Abstract
The immune microenvironment plays an important role in the regulation of diseases. The characterization of the cellular composition of immune cell infiltrates in diseases and respective models is a major task in pathogenesis research and diagnostics. For the assessment of immune cell populations in tissues, fluorescence-activated cell sorting (FACS) or immunohistochemistry (IHC) are the two most common techniques presently applied, but they are cost intensive, laborious, and sometimes limited by the availability of suitable antibodies. Complementary rapid qPCR-based approaches exist for the human situation but are lacking for experimental mouse models. Accordingly, we developed a robust, rapid RT-qPCR-based approach to determine and quantify the abundance of prominent immune cell populations such as T cells, helper T (Th) cells, cytotoxic T cells, Th1 cells, B cells, and macrophages in mouse tissues. The results were independently validated by the gold standards IHC and FACS in corresponding tissues and showed high concordance.
Flow cytometry or immunohistochemistry routinely used for tissue immune cell phenotyping are costly, labor intensive, and limited by the availability of suitable antibodies. For humans, rapid RT-qPCR-based approaches exist. However, they are currently lacking for mouse tissues and complicate the quantification of prominent immune cell populations.
We have developed a robust, rapid RT-qPCR-based approach to determine and quantify the abundance of prominent immune cell populations such as T cells, helper T cells, cytotoxic T cells, Th1 cells, B cells, and macrophages in mouse tissues.
We provide an independently validated, rapid, simple, and cost-effective approach for quantitative analysis of key immune cell populations in mouse tissues.
Introduction
Cell-mediated immune responses against pathogens and diseases are an important part of the host defense. Determination of the relative abundancy of different immune cell populations in tissues is mandatory to understand the functions of the immune response in different diseases. For example, immune cell phenotyping of diseased tissues can provide useful information to stratify patients for immuno- and/or chemotherapy.1,2 Moreover, T-helper cells (CD4+), Th1 cells (T-bet+), and cytotoxic T cells (CD8+) have been reported to contribute to antigen-specific immunity against pathogens.2–5 Macrophages (F4/80+) and B cells (CD79+ or CD19+) play key roles in immune responses related to the prognosis of colitis and colorectal cancer.6–8
Various techniques have been established to conduct immune cell phenotyping. Fluorescence-activated cell sorting (FACS) is widely employed to identify different phenotypes of immune cells in hemato-oncology directly from the blood. 9 Immune cell phenotyping from tissues is more laborious and, reliable quantitative results are more difficult to achieve. 10 For tissue immune cell phenotyping immunohistochemistry (IHC) is the gold standard, but is frequently limited by the availability of antibodies compatible with routine tissue specimens. 11 FACS is also applied for immune cell phenotyping from dissociated tissues but is similarly hampered by the availability of compatible antibodies, associated with high costs and critically dependent on the successful dissociation of the tissue and preservation of antigens.12,13 Some reports described reverse transcription-quantitative polymerase chain reaction (RT-qPCR) protocols that were used to quantify various immune cells infiltrating human tissues.14,15 However, validated RT-qPCR-based immune cell phenotyping methods in mouse tissues are still lacking. To close this gap, we established a rapid RT-qPCR-based protocol to determine the frequency, distribution, and phenotype of immune cells in mouse tissues and validated this by corresponding IHC stainings and FACS measurements.
Materials and methods
Mouse tissue preparation
The animal use for all experiments was reviewed and approved by the government of Middle Franconia according to the applicable legal guidelines.
IHC/qPCR correlation
C57BL/6 mice (n = 12) were treated with 2.5% dextran sulfate sodium (MP Biomedicals, #160110) in the drinking water for 1 week, 16 and mice were sacrificed on day 9 or 10. The spleen, thymus, mesenteric lymph nodes (mLN), and colon were harvested from each animal. All organs were divided into two parts for RNA extraction and were correspondingly embedded in formalin-fixed paraffin-embedded (FFPE)-blocks for IHC. Tissues for IHC were fixed in 10% formalin solution (Sigma Aldrich, #HT501128-4L) at 4°C for 18 h. Then, the tissues were embedded in paraffin for further processing. Tissues used for RT-qPCR were immediately frozen in liquid nitrogen and stored at −80°C until they were further processed.
FACS/qPCR correlation
C57BL/6 mice were either untreated (n = 4) or immunized (n = 4) by applying a mixture of acetone, olive oil, and trinitrobenzenesulfonic acid (TNBS) to their abdomen. On day 5, the mice were challenged by exposing them to a mixture of ethanol and TNBS, by catheter insertion in the colon. The well-being of the mice was monitored via weight and endoscopic measurements. The mice were sacrificed 2 days after TNBS challenge and tissue samples were collected for FACS and RT-qPCR analysis.
Immunohistochemistry
Tissue blocks were cut into 4 µm sections, dewaxed using xylol and rehydrated in ethanol (100%–70%). Tissues were subjected to antigen retrieval (pH 9, DakoCytomation, #S2367) in a 98°C water bath for 30 min and cooled down at RT for 30 min. Next, the tissue slides were incubated in 7.5% H2O2 for 10 min (Sigma Aldrich, #1.0721.0250), followed by using the Avidin/Biotin Blocking Kit (Vectorlabs, #VEC-SP-2001), and diluted normal goat serum for 20 min (Vectorlabs, #PK-6101). The following primary antibodies were used: rabbit anti-mouse CD3 (1:2000; Abcam #ab5690), rabbit anti-mouse CD4 (1:2000; Abcam #ab183685), rabbit anti-mouse CD8 (1:2000; Cell Signaling #98941), rabbit anti-mouse T-bet (1:2000; Cell Signaling #97135), rabbit anti-mouse F4/80 (1:2000; Cell Signaling #70076), and rabbit anti-mouse CD19 (1:2000; Cell Signaling #90176). Antibodies were diluted in Antibody Diluent (Zytomed #ZUC025-500) and incubated overnight at 4°C. Rabbit IgG was used as an isotype control at the corresponding concentrations (R&D Systems #AB-105-C). The primary antibodies were detected using the Rabbit Vectastain Elite ABC-Kit (Vectorlabs #PK-6101) according to the manufacturer’s instructions. Substrate (NovaRED Kit, Vectorlabs # SK-4800) was developed for 12 min. Slides were counterstained with hematoxylin (Merck # 1.05174.0500) for 1 min, destained in tap water for 5 min and mounted with mounting medium (Vectorlabs #H-5000). The quantification of positive cells in the obtained images was performed using ImageJ using six random areas per slide at 10× magnification.
RNA extraction
For RNA extraction, tissue was disrupted with the TissueLyzer (Qiagen) in cold RLT buffer (RNeasy Mini Kit, Qiagen #74104) with 1% β-mercaptoethanol (Sigma-Aldrich, #M6250) using a grinding ball (Retsch, #412-0249) at 30 Hz/min for 4 min followed by brief centrifugation (IHC/qPCR correlation) or immune cells were isolated as described below for FACS analysis (FACS/qPCR correlation). Afterwards, the supernatant was passed through a QIAshredder column (Qiagen, #79654). The eluate was precipitated using 70% EtOH, passed into a mini column (Qiagen, #74104), and washed using RW1 and RPE buffer, followed by elution using ribonuclease (RNase)-free water and deoxyribonuclease (DNase) I digest (1 U for 30 min, Ambion, #AM2222). Glycogen purification was conducted with a final glycogen concentration of 0.108 µg/µl (Thermo Scientific, #R0551) diluted in RNase-free water. The RNA eluates were precipitated overnight with 110 µl isopropanol (Merck, #1.09634.2511). Pure RNA was obtained by centrifuging, washing with 70% EtOH and resuspending in RNase-free water. Integrity and quantity were assessed by agarose gel electrophoresis and measurement using a Nanodrop 2000 (Thermo Scientific).
Primer design and quality control
Primers were designed using PrimerBLAST (NCBI). The primer parameters applied were as follows: product size: 50–300 bp; melting temperatures: 60–63°C, optimal 61°C, max Tm difference: 1; intron inclusion; intron length range: 2000–10,000 bp; allow splice variants; GC content (%): 40%–50%. Primers were ordered from Eurofins Genomics. Serial dilutions of cDNA from positive tissue samples were used to test primer efficiency (Table 1).
Primer pairs for quantitative immune cell phenotyping in mouse tissues with RT-qPCR.
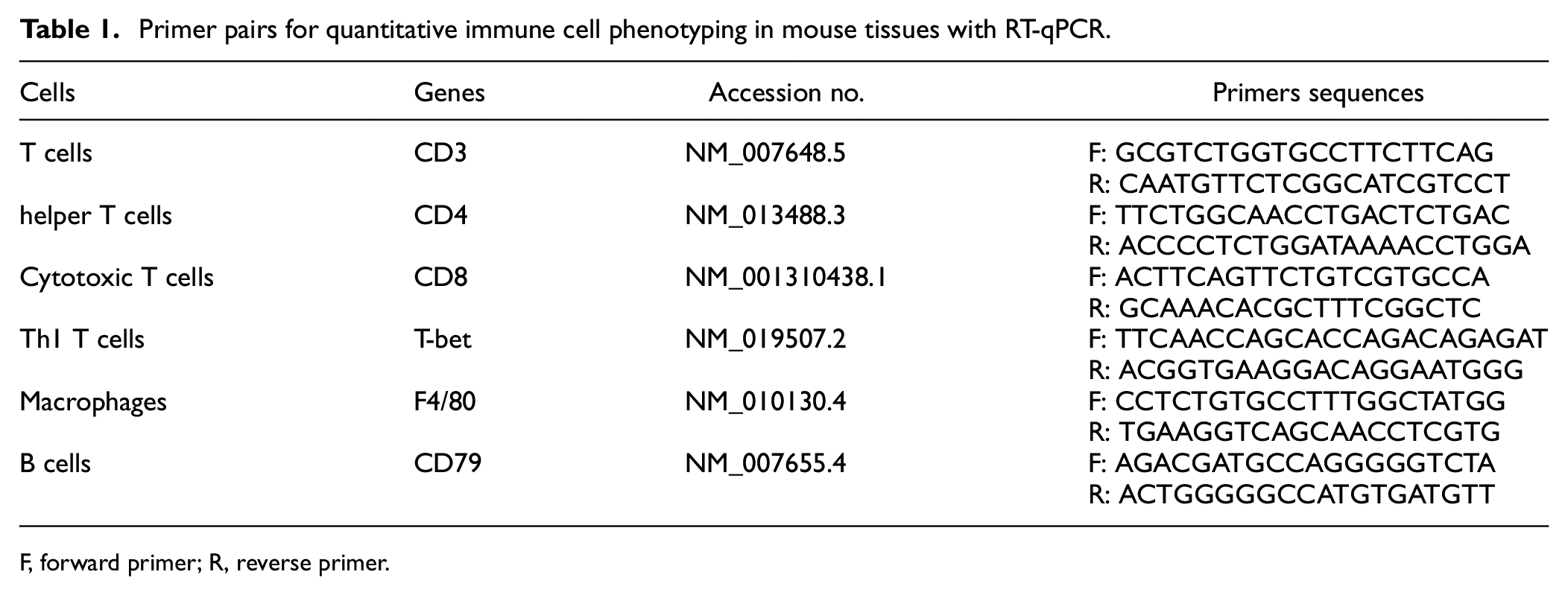
F, forward primer; R, reverse primer.
Immune phenotyping by quantitative real-time PCR
RNA (5 µg) was transcribed to cDNA using the SuperScript III First-Strand Synthesis System (Thermo Fisher, #18080-051) following the instructions of the manufacturer. Quantitative RT-PCR was carried out on a Bio-Rad CFX-Connect machine using RT-qPCR MasterMix Plus for SYBR Assay ROX (Eurogentec, #RT-SN2X-03+). Reactions were run in triplicate using a two-step protocol (2 min at 50°C, 10 min at 95°C, 39 cycles of 15 s at 94°C and 1 min at 60°C). Expression of the target genes was normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH). Differences between samples were determined by the 40-ΔCt method. Statistical significance was determined by using the unpaired two-tailed t-test. The results were visualized with GraphPad Prism.
Fluorescence-activated cell sorting
For immune cell isolation from the mLN, spleen, and thymus, tissue samples were collected and dissociated between glass slides. The cells were washed once and strained to recover the leukocytes. The splenocytes were lysed (ACK buffer) to remove the red blood cells before filtering through strainers. For leukocyte isolation from colon tissue samples, commercially available kits were used (Miltenyi Biotec, lamina propria isolation kit). Once cells were isolated, a fraction of cells was transferred to FACS tubes and washed once with FACS buffer (phosphate-buffered saline [PBS] + 1% FBS). The cells were then resuspended in 100 µl FACS buffer with α-CD16/α-CD32 and incubated at 4°C for 10 min, to block Fc cell receptors. For cellular staining, a master mix with the antibodies of interest was prepared. The cellular antibodies (CD4 Super Bright 600, eBioscience, #63-0042-82, 0.125 µg/100 µg sample; CD8a BV510, BD Bioscience, #563068, 1:100; F4/80 PE, Invitrogen, #12-4801-82, 0.125 µg/100 µl sample; B220 VioBlue, Miltenyi Biotec, #130-110-851, 1:50) were then added to the cellular suspensions and the samples were incubated for 20 min at RT in the dark. Subsequently, the cells were washed with FACS buffer, resuspended in FACS buffer, and measured (BD LSR Fortessa, BD Bioscience).
Results
RT-qPCR-based immune cell phenotyping is more rapid and broadly applicable than classical IHC and FACS
RT-qPCR is usually a more rapid approach than IHC or FACS. The standard IHC workflow includes six different steps: tissue collection, fixation and embedding of tissues, sectioning, staining, imaging, and quantification of positive cells. The FACS workflow consists of a similar number of steps which include tissue collection, tissue dissociation, cell isolation, staining, FACS measurement, and quantification of positive cells. In contrast, the typical RT-qPCR workflow consists of only three steps which include tissue collection, RNA extraction, and qPCR analysis (Figure 1).
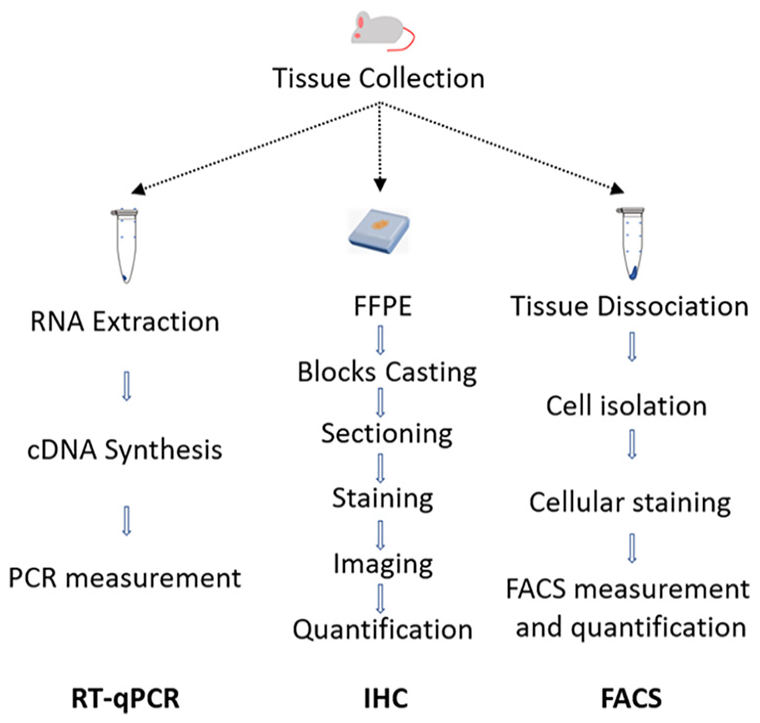
RT-qPCR-based immune cell phenotyping is more rapid than classical IHC or FACS. The IHC workflow consists of six steps: tissue collection, generation of FFPE blocks, sectioning, staining, imaging, and quantification. The FACS workflow consists of six steps: tissue collection, tissue dissociation, cell isolation, staining, FACS measurement, and quantification of positive cells. The qPCR workflow consists of three steps: tissue collection, RNA extraction, and PCR measurement.
Quantitative detection of different T-cell subpopulations, B cells, and macrophages by RT-qPCR in murine tissues
The aim of this study was to establish a rapid and easy RT-qPCR-based method for quantitative determination of the major immune cell populations in mouse tissues. To this end, tissues from mice with prominent infiltration of different immune cells in different organs such as spleens, thymuses, lymph nodes, and colons after induction of colitis were comparatively analyzed by RT-qPCR, and the results were validated by classical IHC and FACS.
RT-qPCR primer pairs were designed for well accepted marker genes known to be specific for the cell type of interest and with minimal expression in other cell types. Accordingly, the markers used to detect the target immune cell populations were as follows: CD3 for total T cells 17 CD4 for T helper cells,18,19 CD8 for cytotoxic T cells,18,19 T-bet for Th1 cells, 20 CD79 for B cells, 21 and F4/80 for macrophages 22 (Table 1). Next, tissues from different organs such as the spleen, thymus, lymph nodes, and colon with either high, medium or low infiltration of T cells and subpopulations were chosen as target tissues for the development and validation of the method. Quantitative analyses of RT-qPCR (Figure 2(a)–(d), upper left panels), and IHC (Figure 2(a)–(d), lower left panels) showed that the results obtained with each method correlated strongly for the detection of total T cells using CD3 (Figure 2(a)), cytotoxic T cells using CD8 (Figure 2(b)), T helper cells using CD4 (Figure 2(c)), and Th1 cells using T-bet (Figure 2(d)) as marker molecules. The same analyses were conducted for macrophages using F4/80 (Figure 3(a)) as well as for B cells using CD79 (RT-qPCR) and CD19 (IHC) as markers (Figure 3(b)). In all cases, the results obtained with both methods were significantly and correlated highly as determined by regression analyses (Figure 4).
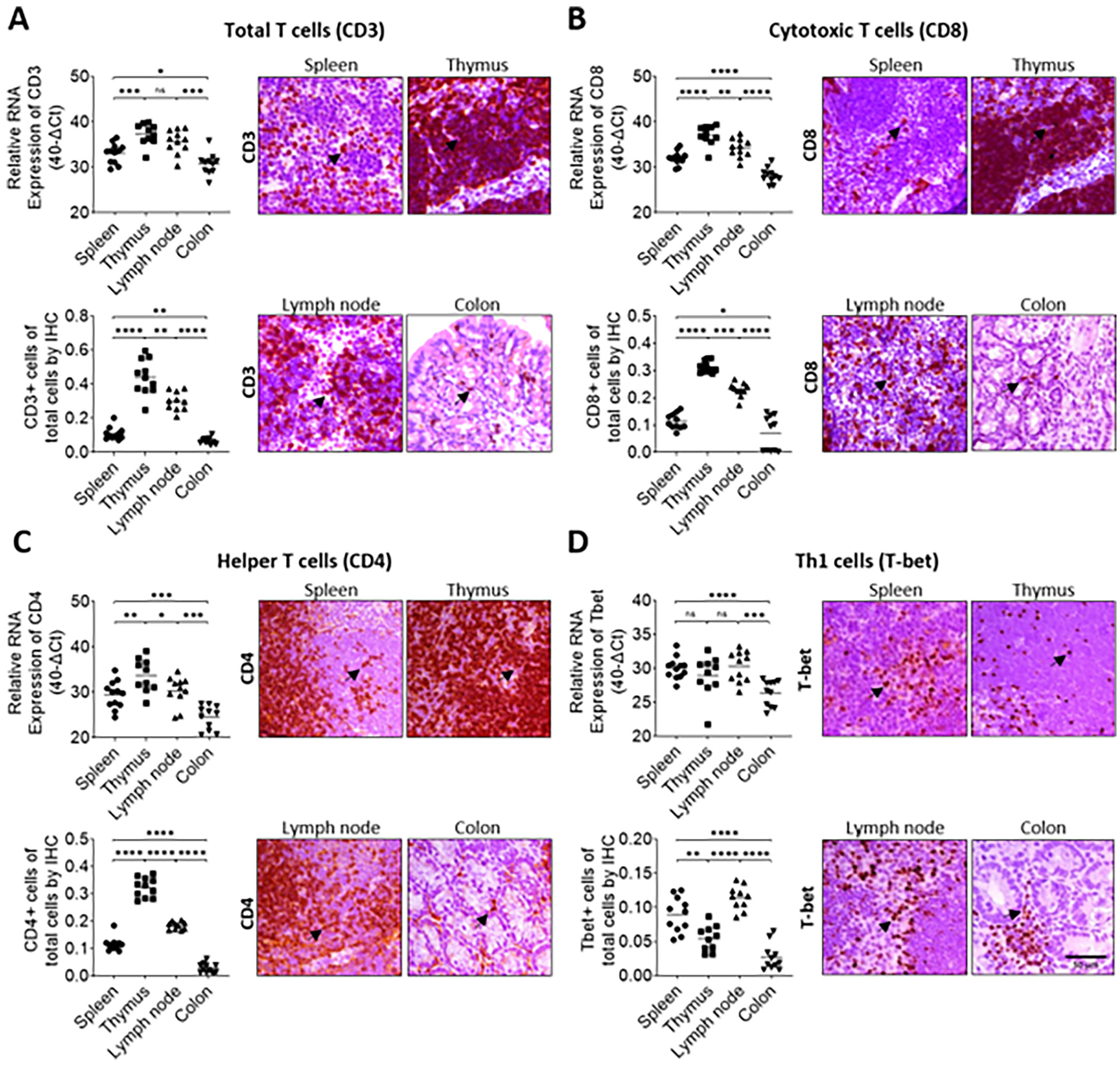
Total T cells, cytotoxic T cells, helper T cells, and Th1 cells can be quantitatively determined in mouse tissues by RT-qPCR as validated by corresponding IHC. Spleens (n = 12), lymph nodes (n = 11), thymuses (n = 10), and colons (n = 11) from mice were analyzed by corresponding RT-qPCR (upper left panels) and IHC (right panels, arrows = positive cells) to quantify the abundance of (a) total T cells (CD3+), (b) cytotoxic T cells (CD8+), (c) helper T cells (CD4+), and (d) Th1 cells (T-bet+) using the characteristic markers indicated in the brackets. Quantitative evaluation of the IHC results is shown in the lower left panels. Statistical significance was determined by Student’s t-test. Lines indicate the mean value. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Scale bar represents 50 µm.
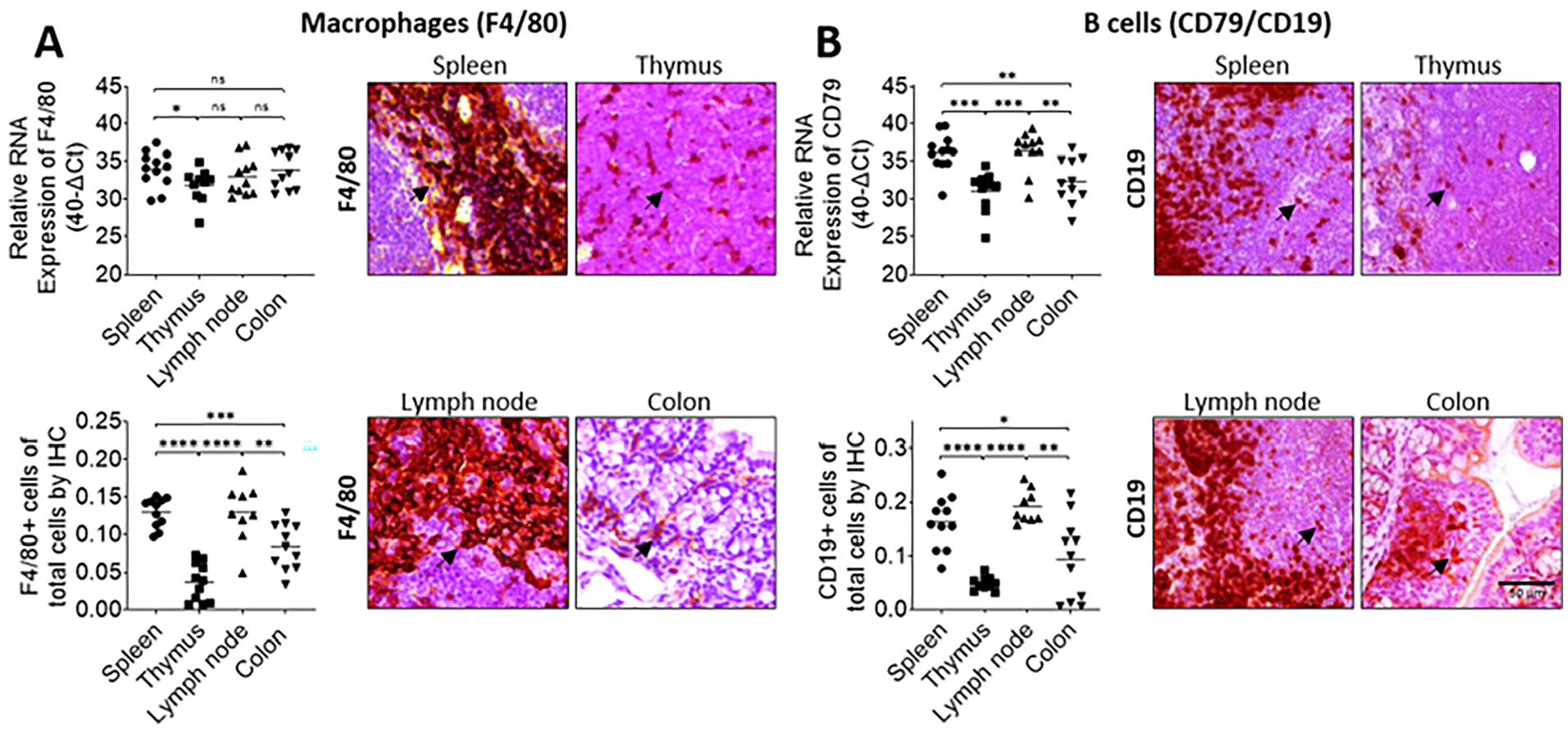
Macrophages and B-cells can be quantitatively determined in mouse tissues by RT-qPCR as validated by corresponding IHC. spleens (n = 12), lymph nodes (n = 11), thymuses (n = 10), and colons (n = 11) from mice were analyzed by corresponding RT-qPCR (upper left panels) and IHC (right panels, arrows = positive cells) to quantify the abundance of (a) macrophages (F4/80+), and (b) B-cells (RT-qPCR: CD79+; IHC: CD19+) using the characteristic markers indicated in the brackets. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001; quantitative evaluation of the IHC results is shown in the lower left panels. Statistical significance was determined by Student’s t-test. Lines indicate the mean value. Scale bar represents 50 µm.
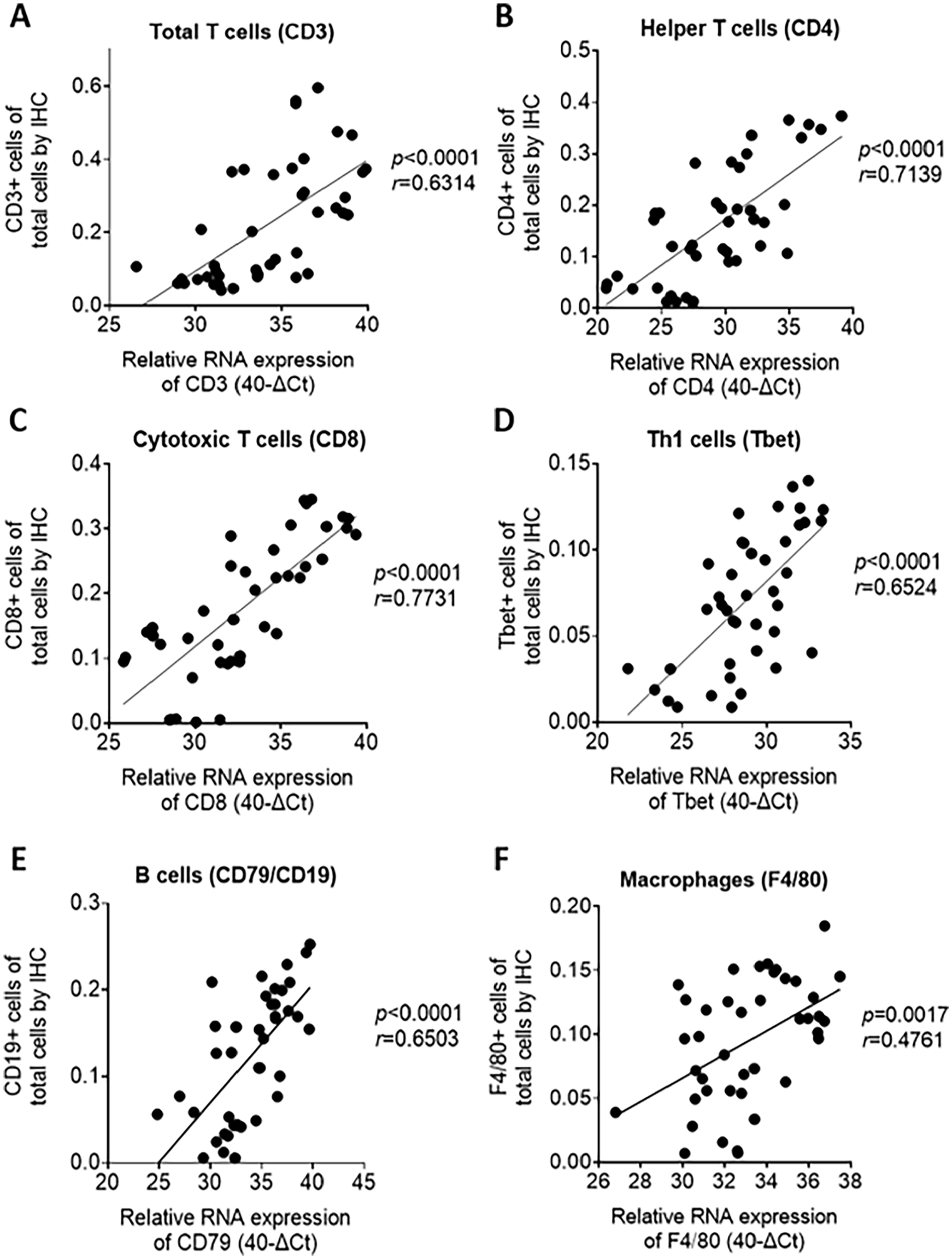
RT-qPCR-based immune cell phenotyping correlates with corresponding IHC analysis in mouse tissues. Correlation plots comparing the corresponding RT-qPCR and IHC results for (a) CD3, (b) CD4, (c) CD8, (d) T-bet, (e) CD79/CD19, and (f) F4/80 analysis.
Moreover, a correlation analysis of the qPCR approach with classical FACS was performed in parallel. Helper T cells (CD4+), cytotoxic T cells (CD8+), macrophages (F4/80+), and B cells (B220+) were analyzed by FACS in different organs of the mice. The obtained results revealed a significant positive correlation with the respective qPCR measurements of the same samples (Figure 5). Of note, primers for regulatory natural killer (NK) cells (NKp46+), M1 macrophages (CD86+), Th2 cells (Gata3+), and Th17 cells (Rorc+) were also developed, but corresponding FFPE-compatible antibodies to validate the RT-qPCR results were not available. Accordingly, we refrained from presenting the respective RT-qPCR results here (data not shown).
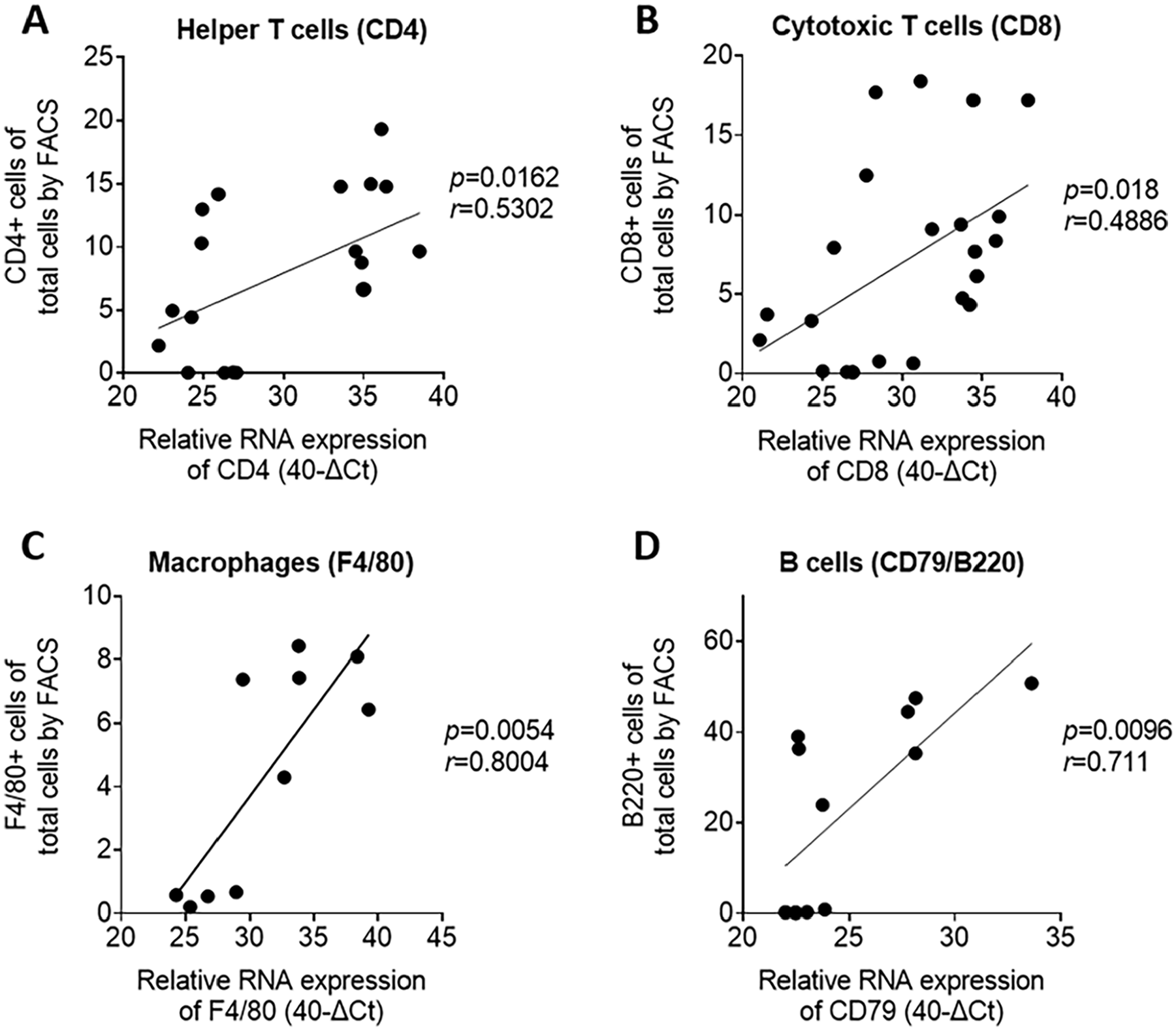
RT-qPCR-based immune cell phenotyping correlates with corresponding FACS analysis in mouse tissues. Correlation plots comparing the corresponding RT-qPCR and FACS results for (a) CD4, (b) CD8, (c) F4/80, and (d) CD79/B220 analysis.
Discussion
Immune responses regulate the pathophysiology of diseases.23,24 For example, CD3+, CD8+, CD4+, and T-bet+ cells are associated with positive responses against pathogens,3–5 whereas macrophages and B cells were reported to be associated with anti-immune responses.6–8,25 Therefore, there is a constant demand for methods allowing rapid and easy quantitative analyses of immune cells in tissues.
IHC is the most commonly used and gold standard method for this task. 11 IHC staining provides loco-regional information about immune cells by detecting characteristic antigens of distinct immune cells (Figures 2 and 3), but the disadvantages of IHC are obvious in terms of time and cost. Moreover, specifically for tissues not of human origin, IHC-compatible antibodies are frequently not available. This is specifically a problem for the analyses of mouse tissues, which are the most commonly used in vivo model systems of human diseases. 26 Moreover, representative quantification with IHC requires staining of multiple sections of a tissue block. The latter is a specific drawback when material is limited. In addition, unspecific background signals can vary between different tissue types, and accordingly the IHC protocols have to be adjusted to each tissue and each target. In contrast, RNA is readily available, and specific primers can be designed for almost any target. The quantification is less time-consuming and can be applied to the whole tissue as a batch method instead of single sections to be sampled and representing only small parts of the whole target tissue. Moreover, nucleic acids are a uniform target allowing the method to be applied to any tissue without modification, reducing bias and background problems. Of note, potential degradation of RNA is a drawback of the RT-qPCR method as well as the requirement of unique and specific cell markers to detect the target cell. 27
Other alternatives, such as FACS, and omics approaches such as bulk RNAseq or scRNAseq partly share the same disadvantages as IHC.28–30 For example, FACS is widely used in many fields 9 but is also laborious and cost-intensive to phenotype cells from tissues. Transcriptome approaches, such as bulk RNAseq or scRNAseq, are gaining attention in the analysis of tumor-infiltrating immune cells 1 but are cost-intensive for high throughput studies and require specific instruments and expertise. Accordingly, the rapid and inexpensive RT-qPCR protocol described here may be used to validate the reliability of such omics approaches. 31 Our rapid qPCR-based method allows specific phenotyping of T cells and subpopulations as well as B cells and macrophages from bulk tissues. The process is fast, requiring only three processing steps, including RNA extraction, cDNA synthesis and PCR. Furthermore, stable cDNA material from a small piece of tissue is sufficient for a large number of subsequent PCR tests. 32 Moreover, RT-qPCR is a broadly available technique well established in many laboratories worldwide,33,34 and specific primers can be designed for almost every target. Here, a panel of commonly used markers was exploited as targets, including CD3, CD4, CD8, and T-bet, for the identification of total T cells, T helper cells, cytotoxic T cells, and Th1 cells.18,19 In addition, F4/80 was used to identify murine macrophages, and CD79 (RT-qPCR) is a specific marker for B cells.21,22,35 In addition to the cell types mentioned above, other immune cell types are also relevant for different diseases.36–41
Of note, the described PCR analysis was developed to obtain a rapid overview of the general cellular composition of the immune microenvironment. The method was not designed to discriminate accurately between different functional states or activation profiles of the different immune cell populations. Moreover, we succeeded in developing amplification primers for murine NK cells (NKp46+), M1 macrophages (CD86+), Th2 cells (Gata3+), and Th17 cells (Rorc+) cells, but IHC-compatible antibodies to validate the RT-qPCR results in mouse tissues are currently not available. Accordingly, we refrained from presenting the results here. In summary, we provide a rapid, simple, and cost-effective approach to quantitatively analyze major immune cell populations in mouse tissues.
Footnotes
Acknowledgements
We thank K. Petter, C. Flierl, and T. Gass for technical help and N. Britzen-Laurent for administrative help.
Author contributions
EN designed the experiments; JH, MMA, and OMT performed the experiments; JH, RD, OMT, MJW, MS, and EN analyzed data; BW contributed data; JH and RD wrote the initial draft of the manuscript that was finalized by MS and EN. All of the authors approved the final manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work of the authors was supported by grants from the German Research Foundation (DFG) FOR 2438 (subproject 2 to EN and MS, project number: 280163318); SFB/TRR 241 (subproject A06 to MS, project number: 375876048), STU 238/10-1 (to MS, project number: 437201724), TRR 305 (subproject B08 to EN, project number: 429280966); the Interdisciplinary Center for Clinical Research (IZKF) of the Clinical Center Erlangen (to MS); the W. Lutz Stiftung (to MS), and the Forschungsstiftung Medizin am Universitätsklinikum Erlangen (to MS).
