Abstract
Harnessing intelligence from brain cells in vitro requires a multidisciplinary approach integrating wetware, hardware, and software. Wetware comprises the in vitro brain cells themselves, where differentiation from induced pluripotent stem cells offers ethical scalability; hardware typically involves a life support system and a setup to record the activity from and deliver stimulation to the brain cells; and software is required to control the hardware and process the signals coming from and going to the brain cells. This review provides a broad summary of the foundational technologies underpinning these components, along with outlining the importance of technology integration. Of particular importance is that this new technology offers the ability to extend beyond traditional methods that assess primarily the survival and spontaneous activity of neural cultures. Instead, the focus returns to the core function of neural tissue: the neurocomputational ability to process information and respond accordingly. Therefore, this review also covers work that, despite the relatively early state of current technology, has provided novel and meaningful understandings in the field of neuroscience along with opening exciting avenues for future research.
Keywords
Introduction
Biological brains exist as the most data- and energy-efficient information-processing systems currently known. To date, these complex neural systems are the only identified platform to produce generalized intelligence capabilities. Exploring and even harnessing neural systems in vitro would offer a powerful model to understand how complex features such as intelligence arise from their cellular basis and may even be leveraged as a novel information-processing platform with advanced capabilities. Yet, harnessing intelligence from brain cells in vitro requires a multidisciplinary approach integrating wetware, hardware, and software—previously termed synthetic biological intelligence (SBI; Kagan, Gyngell, and others 2023). Wetware comprises the in vitro brain cells themselves. Early work in this area almost exclusively used primary sources of neural tissue, typically from rodents (Bakkum, Chao Zenas, and Potter 2008b; Jimbo and others 1998; Shahaf and Marom 2001; Tessadori and others 2012). More recently, however, generating neural cells via synthetic biology methods from pluripotent stem cells (PSCs) has been positioned as a more controllable, customizable, and scalable approach: features vital for the successful expansion of the technology (Kagan, Gyngell, and others 2023). Hardware typically involves a life support system and a setup to record the activity from and deliver stimulation to the brain cells. Most commonly, hardware involves microelectrode arrays (MEAs; also referred to as multielectrode arrays) (Mierzejewski and others 2020), although optical and chemical approaches have been explored. Finally, software is required to control the hardware and process the signals coming from and going to the brain cells. Ideally, the software can function in a real-time closed-loop manner, but open-loop and reservoir computing approaches have also been adopted. Perhaps because of the challenge of integrating these three essential components, relatively few peer review studies have developed systems capable of showing meaningful information-processing capabilities of in vitro neural tissues, even in technically simpler open-loop paradigms (Cai and others 2023; Isomura and others 2015; Sumi and others 2023). Work demonstrating learning in closed-loop systems has likewise been limited (Bakkum, Chao Zenas, and Potter 2008a; Reger and others 2000; Tessadori and others 2012), with fewer publications reporting rigorous real-time closed-loop systems (Kagan, Kitchen, and others 2022). This stands in sharp contrast with the decades-long endeavors of developing each technology separately, lacking a holistic approach (Servais and others 2024). Yet, as accessibility to software and hardware platforms to support closed-loop systems grows (Elyahoodayan and others 2019; Jordan and others 2024; Martinoia and others 2004; Müller and others 2013; Newman and others 2013; Zhang and others 2024), coupled with increasing synthetic biology and bioengineering capabilities (Tanveer and others 2024), it will become increasingly important to understand the interrelated component aspects of this technology. Therefore, this review aims to provide a contemporaneous high-level overview of the foundational technologies for eliciting intelligence from biological neurons.
Need for a Scalable Production of Neural Cells
Early demonstrations of in vitro neural plasticity were achieved in cultures of primary neurons taken from embryonic rodents (Jimbo and others 1998; Shahaf and Marom 2001). Some studies did attempt to use ex vivo tissue-mounted approaches, but the poor scalability of these methods appears to have limited result interpretability (Reger and others 2000). Improvements to methods of primary neural culturing (Potter and DeMarse 2001; Wagenaar and others 2004) did result in meaningful explorations of neural plasticity and even early learning protocols in primary rodent neurons, mostly cortical cultures (Bakkum, Chao Zenas, and Potter 2008a; Bakkum, Chao Zenas, and Potter 2008b; Chao and others 2007; Tessadori and others 2012) but also hippocampal cultures (Brewer and others 2013; Pimashkin and others 2013). Despite some success with these approaches, primary cultures have some key limitations. First, the life span of these cultures is at best limited to months (Potter and DeMarse 2001) but typically weeks (Björklund and others 2010; Kaech and Banker 2006; Lesuisse and Martin 2002). Second, it is ethically difficult to justify the use of a culture that involves killing animals to generate cells when alternatives exist (Lossi and Merighi 2018). Moreover, animal primary cultures do not show the same properties as human neurons, and while human primary culture techniques exist (Schukking and others 2018), this is only linearly scalable. Finally, while some neural subtype specificity is possible with primary cultures, it is technically challenging, difficult to consistently reproduce—especially between research groups—and ultimately limited. Fortunately, the ability to differentiate PSCs into neural cells provides an attractive alternative pathway to generate the wetware component (Buckberry and others 2023; Hergenreder and others 2024). In particular, advances in stem cell engineering have allowed for PSCs to be efficiently differentiated into active neurons that display mature functional properties. While other stem cell sources may be used, particularly for nonhuman cell lines, PSCs from a diverse range of human genotypes will enable early applications of SBI technology that focus heavily on improving disease modeling and drug testing (Kagan, Gyngell, and others 2023; Smirnova 2024). The generation of neurons from human-induced PSCs, a stem cell line typically generated from the blood or skin of adult donors, provides an ethically sound, scalable, and specific production required for SBI.
Differentiation of Neural Tissue from PSCs
In vitro generation of neural tissues from human iPSCs that mimic the stepwise processes of neurogenesis by adding extrinsic factors—usually cocktails of small molecule inhibitors, activators, and growth factors—at a specific time point is time-consuming. For example, PSCs can be differentiated into neural stem cells through the inhibition of critical pathways such as Lefty, activin, and tumor growth factor β pathways and bone morphogenetic protein signaling, which is commonly termed dual SMAD (suppressor of mothers against decapentaplegic); the inhibition of the dual SMAD pathways yields PAX6+ neuroepithelial cells capable of forming rosettes, a classical in vitro representation of the neural tube in two dimensions (Fedorova and others 2019; Hu and others 2010; Miotto and others 2023). Fibroblast growth factor signaling is crucial for rosette polarization and in the expansion and maintenance of the neuroepithelial cells, which are neural stem cells with the potential of differentiating into several neuronal lineages (e.g., dopaminergic or motor; Chang and others 2021). These neuronal lineages have positional identity, which is dependent on the timing and concentration of exposure to morphogenic factors patterning them along the rostral-caudal and dorsal-ventral axes. Sonic hedgehog signaling plays a key role in patterning these precursor cells along the ventral axis; thus, inhibiting sonic hedgehog signaling promotes dorsal patterning. Wnt signaling and retinoic acid contribute to positional patterning along the caudal-rostral axis (Barak and others 2022; Chang and others 2018). Time- and dose-dependent strategies have been used to optimize the differentiation of cortical and subcortical neurons, such as dopaminergic, hippocampal, motor, and hypothalamic (Du and others 2015; Gantner and others 2020; Gantner and others 2021; Hunt and others 2023; Niclis and others 2017).
These differentiation protocols generate a spontaneous and mixed population of neurons and nonneuronal cells, such as glial cells. Fluorescence-activated cell sorting (FACS), which utilizes cell surface markers, can be used to separate the neuronal population from the nonneuronal cells to generate pure populations (Martin and others 2017). Recent studies with single cell transcriptomics have identified various cell surface markers in developing neural stem cells that are lineage specific to aid in isolating specific populations of neuronal precursors (excitatory and inhibitory lineages) and glial progenitors (toward astrocytes, oligodendrocytes, and radial glia; Liu and others 2023). Combining these technologies allows better understanding of the developmental trajectory and spatiotemporal gene expression of neural stem cells, aiding in the optimization of the differentiation protocols. This includes optimizing the exposure time to, and defining the optimal concentrations of, small molecules needed to drive the neuronal lineage of interest.
To circumvent the lengthy stepwise process of neuronal differentiation via extrinsic factors, a directed neuronal differentiation approach has been widely adopted to rapidly generate neuronal cells for disease modeling and drug screening. This approach uses the overexpression of lineage-specific transcription factors (TFs) to directly differentiate cells to specific neuronal cell types, often bypassing the progenitor stage. The lineage-specific TFs are either transfected or transduced into cells by plasmids, synthetic mRNA, or viruses. In addition, with the advent of programmable nucleases, stable or inducible cell lines expressing these TFs have been generated (Ho and others 2016; Hulme and others 2022; Pak and others 2018).
Neurogenin 2 (NGN2) and achaete-scute homolog 1 (ASCL1) have been widely studied as master regulators of neurogenesis during central nervous system development, which specifies neuronal fate (Chouchane and Costa 2019). These TFs have been used to generate a diverse range of functional neurons from human-induced pluripotent stem cells with high reproducibility. Combining the overexpression of NGN2 with other TFs and/or extrinsic factors or small molecules has streamlined the generation of highly specific neuronal population: dopaminergic, glutamatergic, motor, peripheral, and serotonergic neurons. Yet, combining other TFs such as distal-less homeobox 2 and small molecules with ASCL1 has been used to generate GABAergic-induced neurons (MacDonald and others 2013; Yang and others 2017). However, while these protocols can be useful in rapidly generating highly specific cell types, almost pure populations of single cell types have limited physiologic relevance and may not recapitulate phenotypical neural dynamics that arise from an interaction of multiple cell types. Identifying the minimal neural subtypes that give rise to different information-processing capabilities—an effort toward a minimal viable brain—should be a key focus of future work in this space. Multiple minimal viable brains will likely exist for each set of functions of interest, but benchmarking each system will be highly informative.
Morphologic Configurations of Neural Cell Cultures
Stem cells have the intrinsic ability to not only self-renew but also self-organize into various morphology, ranging from simple layers to more complicated arrangements that mimic in vivo tissues and organs (Figure 1).
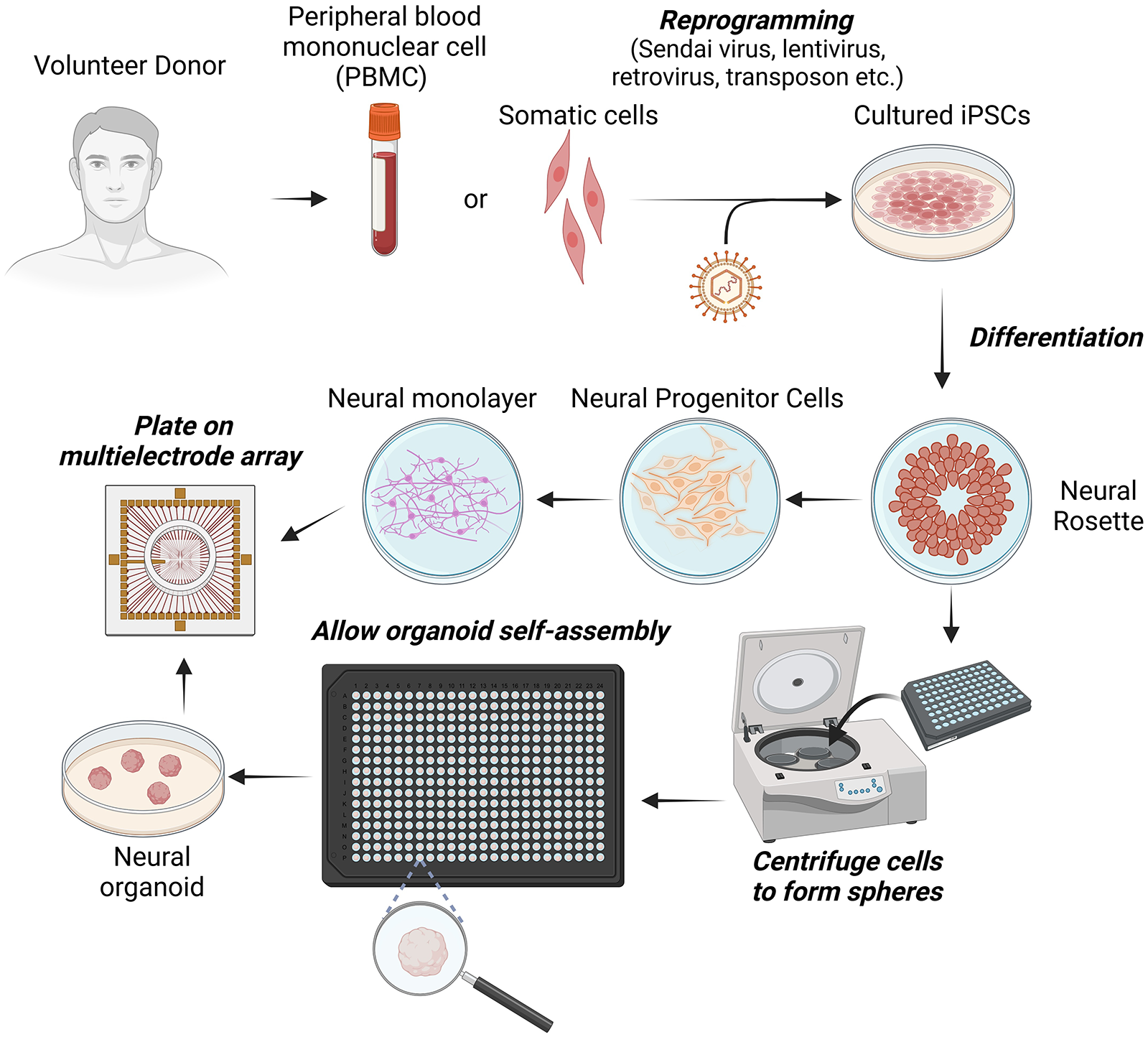
Key steps from donation of genetic material from a consenting volunteer to generation of induced pluripotent stem cells (iPSCs) and either neural monolayers or organoids for plating onto microelectrode arrays (MEA).
Broadly, neural tissue can be grown in two key configurations: as monolayers, often referred to as two-dimensional, or with more complicated structure, often referred to as three-dimensional (3D). Monolayers may be unpatterned or patterned. Structured two-dimensional cultures may be achieved via micropatterning the culture surface with key molecules that influence cell adhesion (Chen and others 1998; Gabay and others 2005; Gilles and others 2012; Klein and others 1999; Koroleva and others 2021; Macis and others 2007; Marconi and others 2012; Shein Idelson and others 2010; Shein-Idelson and others 2011; Vogt and others 2005) or through structures that constrain the location and growth of neural cells, with polydimethylsiloxane being the most commonly used biocompatible material for these structures (Duru and others 2022; Duru and others 2023; Girardin and others 2022; Girardin and others 2023; Le Feber and others 2015; Mateus and others 2024; Oiwa and others 2016; Peyrin and others 2011; Sato and others 2023; Yamamoto and others 2018). More 3D geometry of neural cultures can be achieved with polydimethylsiloxane devices and similar microfluidic approaches (Koroleva and others 2021). However, 3D cell culturing usually refers to the creation of organoids. Brain organoids are a more recent development as compared with monolayers, where the intrinsic developmental pattern of a modeled organ is targeted with a specific subset of cells or tissues (Lancaster and Knoblich 2014; Lancaster and others 2013). Neural organoids can be grown by the variety of methods previously described and can display more complex morphology and electrophysiology than unstructured monolayers (Fitzgerald and others 2024; Osaki and others 2024; Sakaguchi and others 2019). The generation of human cerebral organoids employs similar methods to that used in the aforementioned monolayers. This provides a similar capacity to induce specific neural subtypes, allowing brain organoids such as cortical, cerebellar, hippocampal, and midbrain to be generated. Interfacing multiple organoids can give rise to a structure referred to as an assembloid that displays interconnected properties and may help model interconnected but otherwise discrete neural systems (Birey and others 2017; Miura and others 2020; Reumann and others 2023). While unpatterned monolayers have been used to demonstrate neural plasticity and a responsiveness to stimulation (Bakkum, Chao Zenas, and Potter 2008a; Chao and others 2007; Hattori and others 2021; Jimbo and others 1998; Shahaf and Marom 2001), it is likely that future work will increasingly focus on either patterned monolayer cultures or an organoid approach (or some combination of both).
Long-term Neural Maintenance
The long-term maintenance of neuronal cells is crucial for understanding their behavior, functionality, and responses to external stimuli when focusing on complex phenomena such as criticality, plasticity, and learning. Given the technical challenges and cost of maintaining large quantities of complex cell cultures, macro- and microfluidic approaches have been developed. Traditional methods usually involve storing cell cultures in specialized dishes in controlled incubator-based environments. Features such as temperature and humidity, with gases such as CO2 and often O2, are strictly controlled (Yazdani 2016). To reduce the risk of contamination, cell culture media is typically removed manually by researchers using micropipettes under sterile conditions within biosafety cabinets. Membranes to reduce evaporation and contamination risk have been trialed but with little widespread adoption (Potter and DeMarse 2001). While these established methods are suitable for typical cell culturing experiments, long-term interactions and monitoring of neural cultures will be required to realize the full potential of SBI protocols. Automating the fluid exchange process via a perfusion circuit is needed to address technological and practical limitations, offering a more consistent and controlled environment for neuronal cultures. Microfluidic approaches and organ-on-a-chip models, have aimed to integrate similar approaches but still lack integrated workflows and require technical skills to operate (Alver and others 2024).
In any case, SBI protocols will likely require a larger-scale millifluidic approach, with an automated perfusion system and gas exchange. Such systems may be open loop (Figure 2A) or closed loop (Figure 2B). By implementing such a hybrid system, it is possible to reduce the manual intervention required for media exchange and provide a more physiologically relevant environment that mimics in vivo conditions. However, long-term recordings may benefit from more elaborate setups that ensure adequate filtration and more dynamic control of metabolites, including gas (Figure 2C). Recent advances in ex vivo organ perfusion systems underscore the potential of such technologies for long-term maintenance of complex tissues. For instance, a study demonstrated the use of an organ perfusion system to sustain a whole kidney for up to seven days by employing mathematical modeling to control gas exchange and maintain a self-contained environment conducive to organ preservation (Won and others 2022). Although this system was not designed for neurons, it exemplifies the potential to adapt similar strategies for neuronal tissue, where prolonged maintenance is frequently challenging due to its metabolic demands and sensitivity to environmental changes.
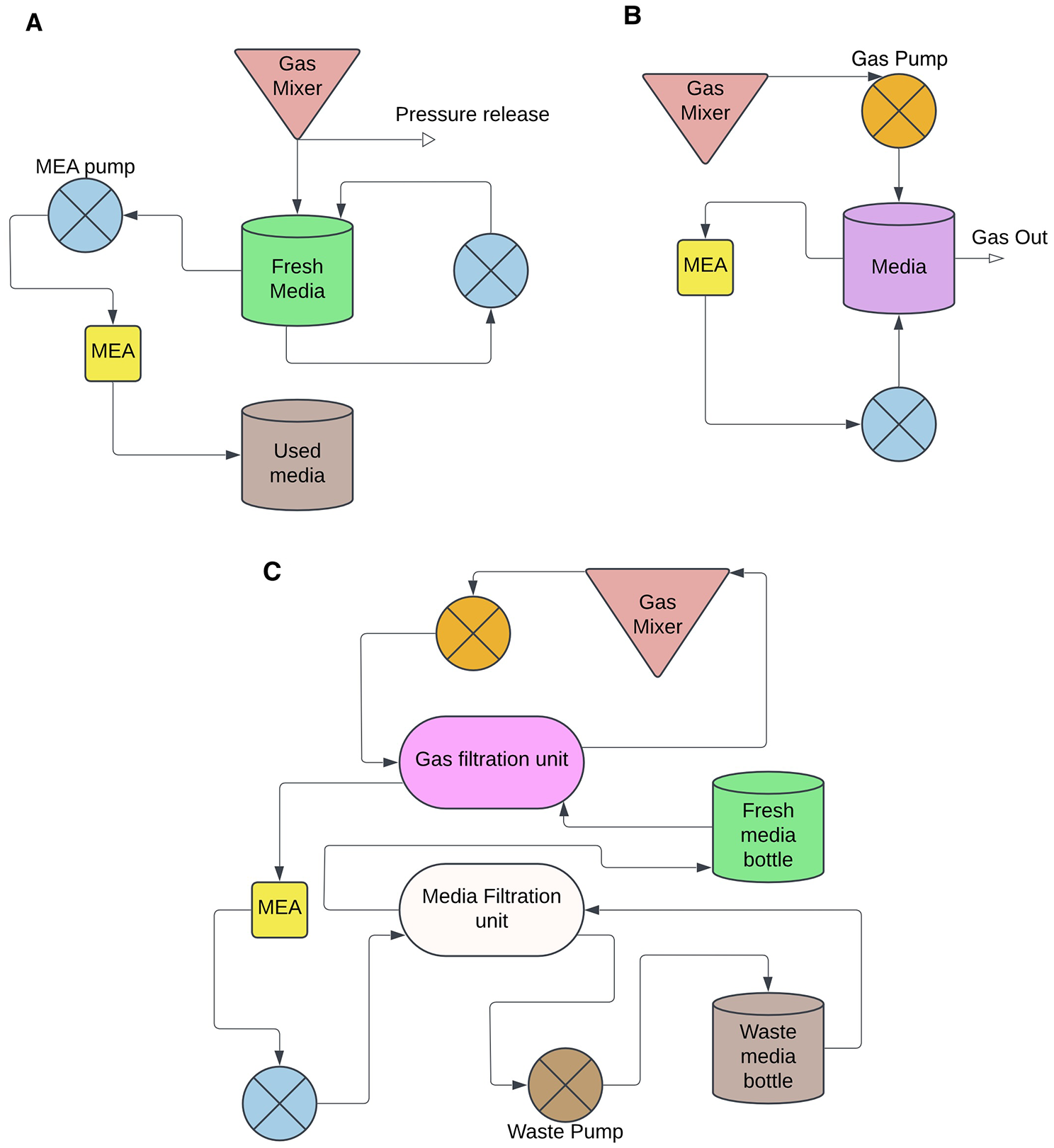
Schematic of integrated system for controlled media change. A comparison of perfusion circuit layouts: (A) and open loop system where the aim is to automatically feed cells without filtration yet separate used media from fresh media; (B) a closed loop system where the media is reused by cells in a limited number of cycles before the media reservoir is refreshed; and (C) a closed loop configuration that also includes multiple filtration units. MEA, microelectrode array.
To establish an effective perfusion system for neuronal cultures, several critical factors must be considered. The system should enable the continuous provision of fresh nutrients and the removal of metabolic waste while maintaining stable pH and osmolarity. This can be achieved through a slow continuous flow of culture media, designed to mimic physiologic conditions. Incorporating a gas-permeable membrane or direct gas perfusion is essential to ensure adequate oxygenation throughout the culture, which is vital for neuronal survival and function. Additionally, the system should incorporate real-time monitoring capabilities for key parameters such as pH, oxygen levels, and metabolite concentrations, providing a comprehensive understanding of the cellular environment (Kim and others 2020). Despite the lack of commercially available systems that integrate all these features in a closed-loop setup for long-term neuronal maintenance, significant progress has been made in developing custom systems that address many of these needs. For example, a study described the development of a perfusion system that combines long-term perfusion, imaging, and electrical interfacing of brain tissue in vitro, enabling the observation of neuronal activity over extended periods (Killian and others 2016). Such advancements highlight the feasibility of creating a perfusion culture system that supports not only neuronal viability but also functional and physiologic monitoring.
Interacting with Neural Cultures
Demonstrating information processing and eliciting intelligence from biological neurons requires a means to record activity from and deliver stimulation to the neural cultures. Despite advancements in microfluidics and microfabrication in the last decade, methods for interacting with in vitro neurons at the temporal and spatial resolutions required for SBI protocols are mostly limited to standard drug-screening platforms (Servais and others 2024). The rise of bespoke platforms and initiatives for interacting with neural cultures does offer feasible pathways forward (Elyahoodayan and others 2019; Jordan and others 2024; Kagan, Gyngell, and others 2023; Newman and others 2013; Smirnova and others 2023; Zhang and others 2024). Here we explore the two primary methods currently used to dynamically interact with neural cultures: optical and electrical. While each method can be used in isolation, combinations are common and likely to achieve the most useful outcomes. Likewise, while chemical inputs (i.e., drugs or neurotransmitters) are possible in conjunction with these methods (Jordan and others 2024), the technical challenges of a fully chemical system for dynamic interactions with neural cultures are yet to be fully overcome, despite some exciting progress in this space (Malekoshoaraie and others 2024; Park and others 2024; Parodi and others 2024).
Optogenetic Approaches for Detecting and Visualizing Neural Tissue Activity in Real Time
Real-time optical visualization of neuronal activity is generally based on changes to either membrane potential (voltage) or cationic concentrations that occur during an action potential (AP). Basic calcium imaging, for example, involves the use of a fluorescent dye that binds specifically to calcium ions. The underlying idea is that when an AP is successfully propagated, the resulting spike in intracellular calcium can be observed as increased fluorescence intensity (Chen and Lui 2024; Mertes and others 2022; Sun and others 2024). In reality, low signal-to-noise ratios, lack of cell-type specificity, and slow decay times often result in poor spatial and temporal resolution, especially in dense cultures or in vivo (Tada and others 2014). Optogenetic approaches—particularly, genetically encoded voltage indicators or calcium indicators—have been broadly implemented in the pursuit of rapid, highly localized tracing of neuronal activity (Figure 3). In these systems, genetic instructions for fluorescently tagged voltage- or calcium-sensitive proteins can be targeted for expression in desired cell types.
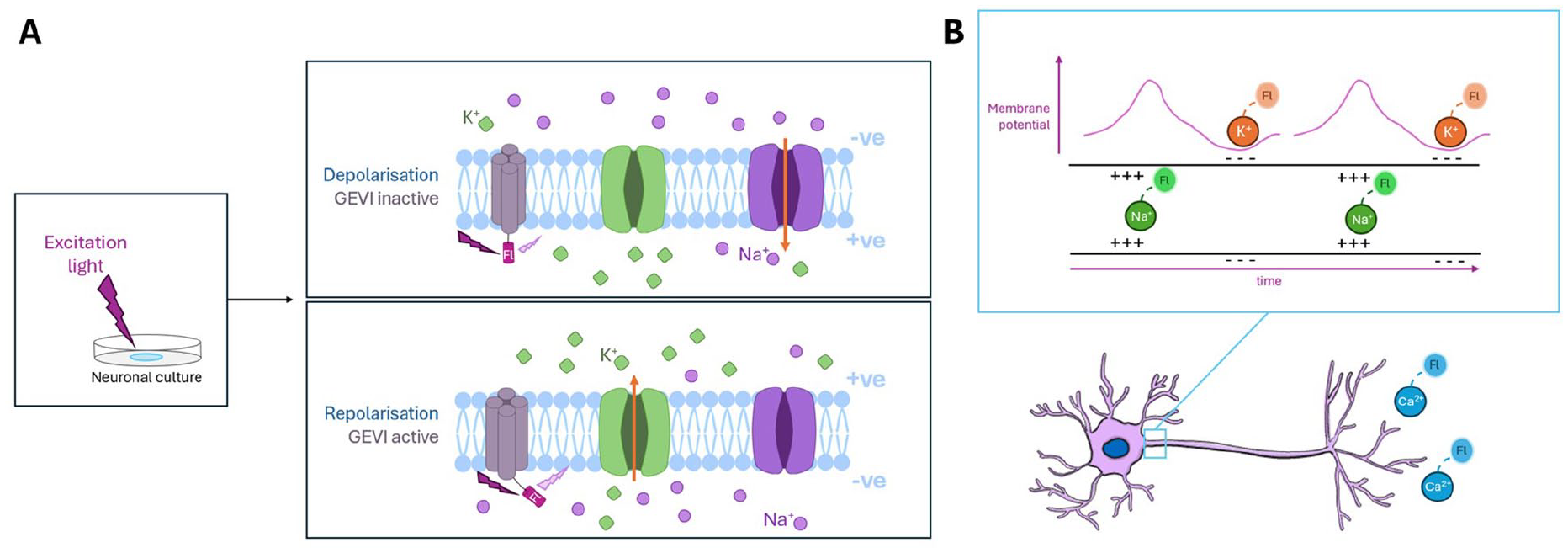
Detection and visualization of neural tissue activity through optogenetic approaches. (A) Light stimulation of genetically encoded voltage indicator (GEVI) fluorescent tags in neuronal membrane during action potential. (B) Sodium ion influx and potassium ion efflux corresponding to action potential propagation along a neuron can be concurrently visualized with targeted fluorescent (Fl) molecules at discrete wavelengths. The accumulation of calcium ions is visualized at the axon termini upon successful firing.
While promising, genetically encoded voltage and calcium indicators are still broadly limited in their ability to fully resolve the 200- to 300-Hz APs exhibited by neurons in vitro (Abdelfattah and others 2016). These systems rely on high fluorophore brightness to improve the signal-noise ratio but demand rapid formation and decay of the fluorescent structure to improve temporal resolution (Han and others 2023; Piao and others 2015; Storace and others 2019). Improvement to these systems has been expedited in the last decade by advances in our ability to genetically fine-tune fluorophore and voltage-sensitive structures. The current generation of genetically encoded voltage indicators has introduced systems capable of resolving APs up to 100 Hz (Alich and others 2021). More recently, the optimization of hybrid voltage indicators (genetically engineered proteins that utilize dyes) has allowed for AP resolution at >400 Hz, although photostability and phototoxicity remain challenges to be overcome (Liu and others 2021).
Stimulation of neural cultures is possible via optogenetic neural modulation (Sun and others 2019; Thompson and others 2014). To achieve this, optical-based stimulation of light-sensitive ion channel proteins on the membrane of neural cells is typically employed (Lin and others 2017). The use of light-activated channels such as channelrhodopsin 2 (ChR2) and halorhodopsin (NpHR) are examples of such a protein and can be selectively expressed in specific cell types (Cardin and others 2010; Gradinaru and others 2008). The use of different proteins can allow selective excitation and inhibition of neural cell types, enabling a fine degree of functional control over the cells stimulated (Tønnesen and others 2011). Optical-based methods have allowed explorations of effective connectivity and a better understanding of how key populations of cells may affect a network (Renault and others 2015; Robinson and others 2016). Despite these successes, there are some limitations to optical methods. Thus, while they can have lower noise than electrical methods, these optical methods cannot allow simultaneous recording and stimulation of the same areas, and prolonged use can result in bleaching of photosensitive channels.
MEAs for Electrophysiologic Stimulation and Recording of Neural Cultures
While there are several options available for electrode interfacing, there is currently no consensus on which is most appropriate for applications eliciting intelligence within in vitro systems. There are several technological reasons for this: behavior of electrodes during stimulation and recording, cost of development, variable difficulty in interfacing with microfluidic structures, and data readout. Passive planar electrode-based MEAs have long been used to record activity from and deliver stimulation to neural cultures (Jimbo and others 1998; Shahaf and Marom 2001). More recently, active complementary metal oxide-semiconductor (CMOS)–based approaches have offered a higher-density platform to achieve similar results (Gosselin and Sawan 2008; Heer and others 2004). Passive planar and active CMOS-based MEAs are limited by interfacing mostly with the adherent surface-based cells, thus limiting their applicability for thick brain slices and 3D organoids. Electrodes with pyramidal shapes have been commercialized to assist in penetrating brain slices. While this approach allows for the electrodes to sense beyond the damaged layers at the top or bottom of a slice, it is currently not possible to accurately assess 3D electrophysiologic activity. Work to overcome this limitation is ongoing and has explored penetrating electrodes (Abu Shihada and others 2024; Mateus and others 2019; Muzzi and others 2023; Steins and others 2022), self-rolling MEAs (Kalmykov and others 2021), flexible shell electrodes (Cools and others 2018; Huang and others 2022), and stretchable mesh-based electrodes (Le Floch and others 2022).
Overall, high-density active CMOS-based MEAs and low-density passive planar MEAs gather electrophysical information but have different advantages and limitations. High-density CMOS MEAs provide for multiple electrodes across a single cell, with some densities even allowing the recording of subcellular features, tracking AP flow along an axon, and permitting detailed mapping of individual cells within a network. For applications that do not require high-volume online data interpretation or highly controlled closed loops, this large amount of recorded data can be useful, especially for neurodevelopmental research. This can aid the identification of more complex electrophysiologic patterns of activity linked to neurocomputationally relevant metrics, such as neural criticality or functional connectivity (Cai and others 2023; Habibollahi and others 2023). While workarounds do exist, processing high-density electrophysiological data at a suitable resolution in real time is highly challenging. In contrast, low-density passive planar MEAs can provide only general data of the cells in their vicinity, with subcellular features out of reach. Some approaches restrict axons through microfluidic structures such that only one can cross multiple electrodes. This would permit tracking some axonal signal propagation, albeit in low throughput due to the limited number of electrodes available. Despite lower-density planar arrays being limited to mapping the network-level activity and connectivity, they can typically stimulate at more electrodes than CMOS MEAs (Newman and others 2013; Ronchi and others 2019). Likewise, the lower density can allow for a more nuanced handling of waveforms in real time. Additionally, stimulation from the smaller electrodes found in CMOS MEAs can result in higher-current densities for a given stimulation current, with potentially higher impedance, both of which increase the risk of electrolysis at the electrode surface relative to planar MEAs (Heer and others 2004; Jochum and others 2009; Ronchi and others 2019).
While most electrode development has focused on recording undistorted signals, biohybrid systems require stimulation capabilities through the same electrodes. Several types of electrode materials are commonly used, but these potentially have very different responses to a specific voltage and can show different capacitive behavior. Most electrodes accumulate charge at their interface that is depleted during stimulation. Excess stimulation times or lack of charge balancing during the stimulation can cause permanent damage to the electrodes. While these effects can be subtle, they do cause noticeable drift during reuse or for long-duration experiments. In addition, the ability to modify or develop MEAs can vary wildly. CMOS-based MEAs offer a high density of electrodes with an active circuit that amplifies the signal at the source. The benefit is that many more electrodes can be accessed for recording, providing more flexibility in observing electrophysiological activity over a given array. However, these improvements in signal resolution require dedicated stimulation electrodes out of the wider subset for any given time. The result is that CMOS-based MEA fabricators must typically make a choice: read out a subset of recording electrodes at a time or encounter additional noise in the recordings. This added complexity of fabrication and circuitry often limits bespoke solutions at scale. In contrast, passive low-density MEAs have been used fir decades in research partly because they can be more easily modified to specific structures. Not only are they easier to modify and to interface with microfluidic structures, they also provide more manageable data, but due to the lower density of readout. Overall, the choice between active high-density and passive low-density MEAs should be based on the specifics of the research question and the surrounding technical and methodological approaches.
Algorithms for Exploration of Information-Processing Dynamics and Intelligence in Neural Cultures
Specific software requirements exist to achieve neural cell–computer interfacing and support algorithms intended to extract features of neuronal activity from the data. Software setups should ensure that a high degree of specificity is possible, with tightly regulated spatial and temporal control of whatever hardware system is used to interact with neural cultures. Issues such as jitter are likely to be problematic regardless of the algorithms or hardware that is adopted. Given the inherent variability within neural systems, it is vital that software controls introduce as little additional variability into the systems as possible. Other features, such as whether a closed- or open-loop algorithm is implemented, will attract different requirements that will need to be optimized. Indeed, the choice between open- or closed-loop algorithms (Figure 4) is currently a key distinction among literature exploring information-processing dynamics and attempts to elicit intelligence in neural cultures.
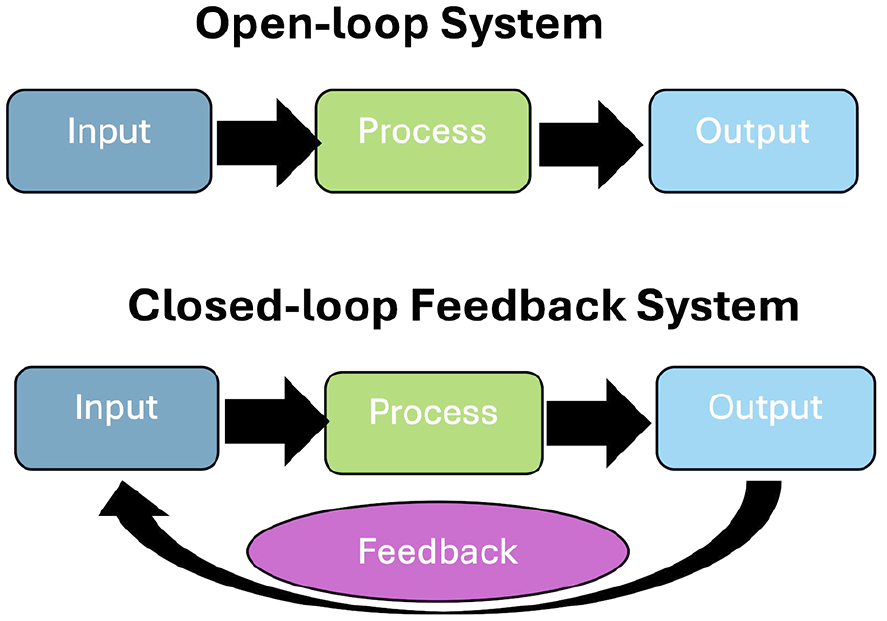
Open-loop systems do not update information inputs based on the output received. Closed-loop systems have a feedback loop that updates future input to a system based on the outputs of the system.
An open-loop algorithm may input any type of information into a system, allow a process to occur, and then monitor an observable output. Importantly, for an open-loop approach, no feedback is provided to the network. Reservoir computing is a common approach that adopts an open-loop algorithm. In reservoir computing, a nonlinear reservoir system, typically a recurrent neural network, acts as a “black box” processing unit into which signals are input, and the inherent nonlinear dynamics of the system project the signal into higher dimensional space (Fu and others 2020; Maass and others 2002; Tanaka and others 2019). The theory is that the original input will be transformed into a richer, more informative signal. As such, unlike artificial neural networks used in machine learning, reservoir computing withbiological neural networks does not require calculation of changes in the weights within a network (i.e., connections between two neurons). Instead, network outputs are trained via a simple linear classification or regression model, and the weights of the network are allowed to update independently according to its dynamics (Sumi and others 2023). This approach has been successful in various classification tasks, by groups using both unpatterned (Lindell and other 2024) and patterned monolayers (Sumi and others 2023) along with organoids (Cai and others 2023), and has shown some success in tasks such as blind-source separation (Isomura and others 2015). It is an advantageous method for testing simple computational paradigms due to its effective information transformation that can be carried out in an “offline” manner. This means that the requirement for hard real-time information processing is absent, as are concerns such as latency. However, it is limited in its application to learning tasks, as the network is not shaped in any way outside the input signal: the neurons do not learn to associate electrical inputs with a desired response, and all learning occurs within external linear models.
In contrast, closed-loop systems are an extension of an open-loop system, whereby the output or the effect of the output on an external environment is transmitted back to the system as revised input, within which the information is altered according to the output. Typically, this may take the form of explicit feedback input, as implemented with the intention to shape the outputs into interpretable behaviors according to some chosen schema. However, provided that any alteration of input signal occurs that is contingent on the system output, this would meet the requirements for a closed-loop system. This does not mean that a classification or similar decoding algorithm like that used in reservoir computing cannot be used in a closed-loop system, only that feedback is also provided to the neural system in some form. Logically, to enable learning within a neuronal network, the loop must be closed, allowing for external feedback to shape the behavior of neurons in an online manner. Dynamic changes may still happen within neural cultures without feedback following open-loop paradigms, but learning for any system requires feedback about the outcome of an output. Ideally, closed-loop algorithms for interacting with neural tissue use physiologically relevant feedback algorithms, with real-time encoding and decoding of data to enable low latency for the loop to ensure that the system can link feedback to the actions that the system has just performed.
For example, a closed-loop interface was introduced to teach cortical cell cultures to improve at a simplified version of the video game Pong (Figure 5; Kagan, Kitchen, and others 2022). In this setup, the cells controlled a paddle by altering activity in counterbalanced predefined regions. The intention was to create a structured information landscape that would shape cell activity in an otherwise arbitrary manner so that activity would move the paddle to intercept a ball. To achieve this, a topographically inspired place-coding method was used to mark the position of the ball on one axis, while a rate-coding method was used to represent the position of the ball on the other axis. The resulting encoding scheme was most similar to how rodent barrel whisker cells encode information (Harrell and others 2020). Through a cycle of iterated structured feedback (predictable on a hit vs random electrical stimulation on a miss), cells “learned” to respond with experimentally desirable outcomes (i.e., hitting the paddle in Pong gameplay). In this case, feedback was designed to test a specific neurocomputational principle of information entropy minimization that has been previously proposed (Friston and others 2006). This framework was based on the free energy principle, which proposes that a neural system has an underlying bias to minimize unpredictable information, either through modifying future predictions or by better controlling the environment around it (Friston 2010; Parr and Friston 2018). However, the results supported that shaping feedback under this framework resulted in rapid improvement in game performance that was seen when the first 5 minutes of gameplay was compared with the last 15 over a 20-minute session. Additional strong support could be seen in the basic electrophysiologic changes concurrent with the performance change observed in the systems. For instance, correlations increased over time between the region in which sensory information was encoded (sensory region) and the regions in which motor information was encoded (motor regions), while the two motor regions became less coupled—a coherent adaptation to facilitate the game action. Despite the supportive results for this work, feedback could have been based on several other principles, although likely with different results; indeed, other work has explored the utility of closed-loop feedback systems from different perspectives (Bakkum, Chao Zenas, and Potter 2008a; Tessadori and others 2012). As such, while significant future work remains to understand how to optimize the features of closed-loop algorithms for interacting with neural cultures to elicit a goal-directed outcome, meaningful understandings in neurocomputational functions with disease modeling and drug testing have already been gained.
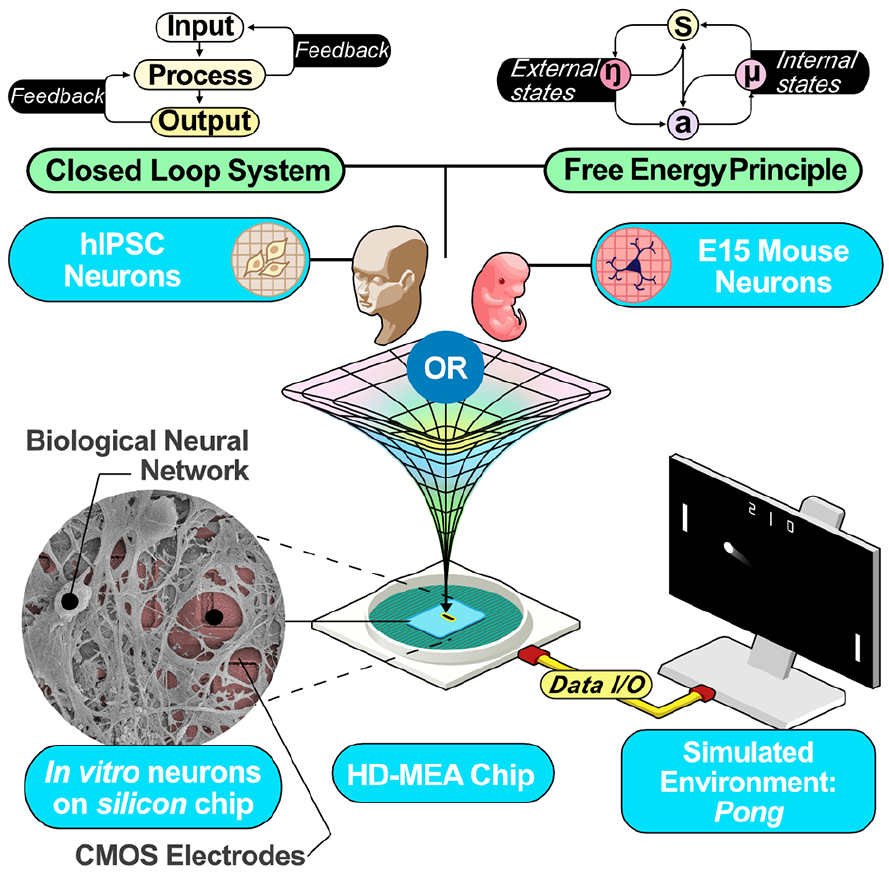
Schematic describing the core components of a real-time closed-loop instance of a simplified version of the game Pong was tested by combining cortical cells taken from embryonic mouse primary cells or differentiated from human-induced pluripotent stem cells (hIPSCs). Implications from the free energy principle were applied as the feedback was based on whether the cultures successfully or unsuccessfully altered their activity to move a Pong paddle to intercept the virtual ball. CMOS = complementary metal-oxide-semiconductor; HD-MEA = high-density microelectrode array; I/O = input/output.
Exploring Neurocomputational Function
Induced Plasticity Changes in Cell Cultures through Functional Connectivity Inference
The activity patterns derived from in vitro neuronal systems provide critical insights into the underlying mechanisms of their function and dynamics. However, the relatively sparse time series data from these cultures (i.e., capturing only occasional neural activity with discrete spatial sampling from larger cultures) require advanced algorithms to identify meaningful patterns beyond underlying stochastic spontaneous activity. A popular method for exploring the population dynamics in neural cultures is to focus on methods such as effective and functional connectivity metrics within the network (Khajehnejad and others 2023; Poli and others 2015; Sharf and others 2022; Yamamoto and others 2018). Understanding these connectivity metrics is crucial for measuring plasticity changes in cultures, which is key to uncovering how neurons work together to process information. Yet, given the sparse spiking activity of these in vitro biological neuron populations, traditional methods, such as cross-correlation or transfer entropy, may struggle to provide reliable connectivity maps, particularly when only a small subset of neurons is active at any given time. Previous studies have adopted simplified models of interconnected neural populations, particularly those based on mean-field approximations, as an effective approach for maintaining the essential dynamic properties of neural networks while significantly reducing computational complexity. These models have been shown to accelerate simulations by several orders of magnitude and offer valuable insights into the emergence of population-level phenomena such as phase transitions (Baspinar and others 2021; Bick and others 2020; La Camera 2022). Additionally, in neural populations, only a small subset of neurons typically shows clear spiking activity at any given time. This sparsity in activity and connectivity allows the network to conserve energy and maximize computational efficiency (Olshausen and Field 2004). Yet, sparsity introduces redundancy, as individual neurons often transmit overlapping or highly correlated signals.
To address the sparsity and redundancy in neuronal firing, dimensionality reduction techniques have become essential. These methods, including principal component analysis and nonlinear embeddings such as t-distributed stochastic neighbor embedding (t-SNE) (Van der Maaten and Hinton 2008) and uniform manifold approximation and projection (UMAP) (McInnes and others 2020), reduce the temporal and spatial dimensions of the data (Figure 6), allowing researchers to focus on the most representative neurons to build more robust functional connectivity networks.
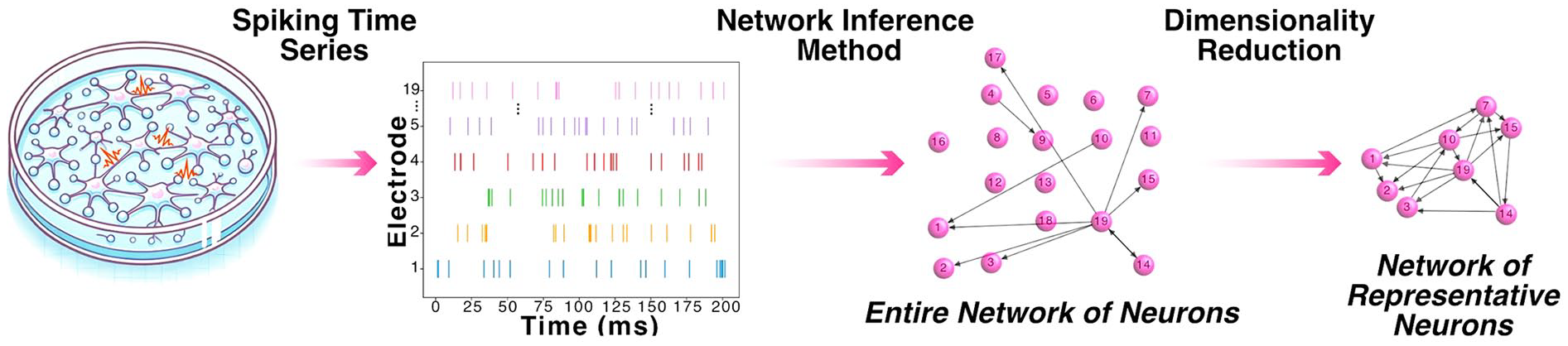
Understanding functional connectivity within in vitro neuronal cultures. Schematic representation of a network of in vitro neurons, highlighting sparse spiking activity and the inference of a functional connectivity network from spiking time series before and after dimensionality reduction.
These algorithms excel at capturing essential features of neuronal dynamics (although it must be acknowledged that other features may not be captured), extracting latent information from high-dimensional and sparse data while minimizing noise and redundancy. For example, recent work (Cai and others 2023; Khajehnejad and others 2023) has identified stark changes in functional connectivity metrics between the presence and absence of structured information by analyzing representative channels from high-density MEAs within in vitro neuronal cultures. Of particular interest is that, for closed-loop environments, deeper analysis has identified that these functional connectivity maps can cluster functionally related regions (i.e., regions in a culture that have similar impacts on an external environment) more closely than the underlying spatial organization of the cells on an MEA (Khajehnejad and others 2024). This approach highlights the adaptability of neuronal networks to external stimuli, demonstrating how population-level dynamics can reveal the plasticity in functional connectivity essential for intelligent behavior.
Phase Transitions and Emergence of Criticality
The ability of biological neurons to process information likely relies on the collective dynamics of neuronal populations (Ebitz and Hayden 2021). Therefore, studying population-wide dynamics provides a broader and equally essential perspective for understanding neural network behavior. Population dynamics encompass various emergent properties that arise from the collective behavior of neurons, including synchronization, phase transitions, and neural oscillations (Bick and others 2020; Moretti and Muñoz 2013). These phenomena highlight how networks can integrate and process information across multiple temporal and spatial scales. When individual spikes fail to capture the full complexity of the system, analyzing population-wide behavior becomes crucial for revealing latent patterns and mechanisms that govern neural function. Among these population-level phenomena, neural criticality stands out as a particularly important concept (Box). Criticality refers to the network’s proximity to a phase transition (Figure 7A), where it optimally balances stability and flexibility for information processing (Cocchi and others 2017; Shew and Plenz 2013). This balance allows the system to be highly responsive to stimuli while avoiding chaotic or overly rigid behavior (Shew and others 2015). While individual spikes may contain latent information, it is the collective behavior, manifested through critical dynamics, that often reveals how the network organizes itself to process structured information. The role of neural criticality was a matter of contention (Cocchi and others 2017; Hesse and Gross 2014), with inconsistent interpretations to the importance and impact of criticality in healthy and diseased function (Arviv and others 2016; Clawson and others 2017; O’Byrne and Jerbi 2022; Toker and others 2022). In vitro neuronal networks recently demonstrated self-organization toward near-critical states when engaged in a structured information landscape, such as playing a simplified version of Pong, but not during the absence of a structured stimulation (Habibollahi and others 2023). This study found that as neural networks processed task-related sensory input, they approached criticality (Figure 7B), with better task performance correlating with proximity to these critical dynamics. However, the study emphasized that criticality alone does not account for learning without feedback about task outcomes. Instead, results demonstrated that criticality is a fundamental organization required for any dynamic system composed of individual units to collectively store, transmit, and respond to structured information in a broader environment. This interpretation would link these population dynamics across scales from neurons to bird flocks and perhaps even to cosmologic scales (Aschwanden and others 2016; Bialek and others 2012).
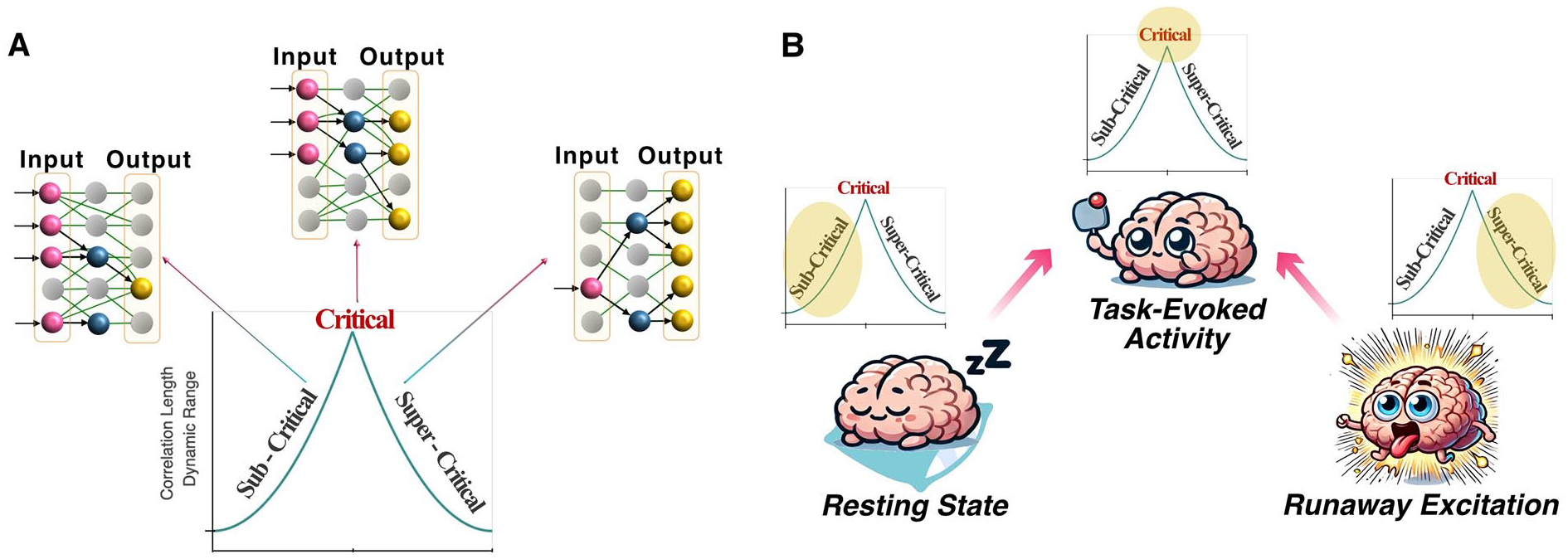
Understanding population dynamics within in vitro neuronal cultures. (A) Comparison of key characteristics between a critical system and systems exhibiting subcritical or supercritical dynamics. Figure adapted from Habibollahi and others (2023). (B) Illustration of how the network adapts to external stimuli within a task environment, transitioning toward a critical state and away from states that are subcritical (resting-state spontaneous activity) or supercritical (runaway excitation).
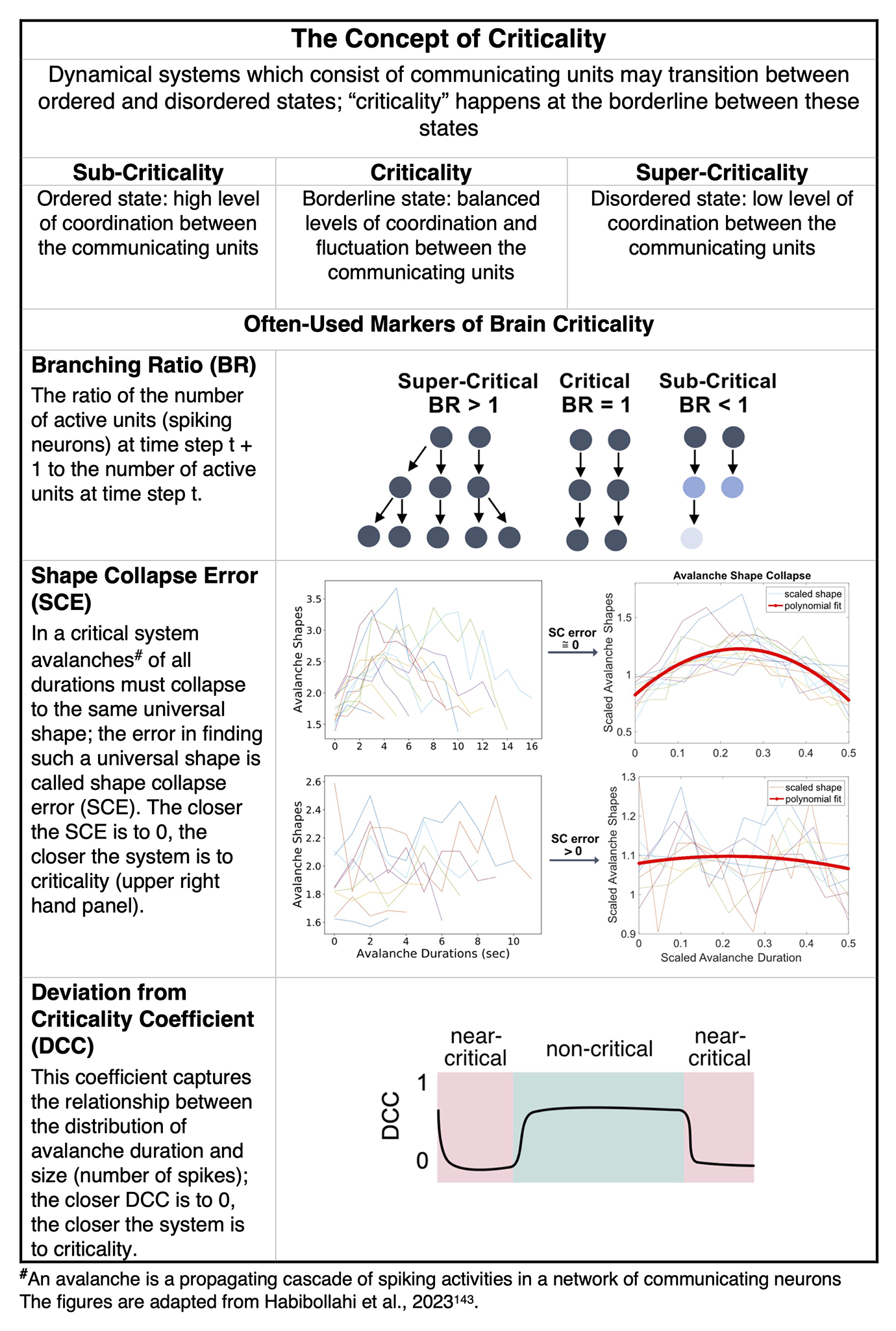
The Concept of Criticality.
Leveraging Biological Intelligence to Evaluate Pharmacologic Interventions
Up to one in three people worldwide are affected by neurologic conditions (Steinmetz and others 2024). In parallel, researchers are facing the issues of outdated preclinical models of drug testing prior to clinical trials (Galanopoulou and others 2012). Pharmacologic compounds are generally added to a neural cell culture model or system to examine their effects on neuronal function, perturb a known cellular or subcellular process, or more broadly identify “hits”—compounds that elicit a desired effect above the threshold for a given measurement and are suitable to continue down the costly pathway of drug discovery and development. Yet, there is a lack of relevant assays to assess therapeutic efficacy; current methods are based on survival and spontaneous activity of the neurons rather than the true function of neural systems—an ability to process and respond to information.
The modulation of basic open-loop information-processing characteristics has already been observed in neural cultures intended to model phenotypically healthy function following pharmacologic interventions (Alam El Din and others 2024; Cai and others 2023). However, the restoration of function in an in vitro disease model within a closed-loop environment designed to elicit a goal-directed learning would be a more compelling demonstration. To this end, SBI paradigms offer a compelling approach for more nuanced disease modeling and drug testing (Figure 8). In a proof-of-concept experiment, induced pluripotent stem cell–derived NGN2 neurons (Pawlowski and others 2017) were rapidly differentiated into highly active cultures and subjected to a real-time closed-loop training environment in a simplified Pong environment. Under control conditions, neurons displayed a hyperactive phenotype reminiscent of that observed in epileptic seizures and did not show improved performance over time. After addition of antiseizure compounds, moderate neural activity was restored, and treated cultures showed significant learning over time and against untreated controls (Watmuff and others 2025).
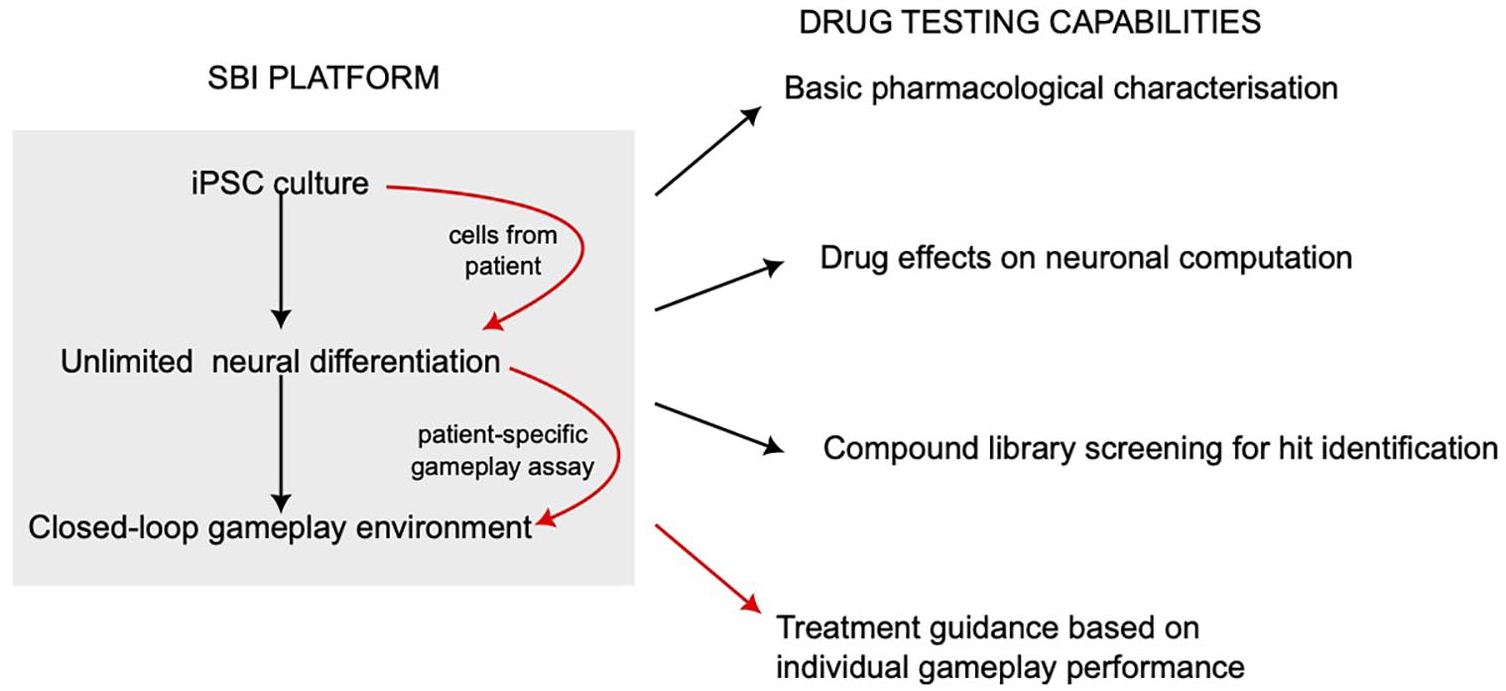
Some possibilities for pharmacologic and translational research via a synthetic biological intelligence (SBI) platform. The gray area on the left represents the components and workflow composing a typical SBI platform capable of interrogating the information-processing capabilities of human neurons. Human neurons are generated in extremely high quantities from induced pluripotent stem cells (iPSCs) for embedding in a closed-loop feedback environment (black arrows), and in the case of a patient with a known disease or mutation, these cells represent an in vitro model of the condition (red arrows). The possibilities for basic drug-testing research and, in the case of patient cells, translational clinical outcomes arising from the SBI platform are listed on the right.
Further afield, there are several directions for future drug-testing research. Compounds that modulate synaptic plasticity might improve learning in cultures by strengthening the conduction of network signaling, helping uncover the deeper mechanisms underlying learning in this cellular context. Compounds with suspected neurotoxicity might be tested safely and accurately in vitro, and while this cannot abrogate the use of in vivo models still necessary to assess how the drug affects the whole organism, it can permit first definition and establishment of possible therapeutic regimens, thereby considerably reducing the use of animals for first-step testing. Last, compounds whose therapeutic benefits are only recently emerging, such as stimulants, depressants, and hallucinogens (e.g., cannabinoids, psylocibin, amphetamine), and whose basic pharmacologic effect has not been tested comprehensively at the cellular level could be tested more relevantly in vitro by assessing how they affect computational processing of neurons as monitored via gameplay.
Lessons, Future Directions, and Prospects
Research in SBI is highly multidisciplinary. Here we have discussed key concepts from a neuroscientific perspective, yet other perspectives have equally important contributions to the growth of this new technology. Indeed, if the early progress in this field reveals anything, it is the inadequacies of any single approach to elucidate the complexity inherent in exploring information-processing dynamics and intelligence within in vitro neural cultures. To understand how different neural organizations affect function, modular vs unpatterned vs organoids will have to be explored. To understand neural function, functional connectivity analyses must be integrated with assessments of population-wide dynamics and single-cell data. This perspective can be applied to many areas in science generally and to all discussed in this article but is particularly true for multidisciplinary work with significant uncertainty. The juxtaposing and integrative approach of multiple methods will provide significantly more insight than any single method in isolation and may help answer some of the most pressing questions (Kagan, Loeffler, and others 2024). While the benefits of this proposal may sound obvious, very few scientific studies are able to incorporate these principles, perhaps due to the inherent difficulty of cross-disciplinary collaboration. Despite these difficulties, impressive progress has occurred in arranging international multidisciplinary collaborations to progress this work (Barros and others 2025). One such example is the creation of organoid intelligence as a collaborative endeavor (Hartung and others 2023; Morales Pantoja and others 2023). Organoid intelligence seeks to harness methods related to those in this review to develop biological computing by using organoids and brain-machine interface technologies (Smirnova and others 2023). A key proposed merit of this approach is to utilize more physiologically relevant cell cultures to better understand mammalian biology for the purposes of biomedicine (Smirnova 2024). Early results and the growing community aiming to establish methods to better access this technology are undoubtedly promising (Alam El Din and others 2024; Cai and others 2023; Jordan and others 2024; Sharf and others 2022).
As with many burgeoning scientific fields, there is a distinct risk that the promise may overshadow the progress. That is not to imply that the future potential of a field is not worthy of discussion—indeed the opposite is true—but care should be taken to ensure that a grounded approach is adopted. This is particularly relevant when the work intersects growing scientific and public interest in this space with a high degree of ethical scrutiny (Hyun and others 2020; Kataoka and others 2023; Lavazza 2021; Lavazza and Reichlin 2023; Sawai and others 2021). Particular ethical concern is often focused on specific cell culture systems such as organoids, even though no evidence currently supports that these systems might have attributes that warrant such concern (Boyd and Lipshitz 2023; Kagan, Duc, and others 2022; Kagan, Gyngell, and others 2023; Koplin and Savulescu 2019). Perhaps the key exacerbating factor to this debate is that semantic usage of specific terms is radically different among fields. Consequently, in public discourse these terms are often poorly defined; therefore, the nuance is just as challenging to communicate as it is to be understood (Boyd and Sugarman 2022). Disagreements about language choices in early work in this field have already affected scientific communication of the findings and future directions (Kagan, Razi, and others 2023; Pereira and others 2023; Rommelfanger and others 2023). This makes establishing methods to facilitate multidisciplinary collaborations a critical endeavor. One effort currently underway is to ease communication across fields and improve public science education by reaching a nomenclature consensus through collaborative processes (Kagan, Mahlis, and others 2024). This approach adopts a multidisciplinary call for collaboration and seeks to work with the full breadth of experts to improve scientific communication in a sustainable manner. Should the various communities be able to come together and follow structured consensus-making processors, this will aid further multidisciplinary explorations and facilitate progress.
Ultimately, exploring the capacity to leverage living neurons for information processing and intelligence has the potential to be a generation-defining technology. The short-term outcomes of improved basic science tools, drug discovery, and disease modeling are promising to overcome barriers that have impeded neuroscientific advancements. Particularly as a tool to test neurocomputational theories, this approach has a unique value. Most neurocomputational theories are limited to describing fairly broad effects, making them difficult to falsify, or are so specific as to have poor predictivity for emergency properties that might arise. As this approach is able to build different layers of complexity, it will allow the elucidation of strengths and weaknesses of any given theory through rigorous testing, theory refinement, and further testing. By exploring these more proximal applications, the long-term potential can be systematically established. Of key interest is the possibility to harness new material capable of significantly more complex and efficient transformations of information, potentially in autonomous and generalized ways—a goal that should be considered inspiring. Yet, the scope of possible applications and technical advancements required. Highlights the fact that no single currently existing field will realize the full potential of this new technology. Only by coming together will we be able to successfully build the foundations for eliciting intelligence from in vitro biological neurons.
Footnotes
Acknowledgements
Figures 1 and 3 were created in BioRender.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: B.J.K., F.H., B.W., A.A., F.D., A.L., S.H.B., C.D., K.D.A., and N.K.d.R. are employees of and may hold shares or another interest in Cortical Labs, a research-focused start-up working in a space and holding patents related to this article. No specific incentive was provided to any author for contribution to this article. B.S. has no conflict of interest to declare.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
