Abstract
Background
As a new class of non-coding RNAs, circRNAs have been recently reported to be involved in the tumorigenesis and progression of human cancers. In the current study, we attempted to explore the potential function of a novel circRNA (hsa_circ_0013290) in hepatocellular carcinoma (HCC).
Methods
Relative hsa_circ_0013290 expression was analyzed by quantitative reverse transcription-polymerase chain reaction (qRT-PCR). The subcellular location of hsa_circ_0013290 was performed by RNA subcellular isolation and fluorescence in situ hybridization (FISH) assays. The effect of hsa_circ_0013290 on proliferation was detected by Cell Counting Kit-8 (CCK-8) assays. The effect of hsa_circ_0013290 on cell cycle distribution and apoptosis was detected by flow cytometry. The invasion and migration abilities of hsa_circ_0013290 were detected by transwell assays.
Results
Hsa_circ_0013290 is significantly upregulated in HCC cell lines and mainly located in cytoplasm of HCC cells. Hsa_circ_0013290 overexpression promotes cell invasion and migration and inhibits cell apoptosis. In contrast, hsa_circ_0013290 knockdown impedes cell invasion and migration and accelerates cell apoptosis. However, hsa_circ_0013290 did not affect cell proliferation.
Conclusions
Hsa_circ_0013290 is overexpressed in HCC cell lines and is mainly located in the cytoplasm of HCC cells. Hsa_circ_0013290 promotes cell invasion and migration, and inhibits cell apoptosis.
Introduction
Hepatocellular carcinoma (HCC) is the most common digestive malignancy and has a poor prognosis. 1 Up till now, the best curative treatment option for HCC patients is surgical resection and liver transplantation. While most patients are usually diagnosed at an advanced stage and are too late to surgery. Thus, HCC has a high recurrence and metastasis probability. 2 Therefore, a deeper investigation of the exact mechanisms that regulates HCC progression and metastasis is crucial for HCC clinical therapy.
Circular RNA (circRNA), a class of special non-coding RNA, is stably present in eukaryotic cells and has a higher tolerance to RNA exonuclease due to its closed loop structure. 3 CircRNAs have been reported to act as either oncogenes or tumor suppressors in the progression of human cancers, 4 including HCC, gastric cancer, and prostate cancer. Huang el al. showed that circMET acts as an oncogene and that circMET overexpression induces HCC development and immune tolerance via the miR-30-5p/Snail/DPP4/CXCL10 axis. 5 Liu et al. demonstrated that circYAP1 suppressed gastric cancer progression by circYAP1/miR-367-5p/p27 Kip1 axis and may provide a prognostic indicator of survival in gastric cancer patients. 6 Feng et al. verified that circ0005276 activated the transcription of XIAP by interacting with FUS binding protein and then promoted the proliferation and migration of prostate cancer cells. 7
Here, we identified a novel circRNA (circAGL, also named as hsa_circ_0013290), derived from the exon region of the AGL gene, which is significantly upregulated in the HCC cells. The functional assays revealed that hsa_circ_0013290 promoted cell invasion and migration, inhibiting cell apoptosis of HCC in vitro. Thus, animal experiment showed that knockdown hsa_circ_0013290 resulted in cell growth rate slower in vivo. Our results indicated that hsa_circ_0013290 may be a novel potential target for HCC treatment.
Materials and Methods
Cell Lines and Cell Culture
Human HCC cell lines (Hep 3B2.1-7, HuH-7, Li-7, PLC/PRF/5 and SK-HEP-1) and the normal human hepatic cell line (L‐02) were kindly provided by Stem Cell Bank, Chinese Academy of Sciences (Shanghai, China). Hep 3B2.1-7, PLC/PRF/5, and SK-HEP-1 cells were cultured in MEM (added with NaHCO3 and sodium pyruvate) with 10% fetal bovine serum (FBS; Cell Sciences, Canton, MA). Li-7 and L‐02 cells were cultured in RPMI‐1640 (Gibco, Thermo Fisher Scientific, Waltham, MA) with 10% FBS. HuH‐7 cells was cultured in Dulbecco’s modified Eagle’s medium (Gibco, Thermo Fisher Scientific) with 10% FBS. All cells were cultured at 37°C in a humidified incubator under 5% CO2 conditions.
RNA Subcellular Isolation
The cells were harvested by trypsin and washed with phosphate-buffered saline (PBS). The nuclear and cytoplasmic RNA were separately isolated following RNA Subcellular Isolation Kit instructions (Active Motif, Carlsbad, CA).
Fluorescence in Situ Hybridization (FISH) Assay
Cells were seeded onto Lab-Tek®II Chamber Slides (Thermo Fisher Scientific) at a density of 1× 104 cells/well and cultured overnight at 37°C in a humidified incubator under 5% CO2 conditions. The next day, cell chamber slide was fixed with 4% paraformaldehyde, permeated with .1% tritonX-100, and washed by PBS. All slides were dehydrated in a series gradient alcohol and dried at room temperature. The hsa_circ_0013290 probe (20 μg/mL) hybridization solution was pre-heated at 73°C for 5 min and was added onto the slides overnight in the 37°C incubator. The second day, the slices were washed three-times using Wash Buffer Solution. Cell nuclei were counterstained with 4,6-diamidino-2-phenylindole (DAPI), and the images were obtained on an inverted fluorescence microscope (Olympus, Japan). The hsa_circ_0013290 probe for fluorescence in situ hybridization (FISH) was 5′‐ATCCATTTACTATTAGCAGCTCAGCTACGTGAAGAGGTG‐3′.
RNA Extraction, RNase R Treatment, and Quantitative Reverse Transcription-Polymerase Chain Reaction (qRT-PCR)
Total RNA was extracted using RNAiso Plus (Takara, Beijing, China) according the protocol. RNase R treatment was processed at 37°C with .1 U/μL of RNase R (Geneseed, Guangzhou, China) for 5 min and inactivated at 70°C for 10 min. Treated RNA was then reverse transcribed to complementary DNA (cDNA) using PrimeScript RT Master Mix (Takara). The cDNA was used as a template for detecting the relative expression of hsa_circ_0013290 with TB Green Premix Ex Taq II (Takara); GAPDH was used as an internal control gene. The primer sequences were as follows: hsa_circ_0013290 forward 5′‐GCCACTCAACACCTCTTCACG‐3′, reverse 5′‐ACAGGAGAAACGCCAAAGTGC‐3′; GAPDH forward 5′‐CAATGACCCCTTCATTGACC‐3′, reverse 5′‐TTGATTTTGGAGGGATCTCG‐3′.
SiRNA Transfection
For hsa_circ_0013290 knockdown, siRNA specifically targeting hsa_circ_0013290 and the negative control (si-NC) were constructed by Genepharma (Suzhou, Jiangsu, China). PLC/PRF/5 cells were transfected with siRNA (20 μM) using LipofectamineTM 2000 (Thermo Fisher Scientific) according to the manufacturer’s instructions. The siRNA sequences were si‐1: 5′‐CUUCACGUAGCUGAGCUGCTT‐3′; si‐2: 5′‐CACGUAGCUGAGCUGCUAATT‐3′.
Lentivirus Packaging and Infection
For hsa_circ_0013290 overexpression, lentivirus of hsa_circ_0013290 (Lv-hsa_circ_0013290) and the negative control (Lv-NC) were constructed and produced by Obio Technology (Shanghai, China). Lentivirus solution (multiplicity of infection = 20) and polybrene were added to Hep 3B2.1-7 cells. After 72 h, the cells with an infection efficiency >90% were selected for subsequent studies.
Transwell Assay
For the invasion assay, the upper chamber was pre-coated with Matrigel (BD Biosciences, San Jose, CA, USA). For transwell assays (migration and invasion), the cells were suspended with serum‐free medium at a density of 3 × 105 cells/mL and 100 μL cell suspension was then seeded onto the upper chamber (8 μm, Millipore). To attract the cells, the appropriate complete medium with 20% FBS was added in the bottom chamber. After incubation for 36 h, cells that stayed in the upper chamber were removed and the ones that passed through the membrane were stained with methanol and crystal violet (Beyotime). The number of cells that migrated or invaded was calculated from 5 random fields by using an optical microscope (Olympus, Japan).
Cell Cycle and Apoptosis Analysis by Flow Cytometry
For cell cycle assay, the cells were harvested and stained by 70% cold ethanol overnight, treated with PI/RNase R Staining Buffer (BD) according to the manufacturer’s protocol. For cell apoptosis assay, an Annexin V/7-AAD Apoptosis Detection Kit (BD) was applied to stain cells. Both experiments were assessed by flow cytometry using FlowJo software (BD).
Cell Counting Kit‐8 Assays
Cell viability was monitored using Cell Counting Kit-8 (CCK-8, Dojindo, Japan). At 48 h after transfection, each group of cells was seeded onto 96-well plates at a density of 3 × 103 cells/well. Cell viability detection was measured at 2 h after addition of CCK-8 reagent every 24 h under OD450.
Animal Experiment
Specific pathogen‐free grade male nude mice (4–5 weeks old; purchased from Experimental Animal Center, Shanghai Institute of Materia Medica Chinese Academy of Sciences) were fed in the Laboratory Animal Center of Nantong University. The procedures were approved by the Animal Experimental Ethics Committee of Nantong University. PLC/PRF/5 cells were harvested by trypsin, washed and re-suspended in PBS for preparation. A total of 1× 107 cells in .1 mL of PBS were injected into each flank area of nude mice. After 20 days of growth, tumor volumes were calculated (.5 × length × width × width) and the mice were randomized into si-NC or si-1 groups for further study. 2′OMe +5′chol-modified si-hsa_circ_0013290 (si-1: 5′‐CTTCACGTAGCTGAGCTGC‐3′) and negative control (si-NC) were purchased from Ribobio for in vivo siRNA delivery. SiRNA (5 nmol) in 50 μL PBS was injected into each tumor mass every 2 days for 2 weeks. After 5 weeks, the mice were sacrificed, and tumors were collected and embedded in paraffin for furtherstudies.
Immunohistochemistry
The mouse tumor specimens were fixed with 4% formaldehyde, and then embedded in paraffin. Ki-67 analysis: After deparaffinization, antigen was retrieved in citrate buffer (pH 6.0). The sections were incubated with the primary and a secondary antibody. Then 3, 3′-diaminobenzidine (DAB, Maixin-Bio, Guangzhou, China) and hematoxylin were applied for staining. Cell apoptosis in situ: Use Proteinase K to permeabilize the deparaffinized sections. After rinsing twice with PBS, incubated the section with TUNEL reaction mixture in a dark humidified box at 37°C for 1 h. Rinse the specimens with PBS, and DAB reagents. The morphological characteristics of apoptotic cells were observed with an optical microscopy and the pictures were taken.
Statistical Analysis
GraphPad Prism 8.0 and SPSS 24.0 were used for data analysis. All data were presented as the mean ± standard deviation (SD). Statistical significance was defined as P < .05.
Results
The Relative Expression and Biological Structure of hsa_circ_0013290
The relative mRNA expression levels of hsa_circ_0013290 were found to be significantly high in 4 of the 5 HCC cell lines tested (Hep 3B2.1-7, HuH-7, Li-7, PLC/PRF/5, and SK-HEP-1) compared to that of L‐02 (the normal human hepatic cell line) (Figure 1A). In particular, the PLC/PRF/5 cells (P < .001) showed the greatest degree of upregulated expression of hsa_circ_0013290, whereas Hep 3B2.1-7 cells (P < .01) expressed the lowest levels of hsa_circ_0013290 compared to others. Herein, we chose PLC/PRF/5 and Hep 3B2.1-7 cells for further experiments to assess the effects of hsa_circ_0013290. The genomic structure indicates that hsa_circ_0013290 consists of 7 exons (947 bp) from the AGL gene (amylo-alpha-1, 6-glucosidase). This circular product was amplified by qRT-PCR with divergent primers and confirmed by Sanger sequencing (Figure 1B). Convergent and divergent primers were designed to amplify linear and circular RNA based on cDNA and genomic DNA (gDNA) by RT-PCR, respectively. As shown in Figure 1C, hsa_circ_0013290 could only be amplified by primers in cDNA, but not in gDNA. In addition, the stabilities of the linear RNA and circular were compared after digestion with the specific exonuclease, RNase R. QRT-PCR analysis showed that AGL gene was 60% degraded with 2U RNase R; in contrast, hsa_circ_0013290 was only 16% degraded (Figure 1D). The relative hsa_circ_0013290 and the parent gene AGL level of PLC/PRF/5 cells after knockdown hsa_circ_0013290 were confirmed by qRT-PCR. The parent gene AGL had no significant change, while the 2 siRNAs downregulated hsa_circ_0013290 level by 72% and 57%, respectively. Both 2 siRNAs had an obvious interference efficiency; therefore, they were used for further experiments (Figure 1E). QRT-PCR analysis also confirmed that the infection of Lv-hsa_circ_0013290 successfully overexpressed hsa_circ_0013290 in Hep 3B2.1-7 cells (Figure 1F), and the product was also confirmed by Sanger sequencing (Figure 1G). The relative expression and biological structure of hsa_circ_0013290. (A) The relative expression level of hsa_circ_0013290 in L-02 and HCC cells were measured by qRT-PCR. (B) The schematic diagram showed that hsa_circ_0013290 was consisted of 7 exons from the AGL gene, and the red arrow indicated its splicing junction. (C) RT-PCR assay demonstrated that divergent primers detected circular RNAs in cDNA but not gDNA. GAPDH was used as the negative control. (D) QRT-PCR was applied to measure of the expression of hsa_circ_0013290 and AGL after RNase R treatment. (E) QRT-PCR analysis was used to verify interference efficiency of hsa_circ_0013290 in PLC/PRF/5 cells, and the relative level of parent gene AGL. (F) QRT-PCR analysis was used to verify overexpression efficiency of hsa_circ_0013290 in Hep 3B2.1-7 cells. (G) Sanger sequence assay showed the back splicing site after overexpressing hsa_circ_0013290. **P < .01, ***P < .001, ****P < .0001.
Hsa_circ_0013290 Is Predominantly Localized in Cytoplasm
To determine the subcellular distribution of hsa_circ_0013290, we performed qRT-PCR using RNA samples isolated exclusively from either cytoplasm or nuclear fractions of Hep 3B2.1-7 and PLC/PRF/5 cells. We found that 87% of hsa_circ_0013290 was distributed in the cytoplasm of Hep 3B2.1-7 cells, while 13% was in the nucleus (Figure 2A). The fluorescence in situ hybridization (FISH) assay showed that hsa_circ_0013290 predominately localized in the cytoplasm of Hep 3B2.1-7 cells (Figure 2B). In line with the findings in Hep 3B2.1-7 cells, qRT-PCR (Figure 2C) assays also revealed that hsa_circ_0013290 was located mainly in the cytoplasm (84%) of PLC/PRF/5 cells, and FISH assay showed that the majority part of hsa_circ_0013290 localized in the cytoplasm (Fig. 2D). Hsa_circ_0013290 is predominantly localized in cytoplasm. (A) The percentage of hsa_circ_0013290 expression level in the cytoplasmic and nuclear of Hep 3B2.1-7 cells. (B) RNA-FISH indicated hsa_circ_0013290 subcellular location in Hep 3B2.1-7 cells. (C) The percentage of hsa_circ_0013290 expression level in the cytoplasmic and nuclear of PLC/PRF/5 cells. (D) RNA-FISH indicated hsa_circ_0013290 subcellular location in PLC/PRF/5 cells.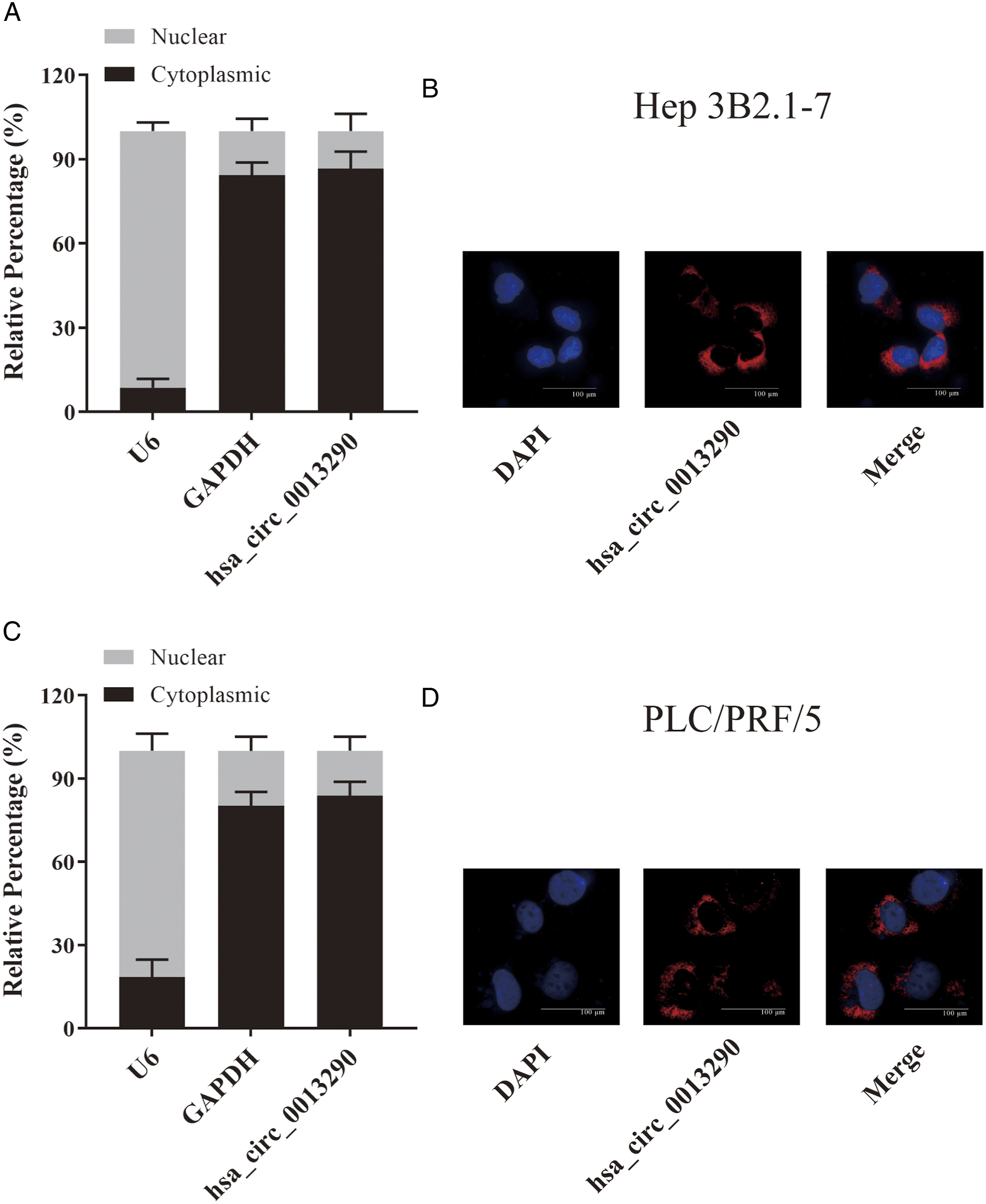
Hsa_circ_0013290 Suppresses Apoptosis in HCC Cell Lines
The cell apoptosis analysis showed that the percentage of apoptotic cells was significantly higher in PLC/PRF/5 cells transfected with siRNAs than in the si-NC group, indicating that knockdown of hsa_circ_0013290 induced apoptosis in HCC cells (Figure 3A and B). Lv-hsa_circ_0013290 resulted in a decrease in the rate of Hep 3B2.1-7 cell apoptosis compared with that detected in the Lv-NC group (P < .0001), including late (P < .05) and early (P < .001) apoptosis (Figure 3C and D). Hsa_circ_0013290 suppresses apoptosis in HCC cell lines. (A&B) Flow cytometry analysis was performed to detect the effect of hsa_circ_0013290 knockdown on cell apoptosis of PLC/PRF/5 cells. (C&D) Flow cytometry analysis was performed to detect the effect of hsa_circ_0013290 overexpression on cell apoptosis of Hep 3B2.1-7 cells. *P < .05, **P < .01, ***P < .001, ****P < .0001.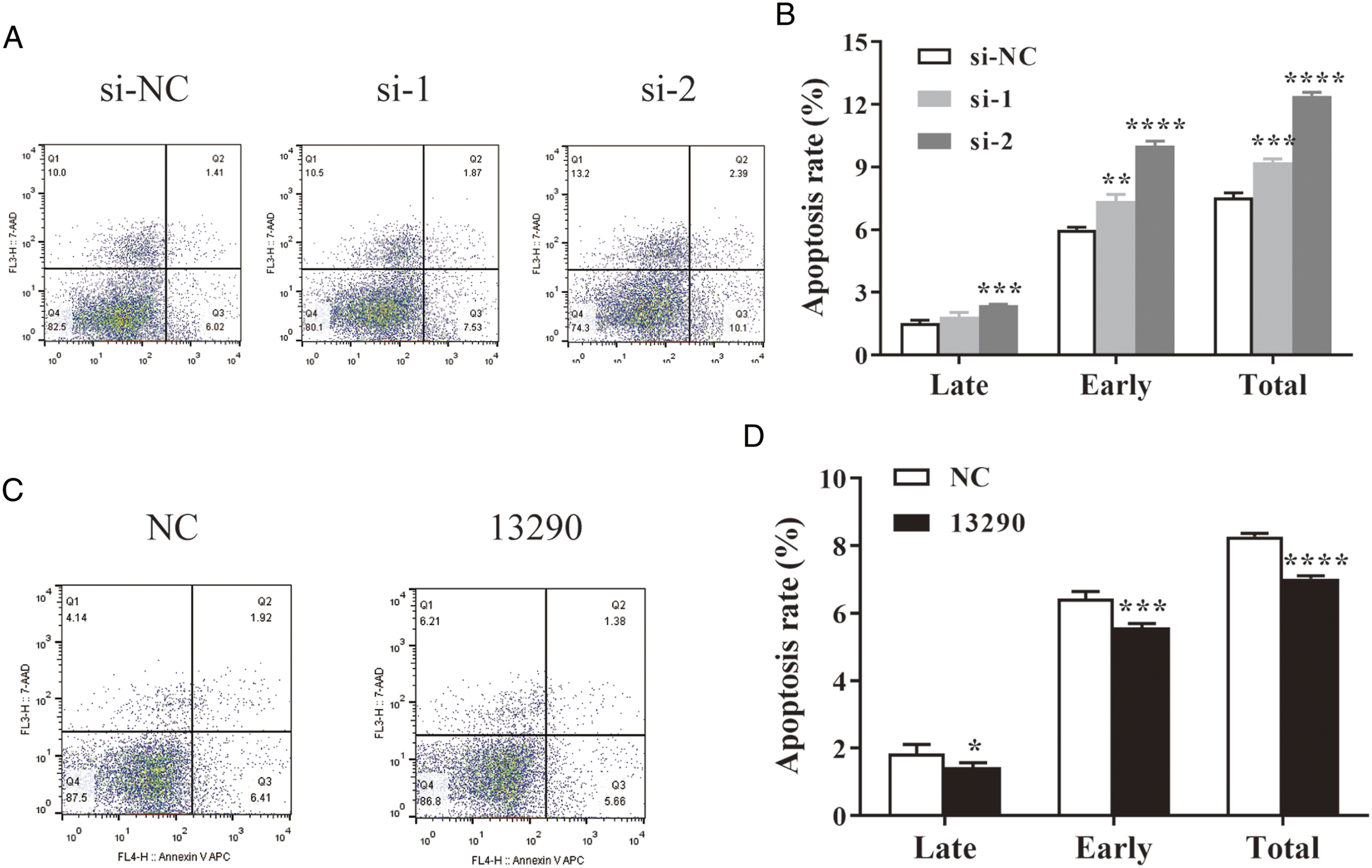
Hsa_circ_0013290 Accelerates HCC Cell Invasion and Migration
The transwell assays showed that when hsa_circ_0013290 knocked down, cell invasion and migration abilities were dramatically impaired compared to the negative control (si-NC) group (Figure 4A and B). In contrast, hsa_circ_0013290 overexpression accelerated the invasion and migration abilities compared to that of the negative control (Lv-NC) group (Figure 4C and D). Hsa_circ_0013290 accelerates HCC cell invasion and migration. (A&B) Transwell assays were used to examine the effect of hsa_circ_0013290 knockdown on cell invasion and migration of PLC/PRF/5 cells. (C&D) Transwell assays were used to examine the effect of hsa_circ_0013290 overexpression on cell invasion and migration of Hep 3B2.1-7 cells. **P < .01, ***P < .001, ****P < .0001.
Hsa_circ_0013290 Does Not Affect Proliferation and Cell Cycle in HCC Cell Lines
CCK-8 assay showed that neither inhibition (Figure 5A) nor overexpression (Figure 5B) could dramatically affect the proliferative capability of HCC cells, compared with the relative control group. Flow cytometry assays revealed that when hsa_circ_0013290 knocked down, there is no clear differences in the percentage of each stage cells compared to the si-NC group in PLC/PRF/5 cells (Figure 5C and D), and similar phenomenon had been observed when hsa_circ_0013290 overexpressed in Hep 3B2.1-7 cells (Figure 5E and F). The above 2 results suggesting that hsa_circ_0013290 does not affect proliferation and cell cycle in HCC cells. Hsa_circ_0013290 does not affect proliferation and cell cycle in HCC cell lines (A) CCK-8 assay was applied to determine the proliferative ability of cells when hsa_circ_0013290 knocked down in PLC/PRF/5 cells. (B) CCK-8 assay was applied to determine the proliferative ability of cells when hsa_circ_0013290 overexpressed in Hep 3B2.1-7 cells. (C&D) Flow cytometry analysis was performed to detect the effect of hsa_circ_0013290 knockdown on the cell cycle distribution of PLC/PRF/5 cells. (E&F) Flow cytometry analysis was performed to detect the effect of hsa_circ_0013290 overexpression on the cell cycle distribution of Hep 3B2.1-7 cells.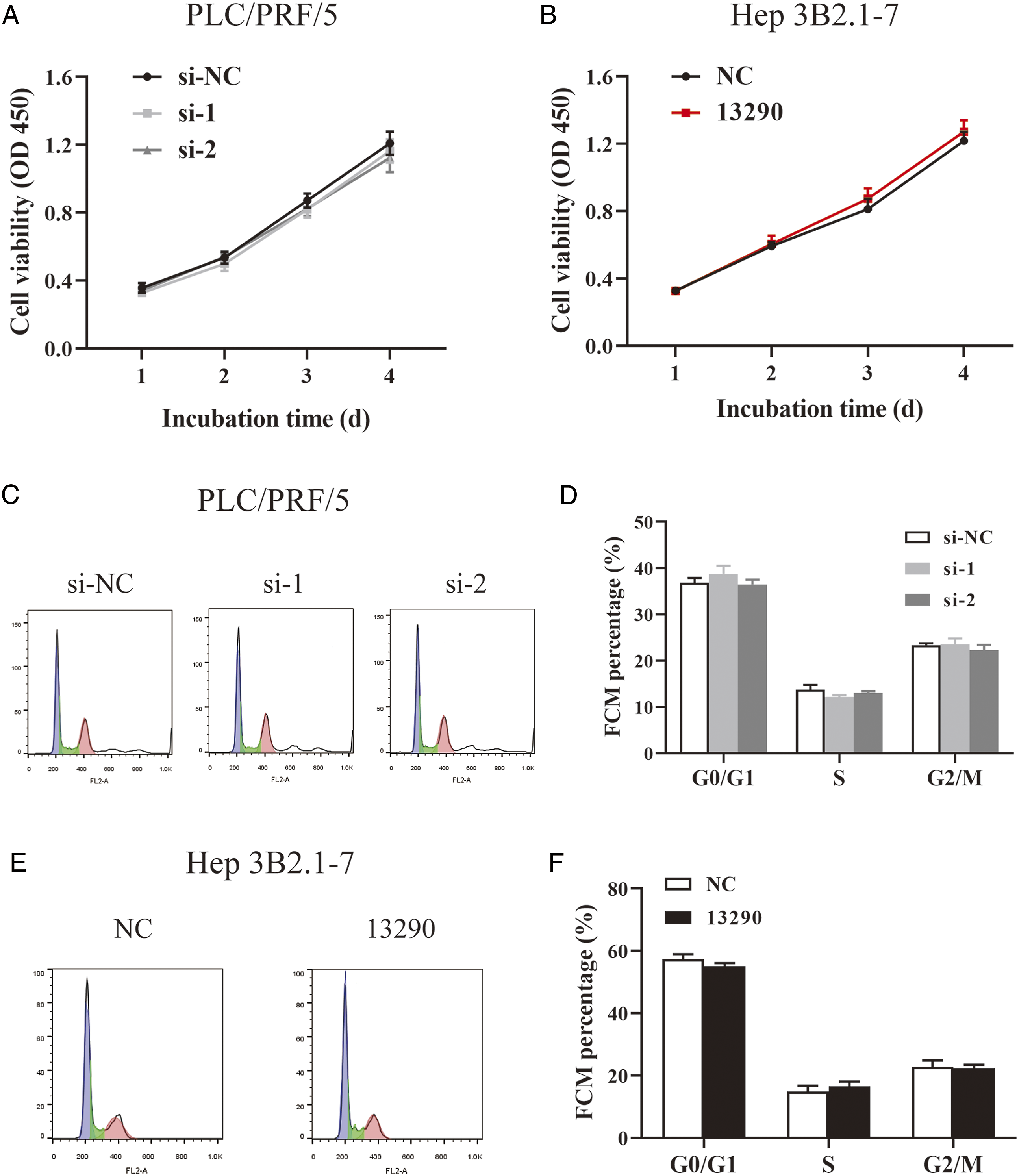
Knockdown hsa_circ_0013290 Suppresses HCC Cell Proliferation in Vivo
Compared with mice in the si‐NC group, tumor growth rates were significantly slower in the si‐1 group, and the tumors had lower mean volume (P < .05) (Figure 6A and B). The mean weight of tumors derived from si-1 group was also significantly less than tumors from si-NC group (Figure 6C). QRT-PCR demonstrated that the relative expression of hsa_circ_0013290 was much lower in si-1 group than in si-NC group (P < .05) (Figure 6D). Hematoxylin and eosin (H & E) staining revealed no visible morphological changes between the 2 groups. Immunohistochemistry assays indicated a decline in Ki‐67 expression and an increase of cell apoptosis in the si‐1 group compared to that in the si‐NC group (Figure 6E). Knockdown hsa_circ_0013290 suppresses HCC cell proliferation in vivo. (A) Growth curvature of the tumors in nude mice. (B) Tumor images for nude mice in the si‐NC and si‐1 groups. (C) The mean weight of tumors derived from si-NC and si-1 group. (D) The relative expression of hsa_circ_0013290 in tumors from si-NC and si-1 groups. (E) Representative photograph of the tumors from the 2 groups. Left, H&E (hematoxylin and eosin) staining; middle, Ki‐67 expression; right, cell death in situ. *P < .05, **P < .01.
The Prediction of miRNAs and Proteins that Binding on hsa_circ_0013290
To further study the mechanism of hsa_circ_0013290, several bioinformatics websites (circular RNA Interactome,
8
miRDB,
9
starbase,
10
and targetscan
11
) were used to select potential miRNAs and their binding-proteins. The miRNAs and proteins were selected and showed in the schematic model (Figure 7A). After statistical analysis, we selected 7 potential proteins (AAK1, AGO1, JAZF1, NOX4, RRAGD, TBC1D15, and ZBTB20). QRT-PCR demonstrated that the relative expression of NOX4 and RRAGD was significantly higher in PLC/PRF/5 cells transfected with siRNAs. (P < .05) (Figure 7B). The prediction of miRNAs and proteins that binding on hsa_circ_0013290. (A) A schematic model showing the predicted miRNAs and proteins targeting on hsa_circ_0013290. (B) Relative expression of NOX4 and RRAGD in PLC/PRF/5 cells transfected with siRNAs.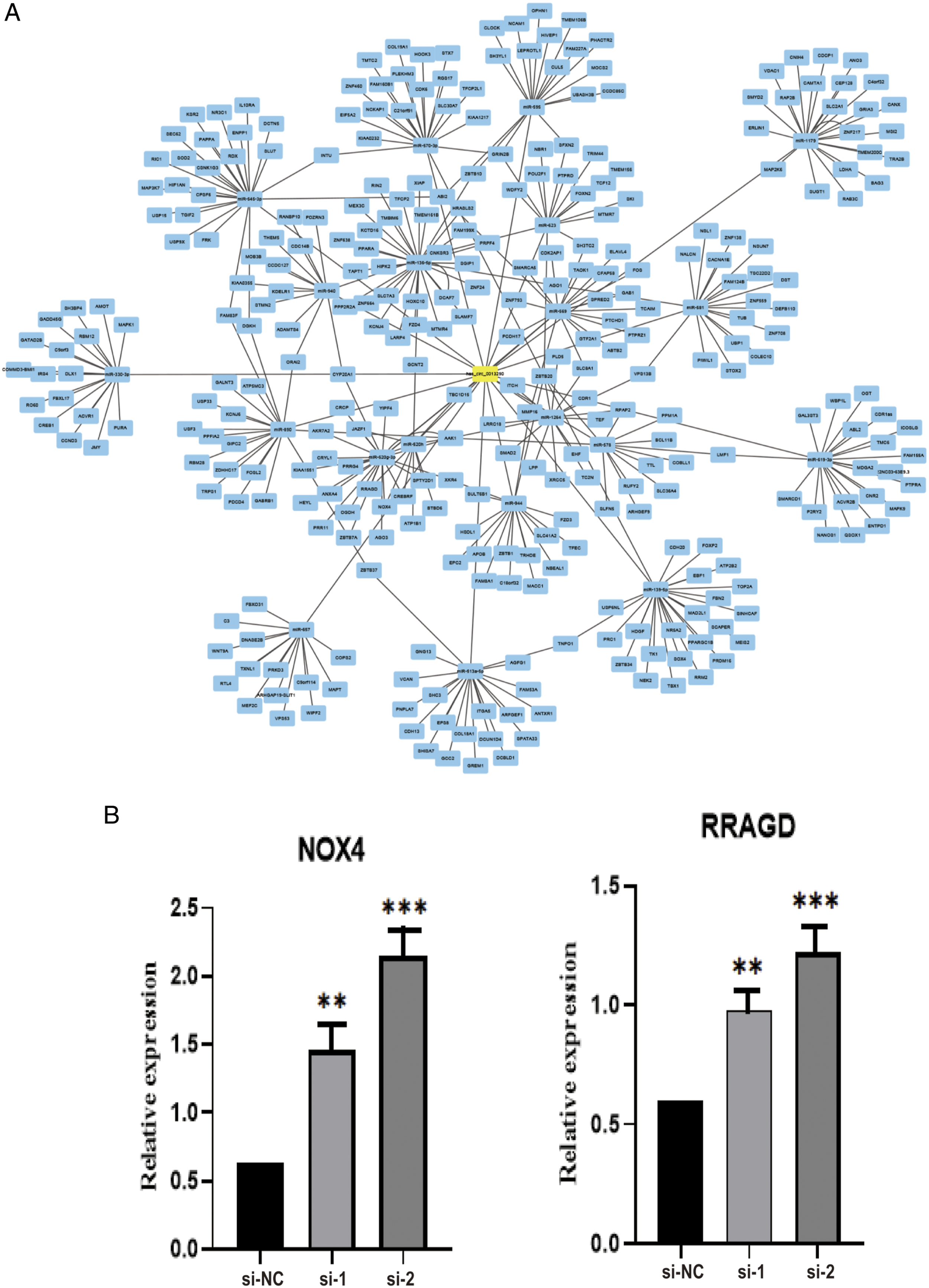
Discussion
Although circRNAs were first discovered decades ago, they were early been mistaken for “trash RNA” without arousing much attention. 12 Since this century, large amounts of studies have resulted in more and more understandings of circRNAs.13,14 CircRNAs can be divided into 4 categories according to its origins: exonic circRNAs (ecircRNA), intronic circRNAs, exon-intron circRNAs (EIciRNA), and intergenic circRNAs. 15 CircRNAs can occur in gene regions or intergenic regions, with the length from few hundred to thousands of nucleotides. Because of lack of 3′ poly A tail and 5′ cap, circular RNAs are resistant to exonuclease and more stable than linear RNA isoforms. 16
With the rapid development of high-throughput and RNA-sequencing technology, the extensive properties and functions of circRNAs have been widely clarified in multiple human cancers.17,18 For instance, Wang et al. reported that circRHOT1 promoted HCC development and progression via TIP60-dependent NR2F6 expression. 19 Wu et al. showed that circ_0006156 was highly expressed in papillary thyroid cancer tissues and cell lines. The survival curve analysis displayed that high level circ_0006156 predicted poor prognosis of patients. Mechanically, circ_0006156 acted as a competing endogenous RNA to sponge miR-1178 and then regulated Toll-like receptor 4 protein in papillary thyroid cancer cells. 20 Compared to healthy people, plasma circ-SLC7A5 expression was significantly upregulated in esophageal squamous cell carcinoma patients, and the total area under the ROC curve was .7717. The ROC and Kaplan–Meier survival curves indicated that circ-SLC7A5 could serve as a novel diagnostic and prognostic biomarker for esophageal squamous cell carcinoma due to its noninvasive feature. 21
In this study, we discovered and identified a novel circRNA, hsa_circ_0013290, in HCC cells. Hsa_circ_0013290 consists of the 6th–12th exons from the AGL gene, which has been confirmed by the Sanger sequencing. The relative expression was measured using qRT-PCR, which showed that hsa_circ_0013290 was significantly upregulated in HCC cells. The subcellular distribution and FISH assays demonstrated that hsa_circ_0013290 was mainly located in the cytoplasm of HCC cells. When downregulated hsa_circ_0013290, cell invasion and migration abilities had been inhibited, and cell apoptosis rate was increased. In contrast, overexpressed hsa_circ_0013290 resulted in more cell invasion and migration and less cell apoptosis rate. However, hsa_circ_0013290 has no effect on proliferation and cell cycle of HCC cells.
Conclusions
To sum up, hsa_circ_0013290 expression is upregulated in HCC cells, and a series of in vitro function experiments demonstrated that hsa_circ_0013290 promotes the metastasis in HCC and inhibits cell apoptosis.
This is the first study to provide evidence of hsa_circ_0013290 as a metastatic promoter in HCC. Further investigations of the specific mechanisms on how hsa_circ_0013290 promotes HCC metastasis depend on more detailed research in the future.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by grants from the Project of Jiangsu Province Youth Medical Talent Development (grant No. QNRC2016400), the six peak talents in Jiangsu Province (grant YY-177), the Project of Nantong Youth Medical Talent Development (grant No. 05), the Nantong Science and Technology Bureau (grant No. MS22018007, MS22018010, MS22019021, JCZ18116, JCZ18057, MSZ18269), the Health Bureau of Nantong City (grant No. WKZL2018057, MB2021057).
Ethical Approval
All study procedures involving animal experiments were consistent with the requirements of the Lab Animal Ethical Committee of Nantong University.
Statement of Human and Animal Rights
All study procedures involving animal experiments were consistent with the requirements of the Lab Animal Ethical Committee of Nantong University, and Ethics Committee reference number is S20210116-201.
Informed Consent
There are no human subjects in this article, and informed consent is not applicable.
