Abstract

Corrigendum to: Obshta O, et al. Oxytetracycline-resistant Paenibacillus larvae identified in commercial beekeeping operations in Saskatchewan using pooled honey sampling. J Vet Diagn Invest 2023;35:645–654. https://journals.sagepub.com/doi/epdf/10.1177/10406387231200178
Previously, we reported 3 that 2 isolates of Paenibacillus larvae (64 and 66; see Suppl. Table 2 3 ) had a discrepancy between oxytetracycline (OTC)-resistant phenotype, with MIC values of 256 and 32 µg/mL of OTC, respectively, and susceptible genotype (absence of tet(L) gene in sequenced genomes).
To address this discrepancy, we repeated MIC determination using a broth microdilution technique as described previously for 27 clinical P. larvae isolates, including:
a) 16 OTC-resistant isolates of P. larvae, including isolates 64 and 66, representing 25% of the OTC-resistant isolates in our study;
b) 8 OTC-intermediate-resistant P. larvae isolates, representing 100% of intermediate-resistant isolates in our study;
c) 3 OTC-susceptible isolates.
As a control, we performed MIC testing on standard strains of P. larvae (ATCC 9545), Staphylococcus aureus (25923), and Pseudomonas aeruginosa (27853) per CLSI 1 guidelines for broth microdilution technique validation for P. larvae testing. MIC testing of standard strains was performed in duplicate using in-house-prepared and commercially available MIC plates (Sensititre NARMS Gram positive plate; ThermoFisher). Broth microdilution tests were set up either according to the CLSI 1 guidelines for an in-house developed test or according to the manufacturer’s instructions for commercially produced plates.
For all of the clinical P. larvae isolates for which MIC determination was repeated, we also validated presence or absence of the tet(L) gene using conventional PCR.2,4 As positive and negative validation controls, we used 3 OTC-resistant and 3 OTC-susceptible isolates, respectively.
Results
Fourteen of 16 isolates of previously OTC-resistant P. larvae isolates were confirmed to have OTC-resistant phenotype by MIC testing and had a positive tet(L) PCR test result. In contrast, 2 isolates previously reported as OTC-resistant (64 and 66), when re-tested, revealed an OTC-susceptible phenotype (Corrigendum Table 1). Moreover, PCR confirmed the absence of the tet(L) gene in both isolates 64 and 66, as reported previously in analysis of whole-genome sequences.
Results of 16 re-tested, previously reported OTC-resistant clinical Paenibacillus larvae isolates.
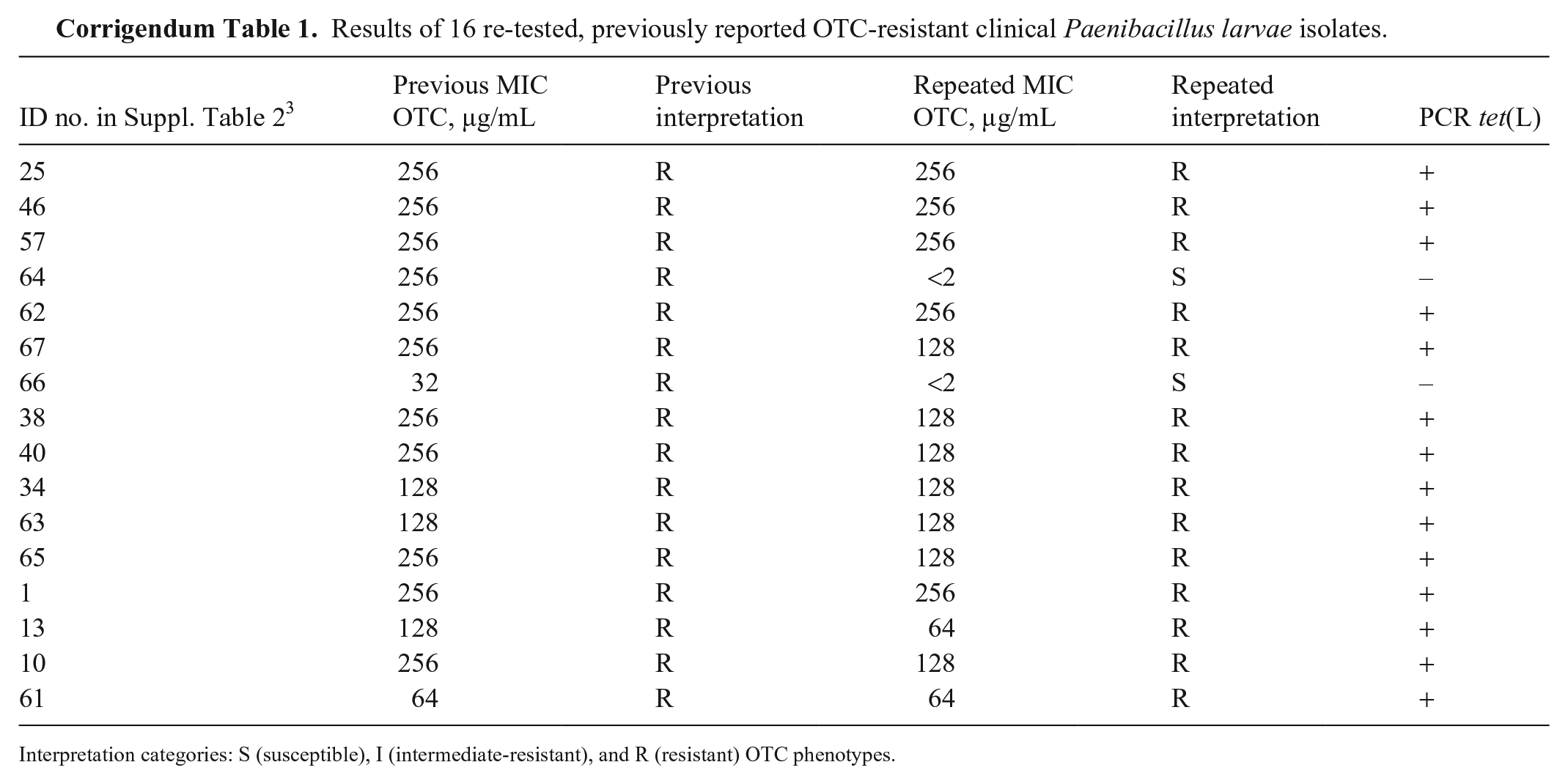
Interpretation categories: S (susceptible), I (intermediate-resistant), and R (resistant) OTC phenotypes.
For the 8 P. larvae isolates that were previously reported to have OTC-intermediate-resistant phenotype, we found 2 to be OTC-resistant, and positive for tet(L) on PCR. The remaining 6 isolates were found to be OTC-susceptible, and negative for tet(L) on PCR (Corrigendum Table 2; Corrigendum Fig. 1B–1D).
Results of 8 re-tested, previously reported OTC-intermediate-resistant clinical Paenibacillus larvae isolates.
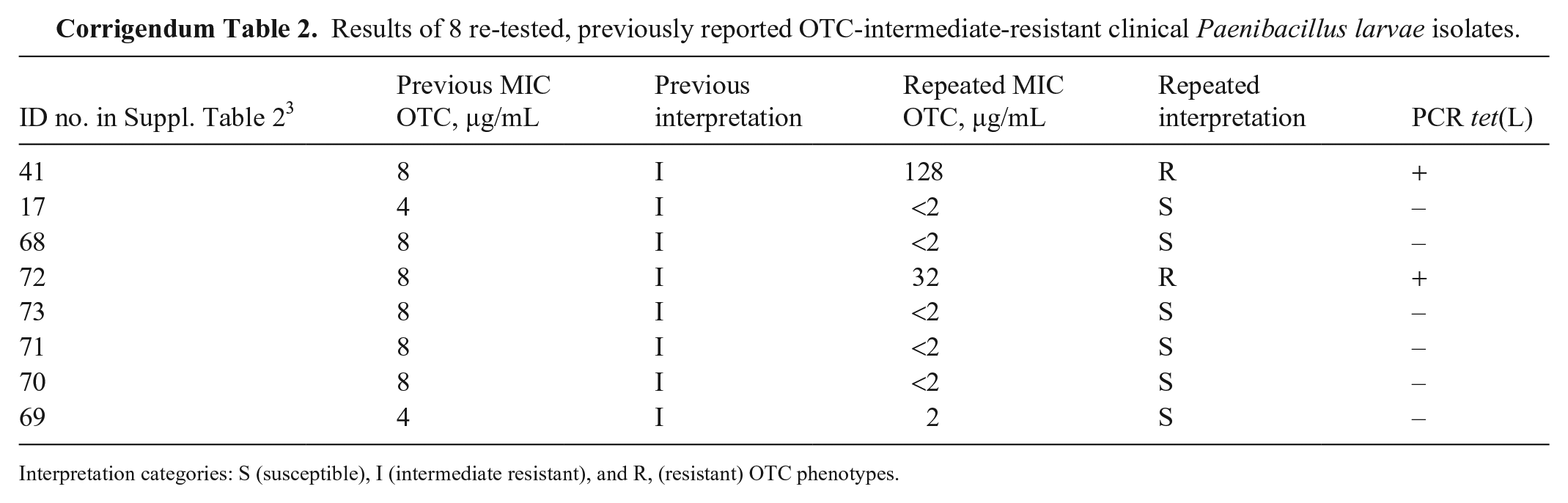
Interpretation categories: S (susceptible), I (intermediate resistant), and R, (resistant) OTC phenotypes.
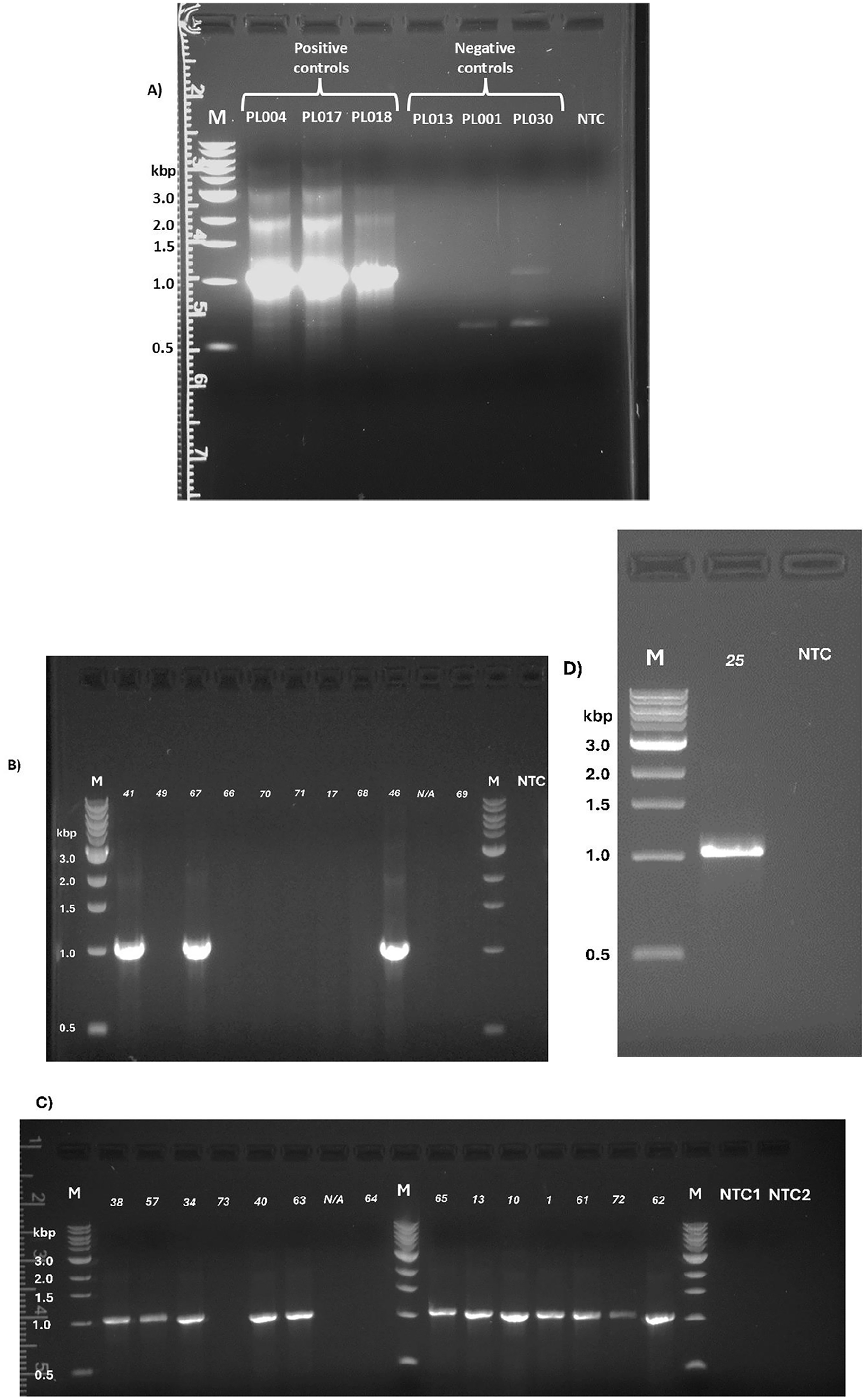
tet(L) PCR of Paenibacillus larvae isolates.
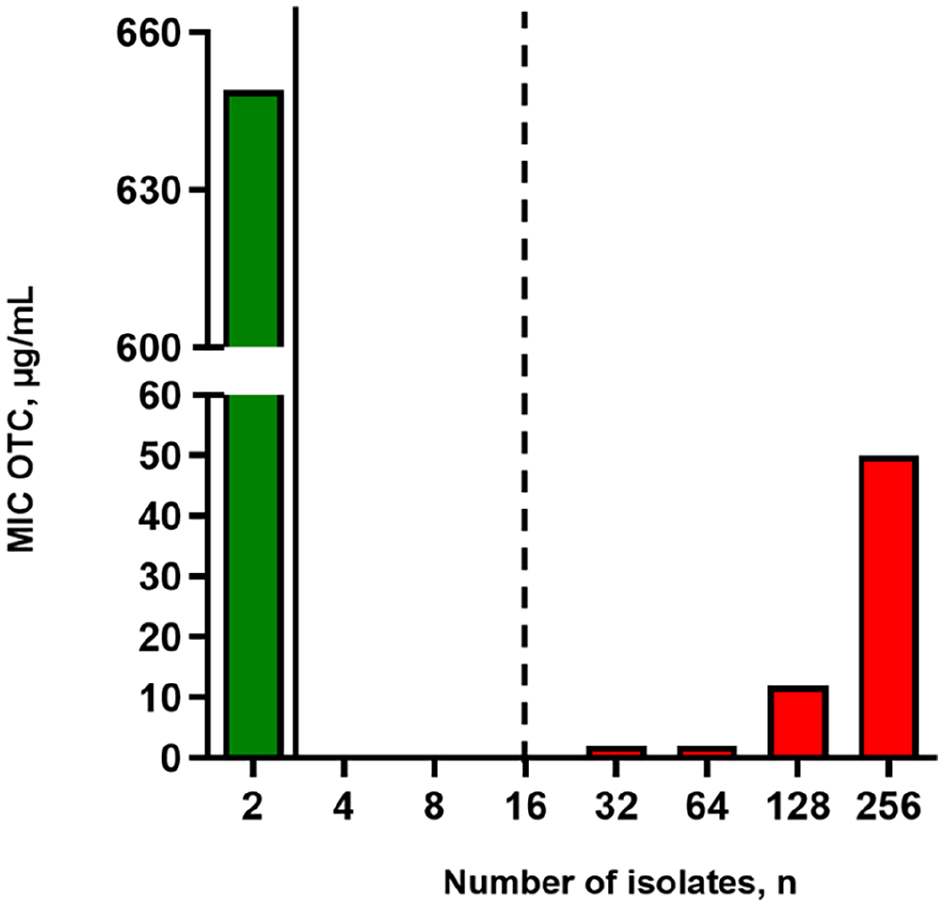
Oxytetracycline (OTC) susceptibility of Paenibacillus larvae isolates from Saskatchewan, Canada, from 52 commercial beekeeping operations in 2019 and 2020. Bars represent number of P. larvae isolates for each MIC value of OTC (µg/mL) for 718 P. larvae samples. Solid and dashed lines indicate the CLSI breakpoints for OTC sensitivity (≤2 µg/mL OTC) and resistance to OTC (≥16 µg/mL OTC), respectively.
The 3 OTC-susceptible isolates that were re-tested were confirmed to have OTC-susceptible phenotype and absence of tet(L) on PCR (Corrigendum Table 3; Corrigendum Fig. 1B–1D).
Results of 3 re-tested, previously reported OTC-resistant clinical Paenibacillus larvae isolates.
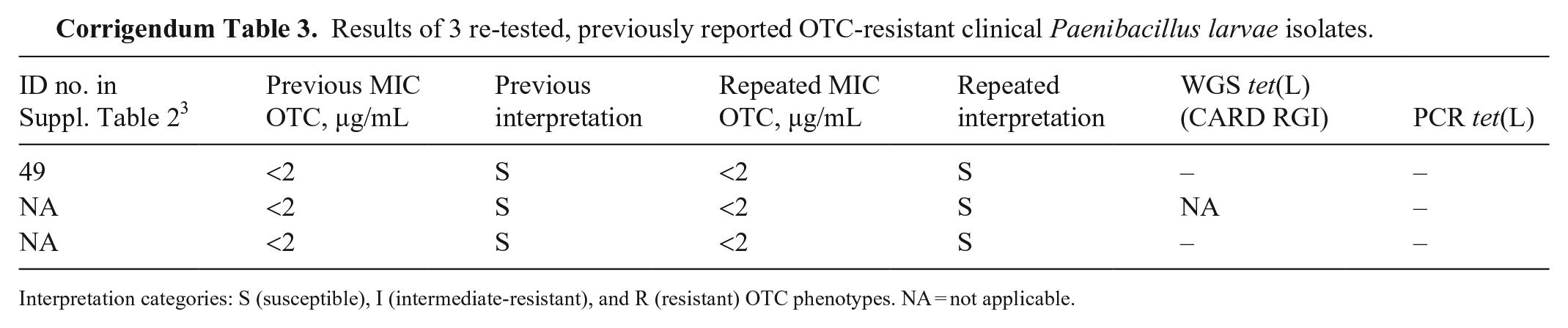

Interpretation categories: S (susceptible), I (intermediate-resistant), and R (resistant) OTC phenotypes. NA = not applicable.
Discussion
Potential explanations for the lack of repeatability in our initial MIC results include
a) Inconsistent cell density. Density of the P. larvae suspensions used for MIC determination were standardized based on optical density. In contrast, MacFarland standards are typically used for inoculum standardization in routine procedures.
b) OTC degradation. The half-life of oxytetracycline in water is only 34 h 5 ; however, MIC determination occurred over 48 h, during which time OTC may have degraded.
c) Operator pipetting errors. This can happen when assay is developed in house; however, these mistakes were ruled out by verification of the assay when tested with standard ATCC strains.
In summary, we found that 2 of 16 previously reported OTC-resistant P. larvae isolates, and 6 of 8 previously reported OTC-intermediate resistant isolates were susceptible upon re-testing and did not carry a tet(L) gene. Additionally, 2 of 8 previously reported OTC-intermediate resistant isolates were found to be OTC-resistant upon re-testing. The remainder of the re-tested P. larvae isolates were confirmed to have the same OTC phenotype and concordant tet(L) PCR results as reported previously in our article (Updated Fig. 2).
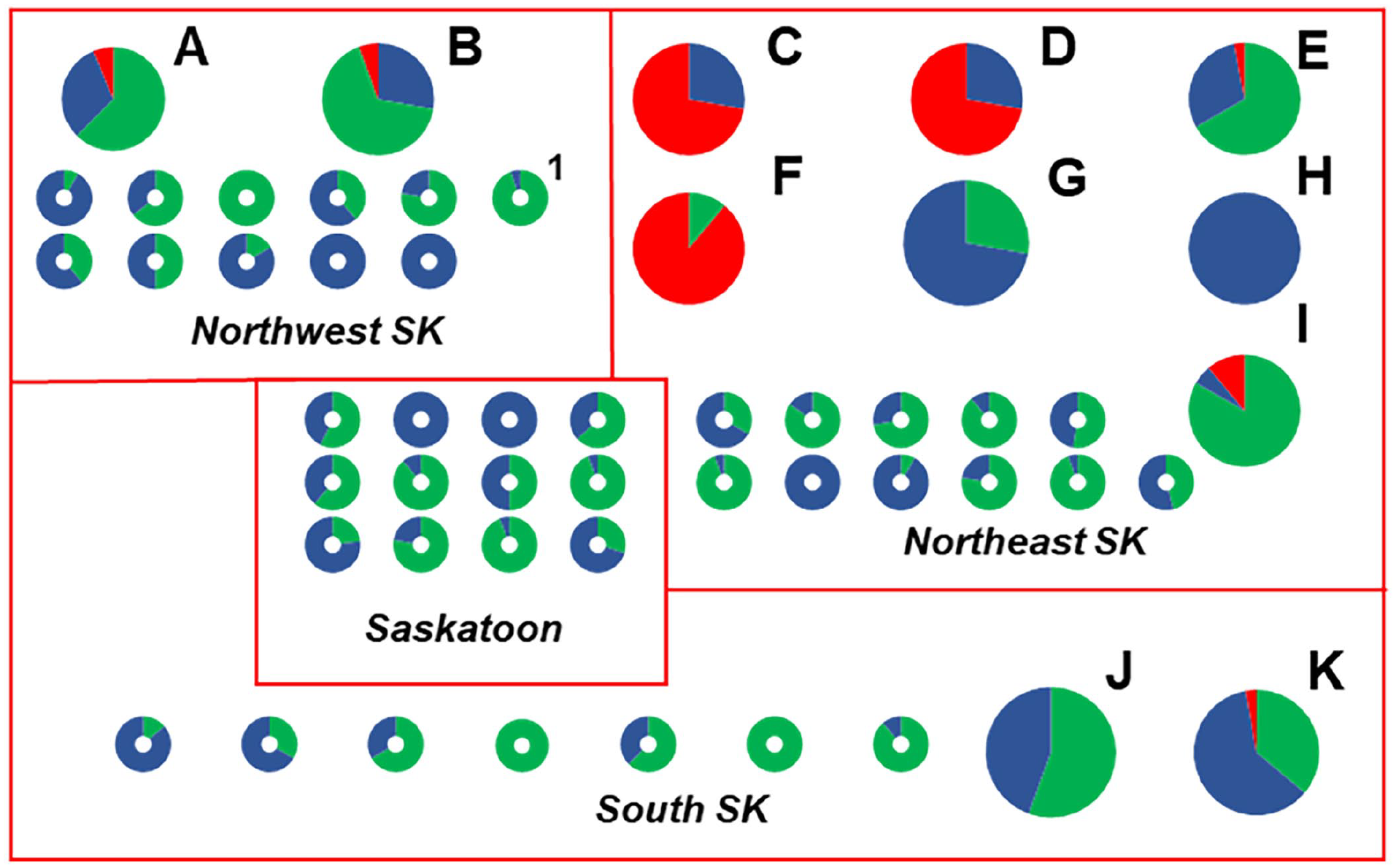
Spatial distribution and oxytetracycline (OTC) susceptibility of Paenibacillus larvae isolates from 1 of 52 Saskatchewan, Canada commercial beekeeping operations from 2019 and 2020. Each pie chart or donut chart represents the OTC susceptibility of P. larvae isolates cultured from the extracted honey in 1 of 4 (northeastern, northwestern, Saskatoon area, southern) regions of Saskatchewan. Lettered pie charts represent beekeeping operations whose extracted honey samples contained resistant (red) isolates of P. larvae. Donut-shaped charts represent commercial beekeepers whose P. larvae isolates were all susceptible to OTC (blue) or whose honey samples did not contain any detectable P. larvae spores (green). Donut chart “1” represents a beekeeper from whom an additional OTC-susceptible P. larvae isolate was characterized by whole-genome sequencing. The size of pie charts was increased relative to donut charts to enhance readability and does not reflect differences in sample size.
