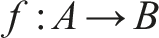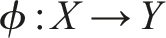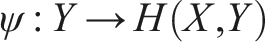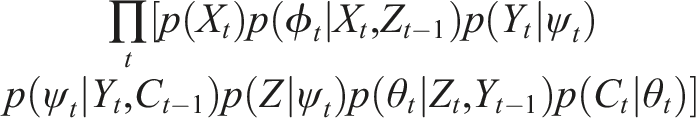Abstract
The notion of self-organisation plays a major role in enactive cognitive science. In this paper, I review several formal models of self-organisation that various approaches in modern cognitive science rely upon. I then focus on Rosen’s account of self-organisation as closure to efficient cause and his argument that models of systems closed to efficient cause – (M, R) systems – are uncomputable. Despite being sometimes relied on by enactivists this argument is problematic it rests on assumptions unacceptable for enactivists: that living systems can be modelled as time-invariant and material-independent. I then argue that there exists a simple and philosophically appealing reparametrisation of (M, R)–systems that accounts for the temporal dimensions of life but renders Rosen’s argument invalid.
The notion of self-organisation plays a major role in enactive cognitive science. In this paper, I review several formal models of self-organisation that various approaches in modern cognitive science rely upon. I then focus on Rosen’s account of self-organization as closure to efficient cause and his argument that models of systems closed to efficient cause – (M, R)–systems – are uncomputable. Despite being sometimes relied on by enactivists (e.g. (Thompson 2010)), this argument is problematic as it rests on assumptions unacceptable for enactivists: that living systems can be modelled as time-invariant and material-independent. I then argue that there exists a simple and philosophically appealing reparametrisation of (M, R)–systems that accounts for the temporal dimensions of life but renders Rosen’s argument invalid. While Rosen’s argument has received valid criticism before (e.g. (Gatherer and Galpin 2013; Palmer et al. 2016)), I present an attack motivated by the philosophical principles of enactive and dynamical systems cognitive science itself: relation biology, Rosen’s modelling approach, in incongruent with dynamical and enactivist concern about temporal, path-dependent unfolding and embodiment. My attack exposes and obvious but surprisingly overlooked disregard for those dimensions in relational biology. Furthermore, my discussion hopefully provides additional insights about the relation between (M, R)–systems and other accounts in theoretical biology and cognitive science such as the dynamical systems approach (Kelso 1995) and the free energy principle (Friston 2013). The paper is structured as follows. In the first section, I review several conceptual models of self-organisation and argue that an account of self-organisation as second-order feedback unifies them in a relatively non-controversial manner. In the second section, I review the role played by these conceptual models of self-organization in various approaches in cognitive science. I argue that enactive cognitive scientists frequently require self-organisation to play an explanatory role similar to the roles computationalists philosophers of cognitive science ascribe to representation. In the third section, I present in more detail Rosen’s relational biology and his account of self-organisation as closure to efficient cause. I then formulate his argument that models of systems closed to efficient cause – (M, R)–systems – are uncomputable. In the fourth section, I provide more historical context to Rosen’s argument and the influence his argument and his way of thinking about living systems had on modern enactivism. Finally, in the fifth section I point out a deep, irreconcilable tension in Rosen’s argument and his relational biology in general: is fails to account for the temporal nature of life. If one assumes living systems to be dynamical systems (as modern enactivists do), the argument turns out to be invalid in a trivial way. Therefore, it should be rejected. I end the paper with some general remarks about the uses and misuses of formal limitative results in philosophy.
Formal models of self-organisation
Self-organisation is among the basic notions of modern systems biology (Bechtel 2007; Hofmeyr 2007). Despite being invoked extensively by the cybernetic grandfathers of cognitive science (McCulloch and Pitts 1943; Ashby 1947), serious philosophical arguments casting self-organisation as an explanans for cognitive science started only with the advent of dynamicism (Port and Van Gelder 1995) and enactivism (Varela et al. 1974; Thompson 2010). Intuitively, self-organisation is usually understood as spontaneous change in organisation of a system or the emergence of novel constraints on the dynamics of a system. There is, however, no single agreed upon definition of self-organisation, which prompts some philosophers to treating it not as a natural kind but acknowledging the existence of multiple concepts of self-organisation linked by family resemblance (Collier 2004).
I will start with seven different conceptual models of self-organisation. The list below is a list of conceptual models (Barandiaran and Chemero 2009) or holistic models (Downey 2018), that is: formalising an idea without a pretence of empirical validity with respect to any class of empirical phenomena. Conceptual models should not be confused with generic models (describing the rules governing certain class of phenomena), functional models (maintaining functional validity) or mechanistic models (maintaining a mapping between observables and variables in the model). However, the usefulness of such a model guides the evaluation of empirical and heuristic validity of conceptual models. Moreover, here I focus on influential historical models, delaying the discussion of their modern incarnations to the next section. (A1) Ashby’s homeostat, a mechanical model of the brain as a self-organising dynamical system, was an aftermath of the first modern and formal notion of a self-organising dynamical system endowed with an attractor in its phase space (Ashby 1947). This model was one of the cornerstones of cybernetics (Wiener 2019) and started thinking about self-organisation as an advanced form of regulation. (A2) Dissipative structures in non-equilibrium thermodynamics, which maintain low entropy and stable steady state due to exchanging energy with the environment. The thermodynamic model of dissipative structure was proposed by Nicolis and Prigogine (1977). It is a model of simple chemical oscillation (such as Belousov–Zhabotinsky reaction or Rayleigh–Bénard convection) but can also be treated as a minimal thermodynamic model of a living system. (A3) McCulloch and Pitts’s (1943) model of recurrent neural networks based on Boolean gates. It was the first attempt of a computational explanation of cognitive processes (Piccinini 2004) by designing a propositional calculus in which a neural event can be represented by a proposition about temporal antecedents of the neural event. Kleene (1956) has later shown that the formal model presented by McCulloch and Pitts is equivalent to finite state automata. A crucial property of this model is the ability to represent network with closed loops, which generate self-sustaining neural activity. As such, it was a predecessor of Hebbian learning. (A4) Varela and Maturana’s model of an autopoietic system (Varela et al. 1974; Maturana and Varela 1980). As a minimal model of a living system, an autopoietic system is characterised by two conditions (i) possessing a spatial boundary and (ii) possessing a network of mutually sustaining processes. This model was a cornerstone of research in artificial life (Froese and Ziemke 2009) and was widely discussed in cognitive science as determining the necessary (and, according to some, sufficient) conditions of being a cognitive system (Thompson 2010). (A5) Rosen’s (1991) model of a system closed to efficient cause. Closure to efficient cause is feature of relational models, that is, models composed of morphisms in the category theoretic sense. A relational model is closed to efficient cause if every morphism included in the model is an endpoint of another morphism in the model. An obvious biological interpretation of this formal model is metabolic activity of a living system, autonomously sustaining crucial processes. The relational way of thinking about life and understanding of self-organisation in terms of closure to efficient cause turned out to be very influential in modern systems biology (Bechtel 2007; Hofmeyr 2007). (A6) Kauffman’s (1969; 1993) model of autocatalytic networks. Using random Boolean networks, Kauffman has shown that for a sufficiently large number of catalytic dependencies, stable and self-sustaining cycles will arise. This allows for formal modelling of autocatalysis, a primary mechanism for metabolism and gene expression regulation. (A7) Dynamical systems theory is an umbrella term for a family of related approaches, including bifurcation theory, fractal geometry and ergodic theory. One philosophically influential model was Bak et al.’s (1987) concept of self-organising criticality: a phenomenon arising in dynamical systems with attractors that are a critical state (a state such that an infinitesimal change in some parameter causes a large change in the dynamics). Self-organising criticality can be applied to model multiple physical and biological systems. Other curious phenomena studied through the lens of dynamical systems include chaos, invariance and metastability. (A8) Gánti’s (2003) chemoton is a minimal model of a protocell that continuously sustains three processes: metabolism, self-replication and a boundary (a bilipid cell membrane). Originally formulated in 1971, it shares many principles with (A2) and (A6): the focus on thermodynamic openness (with (A2)) and on the role of autocatalytic networks of chemical reactions (with (A6)). In contrast with these models and similarly to (A5), Gánti’s account also focused on modelling self-replication (fission of protocells).
What are the common traits of models (A1)–(A8)? Do they all denote the same class of systems? There seems to be one fundamental distinction between different kinds of self-organisation, sometimes cast as a difference between self-organisation and self-assembly. It seems that there are some phenomena that can maintain a steady state through feedback, but lack resilience, that is, ‘the ability to recover from perturbation; the ability to restore or repair or bounce back after a change due to an outside force’ (Meadows 2008). Such phenomena include crystals, tornados, Bénard cells and candle flames. They would instantiate model (A2) and probably also models (A3) and (A7). Other accounts of self-organisation – those requiring resilience, such as (A1) and (A4)–(A6) – would rule them out. This distinction reappears multiple times in the literature with little consistency in terminology. For instance, Friston and Stephan (2007) distinguish self-organising systems (like normal snowflakes) and adaptive systems (like imaginary snowflakes endowed with wings, able to avoid melting – a phase transition). In a similar vein, Bickhard (2009) distinguishes self-maintenant systems (like a candle flame maintaining combustion threshold temperature and inducing convection, which brings fresh oxygen) and recursively self-maintenant systems (that maintain self-maintenance). Furthermore, enactivists distinguish autonomous systems as those ‘composed of processes that generate and sustain that system as a unit’ Thompson (2010). It is not a coincidence that the distinction between self-assembly (self-organisation in the weak sense) and self-organisation in the stronger sense mirrors the distinction between self-organisation in non-living systems and biological self-organisation. In the rest of the paper, by ‘self-organisation’ I will mean the stronger sense of the notion, reserving ‘self-assembly’ for self-organisation in non-living systems.
I am not going to assume a specific formal model of self-organisation. For the purpose of this paper, I will assume a broad, substrate-independent account of self-organisation as second-order feedback or multi-level constraining. These are two complementary views of on what distinguishes self-organisation from self-assembly. According to the first view, there must be at least two nested feedback loops in the system operating on distinct time-scales: one accounting for adaptability to changing conditions and the second accounting for resilience. As an example, consider a sensorimotor loop of a bacterium embedded in a loop of its metabolism. An alternative way of capturing this notion is to say that self-organisation happens when entities from microscopic level of description of a system impose constraints on the dynamics of macroscopic level of description (Pattee 2012; Rączaszek-Leonardi 2012).
Self-organisation in cognitive science
Models (A1)–(A8) are formal models of biological and physical phenomena and lack immediate connections to cognitive models. However, if they are adequate models of biological phenomena, then – assuming that cognition is embodied (Bechtel and Mundale 1999) or that there is a deep continuity between life and mind (Thompson 2010) – they should be adequate to describe at least some aspects of cognition. (B1) Dynamicism. This paradigm was shaped in opposition to the symbolic approach in cognitive science and attempts at building explanations of cognition in terms of dynamical systems theory, focusing on the temporal and situated character of cognition. The cornerstones of dynamicism are Smith and Thelen (2003), Kelso (1995), Buhrmann et al. (2013) and Chemero (2009). Dynamicism draws mainly from (A7), although Chemero and Turvey (2008) explicate the notion of sensorimotor loop and affordances in terms of Rosen’s (1991) model (A5), and cybernetics (A1) is an important frame of reference for dynamically oriented behavioural roboticists (Beer 1990). (B2) Neurodynamics. This approach arose from research on electrophysiology of the brain and explores the notions of synchrony and desynchronisations as its main explanantia (Skarda and Freeman 1987; Kelso 1995; Freeman 2000). It employs mainly (A7) but also (A1) and (A3). (B3) Autopoietic enactivism started out from the biological model of autopoiesis (A4), trying to account for cognition in terms of metabolism (Varela et al. 1974). Later sense-making has become the central notion, which – in addition to autopoiesis – requires adaptivity (Thompson and Stapleton 2009; Thompson 2010). Finally, recently, autopoietic enactivists stress that cognitive systems are essentially precarious and interactive, which entails material and energetic openness (Ruiz-Mirazo and Moreno 2004; Barandiaran and Moreno 2008). (B4) Active inference is an approach in computational neuroscience and artificial intelligence. It assumes cognitive systems are prediction machines employing probabilistic models of them and their environments and trying to minimise prediction errors of these models via perceptual inference, action or learning. The mathematical structure of this model is inspired by (A1), (A2), (A4) and (A7) (Friston 2013; Clark 2016; Buckley et al. 2017).
Most approaches in cognitive science that build on the concept of self-organisation, such as sensorimotor enactivism (O’Regan and Noë 2001), autopoietic enactivism (Thompson 2010), neurodynamics (Freeman 2000) and radical embodied cognitive science (Chemero 2009), distance themselves from computationalism and representationalism. Chemero (2009) claims that self-organisation is a notion able to replace the computationalist notion of planning and model a cognitive system as ‘regular but not regulated’. This regularity would be due to self-organisation (here in the sense of (A2) (Chemero 2008, pp. 274–275). Indeed, in many biological contexts, self-organisation does a good job replacing global control with full information about the system (a classical example may be ants following a pheromone trail (Mitchell 2011) instead of computational mapping, planning and localisation).
One could ask whether there could be a computationalist or mechanistic account of self-organisation. The earliest cybernetics models (A1) and (A3) seem to be exactly that. Historically, however, arguments against mechanism in philosophy of biology were based on notions similar to self-organisation (such as epigenesis, homeostasis, feedback, closure to efficient cause or endogenous activity, cf. (Huneman 2007)). To analyse this apparent disconnect between mechanistic and non-mechanistic accounts of self-organisation in more detail, in the rest of the paper, I will focus on Rosen’s model (A5) and his argument against mechanism based upon this model. This argument received some interest in cognitive science in dynamicist (B1) and enactivit (B3) circles (Thompson 2010; Nasuto and Hayashi 2016) but on a closer look reveals problematic assumptions that are in strike contrast with (B1) and (B3), namely: substrate-independence and ahistoricity of living systems. This exemplifies the more general problem of how to reconcile the teleological (usually rendered as atemporal) and the historical (inherently time-ordered) aspects of self-organisation in a single model. Rosen’s concept of efficient causation (to be introduced in the following section) is designed to capture the teleology but fails to capture the temporal character of living systems, fundamental to (A1). I will argue, however, that it is not a general fact about self-organisation that it implies ahistoricity but a property of a particular modelling approach (A5). This case study should reveal a surprisingly intricate relationship between formal models of self-organisation in theoretical biology (A1)–(A8) and recent approaches in cognitive science (A1)–(B4). In particular, (A5) violates the premises of (B1) and (B3) that frequently build on (A5). The implication of my criticism is that (A5) as a formal model of self-organisation does not live up to the requirements posed by modern enactive and dynamical systems cognitive science while other models, such as (A1) or (A7), do.
Rosen’s argument
Overview
Robert Rosen famously claimed that living systems are not Turing-computable (e.g. (Rosen 2000, p. 325)). Based on category theory, he developed relational biology, an original formal framework for modelling the organisational properties of life and provided a formal argument that a peculiar property manifested by living systems – closure to efficient cause – prevents them from being simulable by a Turing machine. Thus, life cannot be accounted for mechanistically, but only within some theory richer than contemporary physics.
The structure of Rosen’s argument is as follows: • (T1) All living systems manifest the property of closure to efficient cause. • (T2) If a system is closed to efficient cause, at least some of its models are not Turing-computable. • (T3) If mechanism concerning biology is true, every model of a living system can be simulated on a Turing machine. • (T4) Therefore, mechanism is false.
Let us now discuss each premise (T1)–(T3) separately. (T1), though very non-trivial, is now considered Rosen’s crucial contribution to mainstream biology and was recently found valuable in systems biology (Hofmeyr 2007). The concept of closure to efficient cause is to be understood in terms of Rosen’s relational models which is discussed in the next section.
The premise (T2) is the heart of Rosen’s argument. Rosen claimed that ‘organisms possess noncomputable, unformalizable models’ (Rosen 2000, p. 4) and that ‘any material realization of the (M, R)–systems must have noncomputable models’ (Rosen 2000, p. 269). A crucial distinction here is the one between simulation and modelling. For Rosen, ‘simulation’ denotes a formalization of a mathematical system of interest allowing determining the truth or falsity of certain facts about the system in a finite number of steps. More formally, ‘Simulability, is property of a mathematical system, pertaining to the replacement of semantics (external referents) by syntax’ (Rosen 1993, p. 370). Turing machines – the dominant model of computation in theoretical computer science (Turing 1937) – are one of the systems which allow for formalizing the notion of simulability. One can simulate a system if there is a Turing machine that can decide the truth or falsity of some claims about the systems. On the other hand, ‘modelling’ is a broader term denoting a relation of correspondence or homomorphism between an external system and a representation thereof (Rosen 2000, p. 49). Therefore, only some models are simulable. The important claim that Rosen puts forward is that there will be facts about living systems that cannot be simulated (because of some of their properties – closure to efficient cause – cannot be simulated) but can be modelled. Therefore, some models of living systems (e.g. (M, R)–systems) will not be simulable and, in consequence, Turing-computable.
(T3) is Rosen’s definition of a mechanism (a class of phenomena) and of mechanism (a philosophical claim). Rosen understands mechanism (the claim) by analogy to the Church-Turing thesis (Copeland 2020), a hypothesis about the nature of computable functions stating that a function can be effectively calculated by a method if and only if it is computable by a Turing machine. This leads to his definition of a mechanism (a class of phenomena): ‘Let us call a system satisfying Church’s Thesis (i.e., such that every model is simulable) a simple system or mechanism’ (Rosen 1993, p. 370). Elsewhere, Rosen states that: ‘[t]he idea that every model of a material system must be simulable or computable is at least tacitly regarded in most quarters as synonymous with science itself; it is a material version of Church’s Thesis (i.e., effective means computable). […] I call a material system with only computable models a simple system, or mechanism. A system that is not simple in this sense I call complex. A complex system must thus have noncomputable models’ (Rosen 1993, p. 325; emphasis original).
Rosen account of mechanism (the claim) seems to be roughly compatible with modern concepts of mechanism. For instance, according to Bechtel and Abrahamsen (2005). a mechanism is ‘a structure performing a function in virtue of its components parts, component operations, and their organization. The orchestrated functioning of the mechanism is responsible for one or more phenomena’. Though Rosen’s notion of simulability is conceptually different from the notion of mechanism, it seems to be at least a fair operationalisation of the latter. Krajewski (2020) provides a brief discussion of the interrelations between these concepts.
Let us now sup up the argument. (T1) and (T2) jointly imply that at least some models of living systems are not Turing computable. This, in conjunction with (T3), implies that (T4) mechanism is false. In the rest of the paper, I will focus on the most controversial premise, namely, (T2). Following others (Gatherer and Galpin 2013; Palmer et al. 2016), I will show that closure to efficient cause does not imply existence of non-simulable models. Before that, however, I will spend more time explicating (T1) and presenting in more detail the concept of closure to efficient cause and the broader framework in which it is defined, relation biology.
Category theory
Relational modelling is introduced as an alternative to the ‘reductionist account inherited from Newton’ (Rosen 1991, p. 117), which generally means all state-space-based models with some recursively defined transition function. A mechanistic (reductionist) model aggregates constituent particle and provides empirically guided description of law governing the interactions of these. Or, to put in bluntly: ‘In empirical terms, then, the very first step in the analysis of an organized system (e.g. our cell) is to destroy that organization. That is, we kill the cell, sonicate it, osmotically rupture it, or do some other drastic thing to it’. (Rosen 1991, p. 118)
Importantly, Rosen rejects a particular mode of scientific explanation rather than scientific knowledge altogether. It is the relational framework that can be seen as a blueprint for some ideal, richer physics to be developed in the future – a theory that could account for self-organising living systems. While a mechanistic (reductionist) model aggregates constituent particles and provides an empirically guided but organization-insensitive description of laws governing the interactions of these, a relational model focuses instead on the organisational level of a system in question.
According to Rosen, relational models are built of components, instead than particles, where component is to be singled out functionally, in terms of its role in the model. What is more, they can acquire novel (emergent) properties in the context of a system, unlike particles in reductionist models. Formally, a component is to be understood as a mapping
While a mapping is often thought as a binary relation, Rosen notes (1991, p. 221) that this formula actually consists of four ingredients: • A is the domain of the mapping; • B is the range of the mapping; • f is the name of the mapping; • → is the name of a ternary relation between f, A and B.
The influence on the system in which component f is embedded is here expressed in specific arguments in the domain A. Larger relational models can be built through mapping composition, for example, f◦g. Moreover, since a mapping can be easily interpreted extensionally as a set of ordered pairs in the form (argument and value), one can define a mapping whose output is another mapping. Let denote the set of mappings from A to B. Thus, we can define a mapping ϕ◦ ψ composed of
and
Note, however, that this time While solid arrow as previously indicates a mapping, dotted arrow represents that 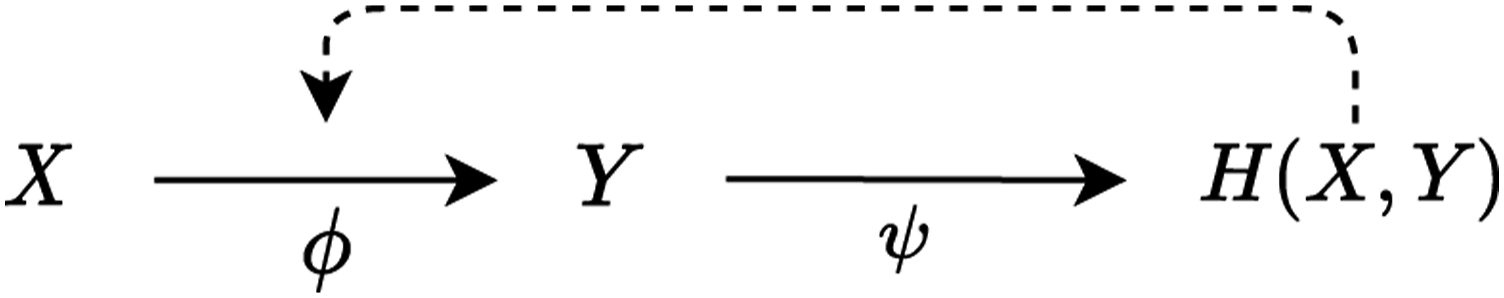
We now see that a graph representation of a relational model can contain cycles.
Rosen grounds his relational biology in category theory – which can be seen as a general mathematical theory of structures and systems of structures, and – possibly – providing an alternate (to set theory) way of formalising the foundations of mathematics (Marquis 2008). 1 A category is traditionally defined as consisting of a class of objects O, and a class of morphisms (mappings and arrows) M between objects of O, such that for every object there exists an identity morphism, and for every two morphisms there exists a composition of these (i.e. morphisms are associative). Sets and mappings between sets trivially meet these requirements, but it should be noted that they are also met by ordered sets and monotonous mappings, groups and group homomorphisms, vector spaces and linear transformations, as well as topological spaces and continuous functions (and so on).
The concept of a mapping on which relational biology is grounded is precisely the category-theoretic concept of morphism. Therefore, Rosen’s argument, if it is sound, is general enough to encompass all kinds of mathematical structures (and formal or computational models). On the other hand, the reader may find Rosen’s argument more comprehensible when understanding it in terms of ZFC set theory; no significant loose in generality will take place. For instance, Chemero and Turvey (2008) use hypersets and mappings between them to explicate the concept of closure to efficient cause. Hyperset theory is a variant of set theory where the axiom of regularity is substituted with its negation; thus, there exists a set Ω such it is itself its only element, that is, Ω = {Ω}. However, the work of Chemero and Turvey has received valid criticism from other authors, for example (Cárdenas et al. 2010). In particular, Chemero and Turvey’s account of closure does not include an output of a self-organising system: it only receives environmental input, but nothing comes out of it. According to Cárdenas et al. (2010), this clearly ‘violates the law of conservation of matter, because the […] cycle is a bottomless pit into which all the reactants disappear, with nothing coming out’. This failure exemplifies a fundamental property of living systems – thermodynamic and material openness – which is indeed a major shortcomings of many purely physical models of living systems. As I will explain later, Rosen’s relational biology sidesteps that problem.
Causation in relational models
Before elucidating what it takes for a relational model to be closed to efficient cause, it is reasonable to ponder how Rosen’s formal models are to be interpreted in the biological domain. Rosen wrote surprisingly little on what biological entities mappings correspond to. Hofmeyr (2007), though, provides an elegant biochemical interpretation in terms of metabolic pathways. On this account, a mapping f: A → B corresponds to a chemical reaction, with A being a reagent, B being a product and f itself being an enzyme catalysing the reaction. Since enzyme can itself be a product, the system depicted in Figure 1 can easily be interpreted as a simple catalytic cycle, an instance of a paradigmatic class of mechanisms in biochemistry. Kauffman’s (1969) model (A3) presents an account or the origins of life in terms of autocatalytic networks very much in the vein of this interpretation.)
An important feature of relational models is that the notion of final cause can be expressed relationally. This is supposed to be unlike state-space-based models, with time as their independent variable; since these are time-asymmetric, the effect must always precede the cause. This constraint is not present in time-invariant relational models, which therefore support multiple modes of causation very much in the vein of Aristotle’s doctrine of four causes. Let us get back to the Figure 1 and ponder the question ‘why Y?’. According to Rosen, there are three answers: Y’s material cause is X, Y’s efficient cause is ϕ and Y’s final cause is H (X, Y). (In the current setting, Aristotle’s formal cause is identical with efficient cause, although Hofmeyr (2007) proposes the information encoded in mRNA to be interpreted as a formal cause for given a biological structure and presents a formalisation of this account.)
While relational models indeed provide an interesting approach to biological functions that recently gains attention in systems biology, it is highly disputable that teleonomy cannot be ascribed in dynamical models unfolding in real time. All it takes to avoid the problem of temporal asymmetry of final causation is to take long-scale system’s dynamics under consideration. This approach was probably pioneered by Rosenblueth et al. (1943), who explicate the concept of purposeful behaviour in terms of negative feedback. Another approach to determine biological function is to consider what given component was selected for in its evolutionary history (Millikan 2002).
Closure to efficient cause
A crucial feature of ϕ is that its efficient cause lies in the system itself, or to use Rosen’s phrase, it is entailed from within the system. ψ on the other hand is only materially and finally caused from within the system, but still requires some external component as its efficient cause; it is not entailed. (This may mean some enzyme produced by some other system that enables reaction ψ to take place). Generally, the more components are entailed by the system itself, the more is the system autonomous. The corner case where all system’s components are entailed from within the system is what Rosen calls the property of closure to efficient cause.
The system depicted on Figure 1 could be extended by defining a third component θ efficiently causing ψ and making sure that this component is efficiently caused by ψ. Such a model is displayed on Figure 2. A relational model made of three mappings, each of which is entailed by a product of one of the others.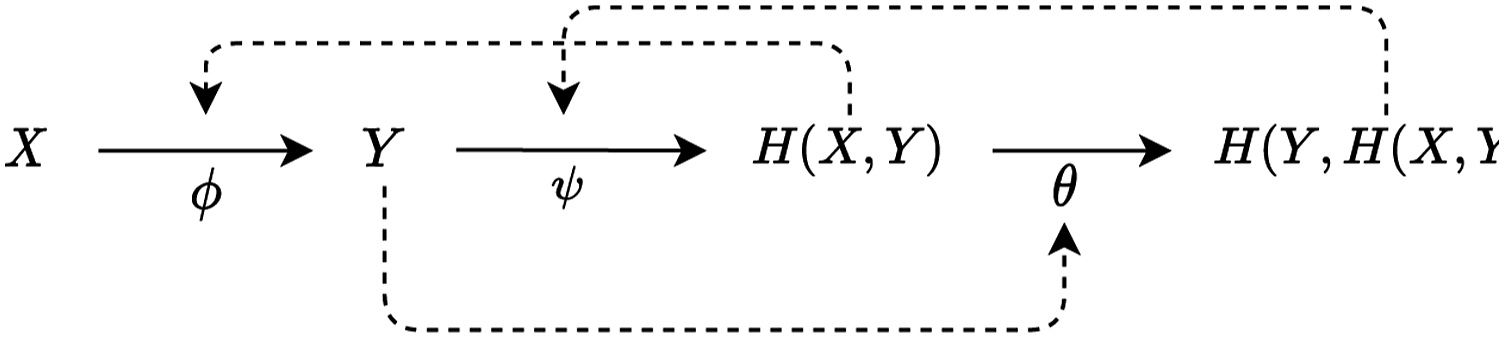
In this system, Y is the material cause of H (X, Y), the final cause of X and the efficient cause of θ. The system consists of three mappings (ϕ, ψ, θ), and each of these is entailed by a product of one of the others. This means that ϕ, ψ and θ, as we selected them, are closed to efficient cause.
Biologically, mapping ϕ can be interpreted as metabolic activity and mapping ψ as activity maintaining ϕ, termed ‘repair’ by Rosen. θ enables the self-replication of the entire system. Thus, systems sharing the relation structure depicted on Figure 2 are called metabolism-repair systems, or (M, R)–systems.
Note also that while the mappings on the Diagram 2 are entailed from within the system, there is still one unentailed object, that is, X. Although Rosen does not emphasise this point, it is quite deliberate that X has to be provided from outside of the system. This corresponds to the fact that living systems are thermodynamically and materially open, that is, they must consume energy to maintain their existence. Indeed, thermodynamic openness is a fundamental property of several models of self-organisation mentioned earlier, for example, (A2), (A4) and (A8).
According to Rosen, systems closed to efficient cause cannot be described in terms of states and transitions between them. Therefore, some of their models are not recursively enumerable, which also means they are not output of any computable function. Those models are not simulable in the sense described earlier that their activity cannot be simulated by a Turing machine.
Instead of narrating Rosen’s circuitous argument in full detail, I will follow Chu and Ho’s 2006 summary. Let us assume that the system depicted on Figure 2 is indeed a mechanism, that is, it can be completely specified in terms of states and a recursively defined transition function between them. Therefore, all of its components (ϕ, ψ, θ) can be specified in three corresponding sets of states J, K, L. Now take ψ and the corresponding set of states L. ψ has at least two functional roles, namely, its output H (X, Y) materially causes
This argument has received serious criticism from numerous authors. For instance, Chu and Ho (2006) accuse Rosen of using an unrealistic assumption that functions are always localised into corresponding organs (i.e. one indivisible component cannot have several functional roles). Consequently, Rosen at least shows that this assumption does not hold for neither mechanism nor living systems, which is quite trivial. Louie (2007) defended Rosen from Chu and Ho’s criticism by accusing them of misinterpreting Rosen by using incorrect definitions (for instance, for the term ‘mechanism’). I do not find Louie’s defense compelling for two reasons. First, a large portion of Chu and Ho’s criticism concerned the biological plausibility of Rosen’s modelling assumptions (e.g. about how to map morphisms to biological activities) not only the formalism itself. Louie claims decidedly that ‘[a]ll Rosen’s theorems have been mathematically proven (although Rosen’s presentations are not in the ordinary form of definition-lemma-theorem-proof-corollary that one finds in conventional mathematics journals). Indeed, no logical fallacy in Rosen’s arguments has ever been demonstrated. Counterexamples cannot exist for proven theorems’. (Louie 2007, p. 294, p. 294)
While it is true that proven theorems have no counterexamples, this only pertains actual proofs, not arguments expressed in a mathematical language. An argument ceases to be a formal proof as soon as it leaves the realms of pure mathematics: it has to be backed up with some mapping from mathematical objects onto empirical ones and the validity of such a mapping is, at least partly, an empirical question. As far as Rosen wants (M, R)–systems to pertain to biological reality rather than just other mathematical objects, he must do the hermeneutic work of translating between mathematical objects and biological entities. This work is not purely formal in nature and can be subject to criticism.
A second problem with Louie’s defense is that counterexamples have been found. There are several demonstrations how (M, R)–systems can be expressed as computable in several models of computation. This includes Gatherer and Galpin (2013) demonstrating the computability of (M, R)–systems in process algebra, Mossio et al. (2009) in lambda calculus and Palmer et al. (2016) using stream X-machines. In this paper, I will take another route in criticising this argument: instead of mounting formal an argument in a particular model of computation, I will attack Rosen’s model on philosophical grounds. Concretely, I will show that abstracting away from time is the crucial and untenable assumption. Before that, however, I will take a broader, philosophical look at relational biology from a historical perspective to trace back what went wrong.
Philosophical origins of relational biology
The concept of living organisms as having peculiar, circular causal structure dates back to Immanuel Kant. Actually, Kant himself took an epistemological stance on the teleonomic character and mechanistic intractability of biological complexity, that is, he considered them as regulative principles of scientific explanation rather than constitutive of some biological reality. The aim of his philosophy of biology outlined in the Critique of the Power of Judgment (Kant 1790) was to defend the notion of natural purpose against critiques of both empiricists (Hume) and rationalists (Spinoza) on the one hand and liberate it out of theological commitments on the other. According to Kant, organism must be conceived as organised and purposive beings; the natural purposes have to be presupposed rather than instantiated in reality. His views, however, inspired an entire research program in 19th century biology and historically, what Kant took as regulative, biologist saw more as constitutive (Huneman 2007). He also emphasised the self-organising character of living organisms: ‘In such a product of nature every part not only exists by means of the other parts, but is thought as existing for the sake of the others and the whole, that is as an (organic) instrument. Thus, however, it might be an artificial instrument, and so might be represented only as a purpose that is possible in general; but also its parts are all organs reciprocally producing each other. This can never be the case with artificial instruments, but only with nature which supplies all the material for instruments (even for those of art). Only a product of such a kind can be called a natural purpose, and this because it is an organized and self-organizing being’. (Kant 1790)
This passage clearly shows the link between Kant’s and Rosen’s accounts of life. The property of closure to efficient cause in relational models can be seen as a formalisation of Kant’s intuition that living organism’s parts reciprocally produce each other. In other words, every constituent process is conditioned by some other process in the system. Thus, the natural purpose of given traits of an organism is to be found in the organism itself, not elsewhere (i.e. in the mind of its creator, divine or not). This way of thinking about biology clearly diverges from both the mechanists (Descartes or La Mettrie) and natural theologists (Paley). Living creatures are autonomous, that is, they themselves shape their organisation rather than relying on some external design.
Kant’s position towards the possibility of a mechanistic explanation of life was therefore ambiguous. He seemed to assume that living systems possess an inherent teleological component that could not be derived from Newtonian physics. Therefore, biologists must add this assumption ad hoc to mount a teleological explanation. Kant famously exclaimed that ‘there will never be a Newton of the blade of grass’. (Ironically, it was Rosen who was posthumously dubbed ‘the Newton of biology’ (Mikulecky 2001).) On the other hand, this limitative claim was probably epistemological rather than ontological: it pertained the limitation of our minds. Curiously, it was just half a century after the Third Critique when a reasonable reductive account of natural purposes in terms of physical processes (natural history) was proposed by Darwin.
An influential continuation of Kant’s account is the idea of autopoiesis (Varela et al. 1974; Maturana and Varela 1980). (Actually, Varela in his last published paper (Weber and Varela 2002) himself dubs Kant as the grandfather of the theory of autopoiesis). Autopoietic system, argued to be a minimal model of living creature, is an organized system characterised by two properties: (a) it continuously realises the networks of processes that maintain its existence and (b) it has a concrete spatial boundary (specified by the topological domain of its realization as a network).
It is often argued that property (a) (sometimes termed ‘operational closure’) is roughly equivalent to the property of closure to efficient cause (Cárdenas et al. 2010). However, Varela and Maturana not only from the very beginning characterise an autopoietic system as a (special kind of) machine, but also introduced the concept of autopoietic systems as a framework for computer simulations of living processes. The concept of autopoiesis later turned out to be very influential in the field of artificial life (Froese and Ziemke 2009). This very fact sheds serious doubts on whether (M, R)–systems are indeed not computable.
What can be considered as the contribution of the Kantian heritage to current biology is the bringing out the concept of (biological) autonomy (Bechtel 2007). According to contemporary enactivists, ‘[a]n autonomous system is a system composed of processes that generate and sustain that system as a unity and thereby also define an environment for the system. Autonomy can be characterized abstractly in formal terms or concretely in terms of its energetic and thermodynamic requirements’ (Thompson 2010, p. 44-45). This concept slightly differs as it stresses the role of environment in the functioning of a system: autonomy certainly does not imply isolation (neither causal, nor energetic or material); this aspect is developed further by Ruiz-Mirazo et al. (2004)). Also, note that the property that every constituent process is conditioned by some other process in the system is to be interpreted recursively: a given process is conditioned by some earlier process and there need not be an infinite loop as far as some external explanation of the origins of the first process is provided. A similar recursive treatment of the concept of self-organization can be found in (Bickhard 2009).
It is, furthermore, worthwhile to point out here that the distinction Rosen makes between reductionist and relational models is hardly congruent with the distinction between dynamical and mechanistic explanation in contemporary philosophy of science. His criticism of reductionist modelling shares some points with early functionalism Cummins (1975); namely: (i) rejection of covering-law model of explanation, (ii) introduction of the concept of function (understood as a causal role in a given system) and (iii) assuming strong substrate neutrality and abstracting away from material constitution of a system under consideration (‘throwing away the matter and keeping the organisation’, to use Rosen’s (1991, p. 120) phrase).
It should also be stressed that the mechanistic (reductionist) explanation Rosen rejects is very unlike contemporary neomechanicism (Machamer et al. 2000; Bechtel 2011)), which at least since (Bechtel 2007) appreciates system-wide organisation level as well. Actually, as Bechtel (2007) points out, it is somewhat ironic for the antimechanist movement of Rosen et consortes to have sprung in the early second half of the 20th century, because philosophers of science recognised the concept of mechanism as an adequate model of biological explanation as late as in the 2000s. Furthermore, Rosen seems to be committed to the view of mechanistic explanation as ‘analysing down to a family of constituent particles’. But if the palette of available bottom-level explanantia is so drastically narrow to encompass only geometrico-mechanical activities (as termed by (Machamer et al. 2000; Bechtel 2011, p. 14), then not only living systems but also an arbitrary material system could not be accounted for mechanistically. In other words, it seems to be Cartesian mechanism Rosen is fighting with and Cartesian mechanism is known to be false (even, or foremost) for physics at least since 19th century, when the existence and role of energetic and electromagnetic activities were recognised.
It is probably not the kind of mistake an educated physicist like Rosen would make, and one could argue ‘the Newtonian picture’ is a deliberately chosen simplistic example of mechanistic explanation and ‘constituent particles’ are to be understood in a broad sense. Still, however, Rosen explicitly assumes that mechanistic explanation must bottom down to some universal basic level (constituent particles ‘are to be further fractionated’, ibidem), while contemporary mechanists argue this basic level is highly domain-dependent and generally a working neuroscientist can, should and actually does abstract away from the level of elementary particles (Machamer et al. 2000). (Note that it is very much in the vein of Rosen’s relational framework.) To sum up, Rosen sees mechanistic explanation as committed to strong explanatory reductionism (‘greedy reductionism’, to use Dennett’s (1995) phrase) and his alternative – relational modelling – shows remarkable similarities to what is known as mechanism today.
Lastly and even more ironically, since dynamical systems are by definition space-state based systems, they are first and foremost to fall under Rosen’s criticism, if it were sound. Thus, the dynamical systems approach in cognitive science (Port and Van Gelder 1995; Kelso 1995) is to be rejected even sooner than computationalism it tries to undermine. Nowadays, however, it is dynamicism that develops the notions of self-organisation, affordance and complexity in a Rosenian way and explicitly draws the notion of autonomy from closure to efficient cause (Chemero and Turvey (2008); Chemero (2008)). It is dynamicism, also, that provides a cogent critique of Rosen’s approach: relational models, being time-invariant, do not capture essentially temporal features of life, such as adaptivity, development and evolution. For instance, Montévil (2020) argues convincingly that living systems, as opposed to non-living physical systems, are inherently historical. They require specific methods and epistemology to accommodate their historicity, even when we study how organisms behave here and now. While physics postulates invariance in order to explain changes, in biology we cannot postulate invariance behind changes because invariances are limited to constraints whose validity can be ascertained only given a particular time and time-scale (Montévil 2020, p. 8). Thus, a time-invariance mode of explanation espoused by relational biology will necessarily miss the essence of living systems.
What is more, as relational models abstract away from the material underpinnings of living systems, they are unable to account for certain constraints on how its components perform their functions (Clark 1997). The latter point could be elaborated as to undermine the concept of abstract, materially neutral modelling altogether: at least in psychology the assumption that cognitive activity can be explained ignoring its neural substrate has turned out to be false (Bechtel and Mundale 1999).
All in all, situating Rosen in contemporary debates in the philosophy of science suggests that he anticipated contemporary neomechanism rather than mounted a successful attack against it. His variety of functionalism is not faultless, but undoubtedly played an important role in recognising the specificity of complex living systems and importance of usually overlooked organisational level of description of living systems. It seems, however, that the anachronism of Rosen’s philosophical commitments does not affect the evaluation of his argument. It rests not on the curiosities of the relational framework, but on a specific (circular) causal structure of certain models and as such may be formulated even if the relational approach is misguided.
It’s about time
There is one crucial dimension of life that relational biology leaves behind: time. Obviously, this is by design: it is time-invariance that allows for final causation as morphism in the direction opposite to material causation. It is this design feature that supposedly makes models of (M, R)–systems uncomputable. I will argue, however, that it is just a feature of a particular formal representation of self-organising systems, not their essential property. Computations over systems composed of interdependent components are omnipresent in computer science and machine learning. It is also quite trivial to compute with self-referring objects. On a machine code level, there is nothing mysterious for a memory location to store its own address. All it takes to compute with self-referring objects is an algorithm that avoids unwanted infinite loops by witless bookkeeping of visited states.
There are numerous algorithms in modern machine learning that exploit self-reference in a non-trivial way to optimise functions defined in a circular way. The usual strategy when dealing with such a circular dependence between two variables, A and B, is to apply an iterative procedure, first pretending that A is fixed and optimize for B, and then the other way around. For instance, such a problem arises in probabilistic inference for latent variable models, when the optimal parameters (i.e. a maximum likelihood solution) depend on the value of a latent variable, and the expectation of the latent variable in turn depends on the parameters. Because the variables and parameters depend on each other, there will be no closed form solution for this distribution. However, it can be computed approximately via an iterative procedure. One theoretically principled scheme is the expectation maximisation algorithm (Dempster et al. 1977). Here, one compute the expected value of latent variables at time-step t based on previous values of parameters from time-step t − 1. Then, one can compute the maximum likelihood parameters at time-step t based on those latent variables and increment the counter t. One other example of an equation with a circular structure is the Bellman equation, recursively linking the value of a given state to other state accessible from it. Here again, the now-classic dynamic programming solution consists of iteratively exploiting dependence in one direction.
The common pattern is parametrising the system in terms of discrete (or continuous) time-steps t to removing the circular dependence. Let us consider a similar parametrisation of an (M, R)–system on Figure 2. First, we index each give variable X, Y, H (X, Y), H(Y, H(H, Y)), ϕ, ψ and θ with an index t. Then, we draw solid arrows between selected pairs of variables sharing an index and dotted arrows between selected pairs of arrows with different indices (such as the input of an arrow is associated with t − 1 and the output of an arrow with t). This guarantees that there is no circular dependence: for a morphism, its input will always have index equal or earlier than its output. This rewriting can be applied recursively, effectively unrolling the circular dependencies of (M, R)–systems into a chain of linear dependencies, as displayed on Figure 3. Note that again there is no input to X, because it stands for the environmental input to the system. Unrolling a cyclic graph of an (M, R)–system into a time-indexed chain.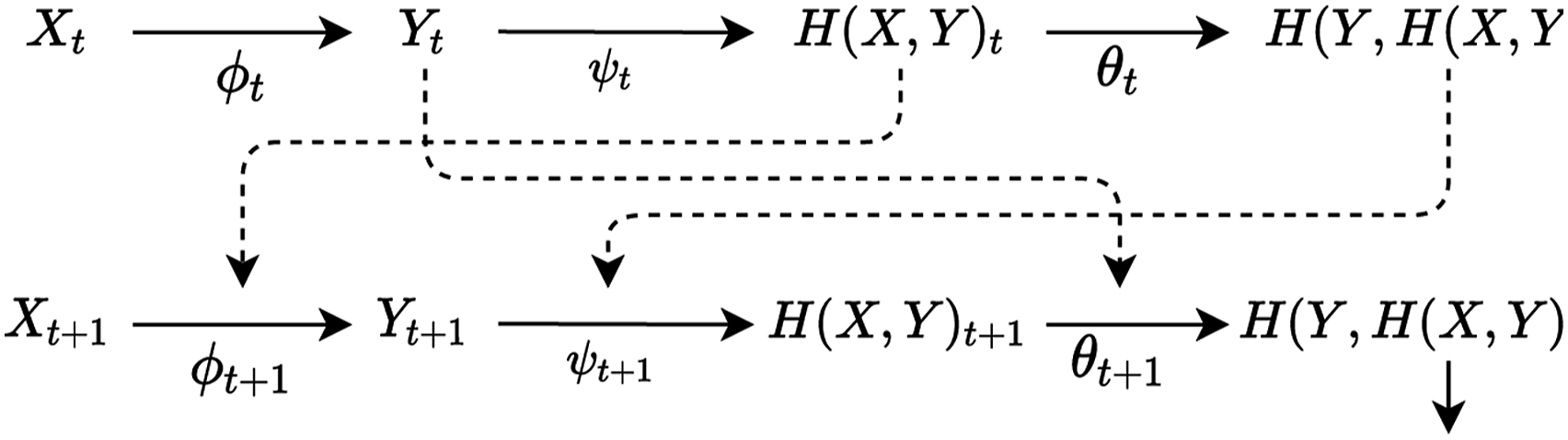
While one may argue that time-indexing does not really capture Rosen’s intended interpretation of these components, there is little justification for such an interpretation. Rosen does not really provide a concrete biological interpretation of these components, and the graphical model I provided seems to capture the essential architecture of self-organisation (nested feedback loops governed by utility) and the philosophical intuitions behind it (autonomy) equally well. There is little justification for deliberately choosing a formal model just because it manifests irremovable, inconvenient properties.
The proposed reparametrisation of (M, R)–systems has benefits on its own as it invites powerful probabilistic interpretations. Highlighting the analogy with expectation maximisation, one can try to reinterpret the (M, R)–system as a graphical model, that is, treating each component as a random variable and each solid line as a random variable and each dotted line as dependence. For notational convenience, let us define Z = H (X, Y) and C = H(Y, H(H, Y)) = H(Y, Z). The corresponding probabilistic graphical model is displayed on Figure 4. A probabilistic graphical model corresponding to the unrolled (M, R)–system from Figure 3.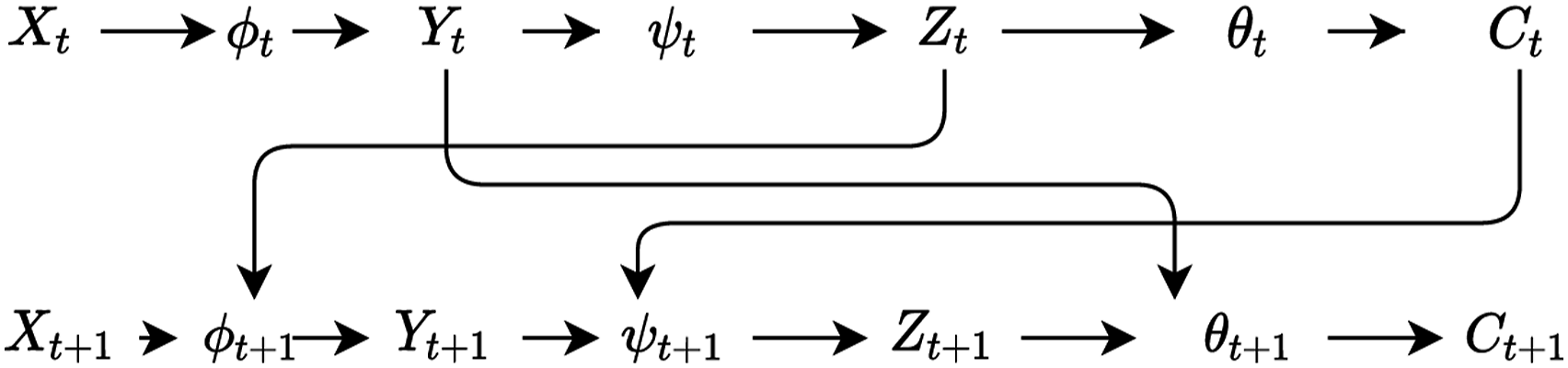
Then the problem of simulating an (M, R)-system reduces to inferring the join distribution p (ψ1, ϕ1, θ1, X1, Y1, Z1, C1, …, ϕ
t
, ψ
t
, θ
t
, X
t
, Y
t
, Z
t
, C
t
). This distribution can then be decomposed as
This particular dependency structure enables using a simple inference algorithm to infer the joint distribution: ancestral sampling. We can just start with an initial distribution over X1 and keep inferring the conditional probability of each variable using its conditioning set. The further reason this probabilistic graphical model reparametrisation is interesting is that it exposes the connection with active inference, a framework described in Section 2. According to inference, living systems are generative models trying to probabilistically model both its internal states and its environments. Mathematically, they can be seen as probabilistic graphical models with a particular dependency structure known as a Markov blanket. The Markov blanket of a random variable A is the set of random variables given which A is conditionally independent from all other random variables in the system. This property has recently attracted much philosophical interest leading to accounts of self-organisation in terms of Markov blankets developed assuming (B4), for example (Friston 2013; Clark 2017; Korbak 2021). The dependence structure embodied in the probabilistic reparametrisation of (M, R)–systems induces three such Markov blankets: 1. The set {ψt−1, θt−1, ϕ
t
, Zt−1} is a Markov blanket of Z. 2. The set {ψt−1, ϕt−1, θ
t
, Yt−1} is a Markov blanket of Y. 3. The set {θt−1, ψ
t
, Y
t
} is a Markov blanket of C.
This fact once again reveals a deep similarity between different formal models of self-organisation, built using different sets of assumptions.
To summarise, Rosen claimed that (M, R)–systems are not computable because each component of the system depends on other components. I have argued that this argument is misguided because a simple reformulation of (M, R)–systems is possible that both (i) captures all the biological and philosophical intuitions regarding the notion of self-organisation and (ii) can implement computations under an iterative scheme. In other words, (M, R)–systems are not only computable but also tractable (i.e. computable in practice). Finally, I have shown that – via the graphical model formulation – (M, R)–systems are linked in important family of models of self-organisation assuming the free energy principle – a paradigm widely discussed in contemporary theoretical biology and cognitive science.
Closing thoughts
When Rosen makes a point that biology is more general than physics because it deals with systems that physics cannot account for, he refers to the Gödel’s incompleteness theorem (Rosen 1991, p. 7-10). Mechanical inexplicability of life is likened to the impossibility of providing a complete axiomatisation for a sufficiently rich and consistent theory, such as Peano arithmetic. The connection, however, is even deeper since Rosen aims at making a philosophical point based on certain metamathematical limitative results. A similar anti-mechanist argument was famously mounted by Lucas (1961). According to Lucas, no machine is equivalent to human mind because the mind can recognize the truth of a Gödel sentence for this machine, but the machine – due to Gödel’s theorem – either cannot recognise it or is self-contradictory (and recognises an arbitrary sentence as true). In both cases, human mind outperforms the machine. This argument, however, is highly questionable because Gödel himself doubted that similar conclusion about mathematical intuition can be derived from his theorem with no further assumptions. Krajewski (2020) offers a compelling analysis of Lucas’ argument, showing that every argument following his way of thinking is either circular or inconsistent.
Both Rosen’s and Lucas’ argument are part of a philosophical project of recognising certain limitations of science or some unusual complexity of certain natural systems, based on their formal properties. This strategy was neatly summarized by Hofstadter: ‘All the limitative Theorems of metamathematics and the theory of computation suggest that once the ability to represent your own structure has reached a certain critical point, that is the kiss of death: it guarantees that you can never represent yourself totally. Incompleteness Gödel’s Theorem, Church’s Undecidability Theorem, Turing’s Halting Problem, Tarski’s Truth Theorem – all have the flavor of some ancient fairy tale which warns you that ‘To seek self-knowledge is to embark on a journey which... will always be incomplete, cannot be charted on any map, will never halt, cannot be described‘.’ (Hofstadter 1979)
On this view, the self-referencing nature of self-organising systems is supposed to impose fundamental limitations on some modes of explanation of such systems. Similarly, Rosen’s argument may be interpreted as using the fact that there are functions that are not computable (a consequence of the undecidability of Turing’s halting problem). While this is indeed true, Rosen fails to show that his (M, R)–systems are indeed to be interpreted as non-computable functions. As I intended to show in the previous section, circular causation does not bring about any actual infinity into system’s description. An (M, R)–system can very well be described in a non-circular way by unrolling along the temporal dimension. Furthermore, I argued that this simple reformulation both (i) captures all the biological and philosophical intuitions regarding the notion of self-organisation and (ii) can implement a computation under an iterative scheme. The flaw in Rosen’s argument is thus remarkably similar to that in Lucas’: circularity is an irreducible property of (M, R)–systems only if it is presupposed from the very beginning.
Footnotes
Acknowledgements
The author thanks Marcin Miłkowski for feedback on earlier drafts of this paper.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was funded by the Ministry of Science and Higher Education (Poland) research Grant DI2015010945 as part of ‘Diamentowy Grant’ programme.
Note
Author Biography
Tomasz Korbak is a PhD student at the Department of Informatics, University of Sussex working on deep reinforcement learning, generative models and computational neuroscience with Dr. Chris Buckley and Prof. Anil Seth. Previously, he was a researcher at the Institute of Philosophy and Sociology, Polish Academy of Sciences working with Prof. Marcin Miłkowski and a research assistant at the Human Interactivity and Language Lab, Faculty of Psychology, University of Warsaw working with Prof. Joanna Rączaszek-Leonardi. He is interested in biologically-inspired and probabilistic approaches to artificial intelligence and recently focuses on control as inference and controllable generative modeling.