Abstract
Kynurenine 3-monooxygenase (KMO) is the pivotal enzyme in the kynurenine pathway and is located on the mitochondrial outer membrane. The dysregulation of KMO leads to various neurodegenerative diseases; however, it is rarely mentioned in cancer progression. Our previous study showed that KMO overexpression in canine mammary gland tumors (cMGT) is associated with poor prognosis in cMGT patients. Surprisingly, it was also found that KMO can be located on the cell membranes of cMGT cells, unlike its location in normal cells, where KMO is expressed only within the cytosol. Since cMGT and human breast cancer share similar morphologies and pathogenesis, this study investigated the possibility of detecting surface KMO in human breast cancers and the role of surface KMO in tumorigenesis. Using immunohistochemistry (IHC), flow cytometry (FC), immunofluorescence assay (IFA), and transmission electron microscopy (TEM), we demonstrated that KMO can be aberrantly and highly expressed on the cell membranes of breast cancer tissues and in an array of cell lines. Masking surface KMO with anti-KMO antibody reduced the cell viability and inhibited the migration and invasion of the triple-negative breast cancer cell line, MDA-MB-231. These results indicated that aberrant surface expression of KMO may be a potential therapeutic target for human breast cancers.
Introduction
Breast cancer is the most common cancer in women worldwide. 1 The treatment approaches for breast cancer include surgery, radiation, chemotherapy, hormone therapy and targeted therapy. 2 Particularly, targeted therapy works by targeting the cancer’s specific genes, proteins, or the tissue environment that contributes to tumor progression and cancer survival. 3 Triple-negative breast cancer (TNBC), a subtype of breast cancer, does not respond to targeted therapy due to the absence of human epidermal growth factor receptor 2 (HER2), estrogen receptor (ER), and progesterone receptor (PR). 4 -7 With the identification of antigens specifically expressed in TNBC cells and the developing technology of monoclonal antibodies (mAbs), chimeric antigen receptors (CAR) and cancer vaccines, immunotherapy has become a novel and promising option for TNBC patients. 8
Kynurenine 3-monooxygenase (KMO) is the second enzyme in kynurenine pathway. It catalyzes the hydrolysis of kynurenine (KYN) to form 3-hydroxy kynurenine (3-HK), which further generates downstream metabolite quinolinic acid, which is an activator of N-Methyl-D-aspartate receptors (NMDA receptors). KMO was previously only investigated in neurodegenerative diseases because its downstream metabolites demonstrate excitotoxicity to the central nervous system (CNS) and act as factors in neurodegenerative diseases, such as Alzheimer’s disease and Huntington’s disease. 9 Recently, one study revealed the role of KMO in promoting cell migration and proliferation in human hepatocellular carcinoma (HCC). 10 In addition, our previous study demonstrated that both gene and protein expression of KMO were higher in stage III–V malignant canine mammary gland tumors (cMGT). This study also confirmed that the proliferation of cMGT cells was significantly inhibited when KMO activity was down-regulated. 11 Surprisingly, we also identified for the first time KMO not only on the outer membranes of mitochondria but also on the cell membranes of cMGT cells (Figure S1). Due to the high similarity in pathogenesis and morphology between cMGT and human breast cancer, 12 we further investigated if human breast cancer had the same phenomenon.
Numerous studies have previously demonstrated that aberrant subcellular localization of proteins may be a factor in tumorigenesis. The transformation from normal cells into cancer cells is mainly regulated by genome instability and results in tumorigenesis via complex mechanisms such as aberrant protein localization. 13 For example, BRCA1, normally localized to the nucleus, is detected in the cytosol of almost all breast and ovarian cancer tissues and cell lines. 14 Nuclear BRCA1 functions in DNA damage repair and cell cycle control, 15,16 whereas cytoplasmic BRCA1 regulates centrosome function 17 and p53-independent apoptosis. 18,19 Thus, understanding the mechanisms of aberrant localization in cancer is an important issue.
The aim of the present study was to identify the locations of KMO in breast cancers and the possible role of surface KMO in cancer development. In concordance with our finding in canine mammary tumors, we found that KMO was also expressed on plasma membranes both in clinical breast cancer tissues and in a variety of breast cancer cell lines. The expression of surface KMO was further demonstrated to promote the cell proliferation and migration of cancer cells. Taken together, the above findings highlight surface KMO as a novel target for cancer tissues and cell lines that might be important for the diagnosis and treatment of human breast cancers.
Materials and Methods
Cell Culture
Human breast immortal cell line MCF10A and triple-negative breast cancer cell lines MDA-MB-468, MDA-MB-453, BT549 and HCC1954 were obtained from Dr. Michael Hsiao at the Genomics Research Center, Academia Sinica. MDA-MB-231 was obtained from Dr. L-M. Tseng at the Taipei Veterans General Hospital. MCF7, HCC1937, MDA-MB-361 and Hs578 T were purchased from Building Construction Resource Center, Inc (Hsinchu, Taiwan). The culture conditions of the cell lines included different supplements, and the cell lines were cultured in suggested growth medium at 37°C with or without CO2 (Table S1).
Antibodies and Reagents
Several antibodies directed against the different parts of KMO were used, as follows: rabbit polyclonal antibodies from Proteintech (#10698-1-AP) (Roosemont, IL, USA), LSBio (#LS-C376014, #LS-C209356) (Seattle, WA, USA) and goat polyclonal antibody from NSJ Bioreagents (#R36453) (San Diego, CA, USA).
Immunofluorescence Assay (IFA)
4 x 105 MDA-MB-231 cells were cultured on slides overnight. The following steps were all performed at 4°C in darkness. The slides were washed with cold PBS 3 times, following by blocking with 1% BSA and 10% goat serum in PBS for 30 minutes. After being washed with cold PBS, the cells were incubated in KMO antibody (1:100, Proteintech) for 1 hour. Alexa Fluor 488 labeled goat anti-rabbit IgG (2 mg/ml, abcam, #ab150077), diluted 500 times in PBS buffer, was incubated for 1 hour with shaking. Plasma membrane was labeled with concanavalin A (Con A) Alexa Fluor™ 647 Conjugate (Thermo Fisher Scientific, #C21421) (Waltham, MA, USA) in the final concentration of 25 μg/ml for 5 minutes. After being washed with cold PBS, the slide was fixed with 2.5% paraformaldehyde at 4°C for 5 minutes. Then the slide was washed with cold PBS 3 times, following by labeling of the nucleus with 1 μg/ml DAPI (Sigma-Aldrich, #D9542) (St. Louis, MO, USA). Slides were imaged with a confocal microscope (Leica TCS SP8 Confocal) (Wetzlar, Germany). Results shown are representative of at least 3 independent experiments. The cell membrane colocalization of KMO was analyzed with MetaMorph Image Analysis Software (Molecular Devices, San Jose, CA, USA).
Cell Surface Staining for Flow Cytometry
Adherent cells were detached from the substrate by trypsinization and 1 × 105 cells were taken per tube. After the cells were washed with PBS once and with FACS buffer once, primary antibodies as indicated were added to the tubes and a pipette was used to gently mix the solutions. Then the tubes were incubated on ice for 1 hour before 1 mL FACS buffer was added to each tube for washing. Pellets were produced by centrifugation for 5 min at 300 xg. After 3 cycles of washing, goat anti-rabbit IgG-FITC (#STAR121F) (Bio-Rad, Hercules, CA, USA) was added and the mixture was incubated on ice for 30 min. Then 1 mL FACS buffer was added to each tube for washing and pellets were produced by centrifugation for 5 min at 300 xg. Results shown are representative of at least 3 independent experiments. All measurements were performed in an LSR Fortessa Cell Analyzer (BD Biosciences, Franklin Lakes, NJ, USA) and analyzed by FlowJo software (Tree Star Inc, Ashland, OR, USA).
Immunohistochemistry Analysis of Human Tissue Microarray (TMA)
The various breast cancer TMAs were purchased from SuperBioChips Laboratories (Seoul, South Korea) and US Biomax (#BC081120d & #BR1141) (Derwood, MD, USA). The TMAs contained 273 paraffin-embedded tumor specimens from 264 individual sarcoma patients. There were 9 normal breast tissues matched to 9 breast cancer tissues. The expression of KMO in the human tissue microarray was determined by immunohistochemistry assay (Paraffin). The three 5 µm paraffin-embedded TMA slides were baked for 15 min at 65 °C, deparaffinized in xylene 3 times for 10 min each, and then transferred through graded ethanol (100%, 95%, 80%, 70%) for rehydration for 3 min for each grade. PBST was used to wash them for 5 min after the endogenous peroxidase was quenched by 3% hydrogen peroxide for 20 min at room temperature. The slides were brought to a boil in citrate buffer, pH 6.0, and held at just below boiling temperature for 30 minutes. The slides were allowed to cool before blocking by goat serum was performed for 1 hour at room temperature. The 3 slides were incubated with rabbit polyclonal antibody to human KMO (#10698-1-AP, 1:200 dilution, Proteintech) at 4 °C overnight in a humidified chamber. Subsequently, bound antibody on the array was detected by UltraVision™ Quanto Detection System HRP DAB (Thermo Scientific). The nuclei of cells were counterstained with hematoxylin to improve the images. Finally, the sections were mounted for long-term preservation.
Immunostainings of the whole slide areas were viewed and scored separately by 3 independent pathologists, who were blinded to the tumor characteristics and core case details. The expression level of KMO was calculated as staining intensity × staining area with positive staining or positive membrane staining. The intensities of expression were divided into 4 levels (0, 1, 2 or 3 levels). The expression areas were defined as follows: 0, 0–5% of the total area in the field (400x magnification); 1, 5–20% of the total area in the field; 2, 20–40% of the total area in the section; 3, 40–60% of the total area in the section; 4, 60–80% of the total area in the section; and 5, 80–100% of the total area in the section. The quick score was the sum of 3 fields per core.
Immunogold-Labeling and Transmission Electron Microscopy (TEM)
1 × 107 MDA-MB-231 cells and MCF10A cells were pelleted by centrifugation for 10 min at 1800 xg. Cells were fixed with solution including 0.5% glutaraldehyde and 4% paraformaldehyde in 0.1M/pH7.3 PBS at room temperature overnight. After being washed with PBS 3 times for 10 min/time, the cells were then subjected to serial ethanol dehydration (25%, 37.5%, 49%, 60%, 65%, 67.5%, 70%, 85% and 92.5%), for 15 min each step, after which the cells were embedded in LR white resin and polymerized at 55°C for 24 h. The samples were then sectioned into ultrathin sections of 90 nm thickness and set for immunostaining. After 1.7% H2O2 was added for antigen retrieval, samples were blocked with 0.3% BSA in PBS for 1 h. The samples were incubated with rabbit polyclonal antibody to human KMO (1:200 dilution, Proteintech, #10698-1-AP) for 1 hour, followed by incubation with secondary antibodies conjugated with 5 nm gold particles (1:20 dilution, Sigma-Aldrich, #G7277) for 1 hour. The grids were dried after being washed with PBST 3 times. Then the grids were stained with uranyl acetate and lead citrate for double contrast before imaging by TEM (JOEL JEM-1400, Japan).
Generation of Anti-KMO Polyclonal Antibodies
Male BALB/c mice (7 weeks old) were injected subcutaneously with 100 µg of peptide emulsified in complete Freund’s adjuvant (Sigma-Aldrich, F5881-10X10ML). Two boosts of peptide emulsified in incomplete Freund’s adjuvant (Sigma-Aldrich, F5506) followed at bi-weekly intervals starting 3 months after the first immunization. One week after the second boost, the serum antibody titer was tested using enzyme-linked immunosorbent assay (ELISA). Two days after the last booster, polyclonal KMO antisera was obtained. Immunoglobulin was collected with affinity chromatography (protein A sephrose) and freeze dried. Poly Abs were kept frozen at -80°C until use. Control mouse IgG was similarly purified from whole normal mouse serum.
Cytotoxicity Assay
3 × 103 MDA-MB-231 and MDA-MB-468 cells were suspended in 100 µl serum-free DMEM with different amounts of polyclonal KMO antisera (25-200 ug/ml) and control mouse IgG for 48 hours. The WST-1 assay was then performed by adding 10 µl WST-1 (TaKaRa Bio, Mountain View, CA, USA), and absorbance at 450 and 690 nm was detected by a microplate reader 4 hours after the addition.
Migration and Invasion Assay
For migration assay, 2.5 x 104 MDA-MB-231 cells were suspended in 200 µl serum-free DMEM with the polyclonal KMO antisera (25–200 ug/ml) and placed into a permeable support insert with 8 mm pores (Corning, Glendale, AZ, USA). As for invasion analysis, 30 µg Matrigel® matrix (Corning) was coated within the support insert, followed by loading the same amounts of cells and antisera. To the lower chamber of the 24-well plate was added 900 µl of DMEM supplement with 10% FBS and 0.1% Antibiotic-Antimycotic, and the plate was incubated in 5% CO2 humidified atmosphere at 37°C. After 24 hours, the insert was washed twice with PBS and the cells were fixed in methanol for 10 minutes. After fixation, the cells were washed twice and stained with 0.05% crystal violet for 1 hour. After being washed twice, the non-migrated cells in the upper chamber were removed with a cotton swab immersed in PBS. The fixed cells in the lower chamber were photographed with an IX83 inverted microscope (Olympus, Japan) at a magnification of × 100, and the ratio was analyzed in ImageJ software.
Bioinformatics Analysis
All data of the cancer cohort was obtained from The Cancer Genome Atlas (TCGA)-PanCancer Atlas Projects. The breast cancer cohort was further retrieved from TCGA provisional data and the UCSC Xena public platform (Caldas 2007). 20 cBioPortal (http://cbioportal.org) was used to analyze the alterations of KMO. 21,22 To analyze the correlation between KMO expression and cancer prognosis, the information about patients with breast cancer were collected from UCSC Xena Hubs and analyzed with UCSC Xena Browser (http://xena.ucsc.edu/), 23 with the browser setting to split the entire group into the upper and lower quartiles for Kaplan–Meier survival analysis.
Statistical Analysis
Prism 8 (Graphpad Software, San Diego, CA, USA) was used to carry out the statistical analyses. Data are expressed as mean ± SEM. Student’s t test was used to analyze the difference between 2 groups in the migration and invasion assay, and 2-way ANOVA was used to analyze the difference between more than 2 groups in the proliferation assay. Statistical significance was achieved when the P-value was < 0.05.
Results
KMO Overexpression Is Related to Poor Overall Survival Rate in Breast Cancer Patients
Large-scale cancer genomics projects, such as the Cancer Genome Atlas (TCGA), generate large amounts of cancer genomics data on different technology platforms. Bioinformatics tools such as cBioPortal, 21,22 Xena browser 23 and MEXPRESS 24 can allow the integration of data for analysis, exploration, and analytics. We first used cBioPortal to analyze the relationships between KMO alterations, including amplification, mutation, fusion, deep deletion and multiple alterations, and cancer. KMO alterations, especially in amplification, are higher in invasive breast carcinoma than in 27 other cancers in the TCGA PanCancer Atlas (Figure 1A). In 185 breast cancer cases with genetic alterations of KMO, the overall survival rate (OSR) was lower than that of 773 cases without KMO genetic alterations (Figure 1B).

The association between KMO alteration/overexpression and the prognosis of breast cancer patients. (A) The frequency of KMO alteration was analyzed in various cancer types using the TCGA database. The arrow indicates that invasive breast carcinoma has the highest KMO alterations. (B) Kaplan–Meier survival curves for breast cancer patients in the TCGA database with or w/o KMO alteration. (C) Kaplan–Meier survival curves for breast cancer patients in the UCSC hub (Caldas, Naderi Gene Exp 2007). Patients’ survival was significantly associated with the decreased KMO mRNA transcripts (P = 3.886e-15).
In addition to validating whether KMO gene alterations are related to the survival outcomes of breast cancer patients, we also investigated the association of KMO expression with the prognosis of BC patients. Using a cohort database from the University of California, Santa Cruz (UCSC) (Caldas 2007) and analyzing the BC patients with different levels of KMO expressions by Xena browser, 20 we found that breast cancer patients with higher KMO mRNA expression had a much poorer overall survival time (Figure 1C). Interestingly, there was no significant difference in overall survival time between cases with and without KMO alteration; however, high KMO expression was significantly associated with poor overall survival time in breast cancer patients.
KMO Is Highly Expressed Both in Cytosol and on Cell Membranes in Clinical Breast Cancer Tissues
To further confirm the results obtained in the TCGA and UCSC databases, in which high KMO expression was correlated with poor prognosis in human breast cancer patients, we then analyzed KMO protein expression in breast cancer tissues. In concordance with the result in Figure 1C, significantly higher levels of KMO were detected in both non-TNBC and TNBC samples (Figure 2A and B) than in normal breast tissues. Moreover, though it has previously been identified as a cytosolic enzyme related to tryptophan metabolism, we revealed for the first time that KMO is also strongly expressed on the cell surfaces of breast cancer tissues (Figure 2A and C). These data suggest that KMO is not only overexpressed within cytosol but also that its expression is enriched on the cell membranes of breast cancer specimens. Moreover, the overexpression of cytosol and surface KMO was found in tissues from both non-TNBC and TNBC patients.

Up-regulation and aberrant localization of KMO in human breast cancer tissue. KMO expression was first analyzed in breast cancer tissue microarrays by immunohistochemistry. (A) Low expression of KMO in normal breast cells (n = 12); medium expression of KMO in non-TNBC tissues (n = 148); highest expression of KMO in TNBC tissue (n = 83). (B) Significantly higher expressions of KMO (***P < 0.001) were observed in non-TNBC tissues and TNBC tissues. (C) Higher expressions of surface KMO were observed in non-TNBC tissues (***P = 0.001) and TNBC tissues (**P = 0.0016).
Aberrant Surface Expression of KMO in Triple-Negative Breast Cancer Cells
We found surface KMO expression in clinical breast cancers, and the aberrant expression pattern was unlike that of normal tissues. Further investigations were then conducted by flow cytometry to detect the surface localization of KMO using a variety of human breast cancer cell lines. MCF10A is a normal breast cell line. MDA-MB-231, Hs578 T, HCC1937, BT549, MDA-MB-453, and MDA-MB-468 are triple-negative breast cancer cell lines, and MCF7, MDA-MB-361, and HCC1954 are non-triple-negative breast cancer cell lines. As shown in Figure 3A-C, surface KMO was not displayed on immortal mammary gland epithelial cells or non-TNBC cells; however, it was found that 7 to 86% of KMO was specifically detected on the cell surfaces in the BC cell lines (Figure 3D). We filtered cell lines with the average value of surface KMO greater than 5% for further analysis by mean fluorescence intensity. The MFI of FITC, which indicated the protein expression of surface KMO, indicated a range of 250 to 4020 (Figure 3E).

Detecting surface KMO of breast cancer cell lines by flow cytometry. (A) normal breast cell line, MCF10A; (B) non-TNBC cell lines, MCF7, MDA-MB-361, and HCC1954; (C) TNBC cell lines, MDA-MB-231, MDA-MB-453, MDA-MB-468, Hs578 T, HCC1937, and BT549. Y axis: cell count. X axis: intensity of fluorescence. (D) Bar chart for the ratio of surface KMO expression detected by flow cytometry. (E) Bar chart for mean fluorescence intensity (MFI) of FITC. At least 3 independent experiments were performed, and standard deviation (SD) is expressed.
We further verified the KMO locations in the TNBC cell line MDA-MB-231 and immortal mammary gland epithelial cell MCF10A by immunofluorescence assay (IFA). In cells without permeabilization, KMO was detected on cell membranes of MDA-MB-231, but it was not expressed in MCF10A (Figure 4A). The membrane colocalization of KMO within a cell was quantified as shown in Figure 4B and C, which display the significantly increased expression of surface KMO in MDA-MB-231.
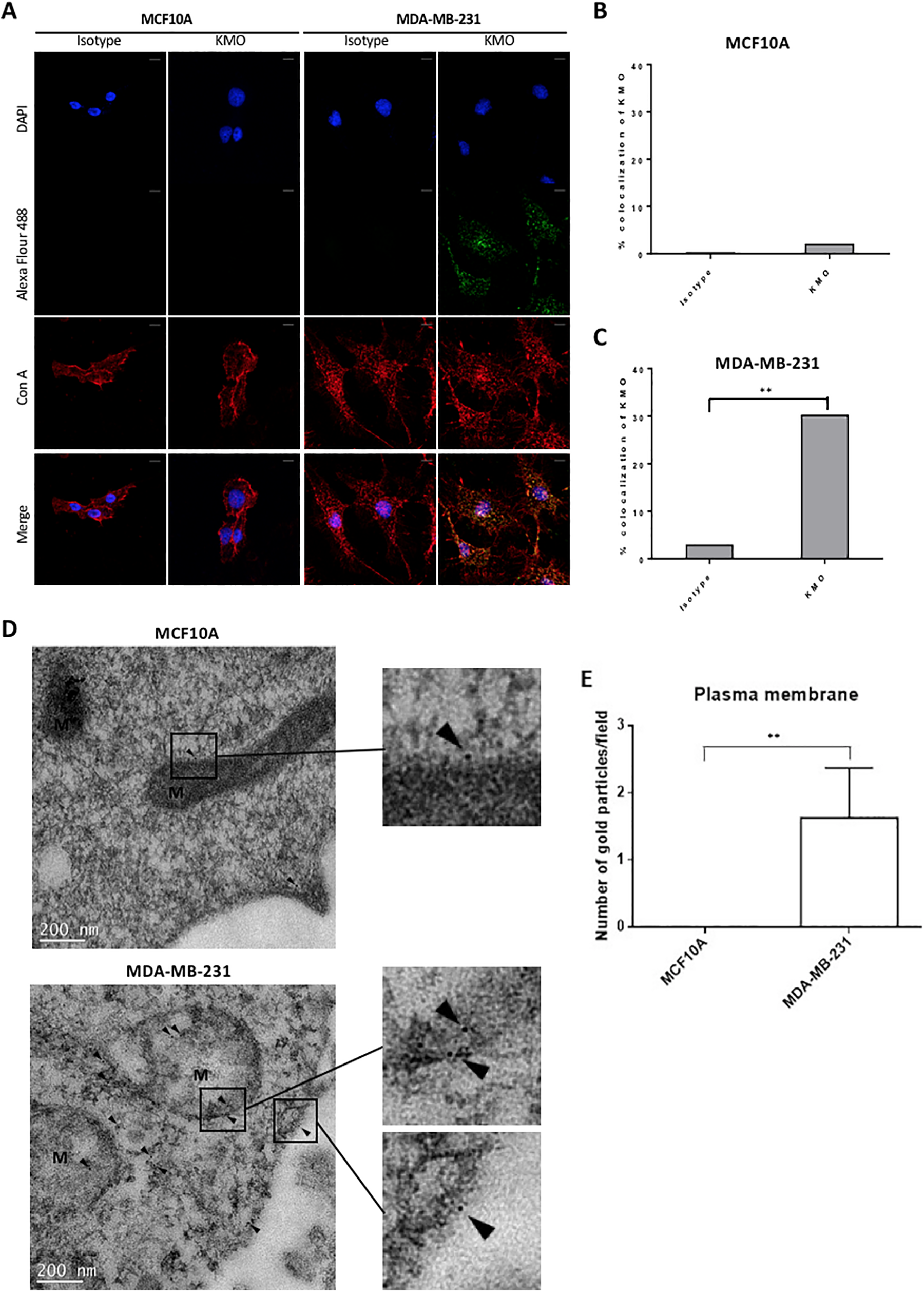
Cellular localization of KMO in breast cancer cell lines. (A) MCF10A and MDA-MB-231 were fixed and stained with rabbit IgG or KMO primary antibody. Con A was labeled with Alexa Flour 647 as the plasma membrane control. DAPI is used as a nuclear counterstain. Bars represent 20 μm. (B, C) Percentage colocalization of KMO in MCF10A and MDA-MB-231 (**P = 0.0091). (D) Top panel: cytosol KMO (arrow) was observed in MCF10A, the normal breast cell line; bottom panel: KMO could be easily found on the cell membrane of MDA-MB-231 (as detected by KMO-specific immunogold-labeling and TEM). M: mitochondria. Bars represent 200 nm. (E) The bar chart shows that more gold particles specifically reacting with KMO per field were found on the plasma membrane of MDA-MB-231 (**P < 0.01). N.D.: not detected. At least 5 randomly selected microscopic fields per section were quantified. The results shown are representative of at least 3 independent experiments.
Immunogold labeling in TEM is a very powerful technique for identifying the presence of specific antigens in cells. Therefore, we next verified the KMO locations in MCF10A and MDA-MB-231 using this protocol to specifically address which organelles contained KMO protein. The results showed that the gold particles reacted with the primary anti-KMO antibody, which was apparently localized in mitochondria, in cytosol, and on cell surface membranes in MDA-MB-231, whereas no such gold particles were found on the surface membranes of MCF10A (Figure 4D, 4E & Figure S2). These findings were consistent with the results obtained by flow cytometry and IFA and further confirmed the surface KMO expression in TNBC cells.
Identifying the Topology of Surface KMO on TNBC Cells
After KMO was demonstrated to be expressed also on the cell membranes of breast cancer cells, we further investigated the orientation of surface KMO by epitope typing. We used a panel of commercial antibodies that could recognize different regions of the KMO amino-acid sequence to probe the possible folded 3-dimensional structure of surface KMO. Human KMO is a protein that is 486 amino-acid residues in length. The results revealed that the antibodies against amino acids (aa) 1–486 and 1–260 could significantly bind to surface KMO proteins, while the antibody reacting to aa 300–400 failed to detect it (Figure 5A). Then we used Protter (http://wlab.ethz.ch/protter/start/) to further confirm and predict the topology of KMO. Protter revealed 2 transmembrane domains within the KMO structure (aa 385–406 & aa 426–445) (Figure 5B). Taken together, these findings suggested that aa 1–260 of KMO might possess an extracellular orientation which might contribute to the interaction with tumor microenvironments to promote cancer development.

Orientation determination of surface KMO in MDA-MB-231 cells. (A) Antibodies that specifically react with total and partial amino acid residues (aa 1–486, 300–400, and 1–260) of KMO were used to probe the possible topology of surface KMO by flow cytometry. Surface KMO could be detected only by the antibodies against aa 1–486 and 1–260. (B) Protter (ver 1.0) was used to predict the transmembrane domains of surface KMO. The results showed 2 transmembrane domains consisting of aa 385–406 and aa 426–445.
Blocking Surface KMO by Antisera Inhibited Cell Proliferation and Migration in TNBC Cells
KMO polyclonal antibody was used to investigate the impact of surface KMO on cancer progression in breast cancer. The WST-1 assay indicated that blocking surface KMO with KMO poly Ab decreased the cell proliferative ability of MDA-MB-231 in a dose-dependent manner (Figure 6A). In addition, transwell migration assay and Matrigel invasion assay were used to investigate the effects of blocking the migratory and invasive abilities of surface KMO on MDA-MB-231 cells. The results showed that blocking surface KMO significantly reduced migration (Figure 6B) and invasion (Figure 6C) in MDA-MB-231 cells. Collectively, the results suggest that surface KMO may promote cell growth and metastasis in human breast cancer cells.

Surface KMO is necessary for TNBC cell migration, invasion and proliferation in vitro. (A) Cell proliferation of MDA-MB-231 cells treated with KMO poly Ab was analyzed by WST-1 assay. (B) Cell migration of MDA-MB-231 cells treated with KMO poly Ab was analyzed by transwell migration assay. (C) Invasive activity of MDA-MB-231 cells treated with KMO poly Ab was analyzed by Matrigel invasion assay. Data are presented as mean ± SEM of 3 biological replicates. * P < 0.01, ** P < 0.01, *** P < 0.001.
Discussion
Breast cancer is the most common cancer in women worldwide. 1 TNBC, which accounts for 15–20% of all breast cancers, does not respond to targeted therapy due to the lack of HER2, ER and PR. 4- 7 , 25 Therefore, it is necessary to identify the biomarker of TNBC to facilitate the possibility of targeted therapies.
Herein, we identified surface KMO as a potential target in TNBC. KMO is involved in the metabolism of tryptophan and is located on the mitochondrial outer membrane. The correlation between the expression of KMO and the malignancy or outcome of clinical breast cancer cases was analyzed in TCGA/UCSC databases and tissue microarray. We first determined that KMO was highly expressed both in cytosol and on cell membranes in clinical breast cancer tissues, particular in TNBC tissues. We also provided novel insights on the surface KMO in breast cancer tissues and TNBC cell lines, BT549, HCC-1937, Hs578 T, MDA-MB-453, MDA-MB-468 and MDA-MB-231, by flow cytometry (Figure 3), IFA and immuno-TEM (Figure 4). The molecular mechanisms through which CAMs, which according to the structure can be categorized as integrins, selectin, immunoglobulin superfamily, cadherin and others, may be involved in the TNBC progression. 26,27 We ascertained its role in gene expression changes and the migratory activity of TNBC cells triggered by surface KMO. The protein level of the KMO in the tissue microarray displayed a strong association with the advanced stage, indicating that KP might also contribute to the tumor progression in TNBC patients. The clinical results were correlated with in vitro experimental results. Several studies have correlated KMO expression with different types of primary and metastatic cancers, including breast cancers. 10,28,29 In accordance with these studies, the expression of KMO was linked to a poor clinical outcome, further highlighting the contribution of KMO in the development of breast tumors.
In this study, we demonstrated for the first time the expression of surface KMO in breast cancer. The mechanism and role of surface KMO in tumor development remains unknown. Previous studies have demonstrated that aberrant subcellular localization of proteins may be a factor in tumorigenesis. The transformation from normal cells into cancer cells is mainly regulated by genome instability and results in tumorigenesis via complex mechanisms, such as aberrant protein localization. 13 In other words, when tumor suppressor proteins and oncoproteins have aberrant localizations, the cells may undergo tumorigenesis. Thus, protein localization is very important in mediating cell physiological function. In addition, membrane proteins have recently been mentioned in relation to cancer progression. 30,31 According to our data, we first proposed that KMO is not only expressed in the mitochondria but also on the cell surface in breast cancer, especially in TNBC. Membrane proteins mediate many biological processes, and in cancer cells, they are associated with multiple drug resistance, 32 adhesion and immune response. We revealed the presence, topology, and potentially tumorigenic functions of surface KMO in TNBC. The change in cell morphology is the initiation of migration, invasion and metastasis in aggressive cancer cells. β1 and β3 integrins are essential focal adhesion proteins in various cancer cell types, and they may affect the initiation, proliferation, survival and metastasis of tumors. Previous studies have demonstrated a ratio between these 2 integrins in cancer cells, which have contradictory functions. In breast cancer, the inactivation of β1 integrin causes the strong compensatory expression of β3 integrin. 27 We have identified the protein region exposed on the outer cell membrane of KMO. Further investigation could be conducted to dissect the interaction between surface KMO and extracellular matrix in cancer malignancy. Moreover, we also used KMO poly Ab to block surface KMO and found that the proliferation, migration, and invasion activities of MDA-MB-231 cells were significantly inhibited. The data indicated that the aberrant localization of KMO might play a key role in tumor development.
The KP including KMO usually refers to the involvement in neurodegenerative diseases and brain tumors, but so far, few studies have focused on cancer progression. Recently, more and more articles have shown that nerve cells can affect the progression of various cancers. 33 Based on the close relationship of KMO and neurons, we suppose surface KMO is not only expressed in TNBC cells but also is probably related to invaded nerve cells in the tumor microenvironment.
Surface KMO in breast cancer is a relevant discovery because many studies have noted the relationship between aberrant protein localization and tumorigenesis. 34 Aberrant protein localization, which is caused by mutation, altered expression of cargo proteins or transport receptors, or deregulation of components of the trafficking machinery, can lead to many human diseases such as cancers. Deregulation of protein trafficking results in mislocalization of proteins, and these proteins may function differently from their original roles if they are in the wrong place. 31 Some recent preclinical studies have focused on targeting protein translocalization for cancer treatment. For example, treating selective inhibitors of nuclear export (SINE) to inhibit nuclear exporter chromosome maintenance protein 1 (CRM1) causes nuclear retention of p53 and p21, increases ERK phosphorylation, and reduces retinoblastoma protein (pRb) phosphorylation in acute myeloid leukemia (AML). 35 Small-molecule SINE can also the block receptor activator of nuclear factor kappa-B ligand (RANKL)-induced nuclear factor-κB (NF-κB) and the nuclear factor of activated T-cells, cytoplasmic 1 (NFATc1). 34 Regardless of the unknowns, this study provides an observation of the atypical surface expression of KMO in breast cancer cells, and KMO should be viewed as a potential therapeutic target for TNBC.
Supplemental Material
Supplemental Material, sj-pdf-1-ccx-10.1177_10732748211009245 - Surface Expression of Kynurenine 3-Monooxygenase Promotes Proliferation and Metastasis in Triple-Negative Breast Cancers
Supplemental Material, sj-pdf-1-ccx-10.1177_10732748211009245 for Surface Expression of Kynurenine 3-Monooxygenase Promotes Proliferation and Metastasis in Triple-Negative Breast Cancers by Min-Hua Lai, Chi-Hsun Liao, Nu-Man Tsai, Kai-Fu Chang, Cheng-Chi Liu, Yi-Han Chiu, Kuo-Ching Huang and Chen-Si Lin in Cancer Control
Footnotes
Authors’ Note
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request. Our study did not require an ethical board approval because it did not contain human or animal trials.
Acknowledgments
We thank Dr. Chiung-Hsiang Cheng and the Joint Center for Instrument and Researches, College of Bioresources and Agriculture, National Taiwan University for the technical supports on TEM.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Ministry of Science and Technology of Taiwan (MOST 102-2313-B-002-031-MY3).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
