentration of 5%. The cells were incubated for 18 h at 37°C, fixed, stained with DAPI, and imaged as above.
Screening of clinically active compounds for activity
A library (NIH Clinical Collection) of ~450 pharmacologically active compounds was blindly screened to identify compounds with anti-inflammatory potential. Cells were plated on poly-D-lysine-coated 96-well plates and grown until confluent. Compounds were added to each individual well at 10× concentration and preincubated for 2 h prior to stimulation with 5% PAF. All compounds were screened at a final concentration of 10 µM in 1% DMSO. WFA was used as a positive control. Controls included cells treated with LB media only and cells stimulated with PAF (5%) without addition of WFA. A compound was considered a “hit” when it inhibited PAF-induced reporter activity by 50% or more based on the assay window. The hit rate was ~3% for the entire collection. The hits were further characterized using concentration-response experiments. Concentration-response studies were done by pretreating cells for 2 h with 6 different concentrations of each compound starting at 10 µM and reducing the concentration by half-log. Cells were subsequently stimulated with 5% PAF. Groups of cells were pretreated with WFA (positive control) or DMSO (vehicle control). Data were fitted using nonlinear regression analyses, and IC50 values were used to generate a rank order of potency (Prism Software; GraphPad, La Jolla, CA).
Results
Development of an NFκB reporter assay in an airway epithelial CF cell line
Chronic and uncontrolled stimulation of cellular signaling by bacterial products through TLRs leads to hyperactivation of the transcription factor NFκB and overexpression of a number of proinflammatory cytokines in CF.
20,21
As such, a disease-relevant in vitro model of inflammation was developed wherein an immortalized CF airway epithelial cell line (KKLEB) was engineered to stably express a GFP-based NFκB reporter. We performed extensive characterization of these cells using various known activators of the NFκB pathway, including IL-1β, tumor necrosis factor (TNF)–α, and LPS from P. aeruginosa (data not shown). However, inflammation in CF is a result of chronic stimulation of NFκB by a variety of bacterial products and not by a single agent. Thus, PAF from a clinically isolated CF strain was used as a stimulant in our assay. This represents a more clinically relevant mode of NFκB activation as it contains various proinflammatory factors, including LPS, DNA, lipoproteins, and secreted inflammatory toxins.
7,22
Initial concentration-response studies were conducted (
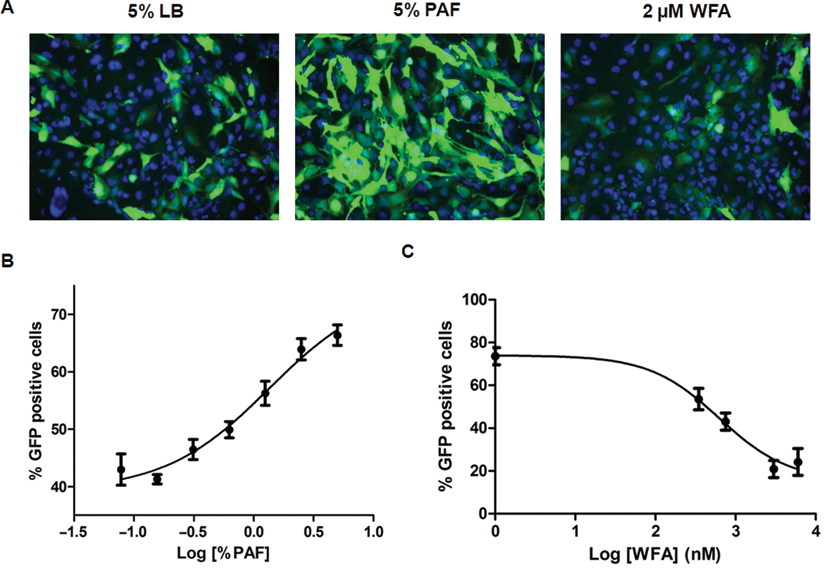
KK-NFκB-GFP cells express green fluorescent protein (GFP) in response to stimulation with Pseudomonas aeruginosa filtrate (PAF). (
Calculation of Z factor
Assay robustness and reproducibility were determined using the method outlined by Zhang et al.
19
In this case, KK-NFκB-GFP cells were plated in 96-well plates on 3 separate days and pretreated with 3 µM WFA or vehicle alone and then exposed to 5% PAF for 18 h. Images were acquired and analyzed as above, and the Z factor was calculated. We obtained a Z factor of ~0.5 in experiments with WFA (
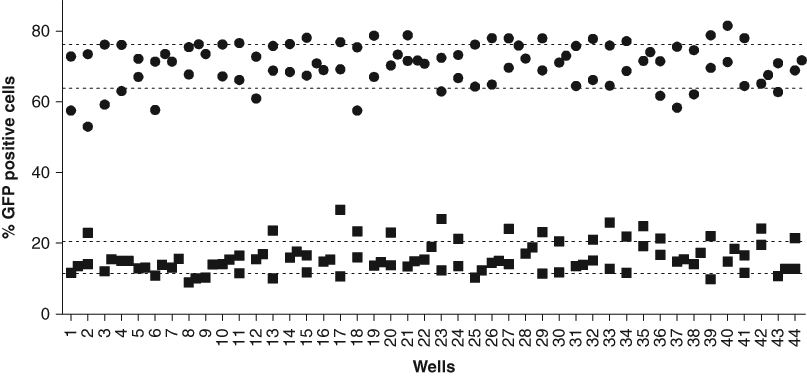
Determination of Z factor. Half of a 96-well plate confluent with KK-NFκB-GFP cells was pretreated with withaferin A (WFA; 3 µM), whereas the other half received vehicle alone. The plate was incubated for 2 h at 37°C, before 5% Pseudomonas aeruginosa filtrate (PAF) was added to all wells. The plate was incubated at 37°C for 18 h, and the cells were fixed, imaged, and analyzed as described in Materials and Methods. The experiment was repeated twice, and a Z factor was calculated. Individual samples are represented by 5% PAF alone (circles) or 3 µM WFA + 5% PAF (squares). The dotted lines represent standard deviation from the means for each group. Data from 2 wells are reported in the figure.
Tolerance of assay to solvents
Tolerance to solvents is often a limiting factor during assay execution. Therefore, we assessed the effect of DMSO, ethanol, or a mixture of 2 on assay performance as these solvents are routinely used as vehicles for in vitro assays. KK-NFκB-GFP cells were treated with PAF in various solvents up to 1% final solvent concentration for 24 h and analyzed using microscopy. As indicated in
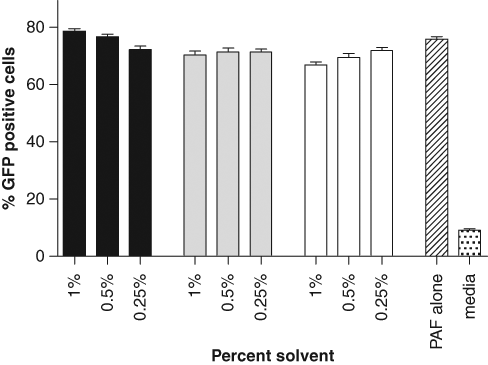
Effect of DMSO and ethanol on assay performance. Confluent wells of KK-NFκB-GFP cells in 96-well plates were stimulated with 5% Pseudomonas aeruginosa filtrate (PAF) in the presence of 3 different concentrations of DMSO alone (black bars), a mix of DMSO and ethanol (gray bars), or ethanol alone (clear bars) to determine the effect of solvent(s) on assay performance. After incubation for 18 h, cells were fixed, imaged, and analyzed as described in Materials and Methods. Data reported as mean ± SD for 3 samples.
Identification of multiple anti-inflammatory compounds
A library of ~450 clinically active compounds (NIH Clinical Collection) was used for assay validation. All compounds were blindly screened at 10 µM final concentration, and compounds that inhibited the number of GFP-positive cells by >50% within the assay window were considered hits. The hit rate was ~3% during the screening process. The majority of hits were either known anti-inflammatory agents or cytotoxic antineoplastic agents. The false-positive cytotoxic compounds were easily identified as they greatly decreased the number of viable cells per image. Generally, compounds that produced a >25% decrease in cell number at the end of the screening campaign were all known antineoplastic agents. In contrast, known anti-inflammatory agents produced little to no cytotoxicity at the cellular level. Furthermore, many of these compounds have been shown to block NFκB activation in other studies, confirming the ability of this assay to identify anti-inflammatory compounds that target this pathway.
Twelve compounds were chosen for further testing based on the strength of the initial response and lack of cytotoxicity. Concentration-response experiments were conducted to further characterize these hits. Representative images with beclometasone are shown in
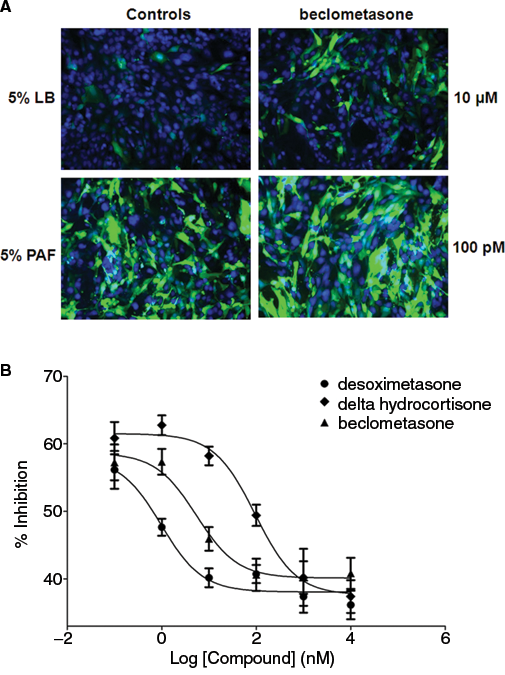
Concentration-response studies with select compounds. KK-NFκB-GFP cells in 96-well plates were pretreated with select compounds at different concentrations for 2 h and subsequently stimulated with 5% Pseudomonas aeruginosa filtrate (PAF) for 18 h. Controls were 5% LB or 5% PAF alone. The cells were fixed, imaged, and analyzed as described in Materials and Methods. (
Concentration-Response Studies with NFκB Inhibitors
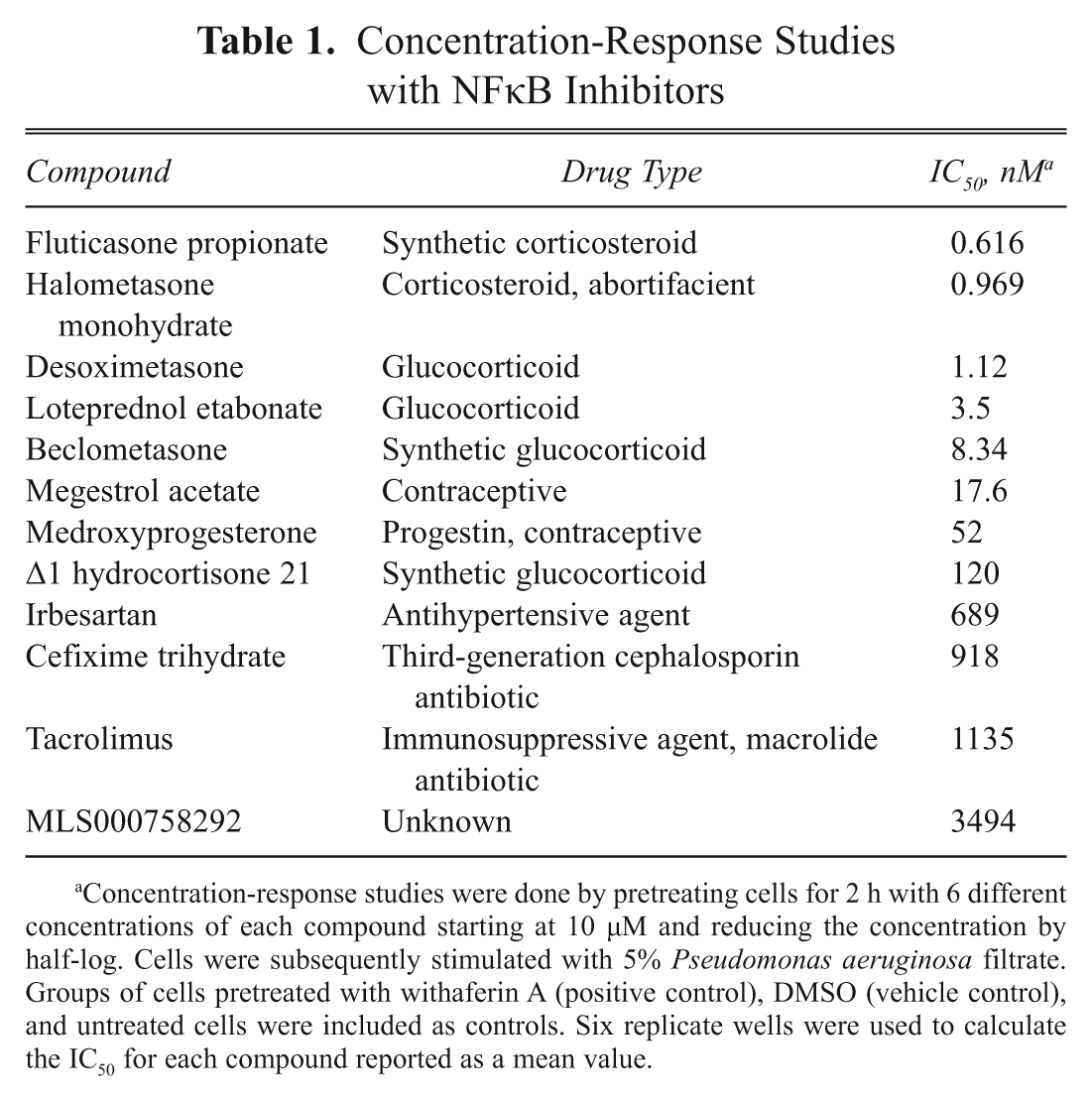

Concentration-response studies were done by pretreating cells for 2 h with 6 different concentrations of each compound starting at 10 µM and reducing the concentration by half-log. Cells were subsequently stimulated with 5% Pseudomonas aeruginosa filtrate. Groups of cells pretreated with withaferin A (positive control), DMSO (vehicle control), and untreated cells were included as controls. Six replicate wells were used to calculate the IC50 for each compound reported as a mean value.
Discussion
The hyperinflammatory state of the airways is one of the major contributors to lung disease within CF patients. Unfortunately, traditional anti-inflammatory agents have limited utility in CF due to undesirable side effects, and new medications are urgently needed. 23 Hyperactivation of the transcription factor NFκB leads to transcription of proinflammatory genes that contribute to lung injury and inflammation in CF. Past studies indicate that inhibition of this pathway is a promising approach to control lung inflammation in CF patients. 15 Furthermore, CFTR itself has been postulated to regulate cellular processes, including antigen presentation, intracellular acidification, and bacterial clearance—functional aspects that are important within the context of inflammatory lung disease in CF, 24 which suggests that screening campaigns to identify anti-inflammatory compounds may benefit from the use of biological models where CFTR function has been inactivated or compromised. In this article, development and validation of a disease-relevant HCS assay are reported that can be used in HTS campaigns to identify anti-inflammatory compounds for use in CF and related disorders that target the NFκB-mediated inflammatory pathway. 24 Furthermore, data from screening a collection of Food and Drug Administration–approved drugs that are currently on the market for anti-inflammatory activity against this pathway are also presented.
To develop a disease-relevant HTS assay to identify NFκB inhibitors that would be active in the lung, an epithelial cell line (KKLEB) derived from human CF lung tissue was altered by transducing it with a vector that inserted a cassette for an NFκB-responsive GFP reporter expression into the cells. The engineered cell line expressed GFP in response to filtrates from a clinically isolated strain of P. aeruginosa, which is a major pathogen in CF. The assay was deemed robust and reproducible, with a Z factor ~0.5, and tolerated vehicle (DMSO, ethanol, or a mixture of 2) concentrations up to 1% without any significant decrease in performance.
The ability of the assay to identify inhibitors of NFκB activation was tested using the NIH Clinical Collection of approved drugs. The screen identified several noncytotoxic inhibitors of NFκB. Of these, ~40% were synthetic or natural corticosteroids. The screen also picked up 2 derivatives of progesterone, and there was weak inhibitory activity with progesterone itself, but it never approached that of the 2 derivatives (data not shown). This is in line with previous evidence that demonstrated that progesterone has an inhibitory effect on inflammation. We did not observe a decrease in NFκB activation with the nonsteroidal anti-inflammatory compounds (NSAID) included in the NIH clinical compound collection. This is not surprising as the concentration used for screening was 10 µM, and previously published data suggest that NSAID concentrations in the mM range are required for inhibition of NFκB activation. 12
The glucocorticoids identified in the screen were able to inhibit NFκB activity at pharmacologically acceptable concentrations, and fluticasone propionate, an anti-inflammatory drug that is used in the treatment of airway inflammation, demonstrated activity in sub-nM range. 25,26 In addition, one of the progesterone derivatives that the screen identified as an inhibitor, megestrol acetate, has been shown to promote weight gain in CF patients. 27 Because CF patients have a high catabolic output due to fighting their ongoing pathogen-induced inflammatory response, NFκB inhibition may eventually contribute to weight gain by promoting a compensatory mechanism upon resolution of inflammation. The identification of megestrol acetate and fluticasone propionate as NFκB inhibitors in these cells also helped with further validation of this assay as these compounds have been tested in CF patients for management of clinical symptoms. Because it is possible to use this cell line for HTS, it is a useful tool in the search for mediators of inflammation for CF and other diseases of the lung.
Footnotes
Acknowledgements
The authors thank Dr. Brian Gilmour for his help with microscopy and Ms. Julianne Tajuba for technical help in the laboratory.
This work was supported by the National Institutes of Health’s Roadmap Initiative through grant R21NS061743-02 to R.M.