Abstract
A cell-based fluorescent protein reporter assay for proteinase activity amenable to high-throughput applications was developed. This assay is based on Förster resonance energy transfer (FRET) between 2 variants of the green fluorescent protein connected by a short cleavable linker and expressed in
Introduction
P
A variety of biological screens have been developed using fluorescent proteins. For example, caspase activation was studied by monitoring the cleavage of a concatenated pair of green fluorescent protein (GFP) variants where cleavage resulted in a loss of Förster resonance energy transfer (FRET) between the 2 variants. 7 A second example, CLiPS, 8 involved mixing a library of bacterial cells expressing peptide substrates on the surface with purified proteinase and a fluorescent tag, which recognized the N-terminus of the peptide. Screening comprised several rounds of (1) prescreening to remove cells not expressing substrate, (2) cleavage by incubating cells with proteinase, and (3) enriching for processed substrates using fluorescence-activated cell sorting (FACS).
The 3Cpro is a cysteine proteinase present in all
Herein, we present a high-throughput method for investigating proteinase specificity based on change in FRET signals of cells expressing a protein substrate consisting of 2 GFP variants. The method was validated by investigating the specificity of the hepatitis A virus (HAV) 3Cpro.
Materials and Methods
Reagents
Oligonucleotides were from Integrated DNA Technologies (Coralville, IA). Restriction enzymes, Antarctic phosphatase, and Taq DNA ligase were from New England Biolabs (Ipswich, MA). T4 DNA ligase was from Fermentas (Glen Burnie, MD). PCR was performed using the expand High Fidelity DNA polymerase system. All other chemicals were of analytical grade. Nucleotide sequencing was performed at the NAPS unit of the University of British Columbia. DNA was propagated and cloned using
Molecular cloning
The
The
The
Fluorescent fusion proteins were generated by inserting a short linker consisting of 2 synthetic semi-complementary oligonucleotides into plasmid pBXCyY-H. The linkers were generated by heating and slowly cooling an equimolar mixture of the oligonucleotides to yield duplexes with 3′ and 5′ overhangs. Plasmid pBXCyY-H was digested with
Plasmids Carrying Genes Encoding Linked CyPet-YPet Fusion Proteins
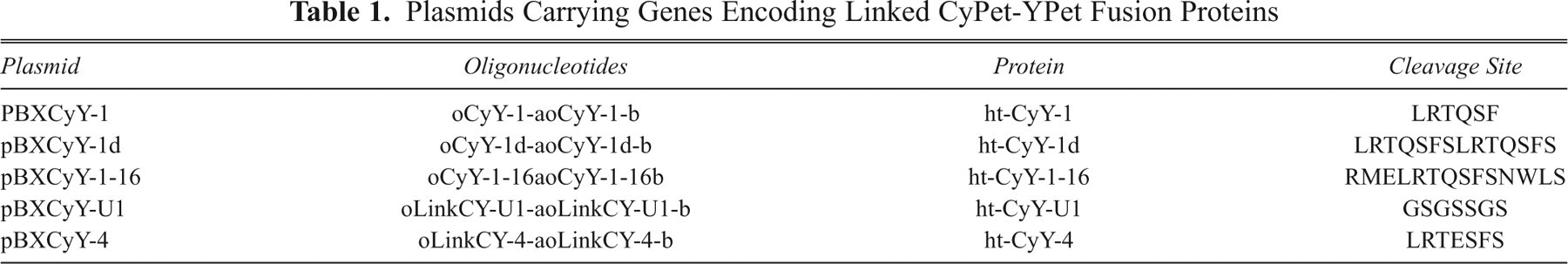
To generate the libraries of plasmids encoding ht-CyPet and YPet connected by potentially cleavable linkers, the
Production and purification of fluorescent protein substrates and proteinase
HAV 3Cpro was produced and purified essentially as described.
11
For fluorescent protein production, a colony of freshly transformed
To purify fluorescent proteins, cell pellets were resuspended in ~35 mL 20 mM sodium phosphate (pH 7.4), 500 mM NaCl, and 20 mM imidazole and lysed using a French press. Cell debris was removed by ultracentrifugation, and the supernatant was passed through a His GraviTrap (GE Healthcare, Piscataway, NJ). Eluted protein was exchanged into 100 mM potassium phosphate, 2 mM EDTA (pH 7.5) by ultrafiltration; concentrated to 5 to 30 mg/mL; frozen as beads in liquid nitrogen; and stored at −80 °C.
Fluorescence-based cleavage assays
Substrate cleavage was monitored fluorometrically in 100 µL 100 mM potassium phosphate, 2 mM EDTA (pH 7.5) at 37 °C using either a Varian Eclipse spectrofluorometer (Cary; Varian, Palo Alto, CA) and a quartz cuvette or a Victor 2 plate reader (PerkinElmer, Waltham, MA) and a 96-well black plate. Using the fluorometer, fluorescence was measured using λex = 434 nm and λem = 477 nm (for CyPet) or λem = 527 nm (for FRET), and initial rates were calculated using the first 5 min of the progress curve. Using the plate reader, fluorescence was monitored using settings for CyPet (λex = 430 nm, λem = 470 nm) and FRET (λex = 430 nm, λem = 535 nm), and initial rates were calculated from the first 45 min of the progress curve.
Screening of fusion protein library
To measure cleavage, 100 µL of the supernatant was added to a black 96-well plate and prewarmed at 37 °C. Initial fluorescence of YPet, CyPet, and FRET was measured. The reaction was initiated with 3Cpro to a final concentration of 1 µM. Changes in CyPet and FRET signals were measured over 2.5 h.
Results and Discussion
Dependence of fusion protein cleavage on linker length
To optimize substrate cleavage for the screen, linkers of 3 different lengths and composition were compared: substrate ht-CyY-1 had linker sequence LRTQ/SFS derived from the 2A/2B cut site of the HAV polyprotein, substrate ht-CyY-1-16 had the extended sequence RMELRTQ/SFSNWLS, and substrate ht-CyY-1d had sequence LRTQ/SFSLRTQ/SFS. Each substrate was purified (>80% as measured by sodium dodecyl sulfate polyacrylamide gel electrophoresis [SDS-PAGE]), and the relative rate of cleavage by HAV 3Cpro (purified to >90% by SDS-PAGE) was determined. Substrate cleavage was confirmed by native-PAGE analysis. At 25 µM substrate, the relative cleavage efficiency of each was ht-CyY-1 (1.0), ht-CyY-1d, (1.6) and ht-CyY-1-16 (1.4), suggesting that linker length did not significantly affect rate of cleavage.
Detecting cleavage of fusion substrate in cell lysates
To test cleavage of fusion protein by exogenously added proteinase in raw extracts and cleared lysates, cells producing either ht-CyY-1 or an uncleavable variant, ht-CyY-U1, were grown in 96-well plates in O2-filled bags. Extra O2 ensured sufficient biomass and fluorescent protein production for the screen. Addition of 3Cpro to either raw extracts or cleared lysates resulted in a steady loss of FRET signal for cleavable compared to uncleavable substrates. Progress curves were similar for ht-CyY-1 in raw extracts and cleared lysates (data not shown).
The ability to use cell lysates rather than purified substrates presents several advantages. First, the library is easily generated using molecular cloning techniques, and by eliminating purification, larger libraries can be screened. Second, substrate cleavage can be measured continuously, facilitating identification of well-cleaved substrates using progress curves. Finally, normalized initial rates (loss of F527) can be used to approximate relative kcat/Km, as reported for CLiPS. 8 Nevertheless, this value is approximate as the screening is not conducted under steady-state kinetic conditions.
Optimizing of 3Cpro concentration for the screen
To optimize the concentration of proteinase used in the screen, up to 1 µM HAV 3Cpro was added to cleared lysates containing either ht-CyY-1 or ht-CyY-U1. Initial rates of fluorescence loss at 527 nm were plotted against the initial F527 signal (approximating fusion protein concentration) (
Fig. 1
). For ht-CyY-1, the initial rate of cleavage, as measured by loss of F527, depended on both substrate and proteinase concentrations. At a fixed concentration of 3Cpro, rates increased nonlinearly with substrate concentration (
Fig. 1
,
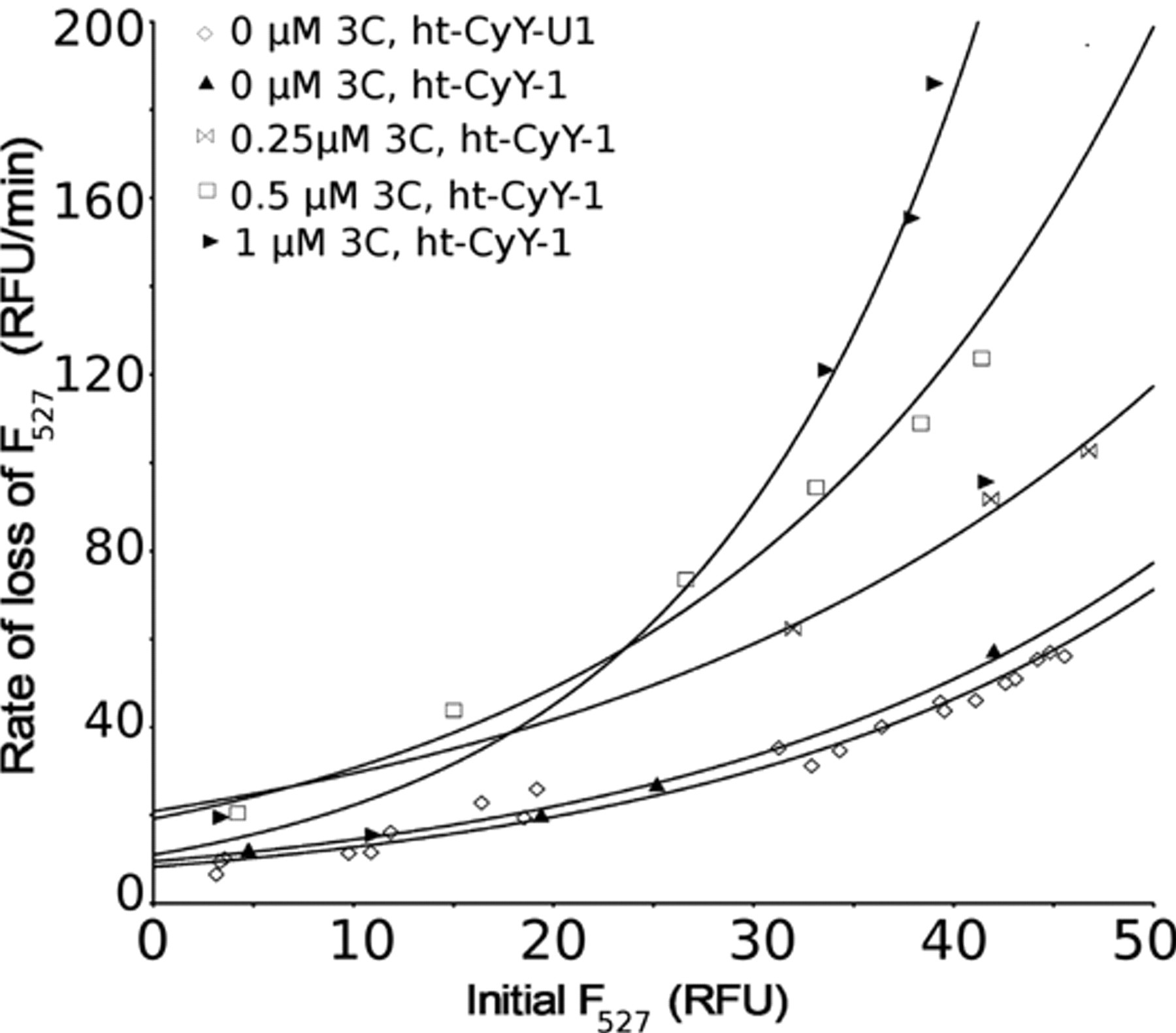
The dependence of cleavage rates of ht-CyY-1 on HAV 3Cpro and substrate concentrations. Initial Förster resonance energy transfer (FRET) signal is proportional to fluorescent protein concentration. Exponential curves were used to highlight trends in the data.
Screening the substrate preference of HAV 3Cpro
To validate the screen, the specificity of 3Cpro was screened using libraries of substrates in which either the residue at the P4 position of the cleavage sequence (XRTQ/SFS) or the P′1 position (LRTQ/XFS) was varied. Random sequencing of 100 clones confirmed libraries’ completeness (results not shown). Based on calculations, the screening of 145 clones provides a 99% chance that the least represented sequence is screened. 12
Library clones were grown in 96-well plates, and initial velocities were calculated using proteinase and cleared lysates as described above. Progress curves displaying obvious irregularities such as jumps in the baseline were eliminated. Initial velocities were plotted versus initial YPet signal instead of the initial FRET signal for 2 reasons. First, the YPet signal is likely to be less influenced by changes in linker structure between variants in the library. Second, measuring YPet signal directly enabled elimination of clones lacking YPet because of the presence of a stop codon in the linker.
In a plot of rate of loss of FRET signal versus YPet signal (
Fig. 2
), cells containing CyPet (ht-Cy-H) lay along the
These initial rates of cleavage were normalized to fusion protein concentration by taking the ratio of rate/YPet: clones with higher normalized rates presumably contain better cleavage sequences. Based on the normalized rate and clone position in the plot ( Fig. 2 ), 14 clones from the P4 library and 8 clones from the P′1 library were selected for sequencing to identify the randomized residue ( Fig. 3 ).
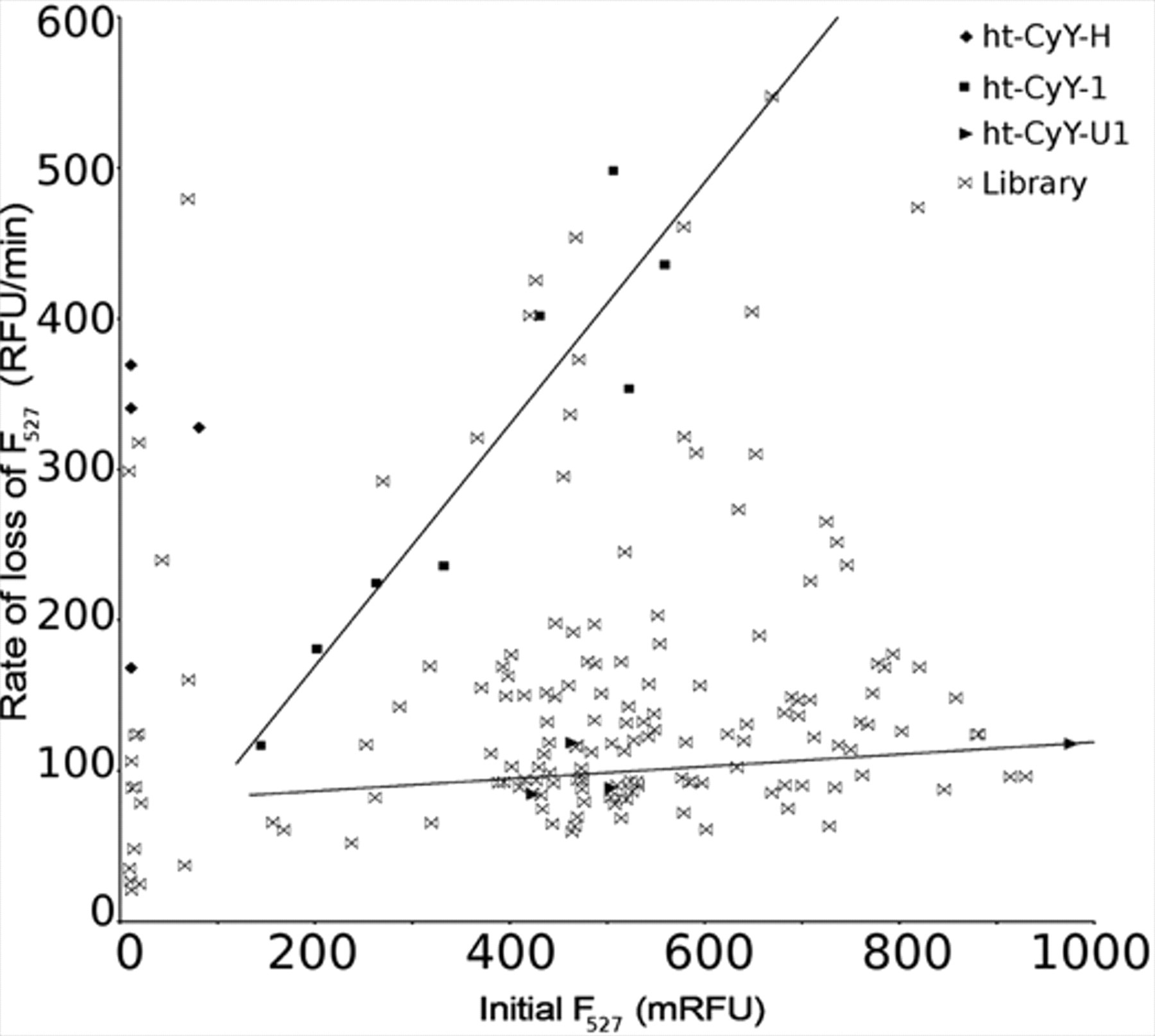
Screening of the P4 library for cleavage by HAV 3Cpro. Initial velocity is plotted against initial YPet signal. Trends for the ht-CyY-1 and ht-CyY-U1 are indicated by best-fit lines.
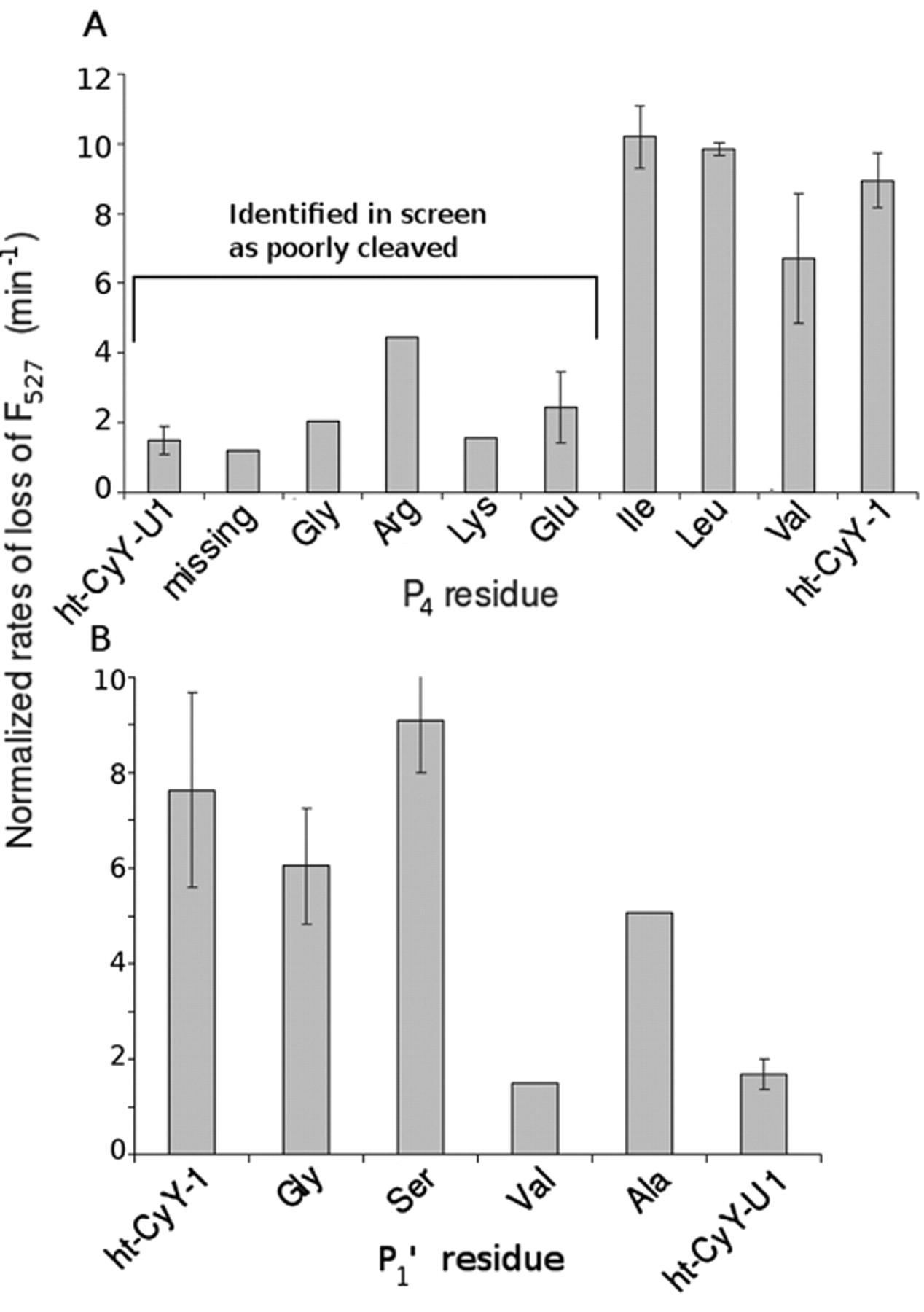
Rate of change in Förster resonance energy transfer (FRET)/YPet signals for selected clones from the HAV 3Cpro screen of cleared lysates of the (
The substrate preference of HAV 3Cpro determined using the screen is consistent with previous studies using peptides and with structural analysis. Specifically, the best substrates had Ile, Leu, or Val at the P4 position and Gly, Ser, or Ala at the P′1 position (
Fig. 3
). Substrates having either charged (Arg or Lys) or small (Gly) residues in the P4 position were poorly cleaved. In a study of 11 synthetic peptides with different natural and artificial amino acids in the P4 position, substrates with Leu, Trp, Val, Ile, and
Kinetic characterization of cleavage of P′1 position substrates
Three P′1 variants (ht-CyY-1A, -1G, and -1V, differing from ht-CyY-1 in having an Ala, Gly, and Val, respectively, at the P′1 position) were purified and kinetically characterized. At a concentration of 4.5 µM, substrates ht-CyY-1A and ht-CyY-1G were well cleaved relative to ht-CyY-1 (relative rates were 0.31 and 0.28, respectively; less than 5% error). The substrate with Val in the P′1 position was cleaved at a relative rate of 0.03, consistent with its identification in the screen as a poor substrate.
Conclusions
This article reports a novel high-throughput screen for analyzing proteinase activity using fused GFP variants. In contrast to peptide screens, libraries are readily generated by PCR amplification of
Four challenges limit the widespread application of the described screen in its current form: (1) potential exosite influence, (2) potential confounding effects from endogenous
Footnotes
Acknowledgements
The
This work was supported by Natural Sciences and Engineering Research Council (NSERC) of Canada Strategic and Discovery grants to LDE. CH was the recipient of an NSERC PGS-D scholarship.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
