Abstract
Systemic delivery of adeno-associated viral (AAV) vectors is a promising approach for brain gene therapy, particularly in combination with emerging techniques such as focused ultrasound (FUS), which can transiently and noninvasively open the blood–brain barrier to facilitate delivery of AAVs to the brain. However, off-target vector accumulation, particularly in the liver, remains a significant safety concern. To address this, we introduced a mutation into variable region 1 of the AAV9 capsid (G266A) and assessed its effect on liver and brain transduction, testing both direct intraparenchymal injection and intravenous injection with FUS. Interestingly, we found that G266A mutation strongly decreased liver transduction while having only a modest effect on transduction of the brain and other tissues. We also found that the G266A mutation had minimal impact on cell attachment and uptake but strongly decreased binding of the capsid to both human and mouse AAV receptor (AAVR), which likely explains the decrease in liver transduction. Overall, our findings suggest that the G266A mutation and modification of AAVR binding could be a useful strategy to mitigate liver toxicity associated with systemic brain gene therapy.
INTRODUCTION
Recombinant adeno-associated viral (rAAV) gene therapy is a promising modality for treatment of neurological disorders. rAAVs are replication-deficient, nonpathogenic, and can deliver transgenes up to 4.7 kilobases in size to a wide variety of cell and tissue types.1,2 Numerous preclinical and clinical studies have demonstrated safe and effective AAV-mediated gene transfer to the liver, heart, skeletal muscle, kidney, brain and other tissues; and to date, seven AAV gene therapy drugs have been approved by the Food and Drug Administration.1,2 Moreover, while the blood–brain barrier (BBB) limits the efficiency of brain transduction upon intravenous (IV) administration of rAAVs, recent advancements in focused ultrasound (FUS) have unlocked the possibility of transiently opening the BBB for enhanced delivery of rAAVs to the brain through minimally invasive IV administration. 3 FUS relies on a combination of (1) low-intensity ultrasound that is applied to a specific brain region, and (2) IV microbubbles that together induce bioeffects in the targeted brain region that open the BBB. 3 This technique has been shown to enhance the delivery of AAVs and other molecules in both rodents and nonhuman primates (NHPs),4,5 and its safety has been demonstrated in human subjects.5–7
However, off-target accumulation of the vector remains a significant obstacle for FUS and other systemic gene therapy approaches. Notably, AAV9, which is typically the capsid of choice for FUS due to its superior ability to transduce the brain, is also highly liver-tropic.8–10 This is problematic in the context of systemic vector administration, in which the liver and other peripheral tissues can accumulate large amounts of circulating vector and prevent efficient targeting of the intended tissue. As a result, excessively high vector doses may be required to achieve the target expression level, which can lead to peripheral injury, exacerbated immune responses, and high vector manufacturing costs. Indeed, liver toxicity has been linked to systemic administration of AAV in several recent studies.11–13 For these reasons, minimizing peripheral vector accumulation is critical to ensure the safety and efficacy for FUS-mediated vector administration and other systemic gene therapy approaches.
One potential approach to achieve peripheral de-targeting is modification of the variable region 1 (VR1) loop of the AAV capsid, which includes a stretch of amino acids from 262 to 272 (VP1 numbering).14,15 For example, mutagenesis of the AAV9 capsid at positions G267, N270, and N272 have been shown to reduce AAV9 liver transduction,16–18 although the underlying mechanism has not been elucidated. N272A-mediated detargeting can likely be attributed to disruption of galactose binding, as N272 is one of several residues that make up the AAV9 galactose binding pocket. 19 In contrast, positions N270 and G267 are not reported to be involved in galactose binding, which suggests that other mechanisms may be at play. Interestingly, the VR1 loop has also been found to interface with AAVR, 15 which is a critical cellular factor required for AAV9 transduction, 20 suggesting that VR1 mutagenesis could impair transduction by interfering with AAVR binding. However, the effect of VR1 mutagenesis on AAV9–AAVR binding has not yet been fully tested. Equally unclear is the potential utility of VR1 mutagenesis for brain gene therapy, as disrupting AAVR binding may affect transduction of other tissues in addition to the liver.
In this study, we introduced a single point mutation at position G266—this site was chosen because it is highly surface exposed within the VR1 loop and is directly adjacent to the G267 “liver toggle” residue, which has been shown to be particularly important for liver tropism. Indeed, we found that mutation of this site to alanine strongly reduced liver transduction. Then, using both direct intraparenchymal brain injections as well as IV injections with FUS–BBB opening (FUS–BBBO), we found that the G266A mutation had only a modest effect on brain transduction and led to a significant improvement in the ratio of brain to liver transduction. Finally, using in vitro binding assays, we found that the G266A mutation potently disrupts AAV9–AAVR binding while having a minimal effect on AAV9–galactose binding and cell/liver uptake. Overall, our findings suggest that the G266A mutation and modification of AAVR binding could be a useful strategy to mitigate liver toxicity associated with AAV gene therapies.
MATERIALS AND METHODS
AAV production
All AAVs were generated by triple transfection of human embryonic kidney (HEK) 293T cells with (1) AAV packaging plasmid; (2) AAV transfer plasmid encoding an expression cassette flanked by inverted terminal repeats (ITRs); and (3) adenoviral helper plasmid (see below for plasmid details) using polyethyleneimine (PEI) Max. Cells were lysed by three freeze-thaw cycles, and AAVs were purified by iodixanol gradient ultracentrifugation as previously described, 21 followed by buffer exchange to PBS with 0.001% Pluronic-F68 using Cytiva VivaSpin Columns (MWCO 100,000). Capsid protein ratio and purity were verified by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and vector titers were measured using a TaqMan qPCR assay for green fluorescent protein (GFP). For vector libraries (used in Fig. 1), vectors were produced individually, titrated, and pooled in equal amounts based on titers.
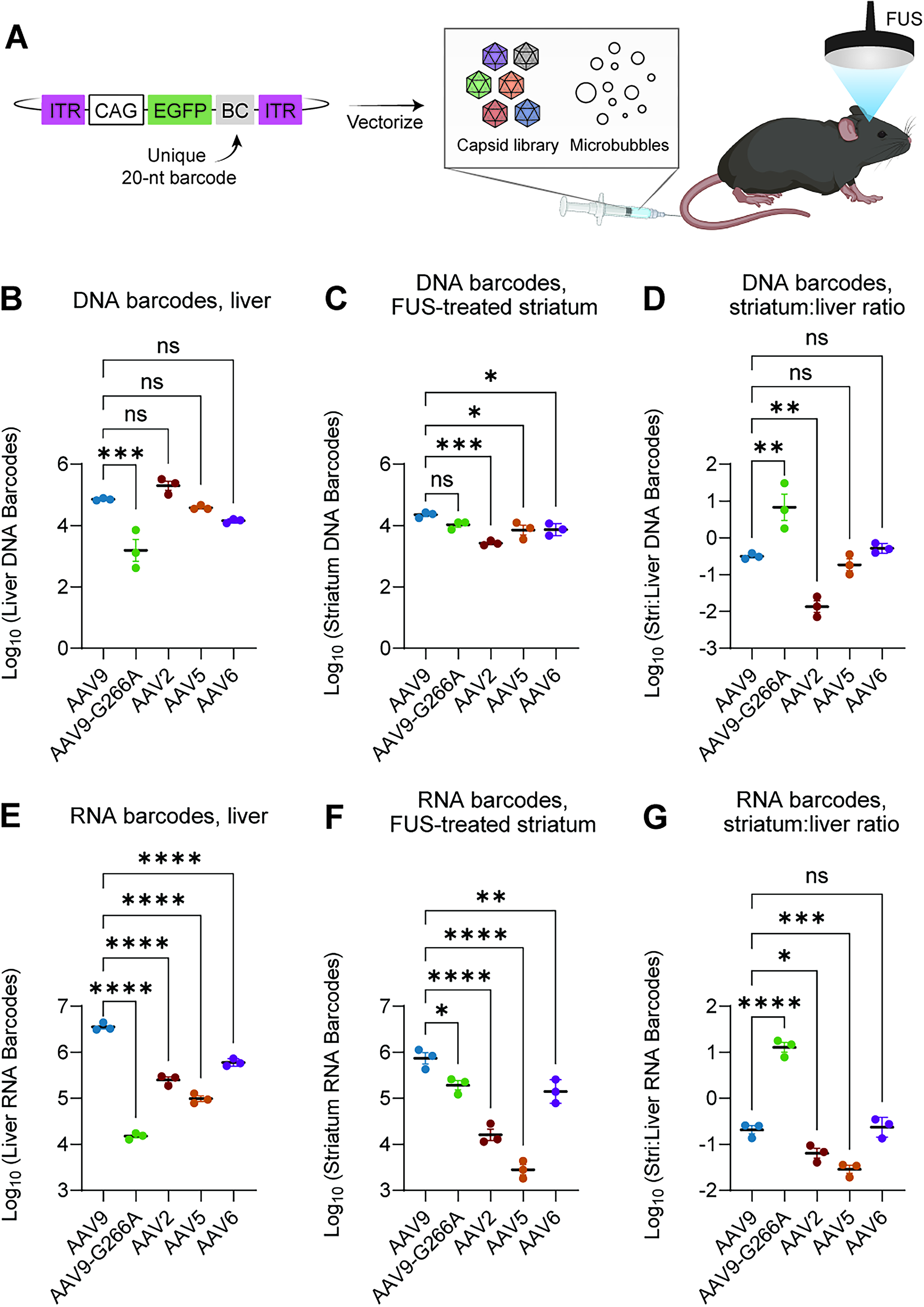
G266A mutation enhances the brain-to-liver transduction ratio with focused ultrasound (FUS).
Plasmids and molecular cloning
Plasmids for AAV production
AAV packaging plasmids encoding a partial p5 promoter, the AAV2 rep gene, and the serotype-specific cap gene were used. Point mutations (G266A, N272A, or W503A) were introduced into the cap sequence by site-directed mutagenesis (cloning performed by Azenta Life Sciences). For production of single capsids (used in Figs. 2, 3, and 4), an ITR plasmid encoding the cytomegalovirus (CMV) promoter, firefly luciferase (FLuc), an internal ribosomal entry site (IRES), GFP, and the SV40 poly(A) signal (purchased from Vector Builder) was used. For production of vector libraries (used in Fig. 1), ITR plasmid encoding the CAG promoter, GFP, and the BGH poly(A) signal was used; each vector within the library was packaged with a unique 20-nt barcode sequence in the 3′ UTR (cloning performed by Aldevron LLC).
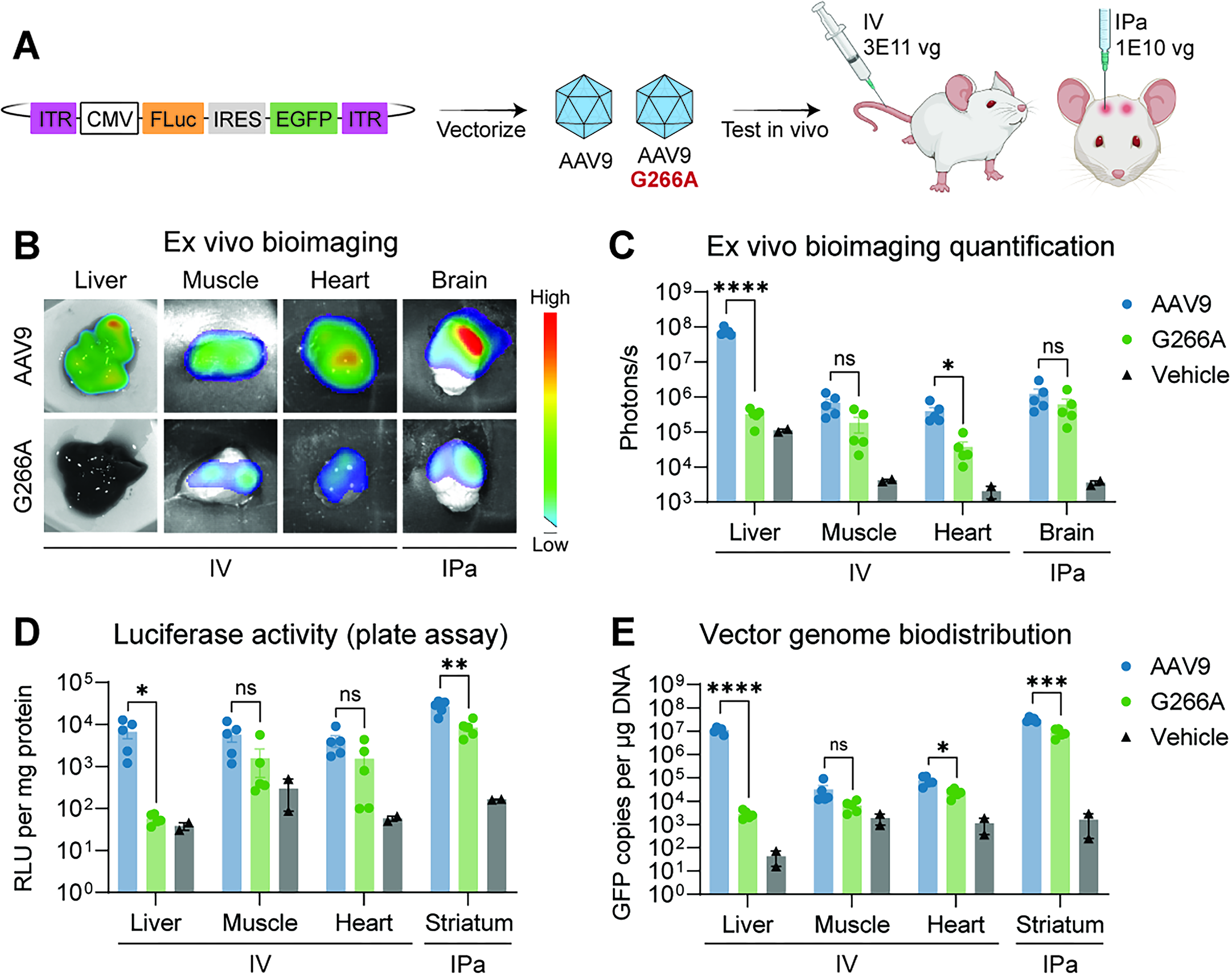
G266A mutation disproportionately decreases AAV9 liver transduction. Assessment of AAV9 and AAV9-G266A transduction profiles in mouse tissues (liver, skeletal muscle, heart, and brain) following IV injections (3E11 vg/injection) or IPa injections (1E10 vg/injection).
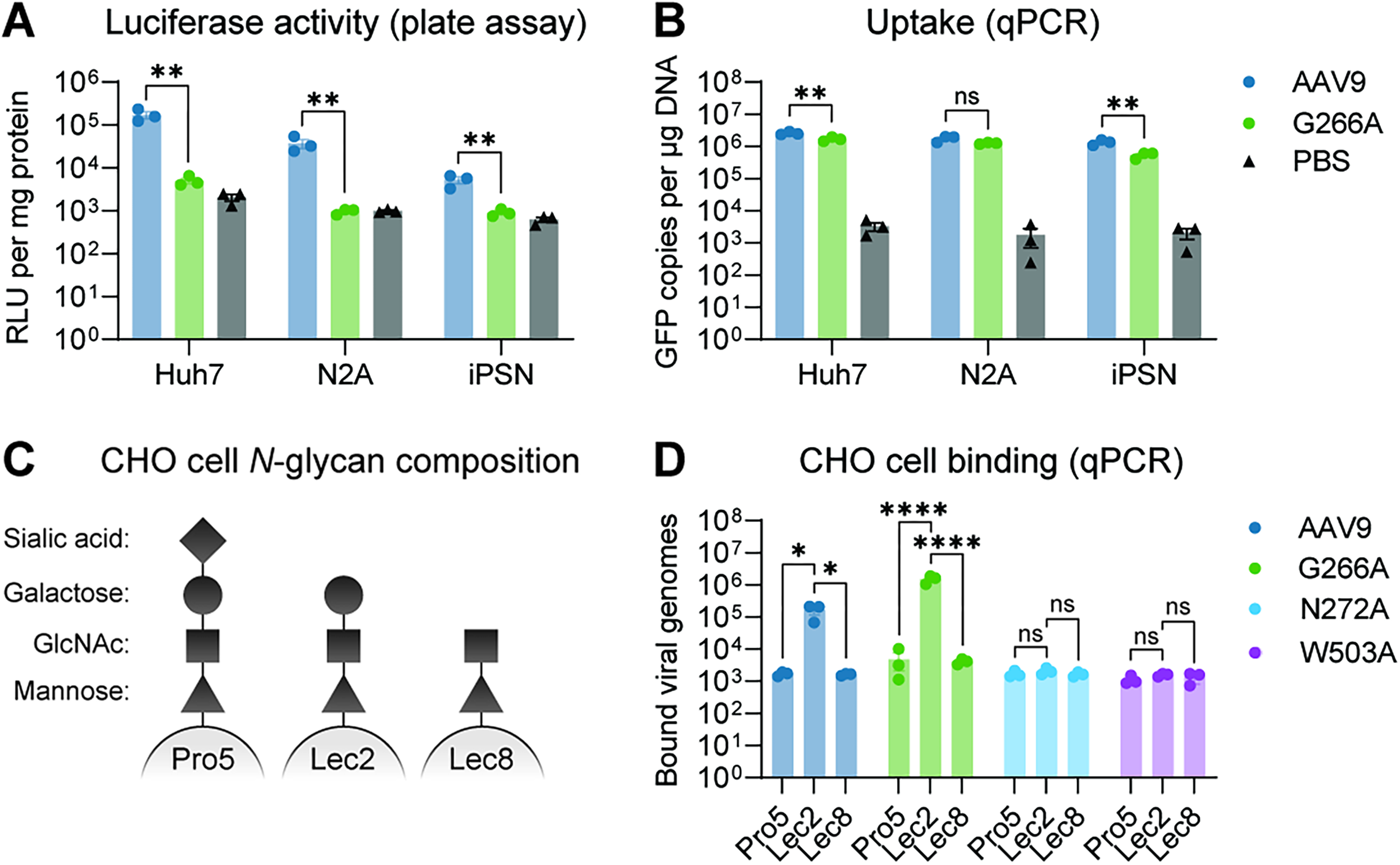
G266A mutation impairs AAV9 transduction in vitro while minimally affecting uptake and cell surface glycan binding.
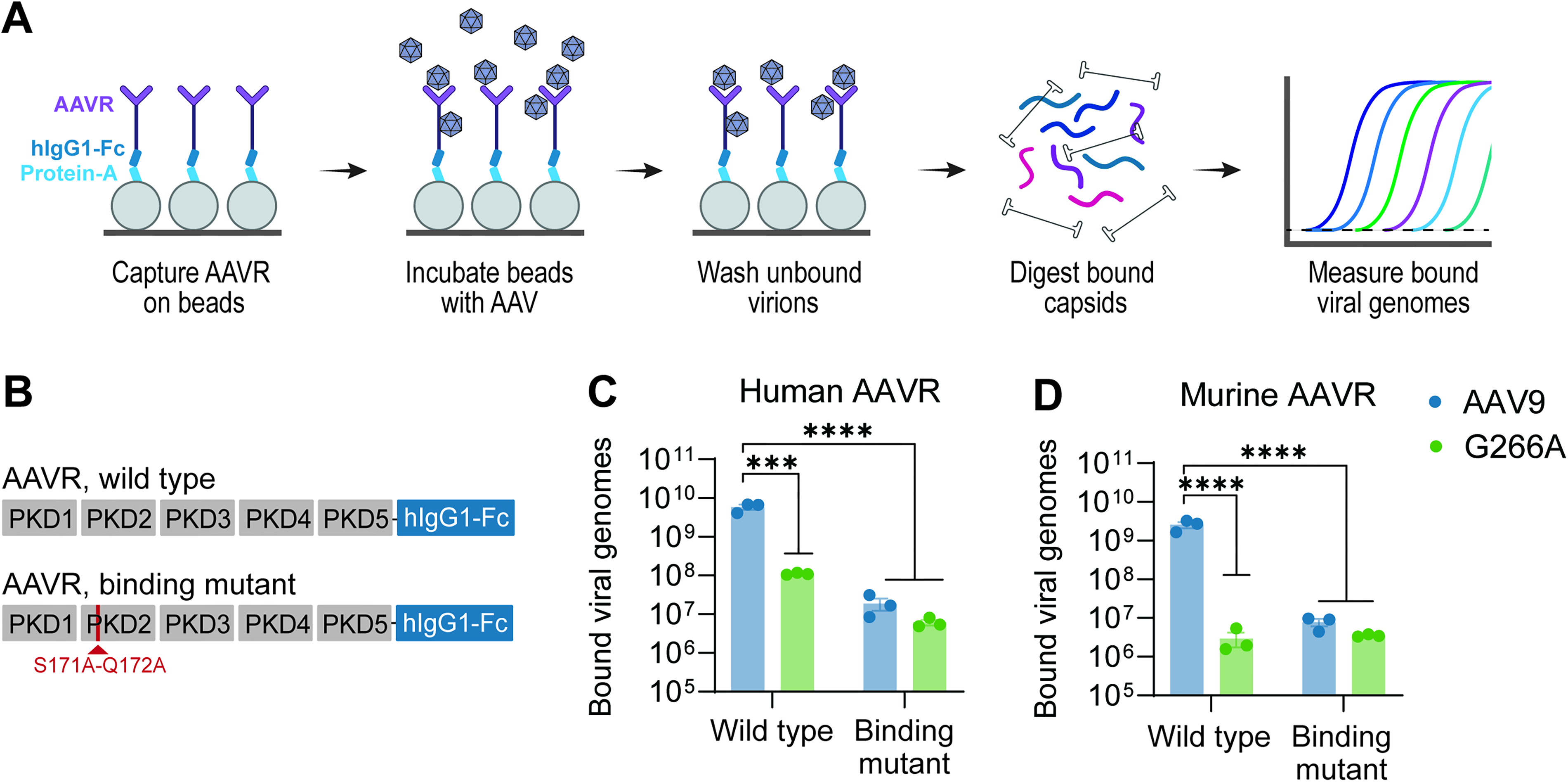
G266A mutation disrupts AAV receptor (AAVR) binding.
Plasmids for hIgG1-Fc-tagged AAVR production
The human AAVR (KIAA0319L) or murine AAVR (AU040320) genes were synthesized by Azenta Life Sciences. The polycystic kidney (PKD) domains 1 thru 5 were amplified by PCR and cloned into an expression vector downstream of the EF-1 alpha promoter and upstream of the hIgG1-Fc sequence. Point mutations (S171A and Q172A) were introduced by Gibson assembly cloning (performed in-house at Spark Therapeutics).
Animals
All animal procedures were carried out in accordance with protocols approved by Spark Therapeutics Institutional Animal Care and Use Committee (IACUC). 8- to 10-week-old B6 Albino mice (Jax: 00058) and/or C57BL/6J mice (Jax: 00064) were used for bioimaging and/or FUS studies, respectively. Animals were maintained on a 12-h light-dark cycle with ad libitum access to food and water.
Intraparenchymal injections
Animals were anesthetized at 3% isoflurane and maintained at 2% for surgical procedures. The head was stabilized in ear bars, shaven, and a midline scalp incision was made to expose bregma and lambda. Using a handheld drill, bilateral craniotomies were made over the striatum (from bregma, AP: +0.65 mm, ML: ± 2.0 mm) and a 33 G Neuros Hamilton syringe (Hamilton 65460) was inserted to a depth of −3.5 mm. 1E10 vector genomes in a volume of 3 µL (formulated in PBS with 0.001% Pluronic F68), or 3 µL of dilution buffer as a vehicle-only negative control, was deposited in each hemisphere at a rate of 400 nL/min with a 5-minute dwell time following complete delivery prior to needle removal. Incisions were then sutured and covered with a topical antibiotic ointment. A subcutaneous injection of carprofen analgesic was given and animals were moved to a recovery chamber for post-operative monitoring.
FUS BBB opening
Animals were anesthetized at 3% isoflurane and maintained at 2% for surgical procedures. A 26 G catheter (ICU medical: SP125-26-14T) was inserted into the tail vein and affixed in place using surgical tape and then flushed with 100 µL of sterile 0.9% saline. Animals’ heads were shaven with electric trimmers and all residual hair removed with depilatory cream. The animals were then moved to the RK-50 (FUS Instruments) placed on a nose cone and stabilized in ear bars. The scalp was sterilized, and a midline incision was made to expose bregma and lambda. The location of bregma was registered in the RK-50 software using a fine-tip metal pointer. Next, the ultrasound transducer cap and tank were filled with degassed water and coupled to the animal skull with sterile ultrasound gel. FUS target coordinates were then selected, consisting of 3 spots in the right striatum. The mice were sequentially injected with 100 µL of purified vector (formulated in PBS with 0.001% Pluronic F68), and approximately 30 µL of DEFINITY microbubbles (approximately 20 µL/kg; Lantheus) followed by a 150 µL flush of sterile 0.9% saline. Immediately after injections, mice were sonicated using the RK-50 FUS system with a 1.5 MHz spherically focused transducer. Ultrasound was applied using 10 ms bursts and a 1 Hz pulse repetition frequency for 60 s. A non-derated peak negative pressure of 0.9 MPa was used for FUS-BBBO based on preliminary tests in our lab. Following pulsing, the incision was closed, and animals were provided analgesia and moved to a recovery chamber for post-operative monitoring.
Ex vivo bioimaging
Animals received a 150 mg/kg body weight intraperitoneal injection of Luciferin (Fisher PI88293). 5 min after the luciferin injection, animals were rapidly euthanized, with liver, heart, skeletal muscle, and brain extracted for imaging. Tissues were then submerged in 15 mg/mL luciferin for 3 min prior to imaging. Tissues were scanned using ex-vivo BLI (AMI HTX, Spectral Instruments). Brain, heart, and skeletal muscle were captured for 5 min using 8 × 8 binning and liver was captured for 30 s using 2 × 2 binning. BLI signal intensities are presented as total flux (photons per second) and quantified using Aura Software (Spectral Instruments).
Plate luciferase assay
For preparation of tissue lysates, 25 mg of frozen tissue was transferred to Eppendorf tubes and homogenized by addition of 350 µL 1X Passive Lysis Buffer (Promega) and a stainless-steel bead followed by shaking at 20 Hz for 1 min using TissueLyser II instrument (Qiagen). For preparation of cell lysates, cells in a 12-well plate were washed once with PBS, then lysed by addition of 200 µL Passive Lysis Buffer directly to the wells; after a brief incubation at room temperature, cell lysates were scraped off the plate and transferred to Eppendorf tubes. Tissue and cell lysates were centrifuged at 18,000 x g for 10 min. 20 µL of clarified lysate was transferred to an opaque white 96-well plate and mixed with 100 µL FLucsubstrate (Promega) and the resulting luminescence was captured immediately using the luminometer setting on the Synergy H1 plate reader (Biotek). Luminescence values were then normalized to total protein concentrations as measured by Pierce bicinchoninic acid (BCA) protein assay (Thermo Fisher Scientific).
Vector genome biodistribution
Bulk DNA was extracted from liver, skeletal muscle (quadriceps), heart, and striatum using the QIAsymphony SP instrument. 200 ng of each DNA sample was analyzed by a TaqMan qPCR assay specific for GFP. A standard curve ranging from 1E7 copies per reaction to 1E1 copies per reaction was generated using a linearized GFP plasmid and used to calculate vector genome copy number per reaction.
Next generation sequencing
AAV vector library characterization
The input vector libraries were characterized via a unique molecular identifier (UMI)-inclusive (quantitative) barcode amplicon sequencing strategy. Briefly, an annealing/extension reaction was performed using a primer with a Truseq R2 Illumina sequencing overhang, containing a UMI and a template-specific sequence (3′ to the barcode). PCR1 was then performed on these products with a primer against the R2 sequencing overhang (which was incorporated in the previous reaction) plus a primer including a Truseq R1 sequencing overhang, containing a template-specific sequence 5′ to the barcode. Following this step, indexing PCR was performed, utilizing the now incorporated full-length R1 and R2 sequencing primer binding sites in the molecule. After sequencing, reads were filtered based on the appropriate flanking sequences and then deduplicated via the UMI. Counts were generated for each known barcode to assess relative input capsid abundance.
Bulk DNA and RNA barcode characterization
Bulk DNA and RNA were isolated from liver and FUS-treated brain regions (striatum) using the miRNeasy Advanced Mini Kit (Qiagen). Bulk DNA was isolated from the gDNA eliminator columns included in the kit by treating them with Proteinase K (Qiagen), washing with buffers AW1 and AW2 (Qiagen), and eluting with nuclease-free water. Barcode counts were generated using the same quantitative (UMI-inclusive) sequencing strategy as the input vector libraries. For DNA, bulk total DNA was input into the R2 + UMI annealing/extension reaction. For RNA, bulk total RNA was input to a reverse transcription (RT) reaction where our RT primer included the R2 sequencing overhang + a UMI + an anchored poly(dT) sequence. Following this step, we used endpoint qPCR to read out relative template abundance (using the same PCR1 primers as in the vector characterization experiment). Ct values from this were used to inform total amplification cycles for the library preparation strategy (which are split between PCR1 and indexing PCR). The same PCR1 and indexing PCR conditions were used for bulk RNA/DNA libraries as for the input vector libraries. For bioinformatics analysis, UMIs were extracted and reads were filtered by primer sequences to make sure the amplicons were valid. Barcode sequences were extracted according to fixed flanking sequences, and reads with the same UMI and barcode sequences were collapsed as PCR duplicates. Deduplicated counts were obtained for each barcode.
Cell culture
Cell lines
HEK 293 T, Huh7, and Neuro-2a cells were obtained from an internal repository at Spark Therapeutics, Inc. and were cultured in Dulbecco's Modified Eagle's Medium (DMEM) supplemented with 10% fetal bobine serum (FBS), penicillin/streptomycin, and L-glutamine. CHO cell lines (Pro5, Lec2, and Lec8) were obtained from ATCC and were cultured in alpha-minimum essential medium (MEM) supplemented with 10% FBS, 1X penicillin/streptomycin, and 1X L-glutamine. All cell lines were maintained using standard cell culture techniques.
iPS cell differentiation into neurons
Human induced pluripotent stem cells (iPSCs) containing a doxycycline-inducible neurogenin2 expression cassette integrated into the AAVS1 locus were obtained from AlStemBio (iP11NA). iPSCs were cultured in mTeSR Plus Basal Medium supplemented with penicillin/streptomycin and RevitaCell Supplement (ThermoFisher Scientific). Media was changed daily, and cells were passaged upon reaching 80% confluency. To differentiate into neurons, iPSCs were seeded onto well plates coated with Matrigel (Corning) and cultured in DMEM/F12 supplemented with 1X non-essential amino acids (NEAA), 1X penicillin/streptomycin, 1X N2 supplement, 1X GlutaMAX supplement, 1X B27 supplement, 10 ng/mL brain-derived neurotrophic factor (BDNF), 10 ng/mL neurotrophin-3 (NT3), and 2 µg/mL doxycycline for 14 days. Half media changes were performed every other day.
Vector genome uptake assay
Cells were incubated with purified AAV for 24 h, washed three times with PBS, and lysed using Genomic Lysis Buffer (Zymo Research). DNA was extracted using the Quick-DNA Microprep kit (Zymo Research). Fifty nanograms of each DNA sample was analyzed by a TaqMan qPCR assay specific for GFP. A standard curve ranging from 1E7 copies per reaction to 1E1 copies per reaction was generated using a linearized GFP plasmid and used to calculate vector genome copy number per reaction.
CHO cell AAV binding assay
The day prior to the assay, Pro5, Lec2, and/or Lec8 cells were seeded onto 96-well plates at a density of 1E4 cells per well. The day of the assay, the plate was prechilled on ice for 30 min and kept on ice for the duration of the assay. Cell culture medium was replaced with ice-cold serum-free alpha-MEM containing purified AAV vector (1E10 vector genomes per well) for 1 h. Cells were then washed three times with ice-cold PBS and lysed using Genomic Lysis Buffer (Zymo Research). Lysates were heated to 95°C for 5 min, then DNA was extracted using the Quick-DNA Microprep kit (Zymo Research). Isolated vector genomes were measured by a TaqMan qPCR assay specific for GFP.
AAVR-Fc bead purification
HEK 293 T cells were seeded into T225 flasks and grown until 85–90% confluent, then transfected with 62 µg of AAVR-hIgG1-Fc plasmids (see above) using PEIpro (Polyplus). The next morning, the cell culture media was replaced with serum-free high-glucose DMEM. Two days later (3 days after transfection), cell culture supernatant (∼50 mL) was collected and supplemented with 0.05% Tween-20 (final concentration) and 65 µL superparamagnetic Dynabeads Protein A for Immunoprecipitation (Thermo Fisher Scientific 10001 D). Tubes were placed at 4°C overnight with end-over-end rotation. The next day, supernatants were briefly centrifuged, and pelleted Dynabeads were resuspended in PBS with 0.05% Tween-20 (PBS-T). Tubes were placed on a paramagnetic block for 2 min, PBS-T was aspirated, and beads were washed twice with PBS-T. An aliquot of beads was analyzed by SDS-PAGE followed by SYPRO Ruby Protein Gel Stain (Thermo Fisher) to verify molecular weight.
AAVR-Fc bead binding assay
Purified AAVR-coated beads were incubated with purified AAV vector (1E10 vector genomes) overnight at 4°C with end-over-end rotation. The next day, beads were washed three times with PBS-T, then resuspended in 30 mM Tris-HCl buffer. Samples were then digested by the addition of Proteinase K and incubation at 56°C for 30 min to release encapsidated AAVR-bound vector genomes. After a 10-min heat inactivation at 85°C, AAVR-bound vector genomes were then quantified by a TaqMan qPCR assay specific for GFP.
Statistical analysis
All data are shown as mean ± standard error of the mean. Raw data were log-transformed to achieve a normal distribution where appropriate. Statistical significance was determined using an unpaired student’s t-test when comparing two groups or one-way ANOVA when comparing three or more groups, with either Tukey’s or Dunnett’s test for multiple comparisons where appropriate.
RESULTS
We began by introducing the G266A mutation into the AAV9 capsid and testing its effect on AAV9 transduction in vivo. To accomplish this, we produced both the AAV9-G266A and parental AAV9 capsids with a CMV-driven dual reporter cassette consisting of FLuc and GFP, separated by an IRES sequence (Fig. 2A). We then tested these vectors in mice via two routes of administration: (1) bilateral intraparenchymal (IPa) injection targeted to the striatum to enable direct assessment of brain transduction, and (2) IV injection, to assess transduction of the liver, heart, and skeletal muscle (Fig. 2A). Three weeks after injection, tissues of interest were harvested for measurement of luciferase activity by ex vivo bioluminescence imaging and plate-based FLuc assay, as well as vector genome biodistribution by qPCR. Strikingly, we observed a consistent and highly significant > 100-fold decrease in liver transduction by the G266A mutation in IV-injected animals (Fig. 2B–E). In contrast, we observed a moderate (3- to 4-fold) decrease in brain transduction by the G266A mutation in IPa-injected animals (Fig. 2B–E). Heart and skeletal muscle transduction in IV-injected animals was also only moderately affected by the G266A mutation (Fig. 2B–E). Therefore, the G266A mutation disproportionately decreases transduction in the liver compared with other tissues—including the brain—which suggests that it would be a strong candidate for systemic administration with FUS-BBBO.
Therefore, we set out to test the performance of AAV9-G266A with systemic administration and FUS-BBBO. As controls, we chose to utilize not only parental AAV9, but also several other naturally occurring AAV serotypes (including AAV2, AAV5, and AAV6). Furthermore, to minimize the number of animals required, we utilized a barcoded AAV capsid library approach, in which each capsid variant was individually packaged with a CAG-driven GFP reporter containing a unique 20-nucleotide barcode in the 3′ untranslated region (Fig. 1A). Variants were pooled together in equal amounts based on vector genome titer. Mice were then coinjected with (1) the barcoded capsid library and (2) microbubbles (a required component for FUS-BBBO) and then immediately treated with unilateral FUS targeted to the striatum (Fig. 1A). A separate cohort of “IV-only” negative control animals received an injection of the capsid library without the FUS treatment. Four weeks later, striatal and liver tissue were harvested for NGS-based measurement of DNA and RNA barcode counts for each variant. Striatal barcode counts were universally lower in the IV-only animals compared with the FUS-treated animals, confirming that the FUS-BBBO treatment was successful (Supplementary Table S1). Additionally, RNA barcode counts for AAV2, AAV5, and AAV6 were all lower than that of AAV9 in both the liver and the FUS-treated brain (Fig. 1B–G), which was expected based on the literature and provided confidence in the robustness of this library-based screening approach. Furthermore, barcode counts in the liver were >30-fold lower for AAV9-G266A compared with parental AAV9 (Fig. 1B, E), once again confirming the liver detargeting phenotype. Remarkably, however, barcode counts for AAV9-G266A in the brain, were just 2- to 4-fold lower for AAV9-G266A compared with parental AAV9 (Fig. 1C, F), with an overall improvement in the ratio of striatum to liver barcodes of 40- to 60-fold (Fig. 1D, G). Therefore, the G266A mutation robustly decreases liver transduction and enhances brain transduction specificity with FUS-BBBO.
We next performed several in vitro studies to assess the underlying mechanism of detargeting by the G266A mutation, and to provide insight into our in vivo findings. We began by measuring both transduction and uptake of AAV9-G266A and parental AAV9 vectors using several liver or neuronal cell lines, including Huh7 cells (a human hepatocyte line), neuro-2A cells (a mouse neuroblastoma line), and human iPSC-derived neurons. In agreement with our in vivo results, the G266A mutation severely disrupted transduction in all cell types, as measured by plate-based luciferase assay (Fig. 3A). Intriguingly, however, the G266A mutation resulted in only a minor decrease in vector uptake over a 24-h period (as assessed by qPCR-based measurement of intracellular vector genome copy number (VGCN); Fig. 3B), suggesting that the mutation primarily affects some stage of the AAV9 infection pathway that occurs postcell entry. To investigate this further, we next compared liver VGCN/uptake in vivo at 4 and 35 days post IV injection. While AAV9-G266A displayed significantly reduced VGCN compared with AAV9 at day 35, no significant difference was observed at day 4 (Supplementary Fig. S1). These data indicate that initial viral uptake remains intact and confirm that the mutation affects a postentry step.
Based on these results, we hypothesized that attachment of AAV9-G266A to the cell surface, which occurs prior to cell uptake, would also be minimally affected. Cell attachment is mediated by binding of the AAV capsid to cell surface glycans—specifically to galactose in the case of AAV9. Therefore, we next tested the effect of the G266A mutation on binding to cell surface galactose. This was done by incubating either AAV9-G266A or parental AAV9 with three different CHO cell lines, each containing different cell surface N-linked glycosylation patterns: (1) Pro5 cells contain terminal sialic acid, which masks underlying galactose, (2) Lec2 cells are deficient in sialic acid, which exposes underlying galactose as the terminal glycan, and (3) Lec8 cells are entirely devoid of galactose (Fig. 3C). Surface-bound vector genomes were then quantified by qPCR. As controls, we also tested the binding of two AAV9 variants with impaired galactose binding: AAV9-N272A and AAV9-W503A. As expected, we observed a robust increase in binding of parental AAV9 to Lec2 cells relative to Pro5 and Lec8 cells and no increase for AAV9-N272A and AAV9-W503A (Fig. 3D). Interestingly, however, the G266A mutant exhibited a similar level of binding to Lec2 cells as that of parental AAV9 (Fig. 3D). These data indicate that the G266A mutation does not affect cell attachment or cell surface galactose binding and further support our conclusion that the G266A mutation affects a postentry step in the AAV infection pathway.
In searching the literature, we noted that the G266 residue falls within a region of the AAV9 capsid that interfaces with AAV receptor (AAVR; KIAA0319L),15,22 a critical cellular factor that is required for transduction by most AAV serotypes. Therefore, we hypothesized that the G266A mutation disrupts this binding interaction. To test this, we used a receptor binding assay, in which paramagnetic beads are coated with a purified receptor and then incubated with purified AAV. Receptor-bound virions are then digested with proteinase K and vector genomes quantified by qPCR (Fig. 4A). Since AAVs bind specifically to PKD domains of AAVR, we purified only these regions for this assay (Fig. 4B, Supplementary Fig. S2). As a negative control, we included a version of AAVR (AAVR-Mut) containing two point mutations in PKD2 (Fig. 4B) that, together, have been shown to disrupt AAVR-AAV2 binding. 22 As expected, with wild-type AAVR, we recovered nearly 100% of AAV9 that was input into each binding reaction (1E10 vector genomes), whereas with AAVR-Mut we recovered less than 1%, thus confirming that this assay can reliably detect changes in AAV-AAVR binding affinity (Fig. 4C). Strikingly, we also recovered nearly 100-fold fewer AAV9-G266A mutant virions with wild-type AAVR compared with that of parental AAV9, indicating that the G266A mutation does in fact disrupt binding of AAV9 to AAVR (Fig. 4C). Additionally, since our in vivo studies were conducted using mice, we repeated this bead-based AAVR binding experiment using the murine orthologue of AAVR (AU040320), which shares >95% sequence identity with KIAA0319L. Again, we specifically expressed and purified only the PKD domains, which were identified by sequence alignment of AU040320 with KIAA0319L, and found that all three AAV9 mutants exhibited clear binding defects to murine AAVR (Fig. 4D). Overall, these findings confirm that the G266A mutation disrupts binding of AAV9 to both human and murine AAVR, highlighting a potential mechanism for the observed transduction deficit.
DISCUSSION
While systemic brain gene therapy holds significant promise for the treatment of neurological disorders, liver toxicity, and immune responses remain a major safety concern. Encouragingly, our findings suggest that mutagenesis of VR1 could offer a potential solution to this obstacle. Hepatotoxicity associated with systemic AAV administration is thought to be driven by presentation of capsid peptides on major histocompatibility complex (MHC) class I molecules and subsequent cytotoxic T-cell response, which peaks around 4–8 weeks postadministration.23,24 Our biodistribution data indicate that by this time point, virtually all AAV9-G266A genomes—and by extension, capsid proteins—have been cleared from the liver. Therefore, it is likely that immune response and hepatotoxicity associated with systemic dosing of AAV9-G266A would be minimal. With a 40- to 60-fold improvement in the ratio of brain to liver transduction with FUS compared with wild-type AAV9, we propose that the G266A mutant could be a viable candidate for systemic brain gene therapy.
Additionally, our findings suggest that the G266A mutation decreases liver transduction specifically by impairing AAVR binding. Indeed, in our bead-based in vitro binding assay, the AAV9-G266A mutant exhibited decreased binding to both the human and murine orthologues of AAVR. In line with this, previous structural data indicate that the G266 residue directly interfaces with AAVR.15,22,25 Additionally, several residues within VR1 of the AAV2 capsid were shown to be required for AAVR binding and functional transduction. 22 As the precise role of AAVR in transduction is not fully understood, the implications of these findings are unclear. Importantly, however, AAVR is not required for attachment of AAVs to the cell surface, 26 which is in line with our finding that the G266A mutation does not affect cell attachment or galactose binding. In contrast, while AAVR localizes to the plasma membrane to some extent, it primarily localizes to the Golgi. 20 Furthermore, deletion of the C-terminus of AAVR not only prevents trafficking to the Golgi, but also blocks AAV2 infection, 20 which suggests that AAVR is directly involved in intracellular transport of AAV to the Golgi. Therefore, we conclude that the G266A mutation disrupts AAV9 transduction by preventing binding of the capsid to AAVR and potentially by interfering with subsequent intracellular trafficking.
Regardless of this AAVR binding defect, we also found that AAV9-G266A retained 25 − 50% of brain transduction, which suggests that AAV9 can transduce the brain in an AAVR-independent manner. Indeed, while almost all natural serotypes are AAVR-dependent, AAV4, AAV6.2, and AAVrh32.33, can transduce AAVR knockout cells, suggesting that AAVs can utilize alternative transduction pathways. 26 Additionally, expression of fusion mutant AAVR carrying the LDL and poliovirus C-terminus, which promotes trafficking through endosomal pathways instead of the Golgi,27,28 partially restores transduction when overexpressed in AAVR knockout cells, 20 further suggesting that alternative intracellular trafficking pathways are possible. Indeed, AAVs have also been reported to utilize various endocytic pathways, including clathrin-mediated endocytosis, CLIC/GEEC, and macropinocytosis.29–32 Additional studies will be required to establish whether AAVR-independent infection pathways are available in the brain, and whether they can be utilized by AAV9.
It is also worth considering the relative advantages of FUS-mediated delivery compared with BBB-penetrant AAV capsids in the context of clinical translatability. Although we did not include a direct comparison in this study, prior work has shown that AAV9 combined with FUS-BBBO can achieve transduction levels comparable to the BBB-crossing variants such as AAV-PHP.B. 33 A key advantage of the FUS approach is its ability to achieve spatially restricted delivery, which may be beneficial for regionally defined neurological disorders by limiting vector exposure in unaffected brain regions and reducing the risk for off-target effects. Additionally, as BBB-penetrant capsids often exhibit heterogeneous transduction patterns, FUS could be used in combination with such vectors to enhance the uniformity and control of central nervous system (CNS) delivery. Lastly, it is possible that IV AAV dosing with FUS might allow for a lower efficacious dose compared with dosing of a BBB-crossing capsid, thus increasing the tolerability and safety of AAV gene therapies.
Several limitations to our findings are worth discussing. First, we found that the G266A mutation had minimal effect on short-term accumulation of vector in the liver. We hypothesize that, after being taken up by the liver and internalized by hepatocytes, the G266A mutant is targeted for degradation due to its inability to bind AAVR. While this is advantageous for mitigating immune responses and liver toxicity, it does not prevent the liver from acting as a sink and means that high vector doses would still be required. Second, our in vivo testing of the AAV9-G266A mutant was limited to mice, and so we cannot be fully confident that our findings will translate to humans. However, we found that the G266A mutation decreased in vitro transduction to a similar degree in both mouse and human cell lines and led to a similar decrease in binding to both mouse and human orthologues of AAVR, which together suggest that the liver de-targeting phenotype would indeed translate. Still, additional testing of this vector in species that are evolutionarily closer to humans (e.g., NHPs) is warranted, but beyond the scope of this article.
Overall, our findings indicate that mutagenesis of VR1 and disruption of AAVR binding could be a useful liver detargeting approach. Future studies will continue to dissect the underlying biology and will further improve upon the ratio of brain to liver transduction—either through additional engineering of the VR1 region or by combining VR1 mutations with additional mutations elsewhere in the capsid—to ensure the safety and efficacy of systemic brain gene therapy approaches.
AUTHORS’ CONTRIBUTIONS
A.T.N., C.T.C., T.W., and E.A.E. designed experiments and interpreted results. A.T.N. performed molecular cloning, generated AAV vectors, and conducted all in vitro work. A.S. and E.K. performed and/or supervised all animal work. C.T.C. provided critical insight and designed a focused ultrasound library study. B.K., A.N.G., D.P., and E.F. provided experimental support. A.T.N. wrote and revised the article. All authors approved the final article.
Footnotes
ACKNOWLEDGMENTS
The authors acknowledge support from the Discovery Research and Medical Communications teams at Spark Therapeutics, Inc.
FUNDING INFORMATION
This work was funded by Spark Therapeutics, Inc.
DATA AVAILABILITY STATEMENT
The data supporting all findings in this study are available from the corresponding author upon reasonable request.
AUTHOR DISCLOSURE
This work was funded by Spark Therapeutics, Inc., a Roche company and was included on the following patent entitled CNS targeting peptides, Provisional Application No. 63/808,863. A.T.N., A.S., C.T.C., B.K., A.N.G., D.P., E.F., E.K., T.W., E.A.E., are either present or past paid employees of Spark Therapeutics, Inc., a Roche company, and may hold equity in the organization.
Supplemental Material
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
