Abstract
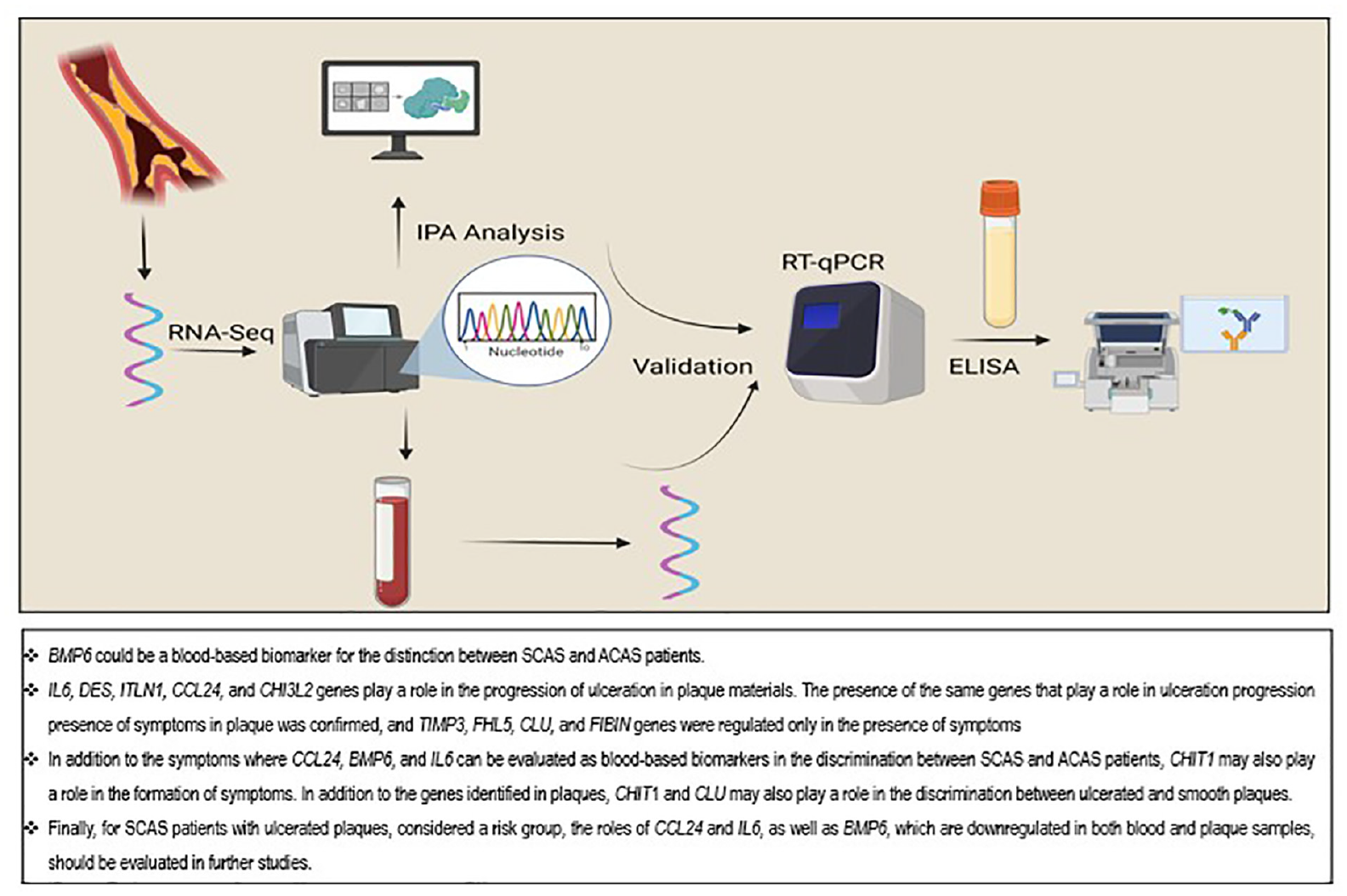
Carotid artery stenosis (CAS) patients are classified as “symptomatic” CAS (SCAS) and “asymptomatic” CAS (ACAS) based on their symptoms. For SCAS and ACAS, complementary biomarkers are needed since the degree of stenosis is insufficient for selecting a treatment or risk assessment. Carotid plaque ulceration represents vulnerable features, and the symptomatic plaque is considered vulnerable. The current study aimed to investigate a blood-based biomarker to determine the presence of ulceration/symptoms in blood by analyzing plaque material based on surface morphology. RNA-Sequencing was performed using the Ion AmpliSeq Transcriptome Human Gene Expression Panel. Additionally, bioinformatic analysis was performed using ingenuity pathway analysis (IPA). Then, gene expression and protein levels were analyzed using Real-Time quantitative polymerase chain reaction (RT-qPCR) and enzyme-linked immunosorbent assay. Based on IPA, immune responses, and reactive oxygen species (ROS) production, lipid transport is activated in SCAS patients. For ulcerated and smooth plaque comparison, it was determined that lipid and sterol transport, and catabolism of lipids, occurred in ulcerated plaque. Based on RT-qPCR, while tissue inhibitor of metalloproteinase 3 (TIMP3), Desmin (DES), ITLN, Chitinase 3-like 2 (CHI3L2), four and a half LIM domains 5 (FHL5), Clusterin (CLU), and fin bud initiation factor homolog (FIBIN) genes were statistically significant in plaque material, C–C motif ligand 24 (CCL24), bone morphogenic protein 6 (BMP6), and interleukin 6 (IL6) were also significant in blood. Interestingly, these genes were downregulated in both plaque and blood. For ulcerated plaque, CCL24 and BMP6 were regulated in both blood and plaque, but only BMP6 was determined to be downregulated in both. Finally, multivariate binary logistic regression shows that BMP6 had the most remarkable ability to predict the SCAS versus ACAS discrimination (odds ratio = 0.553, p = 0.019). Although more studies are warranted, our results suggest that BMP6 seems to be a blood-based biomarker for discriminating SCAS versus ACAS.
Introduction
Cardiovascular disease (CVD) is the leading cause of disability and premature death worldwide. The burden of CVD has increased disproportionately in low- and middle-income countries, where more than 80% of deaths now occur. 1 Atherosclerosis is a CVD (ACVD) characterized by the accumulation of fatty and/or fibrous material in the intima. ACVD also remains the leading cause of vascular disease worldwide, causing many ischemic strokes and transient cerebral ischemic attacks, including coronary syndromes. 2 Carotid artery stenosis (CAS), one of the consequences of systemic ACVD, is characterized by the buildup of plaque in the carotid arteries, causing a wide range of symptoms, from mild to life-threatening paralysis due to stroke. 3
CAS patients are classified as “symptomatic” and “asymptomatic” based on their symptoms, and this classification is major in the management of appropriate treatment. 4 Stenosis >50% in asymptomatic CAS (ACAS) is responsible for ∼10%–15% of all ischemic strokes. Although ACAS can be treated medically or surgically, including carotid endarterectomy (CEA) and carotid artery stenting, treatment approaches are controversial, and treatment is performed at a later stage due to the lack of symptoms. 5 Symptomatic CAS (SCAS) is commonly defined as intracranial or extracranial stenosis in the internal carotid artery, and it has been shown that severe stenosis (70%–99%) confers the highest risk for recurrent stroke or transient ischemic attack (TIA). 6 SCAS plaques are also considered vulnerable and are more prone to plaque rupture than asymptomatic ones. It is known that SCAS has a higher risk of recurrent stroke compared to ACAS. Thus, the proper selection of patients for treatment is of great clinical importance in preventing symptoms, including TIA.7,8 Moreover, since the degree of stenosis is insufficient for the selection of surgical treatment, complementary biomarkers are needed that can contribute to identifying patients in high-risk groups and using carotid imaging techniques. 9
Rupture of the atherosclerotic plaque is responsible for the majority of acute coronary events, including MI and stroke. 10 Plaques are divided into two groups based on their biological characteristics: (i) stable and (ii) vulnerable. Unlike stable plaque, vulnerable plaque, which is a high-risk one, is characterized as having a thin inflammatory fibrous cap, that is, particularly susceptible to rupture. 11 Additionally, carotid plaques are classified according to their surface morphology as (i) smooth, (ii) irregular, or ulcerated. Plaque ulceration is an intimal defect associated with a vulnerable plaque that exposes the necrotic core of the atheromatous plaque. 11 Moreover, the ulcerated carotid plaque surface is an important vulnerability factor associated with neurological symptoms. 12 The studies examining the relationship between atherosclerotic plaque size and rupture risk have found that plaque content increases the likelihood of becoming unstable, and traditional imaging techniques that characterize the plaque according to its appearance and size are thought to be insufficient to predict the risk of rupture or the development of an acute thrombotic event. The ulcerated plaque represents vulnerability, and the symptomatic plaque is considered vulnerable, so determining the relationship between symptom and plaque ulceration is important in diagnostic and prognostic processes. Thus, new biomarkers associated with the high-risk group need to be identified. 13
In the current study, the plaque transcriptome level was evaluated in patients with ACAS and SCAS whose plaque surface morphology was determined by imaging methods (ulcerated vs smooth). Then, biomarkers found to be significant in high-risk groups were confirmed in the blood materials, and a blood-based biomarker that can be used has been investigated in addition to imaging methods. Although there are different biomarker studies to identify the biological character of the plaque and the risk group in CAS patients in the literature,14–17 this is the first study in which blood-based biomarker determination was performed by evaluating the biological character of the plaque in the discrimination between ACAS and SCAS.
Significance statement
Literature demonstrates that the immune system and calcification-related biomarkers may be used to distinguish vulnerable/ulcerated plaques or symptomatic CAS (SCAS)/asymptomatic CAS (ACAS) patients.
Although there are different biomarker studies to identify the biological character of the plaque and the risk group in CAS patients in the literature, this is the first study in which blood-based biomarker determination was performed by evaluating the biological character of the plaque in the discrimination between ACAS and SCAS.
Bone morphogenic protein 6 (BMP6) had the most remarkable ability to predict the SCAS versus ACAS discrimination. Therefore, BMP6 could be a blood-based biomarker for the distinction between SCAS and ACAS patients.
Interleukin 6 (IL6), Desmin (DES), intelectin 1 (ITLN1), C–C motif ligand 24 (CCL24), and Chitinase 3-like 2 (CHI3L2) genes play a role in the progression of ulceration in plaque materials.
In addition to the symptoms, where CCL24, BMP6, and IL6 can be evaluated as blood-based biomarkers in the discrimination between SCAS and ACAS patients, Chitinase 1 (CHIT1) may also play a role in the formation of symptoms.
BMP6 may play a crucial role in the clinical differentiation of SCAS patients and in identifying ulceration associated with the early stages of vulnerable plaque formation.
When combined with imaging modalities, these biomarkers are thought to offer a potentially more reliable approach to the development of targeted therapies and risk stratification for stroke and other cerebrovascular events in high-risk CAS patients.
Materials and methods
The study cohort
In the current study, plaque elasticities of the patients were first determined, and RNA-Sequencing (RNA-Seq) analysis was performed by isolating RNA samples from the plaque materials removed from patients who underwent CEA. Then, candidate biomarker genes were validated in both plaque and blood samples.
Briefly, carotid plaque samples were collected during CEA, a standard surgical procedure to remove atherosclerotic plaques from the carotid artery. All CEA procedures were conducted by an experienced cardiovascular surgeon under sterile conditions. Immediately after surgical removal, plaque samples were rinsed with sterile saline to remove blood residues, transferred to NucleoProtect RNA stabilization solution (Macherey-Nagel, Germany), and stored at −80 °C until RNA extraction. This process ensured that the samples represented the atherosclerotic plaque biology and were suitable for downstream molecular analysis.
Before the analyses, the plaque surface morphologies of the patients were recorded using Doppler ultrasonography and after CEA. An expert (BEG) determined the plaque surface morphology and further confirmed it through post-surgical examination by AY, MB, and MT. Then, plaques were classified as either ulcerated or smooth.
Written informed consent was obtained from all participants for RNA-Seq analyses, and the isolated RNA samples were stored appropriately in the Bursa Uludag University, Department of Medical Biology. This study was approved by Bursa Uludag University Medical Ethics Committee (2021-6/38) and complied with the ethical standards of the Helsinki Declaration.
According to the results of the North American Symptomatic Carotid Endarterectomy Trial (NASCET), 18 CEA provides a durable benefit only in patients with severe stenosis (≥70%), whereas patients with moderate stenosis (50%–69%) gain only limited benefit, and those with <50% stenosis do not benefit at all. Therefore, we included only patients with ≥70% stenosis in our study to ensure clinical relevance and homogeneity. On the other hand, since the risk of CVD, including atherosclerosis, is higher in individuals over the age of 60, and the patients with well-developed atherosclerotic plaques could be diagnosed only after the age of 60, the age of 60 and over was selected as a high-risk group.19,20 Therefore, aiming to determine a blood-based biomarker in the high-risk ACS group, individuals aged 60 and over were included in the study. The inclusion criteria for ACAS and SCAS patients were as follows: (i) age <60, (ii) stenosis <70% according to the NASCET criteria, and (iii) having no history of ischemic stroke within the last 6 months. Exclusion criteria for ACAS and SCAS patients were as follows: (i) age >60, (ii) stenosis >70% stenosis according to the NASCET criteria, and (iii) having ischemic stroke or malignancies. Since the risk of CVD is higher in individuals over the age of 60, and the patients with well-developed atherosclerotic plaques could be diagnosed only after the age of 60, the age of 60 and over was selected as a high-risk group and included in the study.
The effect of sample size and minimum detectable effect size was calculated based on the literature for RNA-Seq analysis by a biostatistics expert, GO. 21 Effect sizes were shown as fold change (FC). Based on the literature, in different scenarios with minimum detectable FC ranging from 1.20 to 2.0, the calculation was carried out with 80% power, and when FC = 2.0, at least six patients should have been enrolled for the discovery group. 21 Therefore, in our study, six SCAS and 10 ACAS patients with no additional vascular disease were selected for RNA-Seq analysis. Then, candidate biomarker genes determined from RNA-Seq results were validated by RT-qPCR in plaque materials. Forty plaques were selected for the validation group, consisting of 24 ACAS and 16 SCAS patients, who were enrolled. Finally, candidate biomarker genes validated in plaque materials were re-analyzed by RT-qPCR in the blood samples. For this purpose, 30 blood samples, 19 of which were ACAS and 11 were SCAS patients, were re-analyzed with RT-qPCR in the current study.
The comparisons for SCAS versus ACAS and ulcerated versus smooth plaque were performed within the same cohort. Specifically, for the discovery group, 16 patients were enrolled in the study as a single cohort and classified according to both their symptomatic status (SCAS n = 6 or ACAS n = 10) and the plaque morphology (ulcerated n = 11 or smooth n = 5). Moreover, these 16 patients in the discovery group were also among the 40 patients in the validation group.
Transcriptome sequencing
Total RNA extraction was performed using the RNeasy Lipid Tissue Mini Kit (Hilden, Germany) from plaque and blood materials obtained from SCAS and ACAS patients based on plaque surface morphology. Red blood cells were lysed by adding a red blood lysis buffer to the blood sample at a ratio 1:10 and incubating at +4 °C for 15 min. After incubation, the samples were centrifuged at 600g for 10 min. The samples were then washed with PBS, Qiazol was added, and shaken vigorously in the same lysis buffer for 1 min. Chloroform was added to the samples and centrifuged at 12,000g for 15 min at 4 °C. The supernatant was removed, and 70% ethanol was added and transferred to the spin column. After washing, 35 µl of RNase-free water was added directly to the column membrane and centrifuged at 10,000g for 1 min. Moreover, for RNA isolation from the plaque materials, tissue samples were lysed with TissueLyser (Qiagen, Hilden, Germany) using beads in 700 µl of Qiazol, then the same protocol was applied. Finally, RNA samples were stored at −80 °C.
RNAs’ quality and concentrations were analyzed using Agilent Technologies 2100 (Agilent Technologies, Santa Clara, CA, USA), and RNA integrity number and distribution value 200 (DV200) scores of the samples were calculated. Samples with DV200 scores above 70% were included in sequencing.
To perform RNA-Seq, mRNA libraries were generated using Gene Studio S5™ (Thermo Fisher Scientific, MA, USA) with the Ion AmpliSeq Transcriptome Human Gene Expression Panel and the Ion AmpliSeq™ Chef-Ready Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions.
First, cDNA conversion was performed by using 10 ng of RNA samples. The generated cDNA was used to construct libraries by performing 31 cycles (based on RNA quality) with the Ion Chef Instrument (Thermo Fisher Scientific). Libraries were diluted to 80 pM concentration, and emulsion PCR, templating, and chip loading were performed using the Ion Chef Instrument (Thermo Fisher Scientific). Sequencing was performed on the Gene Studio S5™ (Thermo Fisher Scientific).
Ingenuity pathway analysis
After sequencing, bioinformatic analysis was performed using QIAGEN CLC Genomics Workbench with FASTQ file(s). Differential gene expression analysis was determined based on a Generalized Linear Model with a negative binomial distribution. For ingenuity pathway analysis (IPA) analysis (https://analiz.ingenuity.com/), RNA expression levels were recorded as reads per million, normalized to the number of sequence reads per sample. Pairwise comparisons of two different groups were performed (ACAS vs SCAS, and ulcerated vs smooth plaque) using IPA. A p value correction was performed using the false discovery rate correction method to eliminate false-positive genes, and adjusted p values were determined for each gene. The adjusted cut-off p value for statistically significant genes was set at 0.05, and genes below this value with ≥2 FC were considered statistically significant as down- or up-regulated.
Validation of candidate genes using RT-qPCR
Validation of 19 genes identified as statistically significant in RNA-Seq results, including lipoprotein lipase (LPL), tissue inhibitor of metalloproteinase 3 (TIMP3), matrix metalloproteinase 9 (MMP9), Chitinase 1 (CHIT1), interleukin 6 (IL6), intelectin 1 (ITLN1), Chitinase 3-like 2 (CHI3L2), four and a half LIM domains 5 (FHL5), C–C motif ligand 24 (CCL24), Desmin (DES), apolipoprotein C-1 (APOC1), APOC2, APOE, bone morphogenic protein 6 (BMP6), fructose-1,6-bisphosphatase 1, neurogranin, stearoyl-CoA desaturase (SCD), Clusterin (CLU), fin bud initiation factor homolog (FIBIN), was performed with RT-qPCR using StepOnePlus Real-Time PCR System (Applied Biosystem, Waltham, MA, USA) with Promega SYBR Green Master Mix (Madison, WI, USA). Gene expression levels were normalized using the GAPDH housekeeping gene. Then, gene expression results were analyzed statistically using the web-based tool, GeneGlobe by Qiagen, at https://www.qiagen.com/tr/applications/geneglobe/qpcr-data-analysis. Based on the results, down- and upregulated genes were determined by FC (≥2 FC, corrected p ≤ 0.05).
Comparison of protein levels between ACAS and SCAS patients
To obtain serum from yellow-capped tubes, the samples were centrifuged at 4000 rpm for 20 min, aliquoted, and stored at −80 °C until analysis.
Serum levels of proteins, including LPL, MMP9, APOC1, APOC2, IL6, neutrophil gelatinase-associated lipocalin (NGAL; Bioassay Technology Laboratory, Shanghai, China), and BMP6 (YL Biotech, Shanghai, China), were studied with enzyme-linked immunosorbent assay (ELISA) kits. Measuring was performed at 450 nm using a multifunctional microplate reader (Thermo Fisher Scientific). Sample amounts were calculated using the appropriate standard curve, and all tests were duplicated.
Statistical analysis
The chi-square (χ2) test and binary logistic regression analysis were used to compare the categorical clinical features in ACAS versus SCAS and ulcerated versus smooth plaque. The significance of clinical features, genes, and protein-independent variables was examined with univariate binary logistic regression analysis, and those found significant were included in the multivariate binary logistic regression analysis. As a result of the analysis, odds ratio and 95% confidence interval values were stated. The descriptive statistics were presented as mean ± standard deviation or frequency and percentage. Differences were considered statistically significant at a p < 0.05. The statistical tests were performed using the IBM SPSS Statistics for Windows, Version 29.0.2.0 (released 2023; IBM Corp., Armonk, NY, USA).
Results
The clinical characteristics of ACAS and SCAS patients
For the discovery group in RNA-Seq analysis, a total of 16 patients with CAS were included in this study. Of these, 6% (n = 9) were male and 44% (n = 7) were female. According to the anamnesis, 43.75% (n = 6) of the patients were diagnosed with SCAS, 56.25% (n = 10) with ACAS, and the mean age of the patients was 67.5 ± 6.4 years. 68.75% of the patients had ulcerated (n = 11), and 31.25% (n = 5) had smooth plaques. Additionally, 76.5% (n = 13) of the patients had hypertension, 47.1% (n = 8) had diabetes, and 23.5% (n = 4) had hyperlipidemia. 5.9% (n = 1) of the patients had peripheral artery disease, and 11.8% (n = 2) had coronary artery disease. The left-sided stenosis of the patients was 56.25% (n = 9), and the right-sided was 43.75% (n = 7).
For the validation group, 40 plaques were selected, and 24 ACAS and 16 SCAS patients were included in the current study. Of these, 62.5% (n = 25) were male and 37.5% (n = 15) were female. According to the anamnesis, 40% (n = 16) of the patients were diagnosed with SCAS, 60% (n = 24) with ACAS, and the mean age of the patients was 67.0 ± 8.0 years. Of the patients, 62.5% had ulcerated (n = 25), and 37.5% (n = 15) had smooth plaques. Additionally, 67.5% (n = 27) of the patients had hypertension, 45% (n = 18) had diabetes, and 20% (n = 8) had hyperlipidemia. Moreover, 22.5% (n = 9) of the patients had peripheral artery disease, and 22.5% (n = 9) had coronary artery disease. The left-sided stenosis of the patients was 62.5% (n = 25), and the right-sided was 37.5% (n = 15). When the SCAS and ACAS were compared, the plaque surface morphology was detected as statistically significant (p = 0.008). Finally, for the blood validation group, a total of 30 patients with CAS were included in the study. Of these, 63.3% (n = 19) of the patients were male and 36.7% (n = 11) were female. According to the anamnesis, 36.7% (n = 11) of the patients were diagnosed with SCAS, 63.3% (n = 19) with ACAS, and the mean age of the patients was 67.5 ± 6.4 years. 67.7% of the patients had ulcerated (n = 20), and 32.3% (n = 10) had smooth plaques. Additionally, 66.7% (n = 20) of the patients had hypertension, 40% (n = 12) had diabetes, and 23.3% (n = 7) had hyperlipidemia. 20% (n = 6) of the patients had peripheral artery disease, and 25.8% (n = 8) had coronary artery disease. The left-sided stenosis of the patients was 73.3% (n = 22), and the right-sided was 26.7% (n = 8). When the SCAS and ACAS were compared, it was observed that the plaque surface morphology was statistically significant (p = 0.04). The clinical features of the patients with CAS are shown in Table 1.
Clinical characteristics of CAS patients for discovery and validation groups.
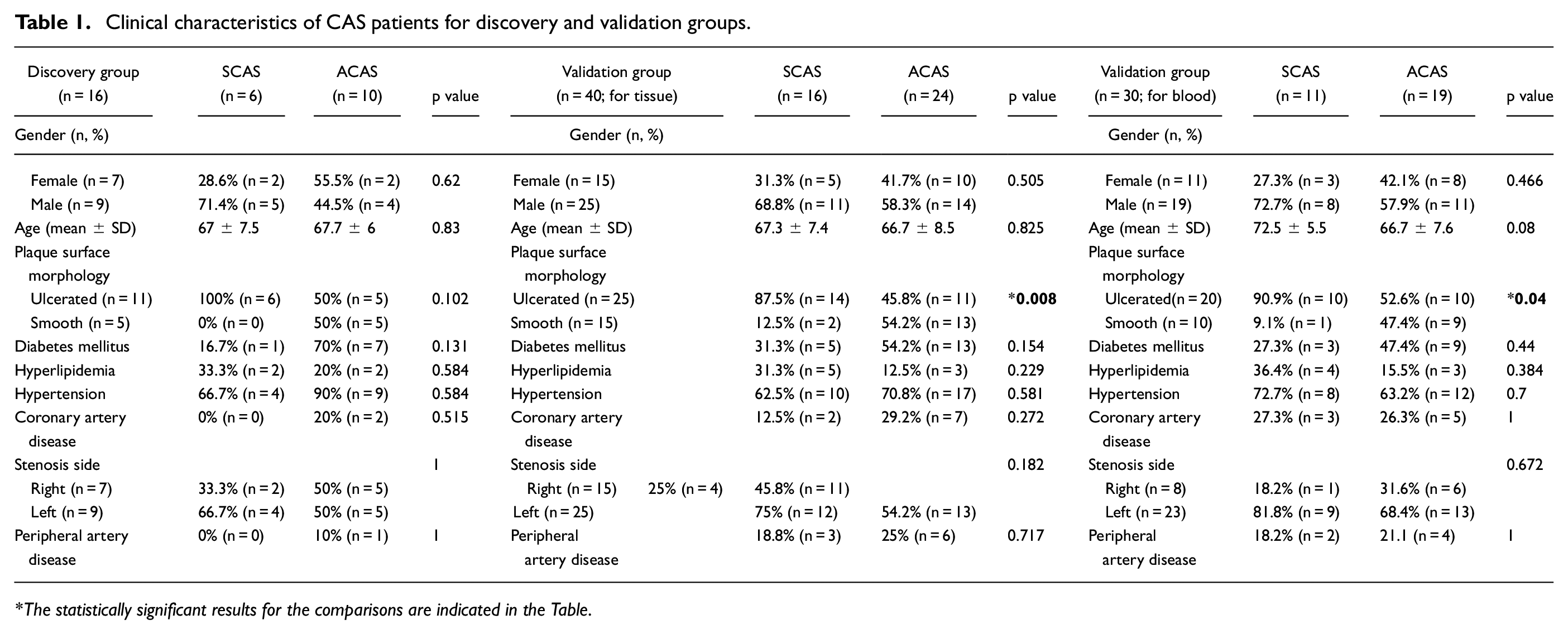
The statistically significant results for the comparisons are indicated in the Table.
Transcriptome sequencing analysis
RNA-Seq analysis was performed to identify genes with significantly different expression levels among the groups and to identify blood-based biomarkers that could be used for discriminating high-risk patients. Analysis was performed in the discovery group using IPA in two different groups: (i) ACAS versus SCAS and (ii) ulcerated versus smooth plaques.
When the canonical pathway and top regulator sections were examined, based on IPA results, it was reported that various mechanisms, including immune responses and ROS production of macrophages, lipid transport system, and sterol transport, are activated in SCAS patients compared to ACAS patients. The genes that can be used in this discrimination were determined to be involved in the pathways (i) atherosclerosis signal, (ii) LXR/RXR activation, (iii) CLEAR signal pathway activation, (iv) macrophage activation signal pathway, and (v) Th1 and Th2 activation signal. All activated and suppressed pathways are indicated in Figure 1(a) to (c). The genes identified in the IPA analysis, the mechanisms in which they function in the pathways, and gene expression FCs are presented in Figure 1(d) and (e). Then, we filtered the genes related to these signaling pathways from the ready molecules with FC >2.0-fold and p < 0.01, and the final DEGs were detected as LPL, IL6, ITLN1, CHI3L2, FHL5, CCL24, DES, BMP6, SCD, CLU, and FIBIN between the two groups.
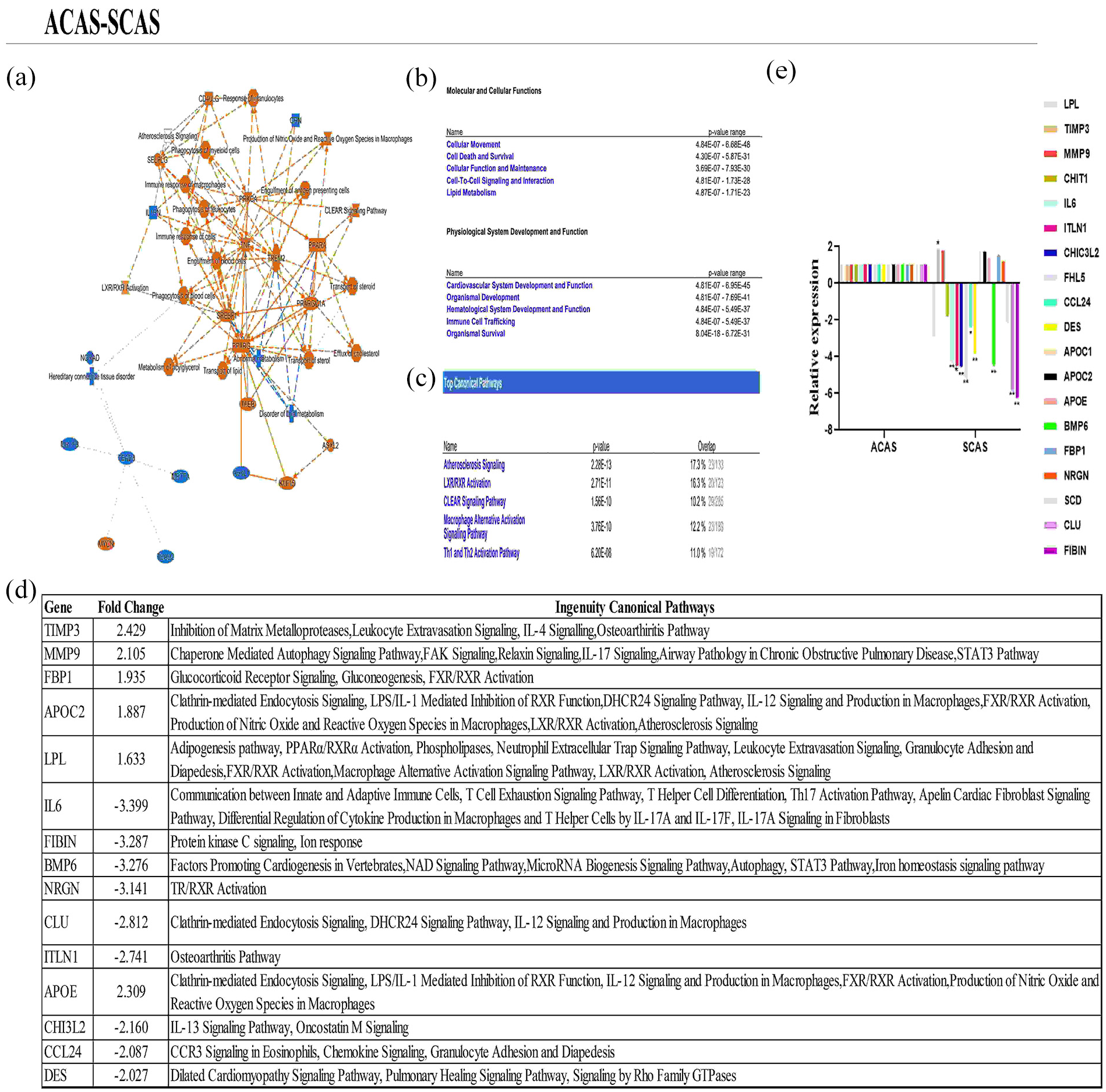
IPA analysis of RNA-Seq results in plaque materials for ACAS–SCAS discrimination. (a) Activated (orange) and repressed pathways (blue) in SCAS patients. (b, c) Molecular/cellular functions of genes involved in this differentiation and the pathways they are involved in. (d) Fold changes and canonical pathways of statistically significant genes involved in the pathways. (e) Validation of RNA-Seq results using RT-qPCR.
Then, the determined FCs were validated in RT-qPCR using 40 plaque materials. Based on RT-qPCR results, LPL, IL6, ITLN1, CHI3L2, FHL5, CCL24, DES, BMP6, SCD, CLU, and FIBIN gene expression levels were determined to be 2.86-, 4.25-, 4.46-, 4.54-, 5.12-, 2.37-, 3.84-, 4.39-, 2.09-, 5.83-, and 6.22-fold downregulated in SCAS patients, respectively. Moreover, TIMP3 gene expression level was determined to be upregulated by 1.9-fold in SCAS compared to ACAS patients. Among the genes, TIMP3, ITLN1, CCL24 (p < 0.05); IL6, CHI3L2, FHL5, DES, BMP6, CLU, and FIBIN (p < 0.01) expression levels were also statistically significant.
Based on IPA results between ulcerated and smooth plaques, especially lipid metabolism (like ACAS vs SCAS), lipid transport system, sterol transport, and catabolism of lipids were re-regulated in ulcerated plaques compared to the smooth ones. The differentially expressed genes between the two groups were determined to be involved in the pathways: (i) LXR/RXR activation, (ii) atherosclerosis signal, (iii) CLEAR signal pathway activation, (iv) macrophage activation signal pathway, and (v) phagosome maturation. All pathways that are activated and suppressed are indicated in Figure 2(a) to (c). The genes identified in the IPA analysis, the mechanisms in which they function in the pathways, and gene expression FC are presented in Figure 2(d) and (e).
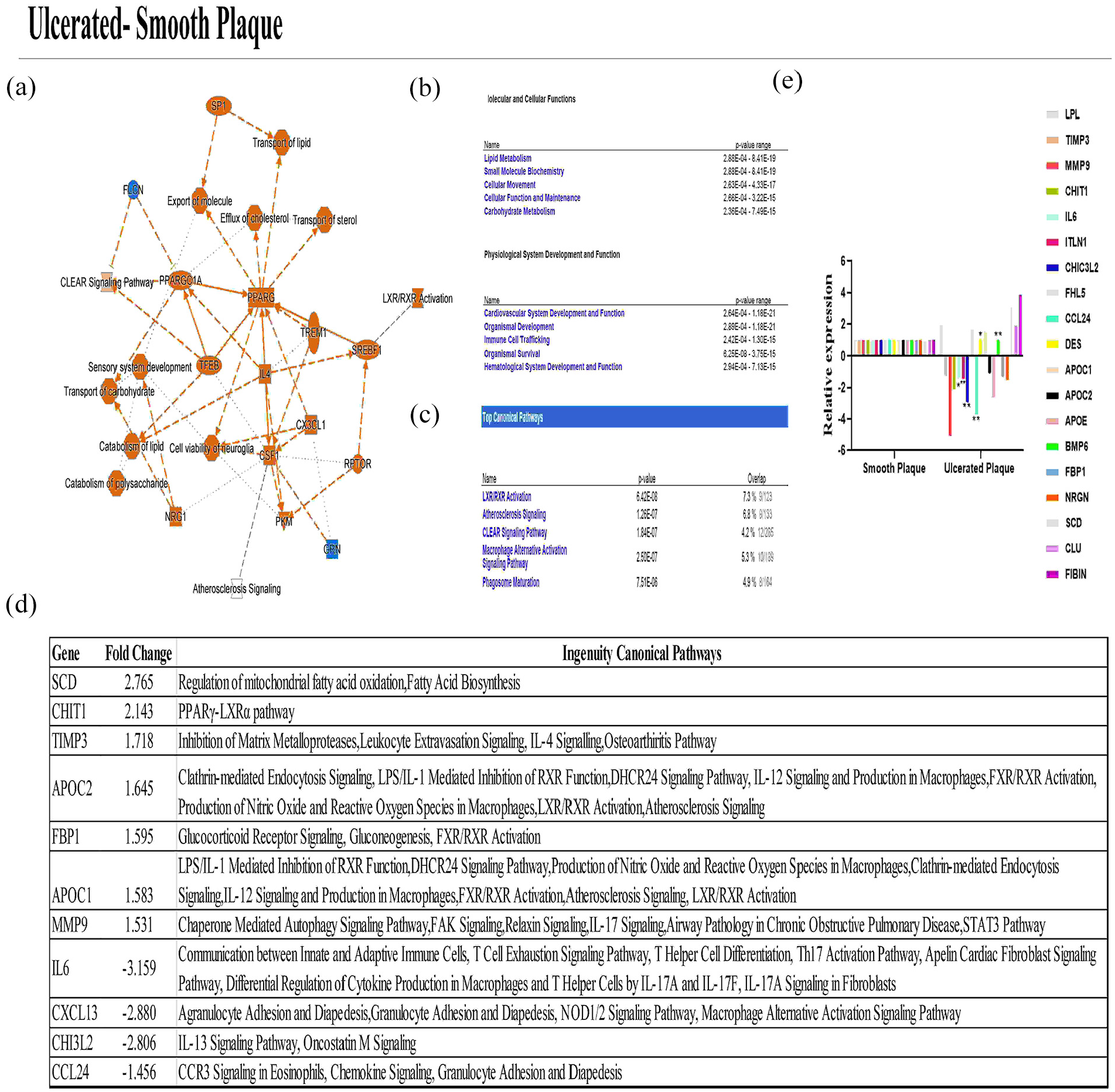

IPA analysis of RNA-Seq results in plaque materials for ulcerated versus smooth plaque discrimination. (a) Activated (orange) and repressed pathways (blue) in ulcerated plaques. (b, c) Molecular/cellular functions of genes involved in this differentiation and the pathways they are involved in. (d) Fold changes and canonical pathways of statistically significant genes involved in these pathways. (e) Validation of RNA-Seq results using RT-qPCR.
Then, the determined FCs were validated in RT-qPCR using 40 plaque materials. Based on RT-qPCR results, when the results from ulcerated plaque were compared with smooth ones, LPL, IL6, ITLN1, CHI3L2, CCL24, DES, APOE, BMP6, and CLU gene expression levels were determined to be downregulated by 2.44-, 2.57-, 3.90-, 4.08-, 3.73-, 1.8-, 2.06-, 2.26-, and 2.19-fold, respectively. However, APOC2 gene expression level was determined to be upregulated by 2.31-fold. Among the genes, IL6 and DES (p < 0.05); ITLN1, CCL24, CHI3L2, and BMP6 (p < 0.01) expression levels were statistically significant. The results of IPA analysis obtained from plaques showed the suitability of common mechanisms and genes for biomarkers that can be used to classify the SCAS patients and the ulcerated plaques.
Comparison of RNA and protein levels for blood-based biomarker analysis
Candidate gene expression and protein levels obtained from plaque materials were investigated in patients’ blood materials to determine blood-based biomarkers. For this purpose, 12 candidate genes were selected after plaque validation, including LPL, CHIT1, IL6, ITLN1, CHI3L2, FHL5, CCL24, DES, BMP6, SCD, CLU, and FIBIN.
Based on RT-qPCR results, LPL, IL6, ITLN1, CHI3L2, FHL5, CCL24, BMP6, CLU, and FIBIN gene expression levels were determined to be downregulated by 1.18-, 3.40-, 1.30-, 1.90, 1.19-, 2.85-, 2.62-, 1.64-, and 1.27-fold. However, CHIT1, DES, and SCD gene expression levels were determined to be upregulated by 3.12-, 1.24-, and 1.1-fold upregulated, respectively, in SCAS compared to ACAS patients. Among the genes, CHIT and CCL24 (p < 0.05), IL6 and BMP6 (p< 0.01) expression levels were statistically significant.
Based on RT-qPCR results for ulcerated and smooth plaque comparison, ITLN1, CHI3L2, BMP6, CLU, and FIBIN gene expression levels were determined to be downregulated by 1.80-, 1.02-, 2.05-, 1.95-, and 3.04-fold, while CHIT1, IL6, FHL5, CCL24, DES, and SCD gene expression levels were determined to be upregulated by 5.64-, 1.05-, 3.19-, 3.55-, 1.41-, and 1.06-fold, respectively, in ulcerated plaques compare to smooth ones. CHIT1, CCL24, BMP6, and CLU (p < 0.05) expression levels were statistically significant.
Considering all gene expression levels, although different candidate biomarkers have up- or down-regulated statistically significant levels depending on plaque ulceration or disease symptoms, BMP6 expression levels were 2.0-fold decreased in the presence of both symptoms and ulceration.
Then, protein levels of the indicated genes were evaluated as being significant at the protein level based on RNA-Seq analysis for SCAS versus ACAS patients, and ulcerated versus smooth plaques. Based on RNA-Seq and RT-qPCR results, the BMP6 protein level was analyzed using the ELISA method. Additionally, LPL, MMP9, APOC1, APOC2, IL6, and NGAL protein levels were examined and associated with oxidative stress and promote cholesterol efflux, which is known to have a strong relationship with atherosclerosis in the literature19–24 in SCAS versus ACAS patients, and ulcerated versus smooth plaques. Interestingly, BMP6 protein levels were not statistically significant in ACAS versus SCAS patients or ulcerated versus smooth plaque discriminations (Figure 3(a) and (b)). Additionally, LPL protein levels were determined as 791.7 ± 55.4 and 1269.14 ± 208 pg/ml (p < 0.05), and MMP9 levels were determined as 1409 ± 101.12 and 2594.02 ± 461 ng/L (p < 0.05) in SCAS and ACAS patients, respectively (Figure 3(a)). However, no statistically significant protein was detected in the discrimination of ulcerated plaque from smooth one (Figure 3(b)).
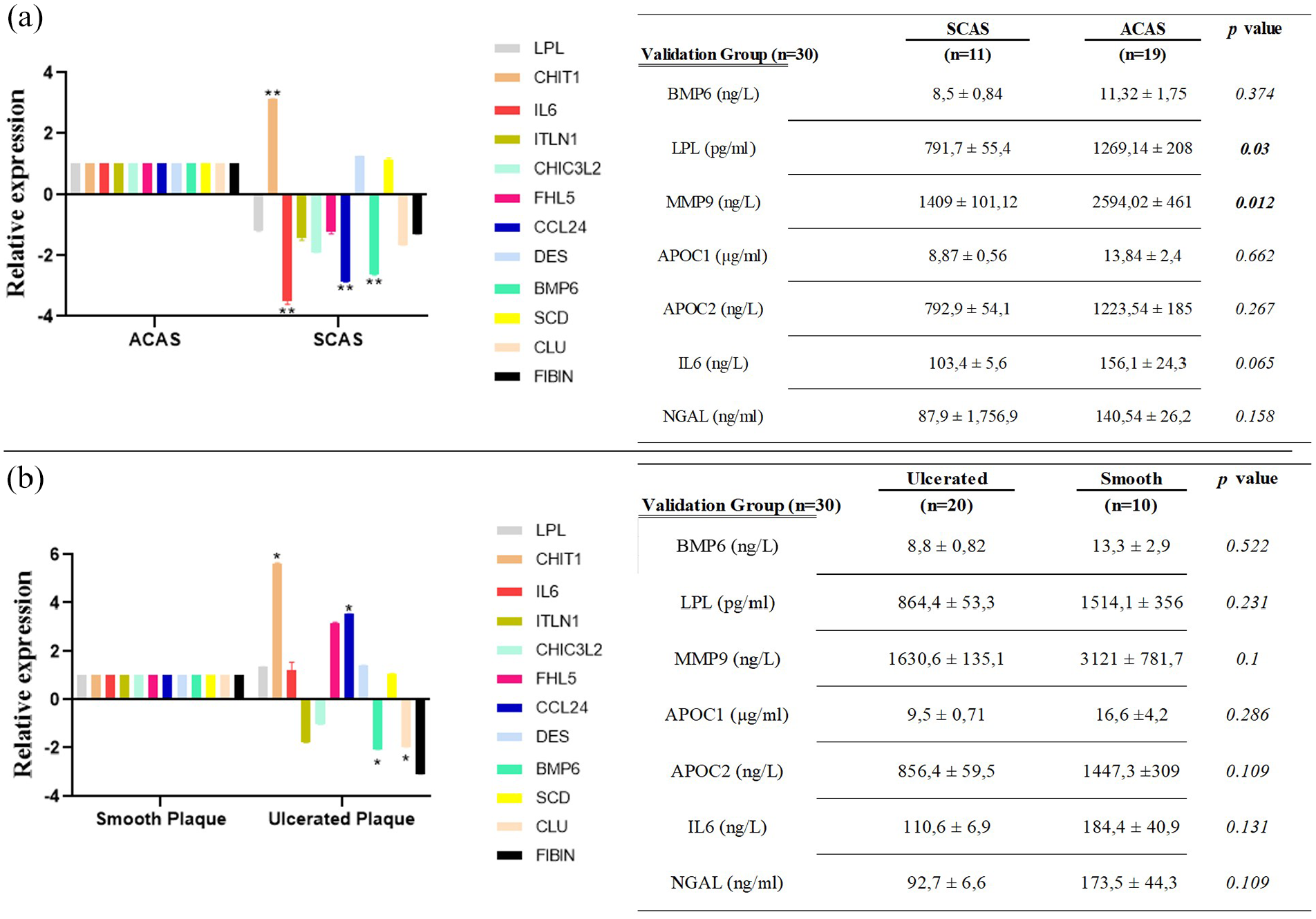
(a) Validation of statistically significant genes detected in plaques for the discrimination between ACAS and SCAS patients, and comparison of serum protein levels. (b) Validation of statistically significant genes detected in plaques for the discrimination between ulcerated versus smooth plaque and comparison of serum protein levels.
We also performed the Pearson correlation test to determine the correlation between each group’s protein and gene expression levels, assigning each biomarker a correlation coefficient. Data showed that LPL and BMP6 were positively correlated at both RNA and protein levels between ACAS and SCAS patients, but only correlated at protein levels between ulcerated and smooth plaques (p < 0.05). Correlation analyses are presented in Supplementary Table 1.
Prediction of blood-based biomarkers for CAS patients
Considering the RT-qPCR results, a diagram was created to evaluate blood-based biomarkers that increase or decrease in the blood and plaque in the presence of symptoms/ulceration (Figure 3).
Common and different genes that could be used in the discrimination of ACAS versus SCAS and ulcerated versus smooth plaque materials are indicated in Figure 4(a). While the IL6, DES, ITLN, CCL24, CHI3L2, and BMP6 genes were determined statistically significant and could be used in the discrimination between ulcerated and smooth plaque, it was shown that, unlike these genes, the TIMP3, CLU, and FIBIN genes, which are independent of ulceration, are regulated in the plaques depending on the symptoms. Interestingly, in blood materials, while the CCL24, CHIT1, and BMP6 genes were determined to be statistically significant and could be used in the discrimination of plaque biology, it was shown that the CLU, independent of symptom, is regulated depending on the ulceration. Additionally, data suggest that IL6 may predict the presence of symptoms independently of ulceration (Figure 4(b)).
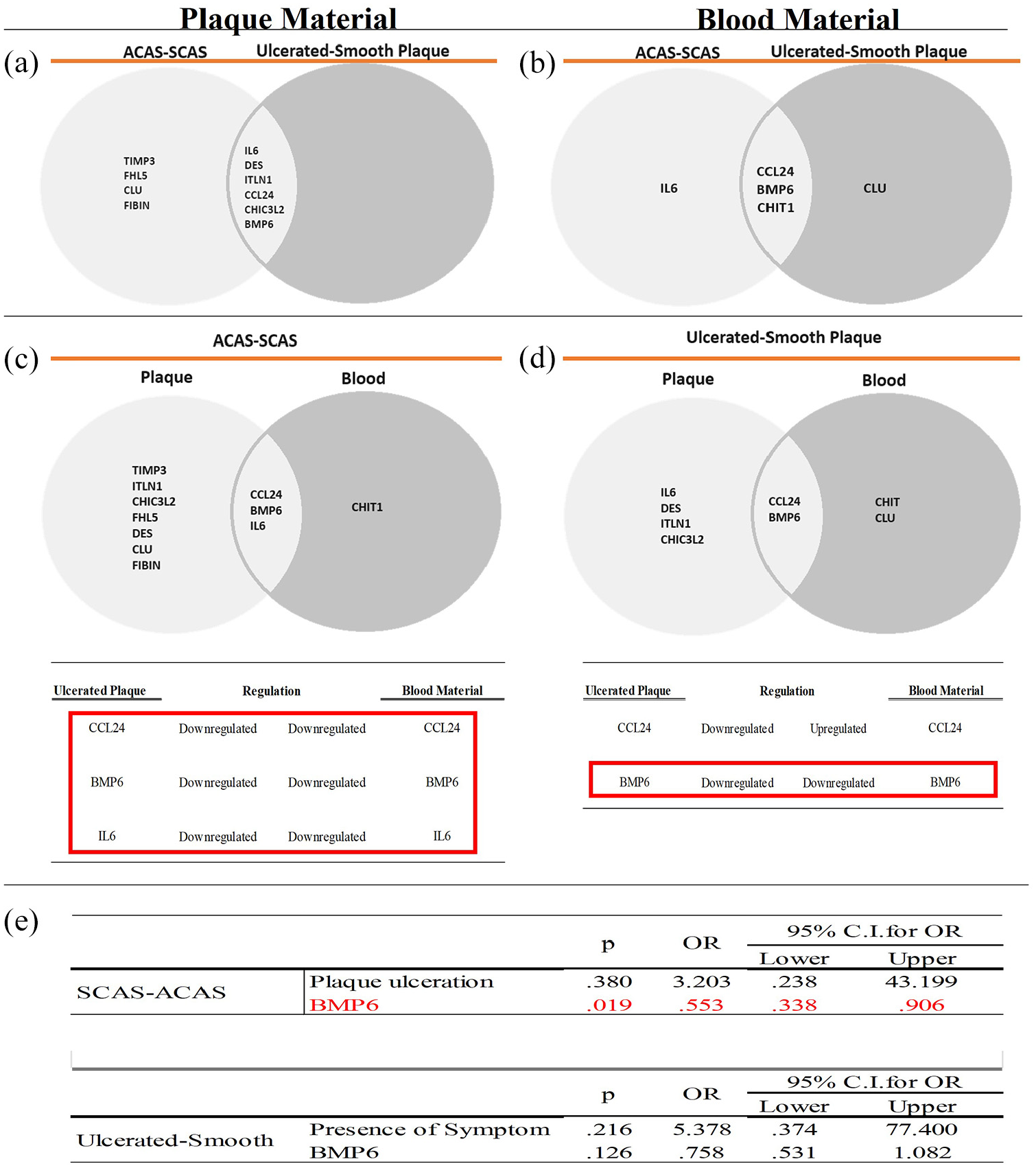
Transcriptomic analysis showed genes with common or different roles in SCAS versus ACAS patients and ulcerated versus smooth groups in (a) plaque and (b) blood materials. Analysis of statistically significant genes commonly up-/down-regulated in both plaque and blood samples to identify blood-based biomarkers (c) ACAS versus SCAS patients and (d) ulcerated versus smooth plaques. (e) Multivariate logistic regression model for BMP6 with CAS patients’ clinical features in ACAS versus SCAS, and ulcerated versus smooth plaque comparisons (statistically insignificant variables were removed from the model).
The genes and their regulations common in blood and plaque materials in the discrimination between ACAS versus SCAS patients, and ulcerated versus smooth plaques are indicated in Figure 4(c) and (d). While TIMP3, DES, ITLN, CHI3L2, FHL5, CLU, and FIBIN genes were statistically significant only in plaque materials, CCL24, BMP6, and IL6 were also significant at the blood level (Figure 4(c)). Interestingly, these three genes were found to be downregulated in both plaque and blood samples. For ulcerated plaque discrimination, CCL24 and BMP6 were statistically significant in both blood and plaque materials, and only the BMP6 expression level was determined to be downregulated in both plaque and blood samples.
Thus, we focused on BMP6 for validation and discussion, as it was the only transcript to meet all of the following criteria: (1) significant differential expression in plaque, (2) measurable and consistent expression change in blood, and (3) association with a key clinical feature of plaque vulnerability—ulceration. Therefore, the relative importance of BMP6 gene expression as a predictive biomarker of ACAS versus SCAS, and ulcerated versus smooth plaque discrimination was evaluated by multivariate binary logistic regression analysis (Figure 4(e)). However, the results showed that BMP6 had the most remarkable ability to predict the SCAS versus ACAS discrimination (p = 0.019), not plaque ulceration.
Consequently, BMP6 gene expression, which has a common role in SCAS versus ACAS, and plaque ulceration, has been determined as a useful biomarker for discriminating SCAS versus ACAS patients, unlike ulceration. Additionally, although further research is needed, blood levels of CHIT1 expression also have the potential to determine ulceration independent of symptoms (Figure 4(d)).
Discussion
The current study aimed to investigate a blood-based biomarker to determine the presence of ulceration and/or symptoms by analyzing plaque materials based on surface morphology using RNA-Seq in SCAS and ACAS patients. Our results suggest that BMP6 gene expression, which has a common role in SCAS versus ACAS and plaque ulceration, has been determined as an ideal biomarker for the discrimination of SCAS-ACAS, unlike ulceration.
IPA results showed that the pathways, including LXR/RXR, which regulate efflux, cholesterol absorption, transport, and immune response, were deregulated in SCAS compared to ACAS patients and ulcerated plaques compared to smooth ones. While genes including LPL, APOE, APOC2, and TIMP3 were upregulated, the IL6, CCL24, BMP6, ITLN1, CHI3L2, FHL5, DES, CLU, and FIBIN genes were downregulated in plaque materials of SCAS patients compared to ACAS. Additionally, while the APOC1 and APOC2 genes were found to be upregulated, the BMP6, IL6, DES, ITLN1, CCL24, and CHI3L2 genes were found to be downregulated in ulcerated plaques. Previous studies stated that MMP9, APOC1, and APOE genes were upregulated in atherosclerotic plaques compared to the control group. Another study has shown that LPL plays a role in plaque development, with the upregulation of mRNA levels expressed in macrophage-derived foam cells.22–26 Mosquera et al. and Kim et al. indicated that inflammatory and foam macrophages have higher expression of lipid genes (APOE) and lower expression of inflammatory genes.27,28 Therefore, the downregulation of genes involved in the immune response and upregulation of genes related to lipid metabolism are thought to be associated with increased foam macrophage cells in SCAS and ulcerated plaques. Additionally, the previous results indicated that BMP6 signaling enhances atherosclerotic calcification, 29 and our data show that the downregulation of BMP6 expression may be a biomarker based on reduced calcification in symptomatic/ulceration. In blood materials, based on RT-qPCR results, the upregulation of CHIT1 and the downregulation of CCL24, IL6, and BMP6 (also LPL and MMP9 protein levels) were statistically significant between SCAS versus ACAS patients. However, only CCL24, IL6, and BMP6 were downregulated in both plaque and blood materials. For ulcerated and smooth plaque comparison, the upregulation of CHIT1 and CCL24, and the downregulation of BMP6 and CLU genes were statistically significant. Only the BMP6 expression level was determined to be downregulated in both plaque and blood materials.
Chemokines play a major role in immune cell recruitment, a critical step in atherosclerosis progression. 30 Among them, the CCL24 downregulation contributes to the recruitment of neutrophils and macrophages. 31 Therefore, the results suggest that the downregulation of CCL24 in blood and plaque samples may be due to the increased accumulation of macrophages related to the increased immune response in SCAS patients. Additionally, IL6 is known to contribute to the development of atherosclerotic plaques through its receptors. 32 It has been stated that the downregulation of IL6 signaling reduces cardiovascular risk. 33 The increased levels of IL6 secreted from lipid-laden macrophages could create a protective response from macrophages to prevent cholesterol accumulation. 34 Thus, our findings show that IL6 levels are decreased in plaque and blood materials of SCAS patients, suggesting that the protective effect of IL6 on lipid-laden macrophages could be reduced in patients. Although the study was based on the blood-based biomarker evaluation of plaque and blood material expressions, there are also interesting gene expression profiles. Chitinases that participate in immunity by medium-sized chitin fragment production are responsible for immunomodulation, switching on. CHIT1 is secreted by activated macrophages, which are responsible for modulating cytokine expression, and its levels are increased in atherosclerotic plaques and patient sera. 35 It has been reported that chitinase activity increases up to 55-fold in atherosclerotic tissue and is related to the severity of atherosclerotic lesions. 36 For ulcerated and smooth plaques and SCAS versus ACAS comparison (in ulcerated and SCAS patients), upregulation of CHIT1 may be associated with increased immune response. Another gene, CLU/Apolipoprotein J, is a chaperone-like protein that exhibits characteristics of innate immunity receptors. A previous study indicated that the distribution of CLU in the aortic wall may increase with the progression of atherosclerosis and may protect against lipid peroxidation. 37 The decrease in CLU expression in blood samples of patients with ulcerated plaque supports the decrease in ulcerated plaque associated with vulnerable features prone to rupture due to the decreased protective effect. Thus, the roles of CHIT1 and CLU genes in the discrimination/progression of ulcerative plaques or their usability as biomarkers need further investigation.
BMPs are members of the transforming growth factor-β pathway and play important roles in organogenesis. BMP signaling contributes to vascular remodeling by determining the fate of vascular progenitors. 29 BMP6 also increases in human atherosclerotic and calcific vascular lesions and regulates cell differentiation and vascular calcification. 38 Studies have found that there was a higher plaque calcification content in ACAS patients compared to SCAS, and rupture is inversely proportional to the number of large calcifications. Furthermore, carotid plaque calcification has been reported to be a protective feature of biomechanical plaque stability.39,40
In our study, multivariate binary logistic regression analysis showed that BMP6 had the most remarkable ability to predict the discrimination of SCAS versus ACAS patients. However, BMP6 expression was not statistically significant for predicting plaque ulceration. Decreased BMP6 expression in the plaques of SCAS patients suggests that there may be reduced calcification in the plaques of SCAS patients, which is associated with a risk of rupture. Moreover, BMP6 expression was not a statistically significant predictor of plaque ulceration or SCAS versus ACAS discrimination at the protein level. These results indicated that there might be post-transcriptional modification to regulate the expression of BMP6 between SCAS versus ACAS and ulcerated versus smooth plaque patients, as reported in the previous study. 41 Thus, unlike protein levels, the change in the regulation of BMP6 at the RNA level suggests that post-transcriptional or epigenetic mechanisms may have played a role in this process. 42
Although statistical results showed that BMP6 could be a blood-based biomarker for the distinction between SCAS and ACAS patients, the study also suggested the following: (i) IL6, DES, ITLN1, CCL24, and CHI3L2 genes play a role in the progression of ulceration in plaque materials. (ii) The presence of the same genes that play a role in ulceration progression presence of symptoms in plaque was confirmed, and TIMP3, FHL5, CLU, and FIBIN genes were regulated only in the presence of symptoms. (iii) In addition to the symptoms where CCL24, BMP6, and IL6 can be evaluated as blood-based biomarkers in the discrimination between SCAS and ACAS patients, CHIT1 may also play a role in the formation of symptoms. (iv) In addition to the genes identified in plaques, CHIT1 and CLU may also play a role in the discrimination between ulcerated and smooth plaques. Finally, for SCAS patients with ulcerated plaques, considered a risk group, the roles of CCL24 and IL6, as well as BMP6, which are downregulated in both blood and plaque samples, should be evaluated in further studies.
This study acknowledges its limitations regarding the need for larger patient populations and the limited number of proteins evaluated to confirm the relationship between expression levels and symptoms/plaque morphology. Moreover, a second limitation is the longitudinal follow-up, which would reveal whether the transcriptomic (especially BMP6 levels) and serological profiles identified at baseline, particularly those associated with plaque morphology and symptom status, have predictive utility for long-term outcomes, including restenosis, stroke recurrence, or cardiovascular events. However, this is the first study to evaluate blood-based biomarkers based on plaque biology to investigate symptoms and ulceration associated with cardiovascular risk at the mRNA and protein levels and to identify risk groups. Therefore, based on the results, future research is needed to evaluate the roles of statistically significant genes (IL6, CCL24, CLU), especially BMP6, in the presence of symptoms and ulceration, and their relationship with cardioembolic/neurological risk.
Conclusions
Consequently, BMP6 gene expression, which has a common role in CAS and plaque ulceration, has been identified as a new biomarker for the discrimination of SCAS versus ACAS patients. Importantly, recent advances in molecular diagnostics have made RNA-based biomarker testing increasingly feasible in clinical settings. RT-qPCR and multiplexed transcriptomic panels are widely used in oncology, infectious diseases, and cardiovascular risk profiling. These platforms are becoming more cost-effective, standardized, and suitable for integration into routine clinical workflows. Given that BMP6 expression changes are detectable at the mRNA level in blood, its inclusion in a future blood-based transcriptomic biomarker panel may allow for practical, non-invasive detection of high-risk patients. RNA biomarkers like BMP6 represent a promising new direction, particularly when protein-level changes are minimal or variable. Thus, while further studies should define the roles of BMP6 in symptoms and plaque biology in larger groups, the potential of using CCL24 and IL6, which are downregulated in both blood and plaque samples, should also be evaluated as blood-based biomarkers for SCAS patients with ulcerated plaques.
Supplemental Material
sj-docx-1-imj-10.1177_10815589251361364 – Supplemental material for BMP6 contributes as a blood-based biomarker to clinical discrimination of patients with carotid artery stenosis: Decoding the transcriptome of plaque biology
Supplemental material, sj-docx-1-imj-10.1177_10815589251361364 for BMP6 contributes as a blood-based biomarker to clinical discrimination of patients with carotid artery stenosis: Decoding the transcriptome of plaque biology by Atif Yolgosteren, Ceyda Colakoglu Bergel, Isil Ezgi Eryilmaz, Unal Egeli, Can Kosukcu, Guven Ozkaya, Basak Erdemli Gursel, Murat Bicer, Gulsah Cecener, Efe Ozoglu, Emre Sarandol and Mustafa Tok in Journal of Investigative Medicine
Footnotes
Acknowledgements
The authors acknowledge support from Aria Biotechnology and the patients who took part in this study.
Author’s note
The article has not been presented elsewhere or reviewed by any other journal. All the authors are aware of and approve the manuscript as submitted to this journal.
Author contributions
CCB, AY, and IEE have given substantial contributions to the literature search, data collection, study design, analysis of data, manuscript preparation, and review of the manuscript. UE, CK, GO, BEG, EO, and ES analyze the interpretation of the data and review the manuscript. GC, MB, and MT revised it critically. All authors read and approved the final version of the manuscript. All authors contributed equally to the manuscript and read and approved the final version of the manuscript.
Ethical considerations
This study was approved by Bursa Uludag University Medical Ethics Committee (2021-6/38) and complied with the ethical standards of the Helsinki Declaration.
Consent for publication
Not applicable.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Bursa Uludag University Scientific Research Projects Coordination Unit, TAY-2022-592.
Data availability statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
