Abstract
Introduction
Proton therapy minimizes damage to healthy tissues while effectively targeting tumors. However, uncertainties in relative biological effectiveness (RBE) calculations remain a challenge, particularly for critical organs at risk (OARs), such as the brainstem and spinal cord. This study introduces ensemble RBE models (eRBE-B and eRBE-SC) developed through a meta-synthesis (MS) approach to improve dose accuracy and stability in proton radiotherapy.
Materials and Methods
A systematic literature review following PRISMA guidelines identified studies utilizing variable RBE models. The MS approach aggregates data on model usage patterns and study quality due to the lack of direct RBE measurements. Three primary models (Carabe, Wedenberg, and McNamara) were weighted on the basis of study quality and case counts to construct ensemble RBE models. Dose calculations and analysis were performed via RayStation 8B, MCsquare, and 3D Slicer.
Results
Compared with the Carabe model, the eRBE-B model resulted in reduced dose underestimation, with mean dose (
Conclusion
Ensemble RBE models offer a robust alternative to standalone models in proton therapy. By aggregating published RBE data, the MS approach enhances dose precision for critical organs, improving the safety and efficacy of proton radiotherapy. These findings underscore the potential of computational tools to optimize treatment planning for head and neck cancer patients.
Plain Language Summary
Proton therapy is a type of radiation treatment that can more precisely target cancer than traditional X-rays. However, predicting how effective and safe the treatment is can be complicated. A key challenge is understanding the “Relative Biological Effectiveness” (RBE), which measures how powerful proton radiation is at damaging cancer or healthy tissue. Currently, doctors often assume a fixed RBE value, but this might underestimate the real effects, especially in sensitive areas like the brainstem or spinal cord. In this study, we combined information from many previous studies to create a new modeling approach called the ensemble RBE (eRBE) model. This method uses data from 3 well-known RBE models and gives more weight to studies with better quality and larger patient numbers. By using advanced computer simulations and treatment planning tools, we showed that this eRBE model gives more accurate predictions of the biological dose, especially in regions where proton beams are most intense. The goal is to help doctors make better decisions in planning proton therapy—reducing risks and improving outcomes for patients with tumors near sensitive organs. Future steps include testing this approach with real patient data to confirm its safety and effectiveness.
Keywords
Novelty and Impact
This study introduces an ensemble RBE (eRBE) modeling framework for proton therapy, developed through a meta-synthesis (MS) approach that integrates and weights data from 3 established RBE models (Carabe, Wedenberg, McNamara) based on study quality and case volume. Unlike fixed RBE assumptions (eg, RBE = 1.1), the eRBE approach provides a biologically adaptive method that accounts for LET (linear energy transfer) and tissue-specific radiosensitivity (α/β ratio), thereby enhancing clinical relevance.
Compared with individual RBE models, the eRBE models demonstrate improved dose accuracy and consistency in critical serial organs such as the brainstem and spinal cord—regions where biological uncertainty is most pronounced due to high LET variations near the distal edge of the Bragg peak. Monte Carlo-based simulations and dose–volume histogram (DVH) analyses conducted in RayStation, MCsquare, and 3D Slicer reveal that fixed RBE models may underestimate biologically effective doses by up to +10% in these regions.
The study also underscores the clinical significance of these deviations: according to QUANTEC guidelines, the brainstem’s maximum tolerated dose
Moreover, the framework highlights the need for uncertainty-aware RBE modeling and paves the way for future validation studies involving patient cohorts and alternative simulation platforms (eg, TOPAS, FLUKA). As such, the proposed eRBE methodology holds promise as a next-generation tool for improving the precision and biological relevance of proton therapy treatment planning.
Introduction
Proton therapy has become a pivotal modality in radiation oncology, offering the advantage of precise tumor targeting while sparing adjacent healthy tissues. A critical parameter in proton therapy is the relative biological effectiveness (RBE), which quantifies the biological impact of proton beams relative to conventional photon radiation. Clinically, a fixed RBE value of 1.1 is widely adopted, as recommended by early studies such as those of Paganetti et al, 1 to simplify dose calculations and minimize the risks of underdosage or overdose. However, this simplification ignores key factors influencing RBE, such as radiation energy, linear energy transfer (LET), and tissue-specific characteristics.2,3 This has led to growing concerns over the adequacy of the fixed RBE approach in accurately representing biological dose distributions.
Several RBE estimation models, including the Carabe, Wedenberg, and McNamara models, have been developed. These phenomenological models incorporate biological and physical parameters, such as LET and the linear‒quadratic (LQ) framework, to provide a more nuanced understanding of RBE variations. 4 Despite these advancements, substantial variability exists among the predictions of these models owing to differences in data fitting, underlying assumptions, and regression methodologies. Studies by Rørvik et al. 5 and McMahon et al. 6 have highlighted these inconsistencies, which are further complicated by the absence of direct RBE measurements or robust clinical benchmarks. Without patient-level RBE data, efforts to validate or prioritize specific models remain limited, leaving clinicians to navigate significant uncertainties.7-11
Given the lack of harmonized quantitative RBE endpoints across studies, traditional meta-analysis—which statistically integrates comparable outcomes—was not feasible. Instead, this study adopts a meta-synthesis (MS) approach that aggregates heterogeneous patterns of model usage, methodological rigor, and theoretical underpinnings. Unlike meta-analysis, MS provides a structured yet flexible framework for combining diverse types of published evidence.12-15 In computational modeling and health systems science, MS has been effectively adapted in scenarios where quantitative pooling is limited by data heterogeneity.12,14
This study adopts such a framework to address the inherent heterogeneity in RBE modeling literature.
By applying weighted contributions based on study quality and case counts, the MS methodology enables the development of eRBE models that reflect prevailing modeling trends and offer probabilistic insights into biological dose estimation.16,17 While exploratory in nature, this approach addresses a key methodological gap in RBE modeling by providing a reproducible synthesis strategy suitable for clinical integration.
Proton therapy for head and neck cancers highlights the need for precise RBE calculations. Critical organs at risk (OARs), such as the brainstem and spinal cord, are often located in close proximity to the clinical target volume (CTV). Even small inaccuracies in RBE-weighted dose calculations can lead to significant deviations in treatment outcomes. For example, excessive radiation to the brainstem can cause severe or fatal complications, whereas spinal cord overexposure may result in motor dysfunction or sensory deficits. Accurate dose estimation for these structures is essential for optimizing therapeutic efficacy while minimizing adverse effects.
This study aims to develop 2 eRBE models tailored for these critical structures: the eRBE-Brainstem (eRBE-B) and eRBE-Spinal Cord (eRBE-SC) models. As a proof-of-concept study, this work is intended to demonstrate the feasibility of integrating heterogeneous RBE modeling literature into an ensemble framework using a structured MS approach. By synthesizing data from published studies, these models provide a more consistent and stable framework for estimating biologically weighted doses, addressing the limitations of standalone models. Although the ensemble models do not directly validate RBE values against clinical benchmarks, they consolidate existing knowledge to increase the precision of proton therapy dose calculations. This work underscores the importance of future clinical and experimental efforts to establish standardized RBE validation protocols while offering an interim solution to current uncertainties in RBE modeling.
Materials and Methods
Study Design and Aim
The primary aim of this study was to establish eRBE models via a MS approach, acknowledging that the lack of direct RBE measurements or robust clinical validation precludes the use of traditional meta-analysis. Instead, the MS approach aggregated published data on RBE model usage patterns and study characteristics to identify trends and develop weighted eRBE models. This methodology does not directly validate RBE models against clinical benchmarks but serves as an exploratory framework for addressing uncertainties in RBE estimation.
A systematic literature search was conducted via the Web of Science, PubMed, and Scopus database, with a focus on studies utilizing variable RBE models to evaluate absorbed dose distributions in the brainstem and spinal cord. Weighted averages for eRBE models were computed by integrating 2 key factors: study quality, assessed via the Joanna Briggs Institute (JBI) checklist, and case counts, reflecting the empirical relevance of each model. Normalization factors were applied to ensure balanced integration of model contributions, avoiding biases from prolific research groups or highly cited studies.
To evaluate dose variations due to different RBE models, treatment plans were created via RayStation 8B (RaySearch Lab, Stockholm, Sweden) and subsequently imported into MCsquare (Université catholique de Louvain, Belgium) for Monte Carlo simulations. These simulations generated spatial distribution files for dose and LET, which were used to calculate RBE-weighted dose distributions on the basis of both “published model-included” RBE and eRBE models. The final biologically weighted dose files were analyzed via 3D Slicer software, which enables detailed visualization and dose‒volume histogram (DVH) generation for the CTV, brainstem, spinal cord, and parotid gland.
This study emphasizes that eRBE models are probabilistic tools designed to address uncertainties in RBE estimations rather than direct replacements for clinical validation. The overall research framework, including the MS approach and computational workflow, is depicted in Figure 1. Overall Research Flowchart. Abbreviation: LET: Liner Energy Transfer, RBE: Relative Biological Effectiveness, MS: Meta-Synthesis, DVH: Dose-Volume Histogram
The MS methodology employed in this study involves aggregating published data patterns rather than statistically integrating validated experimental results, as in traditional meta-analysis. While traditional meta-analysis is designed to synthesize outcomes from multiple studies to estimate an effect size, the MS approach adopted here focuses on evaluating trends in model usage and study characteristics to construct eRBE models. This process supports the development of eRBE models by following an adapted version of the standard six-step process of meta-analysis, as detailed below:
Formulating the Review Question
The primary research goal was to evaluate how variable RBE models have been utilized in proton radiotherapy studies, specifically for assessing doses to the brainstem and spinal cord in head and neck cancer patients. To address this research question, the Patient, Intervention, Comparison, Outcomes, and Study Design (PICOS) framework was used to guide the review.
Defining Inclusion and Exclusion Criteria and Developing a Search Strategy
The inclusion and exclusion criteria were established, and a targeted search strategy was implemented via a partitioned keyword combination method. The search was conducted in major databases, including Web of Science, PubMed, and Scopus, with specific criteria: (1) First-tier constraint: The literature must focus on cancer patients and specifically address brainstem and spinal cord dose assessments. (2) Second-tier constraint: Only studies from established peer-reviewed collections were included. (3) Third and fourth tiers: Studies must incorporate research designs using both a fixed RBE value of 1.1 and variable RBE models.
For example, when searching for literature related to the brainstem, the following search terms were used:
(“Brainstem” OR “Medulla oblongata” OR “Pons” OR “Midbrain” OR “Mesencephalon”) AND (“PT” OR “proton therapy” OR “proton therapies” OR “proton beam therapy” OR “proton beam radiation therapy”) AND (“RBE” OR “relative biological effectiveness”) AND (“variable RBE” OR “variable relative biological effectiveness” OR “RBE model” OR “RBE models” OR “relative biological effectiveness model” OR “relative biological effectiveness models”).
The Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) flowchart was used to document the manual screening process, including title, abstract, and full-text review, to identify studies that met the inclusion criteria. The literature search was limited to studies published up to July 2023. Additionally, this review protocol was registered in the INPLASY database under the registration number INPLASY202540057.
Study Quality Assessment
The JBI critical appraisal checklist for quasiexperimental studies was used to evaluate study quality. Each study was scored on 9 questions, with each “yes” response receiving 1 point, for a maximum possible score of 9 points. 18 This assessment ensured that only methodologically rigorous studies contributed to the eRBE models.
Assessor Calibration and Quality Assessment
Although traditional inter-assessor calibration was not applicable in this study, given that our data sources were published studies rather than human assessors, we addressed potential heterogeneity by implementing a structured weighting system. Specifically, each study was evaluated based on 2 criteria: methodological rigor (assessed using the Joanna Briggs Institute (JBI) critical appraisal checklist) and empirical strength (reflected by case numbers). These 2 metrics were combined and normalized to generate ensemble weights for each RBE model. This systematic approach ensured that contributions from different studies were homogenized and minimized the impact of individual study variability, thereby achieving an effect analogous to assessor calibration in conventional settings.
Data Extraction
The following data were extracted from the selected studies: (1) The specific RBE models employed. (2) Study quality scores based on the JBI checklist. (3) The number of cases evaluated in each study.
These extracted parameters were used to calculate weighted contributions for each RBE model. The weighted values were then applied to construct 2 organ-specific eRBE models: the eRBE-B and eRBE-SC models.
Treatment Planning Simulation and eRBE Model Weight Calculation
Treatment Planning Simulation
This study utilized the MCsquare Monte Carlo simulation toolkit, developed in Python, to process and analyze treatment plans generated by RayStation 8B. The MCsquare software simulated particle interactions within the proton beam, yielding spatial distributions of dose and average LET.19-21
For simulation purposes, a commercial Rando phantom (The Phantom Laboratory, Salem, NY, USA) was employed to model the anatomical structures of head and neck cancer patients, including the brainstem, spinal cord, and CTV. The Rando phantom is widely recognized in radiation oncology research for its anatomical fidelity and tissue-equivalent properties.
To minimize radiation exposure to critical structures such as the brainstem, 2 proton beams were applied at gantry angles of 90° and 270°, replicating typical clinical treatment strategies. The prescribed dose for the CTV was set at 60 Gy (RBE), aligning with real-world clinical dose prescriptions for head and neck tumors.
Although direct experimental phantom validation was not performed within this study, the MCsquare Monte Carlo simulation toolkit, utilized for dose and LET calculation, has been extensively benchmarked against clinical measurements in previous studies, ensuring reliability and clinical relevance of the simulated dose distributions.
The biologically weighted dose distributions generated from the simulations were subsequently analyzed using the constructed variable eRBE models to evaluate their impact on critical organ sparing.
Model Weight Calculation: eRBE Framework
This study utilized an MS approach to develop eRBE models for critical organs by systematically integrating published data on RBE model usage patterns, study quality, and case numbers. Instead of relying solely on model frequency or prevalence in the literature, this approach incorporates study quality assessments and case counts to ensure a balanced and structured model integration.
Three RBE models—Carabe, Wedenberg, and McNamara—were selected due to their well-documented theoretical foundations based on the linear-quadratic (LQ) framework. Each model incorporates dose-averaged linear energy transfer (LETd) to account for the biological effects of proton beam interactions with tissues. While newer datasets such as the Particle Irradiation Data Ensemble (PIDE) database provide additional information, this study focused on these 3 models due to their established reliability and extensive use in prior research. The ensemble modeling approach aims to reduce variability in RBE estimations by integrating multiple models rather than relying on a single theoretical framework.
The eRBE model weighting process is as follows: (1) Study Quality Scores: Evaluated via the JBI critical appraisal checklist, which assigns scores (0-9) on the basis of methodological rigor. (2) Case Numbers: This reflects the empirical data strength of each study.
The weights for each model were calculated as follows (Equation (1)): • F: Total number of RBE models included. • • S: Cumulative quality score across all studies. • • N: Cumulative number of cases across all studies. • Normalization factor:
Weights were normalized to ensure that the sum equaled 1.
For example:
Suppose that we have the following data for 3 models (A, B, C) across 5 studies: Quality scores for the 5 studies: 8, 9, 7, 8, 9 Patient counts for the 5 studies: 4, 5, 6, 7, and 8
Calculate the total quality score S and total case number N: S = 8 + 9+7 + 8 + 9 = 41 N = 4 + 5+6 + 7 + 8 = 30
For Model A (Carabe), 3 studies with quality scores of 8, 7, and 9 and case numbers of 4, 6, and 8 were used. s
f
= 8 + 7+9 = 24 n
f
= 4 + 6+8 = 18
For Model B (Wedenberg), 1 study with a quality score of 8 or 4 cases was used. s
f
= 8 n
f
= 4
For Model C (McNamara), 5 studies with a total quality score of 41 and a total case number of 30 were used. s
f
= 41 n
f
= 30
The weights are normalized to sum to 1: Total weight = 0.59 + 0.16+1.0 = 1.75
The eRBE model can then be represented as:
Biological Dose Weighting
The eRBE model integrates LET and radiosensitivity parameters (α/β to account for variations in tissue responses: • •
The weighting calculation for the eRBE model integrates • •
For example: • A low • In the McNamara model,
The essence of eRBE lies in combining 3 empirical formulas via weighted averaging, balancing individual model biases while capturing their strengths. This provides a comprehensive framework for more accurate dose calculations in proton therapy.
Treatment Plan Loading and Evaluation
This study utilized 3D Slicer to analyze proton radiotherapy plans incorporating eRBE models developed through the MS approach. 22 Dose distribution maps and dose-volume histograms (DVHs) were generated to compare dose assessments across different RBE models. This comparative analysis allowed for the evaluation of how variable eRBE models influence dose estimations for critical anatomical structures, such as the brainstem and spinal cord.23,24
By integrating multiple RBE models, this approach provides a broader perspective on dose variations and potential uncertainties in biological effectiveness calculations. The study highlights the importance of accounting for these variations in treatment planning to achieve a more comprehensive and balanced assessment of radiation dose distribution.
The key evaluations included the following: • Brainstem: Located at the distal end of the proton beam, the brainstem is highly sensitive to radiation. The fixed RBE value of 1.1 was found to underestimate biological doses in this region, especially under high LET conditions. The eRBE model provided a more nuanced and biologically accurate dose estimation. • Spinal Cord: The spinal cord received near-zero doses in this treatment plan, and minor differences were observed between the fixed RBE and variable eRBE models. • Left Parotid Gland: When far from the beam’s end, the left parotid gland showed negligible differences in dose assessment between the fixed and variable eRBE models, reflecting its distance from high-LET regions.
Radiosensitivity Parameters
Radiosensitivity parameters (α/β) were incorporated to refine biologically weighted dose calculations across tissues: • Brainstem (α/β = 2): Increased dose sensitivity, emphasizing the importance of accurate RBE modeling in this critical organ. • Parotid Gland (α/β = 3): Demonstrated moderate sensitivity, with minimal impact observed from variable RBE modeling.
Enhanced Dose Evaluation with eRBE
The eRBE-weighted approach enables precise analyses of biologically weighted dose distributions across critical anatomical structures. For instance: • Biological Dose Accuracy: The eRBE model, derived via MS methodology, mitigated the underestimation observed in fixed RBE (1.1) dose models, especially in high-LET regions such as the brainstem. • Organ-Specific Adjustments: By incorporating tissue-specific radiosensitivity parameters and LET distributions, the eRBE framework improved dose estimation across OARs. • Comparative DVH Analysis: Differences in dose estimates between fixed RBE and eRBE models were visualized through DVHs, offering insights into the potential risks of dose underestimation in critical organs.
Results
Ensemble RBE Modeling
Following the PRISMA guidelines, a systematic screening process was conducted to identify relevant studies for the development of eRBE models. This screening involved the evaluation of titles, abstracts, and full-text articles to ensure compliance with the inclusion criteria. The workflow is depicted in Figure 2, with panel (a) focusing on studies related to the brainstem and panel (b) on studies concerning the spinal cord. Flowchart for the Literature Search for Two Critical Organs: (A) Brainstem and (B) Spinal Cord in the Database. The PRISMA Guidelines Were Followed for the Title, Abstract, and Full-Text Screening Process. Abbreviation: RBE: Relative Biological Effectiveness, OER: Oxygen Enhancement Ratio, PRISMA: Preferred Reporting Items for Systematic Reviews and Meta-Analyses. Note: The Brainstem and Spinal Cord are Both Considered Serial Organs in Radiobiological Terms, Meaning that Even Small Focal Overdoses May Result in Significant Clinical Toxicity
Summary of RBE Models Used, Number of Cases, and Literature Quality Scores in Adopted Studies

RBE Models (Based on the Linear Quadratic Fitted Models: Cell Lines, Data Count, and Model Formulas)

Abbreviation: RBE: Relative Biological Effectiveness.


Models-Included and Weighting Estimation for Two eRBE Models

Abbreviation: RBE: Relative Biological Effectiveness, eRBE-B: Ensemble RBE from Brainstem literature collection, eRBE-SC: Ensemble RBE from Spinal Cord literature collection.
This ensures that the combined model remains consistent with the input RBE models and provides a properly normalized biological dose calculation.
These ensemble models integrate contributions from the Carabe, Wedenberg, and McNamara models to address the variability in RBE estimations. However, these models aggregate model usage trends rather than combining validated patient-derived RBE measurements.
Treatment Plan Simulation
Treatment plans were generated at RayStation 8B and imported into MCsquare for Monte Carlo simulations to calculate dose and LET distributions. Figure 3 shows the dose coverage achieved with 90° and 270° proton beams. The LET distributions revealed sharp increases at the distal end of the proton beam, highlighting potential underestimations in biological dose assessments if a fixed RBE value (1.1) is applied. This finding reinforces the need for variable RBE models to account for the spatial variability in LET and its biological impact. Dose Distribution and Linear Energy Transfer Profiles in MC Square Simulations of Beamlets for Head and Neck Cancer Treatment Plans With the Rando Phantom, including 90° Fields (A and B) and 270° Fields (C and D). Abbreviation: RBE: Relative Biological Effectiveness, OER: Oxygen Enhancement Ratio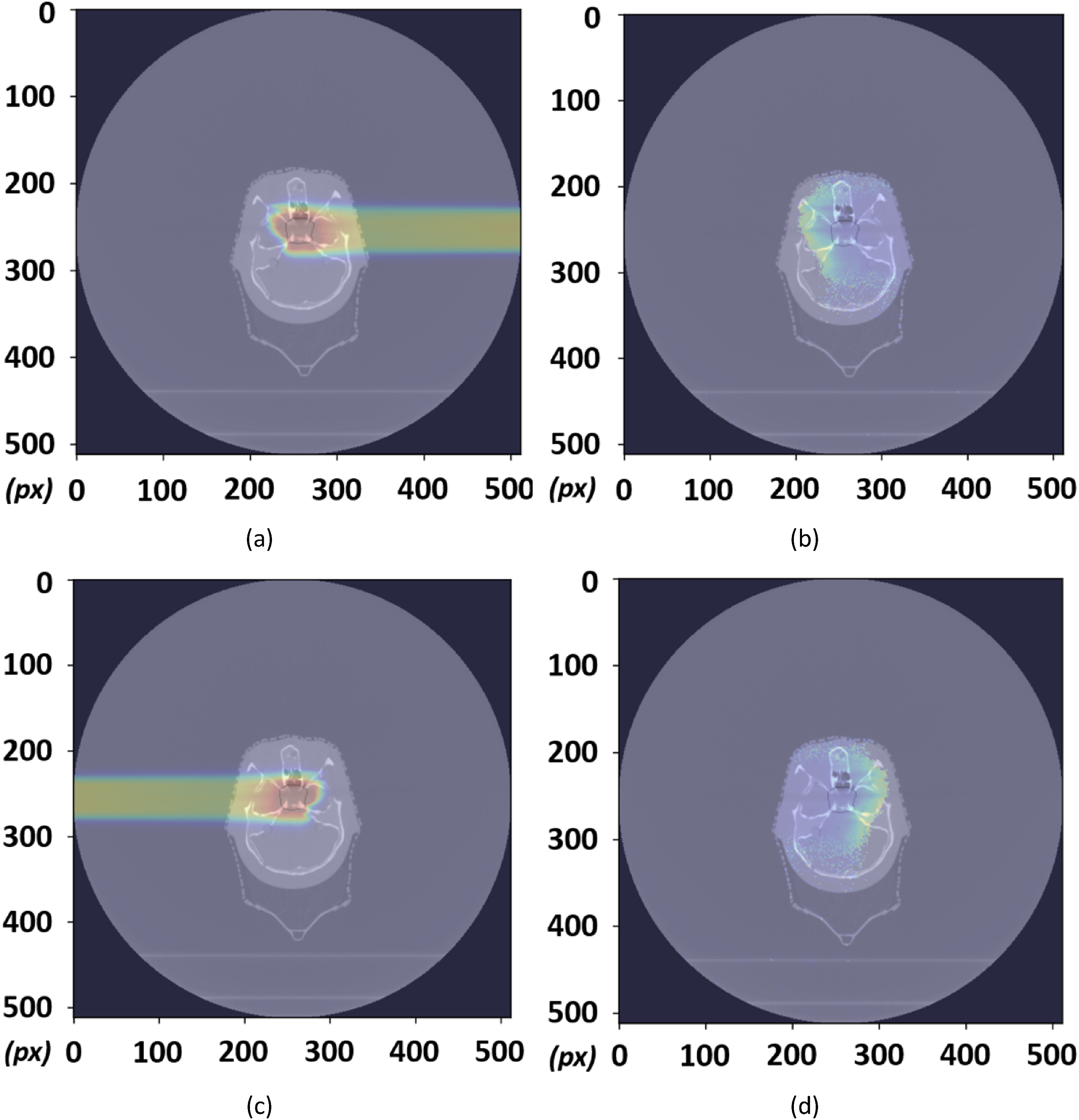
Biological Dose Weighting
Using the fixed RBE (1.1) as a baseline, dose distributions were recalculated for 3 variable RBE models and the 2 eRBE models (eRBE-B and eRBE-SC). These dose distributions, visualized in 3D Slicer, provided detailed insights into dose gradients for critical structures such as the brainstem, spinal cord, and left parotid gland. Figure 4 highlights these structures, with: • Pink for the CTV • Blue for the brainstem • Green indicates the left parotid gland. 3Dslicer Display of Variable RBE Weighted Dose Data in Sagittal, Coronal, and Axial Views for McNamara, eRBE-B, and eRBE-SC Distributions. Abbreviation: RBE: Relative Biological Effectiveness, eRBE-B: Ensemble RBE From Brainstem Literature Collection, eRBE-SC: Ensemble RBE From Spinal Cord Literature Collection

Dose‒volume Histograms (DVHs)
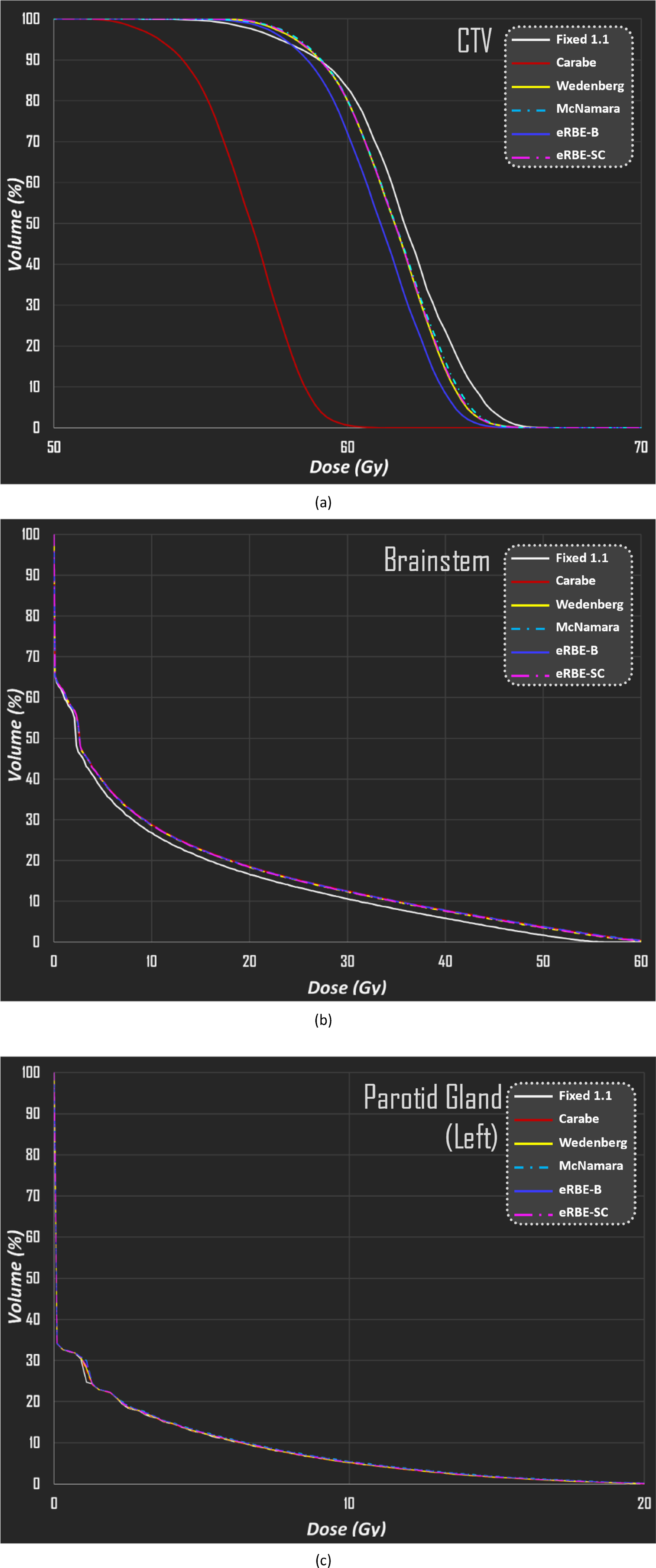
DVHs were generated for the CTV, brainstem, and left parotid gland, and the fixed RBE (1.1) model was compared with the variable and eRBE models. The DVHs (Figure 5) revealed the following: • CTV: The variable and eRBE models estimated slightly lower maximum doses than did the fixed RBE model. The Carabe model exhibited the most significant underestimation. • Brainstem: All the variable RBE models predicted higher doses than did the fixed RBE model, with close alignment among the models, underscoring the consistency of the ensemble models. These findings suggest that a fixed RBE underestimates biological doses near critical structures. • Left Parotid Gland: Due to its distance from the distal end of the proton beam, minimal differences were observed between the fixed and variable RBE models. Dose-Volume Histograms for Dose Assessment Using RBE Fixed at 1.1 and 5 Variable RBE Models (Carabe, Wedenberg, McNamara, eRBE-B, and eRBE-SC) in 3 Organs (Clinical Target Volume, Brainstem, Left Parotid Gland). Abbreviation: CTV: Clinical Target Volume, RBE: Relative Biological Effectiveness, eRBE-B: Ensemble RBE From Brainstem Literature Collection, eRBE-SC: Ensemble RBE From Spinal Cord Literature Collection
Radiosensitivity parameter analysis
The radiosensitivity parameter (α/β) was varied across 3 structures to assess dose differences. For the clinical target volume (CTV, α/β = 10), the Carabe model significantly underestimated doses (P < 0.05), while other models—including eRBE-B and eRBE-SC—showed only modest reductions (<1.6%) in
Dose Assessment With 6 RBE Models (Fixed 1.1, Carabe, Wedenberg, McNamara, eRBE-B, and eRBE-SC) Weighted Doses in 3 Organs (Clinical Target Volume, Brainstem, Left Parotid Gland) and the Difference in Assessment Doses Compared to RBE Fixed at 1.1

Abbreviations: eRBE-B, Ensemble RBE for Brainstem; eRBE-SC, Ensemble RBE for Spinal Cord.
Clinical Implications of eRBE Models
The ensemble models mitigated the variability and outlier behavior observed in single-model approaches. For example, the eRBE-B model closely aligns with the McNamara model for the brainstem, achieving
Discussion
Distinction Between MS and Meta-Analysis
Although MS is traditionally utilized for qualitative research, its application has expanded to include health systems research, computational decision modeling, and evidence synthesis in contexts where direct statistical pooling is precluded by data heterogeneity. In the current study, MS provided a structured and reproducible method to synthesize RBE model usage, theoretical frameworks, and methodological rigor across a heterogeneous body of literature. Unlike traditional meta-analysis—which statistically integrates quantitative outcomes—MS conceptually aggregates existing data patterns. This allowed identification of dominant modeling trends and enabled ensemble model development. Despite being unconventional in radiation oncology, this approach is consistent with methodologies in decision sciences and computational healthcare modeling, where structured synthesis of fragmented data is necessary.
This approach aligns with recent SR-MA methodologies proposed in radiation oncology literature, where systematic synthesis of modeling strategies is increasingly recognized for improving translational relevance (eg, 35,36). Given the lack of harmonized clinical RBE endpoints and model validation data, this synthesis-based methodology serves as a critical bridge between theoretical RBE modeling and future translational validation.
Model Selection and Weighting
Study Quality: Assessed via a standardized critical appraisal checklist to ensure methodological rigor.
Case Count: This reflects the empirical basis of each model.
To address the inherent biases in individual models, this study employed a weighted normalization approach. The normalization factor, defined as the sum of all weights (
Challenges in RBE Model Applications and Concerns on “Popularity Metrics”
Nevertheless, the reliance on reported data highlights the broader issue of variability in RBE model validation across studies.37,38 This underscores the urgent need for standardized clinical benchmarks and experimental validation to establish universally applicable RBE parameters. The eRBE framework is not intended as a definitive solution but rather as an exploratory tool to aggregate current knowledge and inform future research and clinical applications. Moreover, future iterations of the eRBE framework should explore the inclusion of biologically grounded models such as Microdosimetric Kinetic Model (MKM) and Repair-Misrepair-Fixation (RMF), particularly as more experimental and clinical validation data become available. This expansion will help diversify the ensemble and mitigate the current dominance of phenomenological models.
By addressing these challenges and limitations transparently, this study provides a stepping stone for advancing RBE model research while emphasizing the necessity of further clinical and experimental efforts.
Performance of eRBE Models
CTV: Under higher (α/β) ratios (10 Gy), the Carabe model underperformed, yielding RBE values below 1. This suggests potential underestimation of biologically weighted doses.
Brainstem: eRBE-B models consistently reduced dose estimation biases in high-LET regions, highlighting their adaptability.
Spinal Cord: The eRBE-SC model demonstrated stable dose estimations, outperforming fixed RBE models under variable (α/β) ratios.
While eRBE models provide improved consistency, their accuracy remains theoretical without direct clinical validation. This underscores the need for empirical data to validate the biological effectiveness of radiation treatments.
To further illustrate the potential clinical implications, a hypothetical increase of +10% in biological dose—as might result from underestimated RBE in fixed-model assumptions—could push an organ-at-risk such as the brainstem beyond its accepted tolerance limit. For instance, if the clinical tolerance of the brainstem is approximately 64-65 Gy (RBE), an unaccounted 10% increase may elevate the effective biological dose to ∼70-72 Gy (RBE), substantially raising the risk of radiation-induced necrosis, cranial neuropathy, or even life-threatening complications.
According to the QUANTEC guidelines,
39
the maximum tolerated dose
In proton therapy, LET increases near the distal edge of the Bragg peak, and applying a fixed RBE of 1.1 can underestimate the actual biological effect. Paganetti comprehensively reviewed the variability of RBE values and emphasized that RBE can exceed 1.1 in high-LET regions depending on tissue type, dose per fraction, and biological endpoint.
2
Thus, even modest RBE-weighted increases—such as those predicted by eRBE models—could elevate effective
Nonetheless, we acknowledge that the observed dose differences in this study did not reach statistical significance and may fall within the range of simulation or modeling uncertainty. As such, these findings should be interpreted as exploratory and hypothesis-generating rather than confirmatory. Further work is warranted to assess whether these modest dose deviations translate into clinically significant outcomes. Specifically, future studies should incorporate larger patient cohorts, enhanced Monte Carlo modeling accuracy, and retrospective outcome data to validate the clinical significance of RBE-driven dose variations in real-world treatment scenarios.
Future Directions and Methodological Contributions
Future studies should integrate clinical endpoints, such as the tumor control probability (TCP) and normal tissue complication probability (NTCP), to validate RBE models. 42 Additionally, incorporating broader databases (eg, Embase and the Cochrane Library) could increase the robustness of the MS process.
This study demonstrates the potential of eRBE models as an exploratory framework. By aggregating published data and leveraging weighted calculations, eRBE models offer a scalable method for addressing uncertainties in proton therapy dose assessments. However, the findings should be interpreted cautiously, serving as a precursor to experimental and clinical research aimed at achieving precise and clinically validated RBE models.
Limitations
This study has several limitations that must be acknowledged. First, the eRBE models were not validated against actual patient treatment outcomes. In clinical radiotherapy, each patient can receive only 1 treatment plan, making it ethically and practically infeasible to compare multiple biological dose models within the same individual. Therefore, this study relied on phantom-based simulations, which, while anatomically realistic, cannot fully capture patient-specific anatomical heterogeneity. Future studies should incorporate retrospective or prospective clinical datasets to evaluate correlations between eRBE-calculated doses and clinical endpoints, such as TCP and NTCP.
Second, this study focused on the brainstem and spinal cord because of their anatomical location at the distal edge of the Bragg peak, where LET variation is most pronounced. Although other OARs, such as the optic nerve and parotid glands, are clinically significant, the literature lacks sufficient quantitative data for reliable RBE modeling in these regions. Their exclusion reflects data limitations rather than a judgment of clinical importance. Future research should systematically develop RBE models for additional organs to improve the clinical applicability of the eRBE framework.
Third, the robustness of our results depends on the computational platform used. This study employed a single Monte Carlo environment without cross-validation across platforms. Validation using other systems, such as TOPAS or FLUKA, is necessary to ensure reproducibility and generalizability.
Finally, this study did not include formal uncertainty propagation for the Monte Carlo-derived dose and LET distributions. As such, confidence intervals for eRBE predictions were not quantified. Future work should incorporate robust uncertainty analysis—such as statistical convergence thresholds or repeated simulation trials—to improve the reliability and interpretability of biologically weighted dose estimates.
Despite these limitations, methodological safeguards were implemented. Model weights were determined using aggregated study quality scores and case numbers, which helped mitigate assessor bias and enhance the reproducibility of the meta-synthesis process.
Furthermore, we acknowledge that the number of recent publications on organ-specific RBE modeling remains limited. This constraint may reduce the temporal representativeness of the meta-synthesis and highlights the need for continued efforts to generate and include up-to-date clinical evidence.
Addressing biases in RBE model aggregation
The selected RBE models—Carabe, Wedenberg, and McNamara—were included based on their prevalence in the literature and distinct theoretical frameworks rooted in the linear-quadratic (LQ) algorithm. While each standalone model offers valuable insight, all are subject to limitations stemming from data selection, regression assumptions, and radiobiological simplifications. To mitigate these constraints, we developed eRBE models by systematically aggregating model usage patterns, case numbers, and methodological quality scores, applying normalization to ensure balanced contributions.
However, we explicitly acknowledge that combining model popularity, study quality, and case counts may introduce biases associated with publication frequency and the influence of prolific research groups. This aggregation strategy does not inherently guarantee biological or clinical accuracy, nor does it equate to definitive model validation. Rather, the eRBE framework serves as a transparent, exploratory synthesis of existing practices—intended to inform, not replace, future validation efforts.
Our analysis demonstrated that the eRBE approach effectively reduced dose underestimations in critical structures, particularly within high-LET regions. For example, the eRBE-B model corrected systematic biases observed in the Carabe model, leading to improved dose consistency across varying biological conditions. Still, the clinical significance of these improvements remains provisional, as the models are derived from aggregated theoretical data rather than patient-level clinical endpoints.
To further address concerns related to “popularity metrics,” methodological rigor—assessed using the Joanna Briggs Institute criteria—was incorporated into the weighting scheme. This strategy helped mitigate the influence of research output volume. Nonetheless, ensemble weighting strategies must be continually reassessed in light of evolving clinical evidence and emerging radiobiological insights. Ultimately, direct clinical validation and integration of patient-specific outcomes will be necessary to translate ensemble RBE models into routine practice.
Conclusion
This study introduced eRBE models using a MS approach to address uncertainties in proton radiotherapy dose calculations for critical organs, such as the brainstem and spinal cord. By aggregating patterns of model usage and theoretical foundations, the eRBE framework demonstrated improved stability and consistency in dose estimations compared with standalone models.
However, these findings represent a preliminary proof-of-concept, based exclusively on simulation data without validation against clinical outcomes. The eRBE models should therefore be interpreted as probabilistic tools for exploring biological dose variability rather than definitive replacements for clinical RBE assignments.
Future research should focus on validating the proposed models through real patient treatment data, assessing correlations with clinical endpoints such as TCP and NTCP. Such studies are essential for translating the theoretical advancements demonstrated here into practical clinical applications.
Footnotes
Acknowledgment
This study was supported financially, in part, by grants from the National Science and Technology Council (NSTC) of the Executive Yuan of the Republic of China (111-2221-E-992-016-MY2, 113-2221-E-992-011-MY2).
Ethical Considerations
Institutional review board approval was not needed because this study did not involve human participants.
Author Contributions
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Grants from the National Science and Technology Council (NSTC) of the Executive Yuan of the Republic of China (111--2221--E-992--016--MY2, 113--2221--E-992--011--MY2).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.





