Abstract
Mycobacteriosis is an emerging and often lethal disease of aquatic organisms caused by several non-tuberculous mycobacteria (NTM) species. Early diagnosis of mycobacteriosis in aquaculture and aquatic settings is critical; however, clinical diagnoses and laboratory detection are challenging, and the available literature is scarce. In an attempt to fill the gap, here we review the most relevant approaches to detect and characterize mycobacteria in clinical specimens of aquatic organisms. Emphasis is given to recent advances in molecular methods used to differentiate NTM species spanning from targeted gene sequencing to next-generation sequencing. Further, given that there are major gaps in our understanding of the prevalence of the different NTM species, partially because of their distinct requirements for in vitro growth, we also reviewed the most relevant NTM species reported to cause disease in aquatic organisms and their specific in vitro growth conditions. We also highlight that traditional bacterial culture continues to be relevant for NTM identification, particularly in non-automated laboratories. However, for NTM species discrimination, a high level of accuracy can be achieved with MALDI-TOF MS and molecular approaches, especially targeted gene sequencing applied from clinical specimens or from pure NTM isolates.
Aquatic mycobacteriosis is a chronic, progressive, often lethal disease, affecting mostly aquaculture and aquarium settings. The first case of disease caused by mycobacteria in fish was reported in 1882 in a carp (Cyprinus carpio) 86 ; since then, mycobacteriosis has been reported to contribute to the decline of fish populations worldwide. Aquatic mycobacteriosis has received more attention in recent years as a result of the discovery of new species of mycobacteria in piscine hosts, epizootics in wild fish, and the zoonotic potential of some mycobacterial species. Mycobacteriosis is generally chronic and progressive, although on some occasions it can be acute or even clinically silent.
Clinical signs include weight loss, lethargy, epidermal discoloration, dermal lesions, pigmentary changes, stunted growth, exophthalmia, skin ulceration, and slow swimming, which are largely nonspecific signs that vary in frequency and severity in different populations. 16 Gross findings tend to be more specific, consisting of discrete white nodules distributed throughout various tissues, but most apparent in the spleen, liver, and kidney. 19 Given that diagnosis is often based on clinical signs and that several mycobacterial species can cause mycobacteriosis, laboratory confirmation enables rapid implementation of loss-minimization measures as well as disease surveillance.
The few articles published on mycobacteriosis, especially in piscine hosts, have focused on overall disease presentation, pathology findings, diagnosis, and treatment; however, in most of these studies the scope of laboratory methods was limited.15,16,24,30 Therefore, we prepared this review to provide comprehensive and updated information on relevant laboratory techniques applied to diagnose mycobacteriosis in aquatic organisms, including traditional microbiologic methods, in vitro susceptibility testing, and proteomic and molecular approaches. We intend to provide central aspects with the hope of helping laboratory diagnosticians use the most appropriate approaches for fast and reliable diagnosis of mycobacteriosis in aquatic organisms.
Etiologic agents
Mycobacteria (Actinomycetales, Mycobacteriaceae) are aerobic, non–spore-forming, non-motile, rod-shaped, acid-fast, and gram-positive bacilli. The acid-fast feature is attributed to the rich cell wall composed of mycolic acids and complex lipids, giving them the capacity to retain carbol–fuchsin stain when decolorized with acid. To date, there are 188 species of mycobacteria with valid names. Recent studies based on 16S rRNA, 16S-23S spacer regions, and multilocus and whole-genome sequencing (WGS) led to the division of the genus Mycobacterium into 5 genera: Mycobacterium, Mycobacteroides gen. nov., Mycolicibacillus gen. nov., Mycolicibacter gen. nov., and Mycolicibacterium gen. nov. 27
Mycobacteria are classified as tuberculous (M. tuberculosis complex, M. leprae, M. lepromatosis) and non-tuberculous mycobacteria (NTM), which are mycobacteria that do not cause tuberculosis or leprosy. 27 NTM are also commonly referred to as atypical mycobacteria, mycobacteria other than tuberculosis, or environmental mycobacteria, and are often classified by growth rate, either rapidly growing (2–3 d) or slow growing (> 7 d). Three primary mycobacteria have been linked to disease in aquatic organisms: M. chelonae, M. fortuitum, and M. marinum 34 (Table 1). Of these, M. marinum is considered the most important species in aquatic hosts and is an important pathogen in humans. M. marinum is a pathogenic mycobacterium, which sets it apart from the other NTM, which are usually opportunistic pathogens. 2 M. ulcerans is also an important human pathogen closely related to M. marinum. DNA of M. ulcerans has been detected in snails, fish, and amphibians 95 ; however, its contribution to mycobacteriosis in aquatic organisms is uncertain, therefore we did not include M. ulcerans in Table 1. In research facilities, outbreaks of NTM in zebrafish colonies are commonly associated with M. abscessus, Mycobacteroides chelonae, Mycobacterium fortuitum, M. haemophilum, M. marinum, and M. peregrinum. 93
Non-tuberculous mycobacteria species associated with mycobacteriosis in aquatic organisms, described hosts, and growth condition in agar culture.
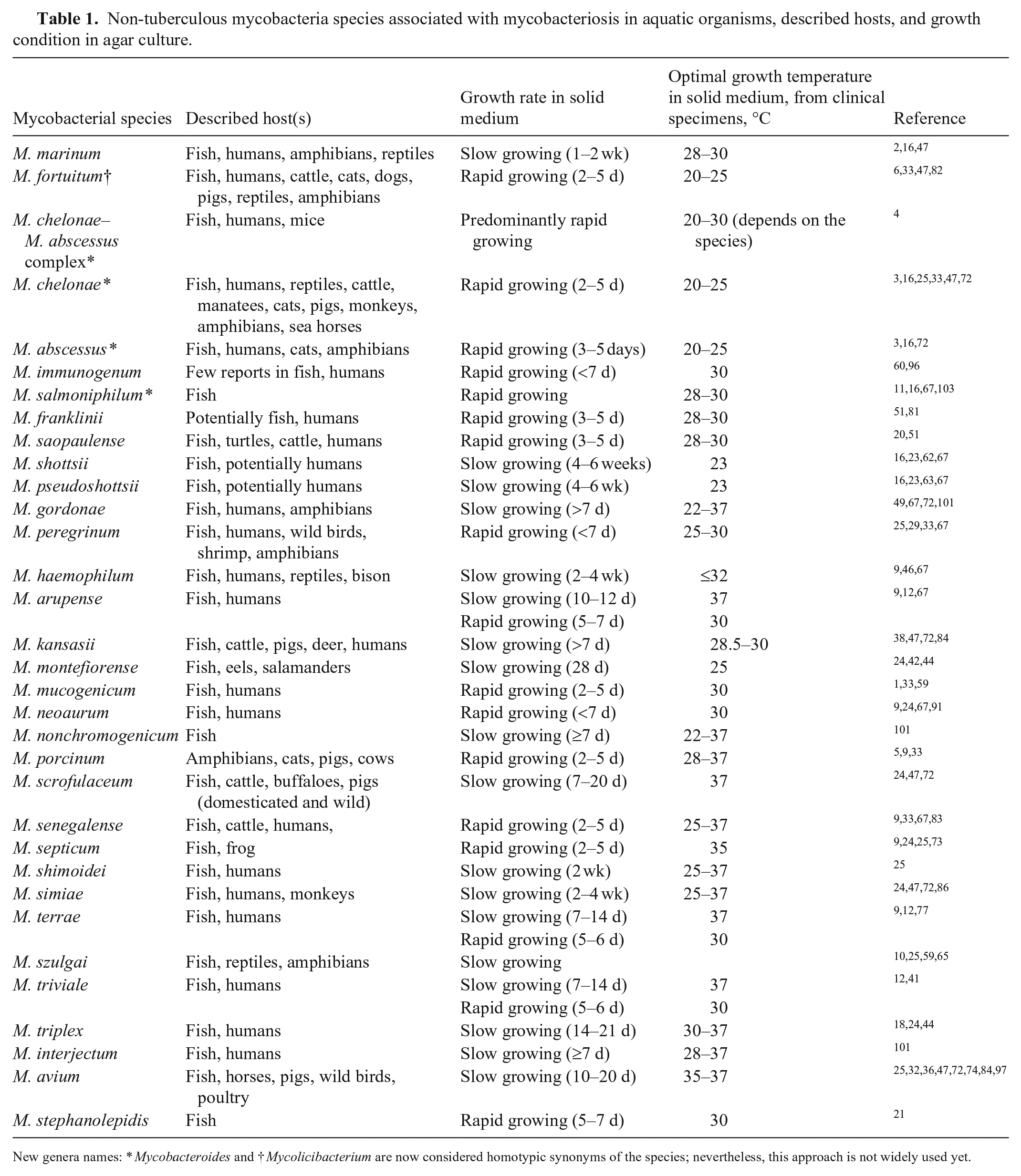
New genera names: * Mycobacteroides and † Mycolicibacterium are now considered homotypic synonyms of the species; nevertheless, this approach is not widely used yet.
Several other species have been associated with mycobacteriosis in aquatic organisms, and, with the advancement of molecular techniques, the list is still growing (Table 1). Additional species identified in diseased aquatic organisms include M. pseudoshottsii and M. shottsii,62,63 M. gordonae, 58 M. haemophilum, 93 M. peregrinum 58 (Table 1), and less frequently M. chesapaeki, M. M. interjectum, kansasii, M. montefiorense, M. phlei, M. scrofulaceum, M. terrae, and M. triplex.31,61,76 In 2017, a new species was reported from the thread-sail filefish (Stephanolepis cirrhifer) and named Mycobacterium stephanolepidis (Table 1). 21
Traditional staining and bacteriologic methods
Specimens
Several different specimens can be collected for laboratory testing of NTM, reflecting the variety of clinical signs and the wide distribution of lesions in different tissues. Lesions may be granulomas found in the liver, kidney, spleen, and occasionally heart. From live animals, the preferred specimens include skin scrapings and biopsies of gills, skin, and fins. Postmortem specimens for culture and molecular methods include fresh samples of lesions, and fixed tissues in 10% neutral-buffered formalin for histopathology. Transport media and preservatives are not usually recommended. Refrigeration of samples at 4°C is preferred if transportation to the laboratory is delayed > 1 h.
Preliminary procedures are necessary to decontaminate the rapidly growing microbiota of clinical specimens from non-sterile sites such as external fish surfaces. Decontamination of postmortem specimens may not be necessary if samples are collected aseptically. Details of decontamination procedures for fish specimens have been described.17,75 Decontamination of specimens can also be performed with commercial systems (e.g., BBL MycoPrep kit; Becton Dickinson). 78 The recommendation of the U.S. Center for Disease Control and Prevention is to use Biosafety Level 2 (BSL2) measures for the manipulation of infected tissues and isolates of NTM (https://www.cdc.gov/labs/pdf/CDC-BiosafetyMicrobiologicalBiomedicalLaboratories-2020-P.pdf).
Direct microscopy from specimens
The Ziehl–Neelsen stain is commonly used to identify acid-fast mycobacteria from isolated colonies, smears, tissues, and other specimens. The Kinyoun stain (cold method) is a variation of the Ziehl–Neelsen method that does not require heating and is therefore more practical and suitable for field conditions. Unlike the Ziehl–Neelsen stain, the Kinyoun method can also stain other acid-fast organisms such as Nocardia spp. and Cryptosporidium spp.87,88 Mycobacteria stained with Ziehl–Neelsen or the modified Kinyoun stains appear as red rods against a blue background. Mycobacteria are often beaded, small, with individual species of mycobacteria differing in size (0.2–0.6 × 1–10 µm), and often unevenly stained, with areas of stain deposition. In specimens with small numbers of bacteria, mycobacteria are seen more easily using fluorescent stains such as auramine, acridine orange, or fluorochrome. Acid-fast staining in tissue sections is helpful because it highlights the presence of bacilli within lesions (Fig. 1). However, mycobacteria cannot always be detected in tissue sections, depending on species, abundance, and/or the growth phase of the pathogen. Therefore, heavy reliance on Ziehl–Neelsen stains in routine histologic samples may result in underdiagnosis of fish mycobacterioses.102,103 In H&E-stained tissue sections, mature discrete granulomas are characterized by central areas of caseous necrosis containing variable numbers of acid-fast bacteria admixed with cellular debris and surrounded by epithelioid macrophages with scattered peripheral lymphocytes (Fig. 1). 19
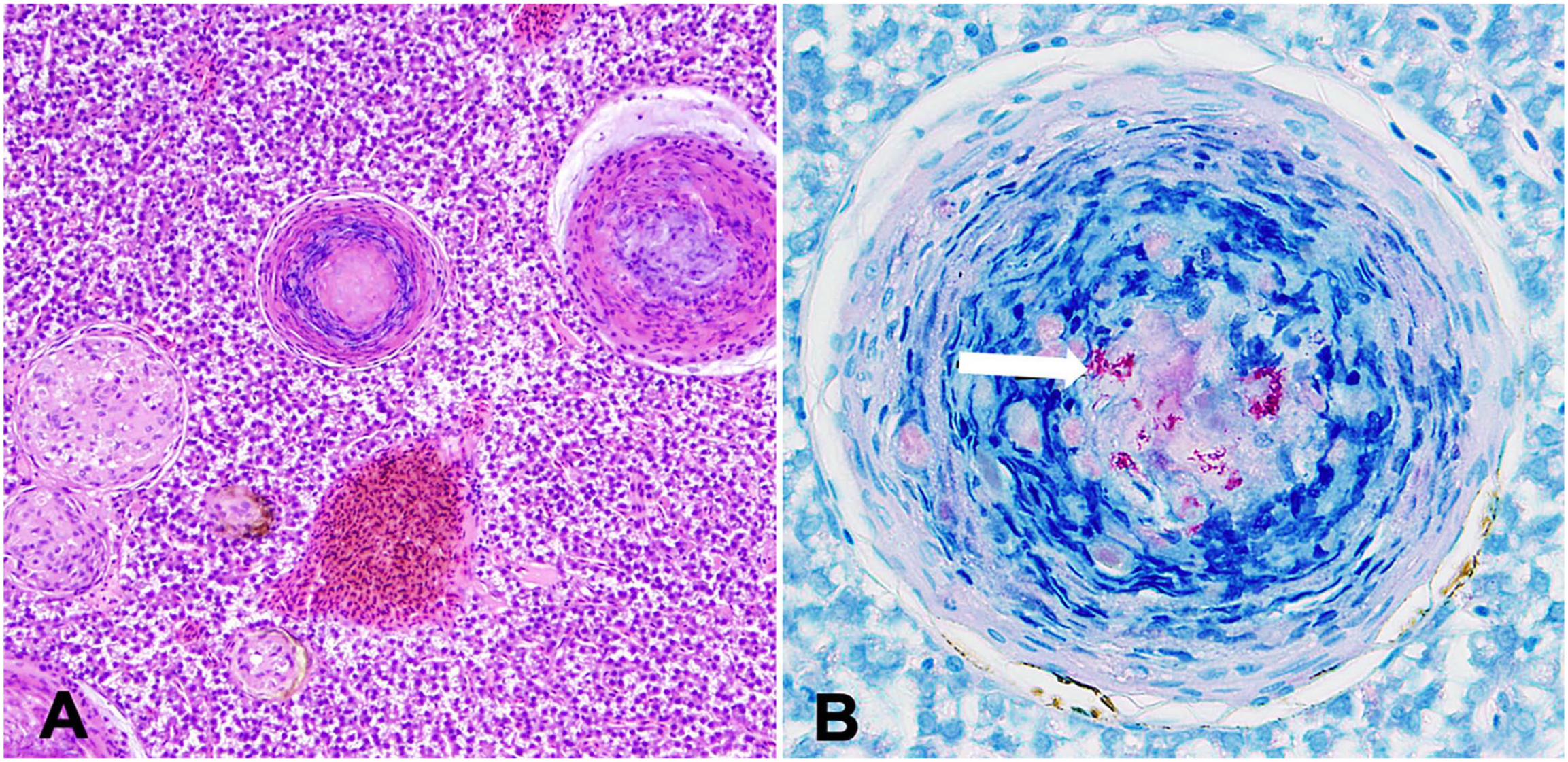
Histologic sections of liver from a tiger barb (Puntigrus tetrazona) infected with mycobacteria.
Isolation of mycobacteria on solid media
Culture of NTM continues to be an important tool in the diagnosis of mycobacteriosis. Culture is also critical for determining susceptibility to anti-mycobacterial agents. Lowenstein–Jensen (L-J) medium, which is commonly used to isolate NTM from clinical specimens of fish and other aquatic organisms, is a glycated, egg–potato medium containing malachite green dye as a selective agent. The incorporation of lincomycin, nalidixic acid, penicillin G, and cycloheximide increases the selectivity of the L-J medium by inhibiting contaminating organisms. Middlebrook 7H11, which is another commonly used selective agar for isolation and culture of NTM from specimens containing mixed flora, is also the recommended medium for isolation of colonies for antimicrobial susceptibility testing.98,99 Incubation of Middlebrook 7H11 agar in a 5–10% CO2 atmosphere may enhance mycobacterial growth. Both L-J and Middlebrook 7H11 media are photosensitive given the presence of malachite green dye, and both media should be incubated for one week with loosened caps to allow for the circulation of CO2 at the beginning of bacterial growth.
The isolation of NTM from clinical specimens can be laborious because of the potential overgrowth of coexisting non–acid-fast bacteria. RGM medium—a new selective medium for rapidly growing mycobacteria—was designed to enhance their recovery from human respiratory specimens. 57 RGM medium is composed of Middlebrook 7H9–based agar supplemented with oleic acid–albumin–dextrose–catalase and 4 antimicrobial agents, which inhibits growth of non-mycobacterial organisms and, therefore, does not require prior decontamination of clinical specimens. 66
The high incubation temperatures frequently used for mammalian clinical isolates can inhibit the growth of fish mycobacteria, which often have an optimal growth temperature of 23–30°C (Table 1). 39 Several NTM of fish importance are slow growers, requiring > 7 d incubation; hence, agar plates must be maintained in incubation for 8 wk before being considered negative. In contrast, colonies of rapid-growing NTM appear on agar plates within 2–5 d of incubation (Table 1).
Colony morphology and phenotypic characterization
Most non-tuberculous mycobacterial species have similar colony morphology on solid media (Fig. 2A); therefore, identification must be made using other criteria, such as growth rate, colony pigmentation, acid-fastness, Gram stain morphology of cells, and biochemical reactions (Table 2). As for clinical specimens, microscopic identification of mycobacterial cells from isolated colonies can be performed using Ziehl–Neelsen and Kinyoun stains, appearing as red-staining rods, often beaded, pleomorphic, small, and unevenly stained (Fig. 3). Gram staining might be useful for morphologic characterization; mycobacteria appear as slender, long, gram-positive rods that are frequently stained unevenly (Fig. 3).
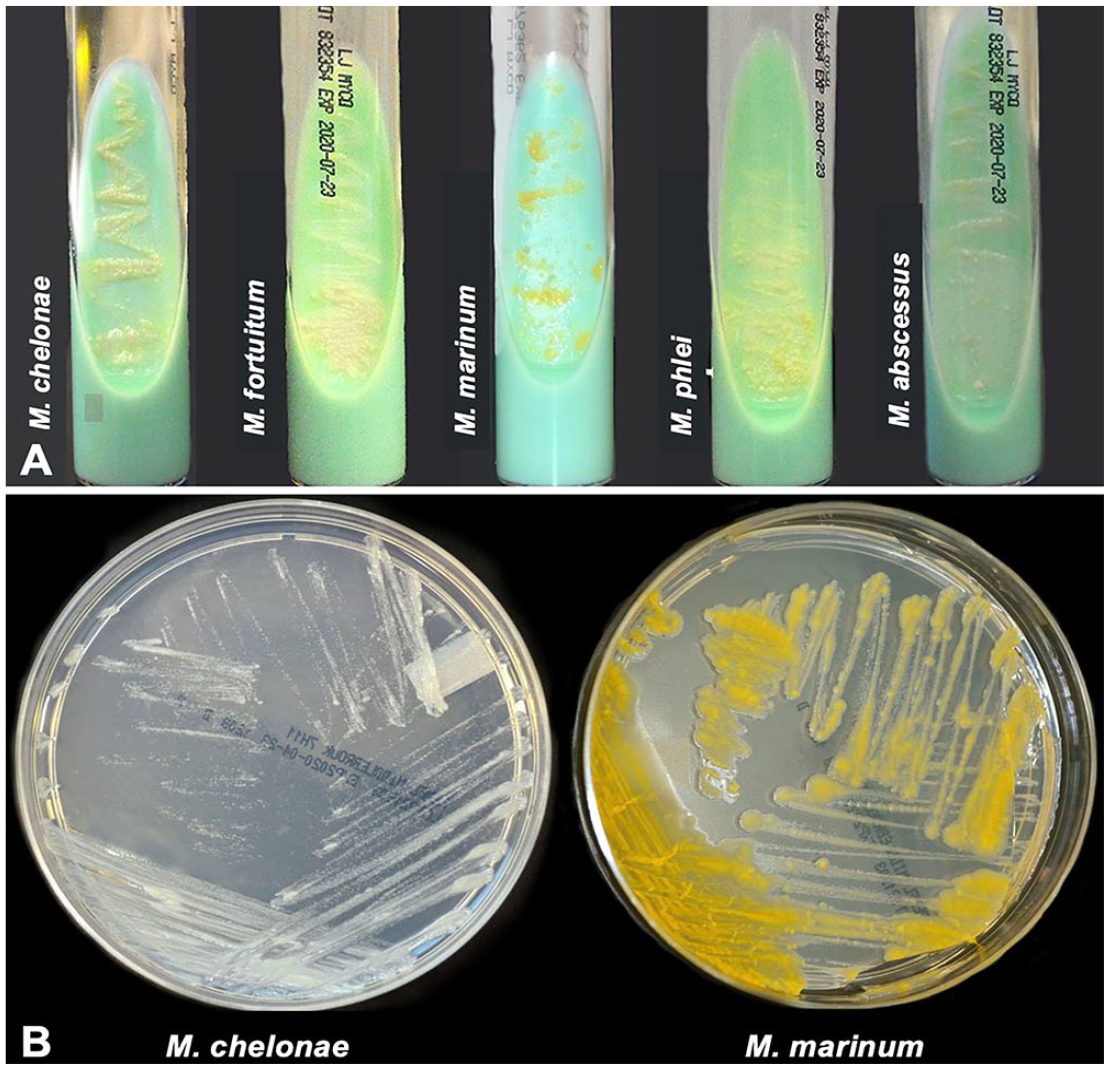
Cultures of non-tuberculous mycobacteria isolated from aquatic organisms.
Phenotypic properties and differentiation characteristics of Mycobacterium marinum, Mycolicibacterium fortuitum, and Mycobacteroides chelonae.
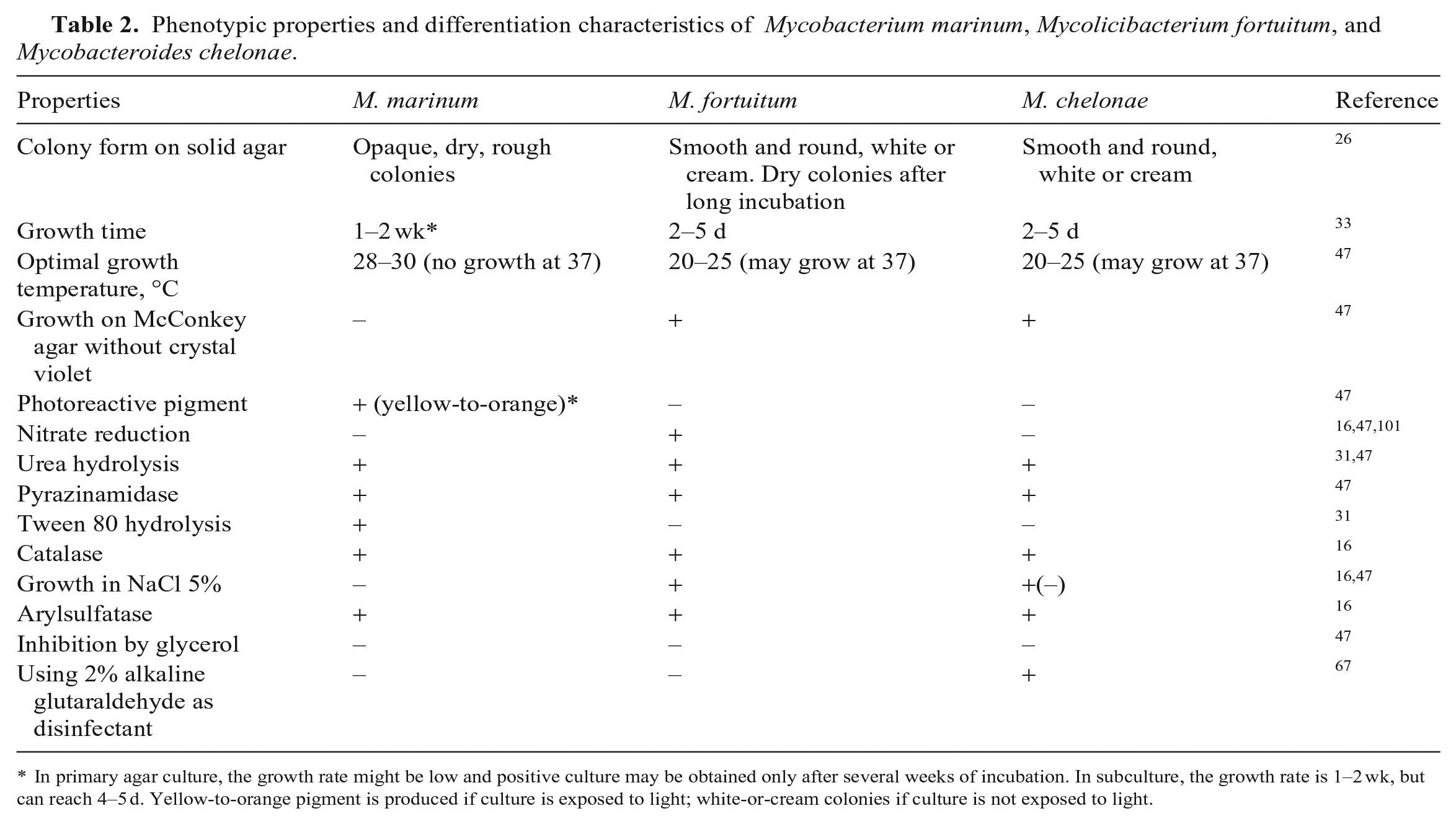
In primary agar culture, the growth rate might be low and positive culture may be obtained only after several weeks of incubation. In subculture, the growth rate is 1–2 wk, but can reach 4–5 d. Yellow-to-orange pigment is produced if culture is exposed to light; white-or-cream colonies if culture is not exposed to light.
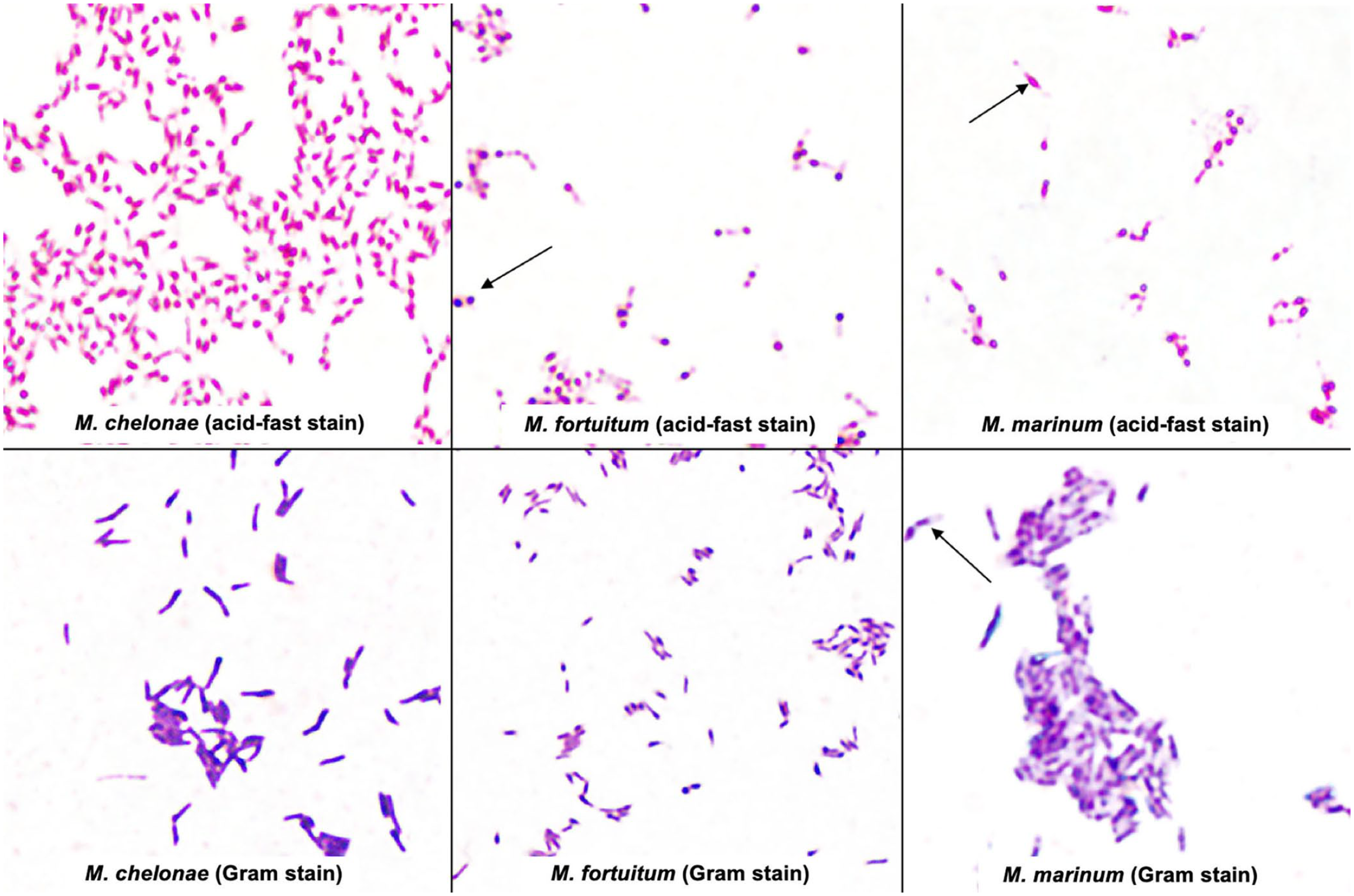

Mycobacteroides chelonae, Mycolicibacterium fortuitum, and Mycobacterium marinum in acid-fast stain (Kinyoun stain, modified cold method) and Gram stain from pure colonies isolated on Middlebrook 7H11 medium. The red-staining rods often appear beaded, pleomorphic, and small on acid-fast stain (arrows). Gram stain may allow morphologic characterization. The mycobacterial cells often appear unevenly stained with areas of stain deposition (arrows).
Pigment production and photoreactivity are important features used to identify and classify mycobacteria. The NTM have been grouped as photochromogens, non-chromogens, and scotochromogens, 68 which is still a useful method of colony identification even though not all recently identified mycobacteria fit into these groups.
Photochromogens, when cultured in the presence of light, produce yellow-to-orange carotenoid pigment, and when cultured in the dark produce non-pigmented colonies (e.g., M. marinum, M. kansasii; Table 2, Fig. 2B).
Non-chromogens produce white-cream colonies devoid of carotenoid pigments (e.g., M. fortuitum, M. chelonae; Table 2, Fig. 2B).
Scotochromogens produce yellow-orange pigment regardless of whether they grow in the dark or the light (e.g., M. gordonae).
Molecular, proteomic, and immunologic methods for NTM detection and characterization
Accurate identification of NTM species is important because treatment, control, and further prevention measurements may differ according to the etiologic NTM organism. 69 Phenotypic characteristics and biochemical tests have been the traditional method of identification of NTM isolates at the genus and species level; however, these tests are complex and require long incubation times. Gene sequencing has become the reference method in the clinical microbiology laboratory for identifying NTM species and subspecies. Gene sequencing has led to the discovery of > 150 species. 27 Matrix-assisted laser desorption/ionization–time-of-flight mass spectrometry (MALDI-TOF MS) and WGS have marked a new era of NTM identification. Here, we briefly describe the methods used most commonly to identify NTM from samples of aquatic organisms at the genus and species level (Table 3).
Molecular, proteomic, and immunologic methods to identify and characterize non-tuberculous mycobacteria (NTM) species in aquatic organisms.
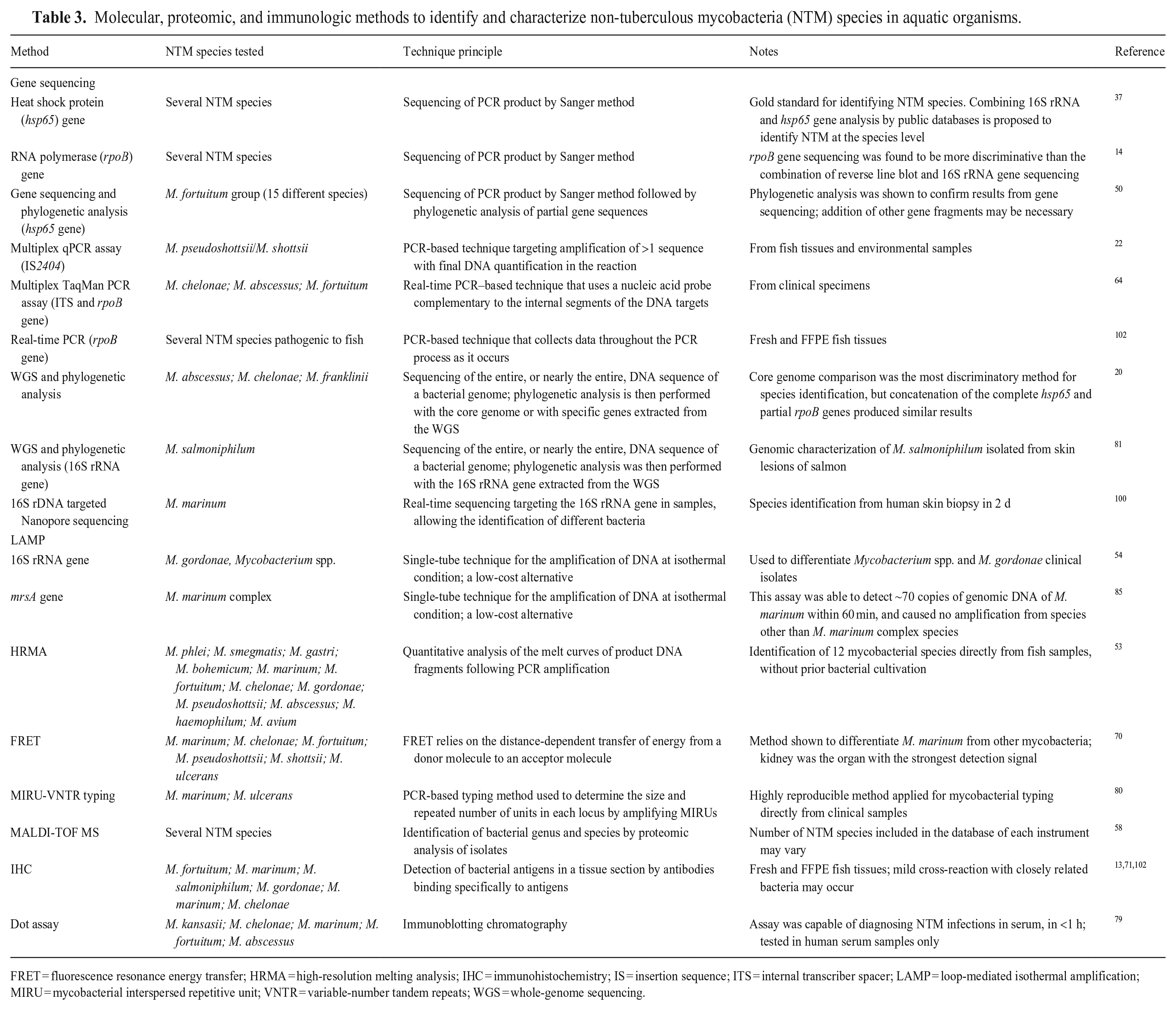
FRET = fluorescence resonance energy transfer; HRMA = high-resolution melting analysis; IHC = immunohistochemistry; IS = insertion sequence; ITS = internal transcriber spacer; LAMP = loop-mediated isothermal amplification; MIRU = mycobacterial interspersed repetitive unit; VNTR = variable-number tandem repeats; WGS = whole-genome sequencing.
MALDI-TOF MS
During the last decade, many clinical laboratories have introduced MALDI-TOF MS to the workflow of mycobacterial identification. MALDI-TOF MS allows the identification of microorganisms at the species level by analysis of total protein.43,45 Two platforms are used commonly in veterinary laboratories, MALDI-TOF Biotyper (Bruker Daltonics) and Vitek MS (bioMérieux). The success of MALDI-TOF MS depends on the quality of harbored spectra; achieving high-quality harbored spectra is challenging because of the complexity of the mycobacterial cell wall, therefore, inconsistent results are common, and may indicate that optimal performance for routine testing has not been reached yet. 45 The MALDI-TOF MS platform was successfully used to differentiate mycobacterial species in aquarium fish including M. marinum, M. peregrinum, and M. abscessus. 58 A combination of MALDI-TOF MS and molecular approaches is recommended to reach a high level of accuracy.
Immunologic techniques
Immunohistochemistry (IHC) is also used commonly to identify NTM species in formalin-fixed, paraffin-embedded (FFPE) tissue sections. 16 In cases of small granulomas, IHC is considered more sensitive than Ziehl–Neelsen staining because it can detect mycobacteria within 1–2 wk of infection. 71 Further, IHC is advantageous in localizing the in situ detection of mycobacteria to lesions. However, in nonspecific inflammatory infiltrates with more active lesions, histologic evidence of M. marinum infection may not be clear. 13 A rapid immunochromatographic serologic assay (dot assay), based on the principle of immunoblotting chromatography, identified NTM species infecting human patients in less than 1 h, including species associated with fish mycobacteriosis such as M. kansasii, M. chelonae, M. marinum, M. fortuitum, and M. abscesus. 79
Molecular methods
Molecular methods have been applied widely to identify cultured isolates or to detect mycobacteria in clinical samples (Table 3). To date, gene sequencing is considered the gold standard for identifying NTM species; however, the lack of complete sequences in databases can result in incorrect alignment and misidentification. Another disadvantage is that many closely related NTM species are undistinguishable using gene sequencing. To date, the heat shock protein gene (hsp65) and the RNA polymerase gene (rpoB) are considered the gold standard gene-sequencing targets.14,37 Other genes used to differentiate closely related species include DNA gyrase subunits A (gyrA) and B (gyrB), and the 16S-23S internal transcribed spacer region (ITS). Sequencing of the 16S rRNA gene is reported to be unsuitable for discrimination of the M. chelonae–M. abscessus complex, and of M. marinum versus M. ulcerans because they show little variability in the 16S rRNA locus.20,33
Gene sequencing has been applied to identify NTM species from isolates or directly from tissues. M. triplex–like mycobacteria were identified by sequencing directly from fish tissues when attempts at agar culture were unsuccessful or unreliable.55,93 Similarly, M. haemophilum was identified as the cause of an outbreak in a zebrafish research facility by DNA sequencing directly from affected tissues. 93 One of the limitations of identifying NTM by direct sequencing from tissue, without culture, is the inability to conduct phenotypic characterization.
In addition to gene sequencing, species-specific PCR assays have been developed for the identification of NTM species. Several of these assays target pathogenic mycobacteria of fish, such as a probe-based real-time PCR assay to detect M. pseudoshottsii and M. shottsii from specimens and environmental samples. 22 Detection and differentiation of M. abscessus, M. chelonae, and M. fortuitum groups can be performed in clinical samples using a recently designed multiplex TaqMan PCR assay (Table 3). 64
Another molecular technique used to differentiate NTM is the loop-mediated isothermal amplification (LAMP) assay, which uses 4–6 primers to recognize 6–8 distinct regions of target DNA resulting in a highly specific amplification reaction. A LAMP assay was shown to differentiate Mycobacterium spp. and M. gordonae isolated from diseased guppies (Poecilia reticulata). 54 Similarly, a LAMP assay was developed for the M. marinum complex based on the detection of the mrsA gene. 85 This assay was able to detect ~70 copies of genomic DNA of M. marinum within 60 min, and caused no amplification from species other than the M. marinum complex species (Table 3).
High-resolution melting analysis (HRMA) is another rapid molecular technique applied to detect 12 different NTM species commonly associated with disease in fish. HRMA is the quantitative analysis of the melt curves of product DNA fragments following PCR amplification, and it is based on the difference in melting behaviors of DNA molecules, according to their sequence, lengths, and GC content. 53 Furthermore, fluorescence resonance energy transfer (FRET) is a highly sensitive technique that has been used to differentiate M. marinum from other mycobacteria even before the appearance of clinical signs in fish. 70 FRET is a distance-dependent physical process by which energy is transferred nonradiatively from an excited molecular fluorophore (the donor) to another fluorophore (the acceptor) by means of intermolecular long-range dipole–dipole coupling (Table 3).
Culture-independent detection of M. tuberculosis complex in human clinical samples can be performed by shotgun metagenomics and WGS workflows. 7 WGS represents an all-encompassing method for identification of closely related mycobacterial species. The comparison of core genomes is one of the most sensitive methods for NTM species identification. In addition, concatenation of the complete hsp65 and partial rpoB genes extracted from WGS produces similar results to core genome analysis. 19 However, NGS workflows is still not performed widely in veterinary diagnostic facilities mostly because of costs.
Strain typing can be performed using the MIRU-VNTR. The acronym MIRU is used to denote a Mycobacterium tuberculosis–specific multiple locus VNTR (variable number of tandem repeats) analysis (MLVA) scheme. MIRU-VNTR is a PCR-based typing method that has been used to determine the size and repeated number of units in each locus by amplifying mycobacterial interspersed repetitive units. 35 This technique is highly reproducible and has been used in mycobacterial typing directly from clinical samples. MIRU-VNTR allows the differentiation of species from the M. marinum–M. ulcerans complex. However, this technique has been reported to not allow the association of M. marinum strain typing and its geographic regions (Table 3). 80
Molecular detection of NTM from FFPE tissues
Detection of mycobacterial DNA from FFPE specimens suspected of being infected with NTM is a routine clinical procedure. FFPE tissues are often the only fish clinical samples available 92 . Molecular approaches have been proved more sensitive, specific, and rapid than traditional histology 56 ; results from a PCR method, which amplifies a fragment of the mycobacterial hsp65 gene, were compared with those obtained by histology and culture using 45 FFPE fish tissue samples. Besides improved sensitivity, the PCR method detected mycobacterial DNA in archival FFPE blocks dating as far back as 1996. 56 Further, quantitative real-time (qPCR) assays based on the hsp65 gene have been developed to detect common NTM of zebrafish research colonies such as M. chelonae, M. haemophilum, and M. marinum, which were moderately detected in paraffin cores. 48 Another study demonstrated that slow- and fast-growing NTM could be detected from both fresh and FFPE fish tissues using a real-time PCR assay based on the rpoB gene. Such an assay provided rapid and specific mycobacterial detection in the major type of fish samples when applied along with IHC. 102
In vitro susceptibility testing for NTM isolates from aquatic organisms
Antimicrobial treatment of NTM in aquatic organisms is not often attempted given that it appears unlikely to eliminate NTM from affected animals. To date, there are no treatments approved by the U.S. Food and Drug Administration for mycobacteriosis in cultured food fish. However, there are a few reports documenting the treatment of valuable ornamental, zoo, or research fish. Treatment, when attempted, is generally administered via injections, medicated feed, or medicated water. The antimicrobial drugs that have been used to treat NTM in humans include clarithromycin, doxycycline, ethambutol, certain fluoroquinolones, minocycline, rifabutin, rifampin, and trimethoprim–sulfamethoxazole. 52 Some of these agents have been used in aquatic organisms with variable success. Careful consideration must be taken if these antimicrobials are used in aquatic organisms because acquired antimicrobial resistance in NTM is a global issue in both human and animal health.
Antimicrobial susceptibility testing (AST) is proposed for NTM that are commonly encountered in clinical practice. The most widely accepted protocol for AST of NTM is published by the Clinical and Laboratory Standards Institute (CLSI), which has recommended the use of broth microdilution as the gold standard for the determination of susceptibilities based on the determination of the minimum inhibitory concentration (MIC) of drugs.98,99 There are no current MIC breakpoints established for NTM of aquatic organisms; therefore, breakpoints established for human isolates are often applied. 9 Moreover, there is controversy about the role of in vitro susceptibility testing for NTM diseases because the correlation between in vitro and clinical outcomes for certain drugs and NTM species cannot always be predicted. 8
Several methods for susceptibility testing have been developed for NTM, including the absolute concentration method, resistance ratio method, proportion method, disk diffusion/disk elution, broth microdilution, the Epsilon test, and the rapid analysis of mycolic acids; however, these methods are not recommended by CLSI for testing antimicrobial effects against NTMs. For instance, the proportion method often yields misleading results for NTM 89 ; the agar disk diffusion method carries the inherent difficulty in the interpretation of zones of inhibition, especially when the amount of drug in the disk is near the breakpoint of the drug 90 ; and the E-test strip test is quite rapid and simple, but suffers from lack of reproducibility and exaggeration of drug susceptibility as determined by other techniques because of tailing of the ellipses.9,40 Broth microdilution, although the method recommended by CLSI, has some disadvantages, including a large volume of reagents, long experimental time, the possibility of false positives given long incubation times, and chances of cross-contamination.9,40 Careful interpretation of MIC results must consider that high susceptibility in vitro may not necessarily imply an effective in vivo outcome, particularly for NTM. CLSI has emphasized the need to ensure that the clinical significance of NTM isolates is related to the pathogenicity of the NTM isolate, host factors, and clinical presentations. Experts generally agree that susceptibility testing of NTM should be performed only for NTM isolates with clinical significance in humans or animals.8,9,16,94
The in vitro susceptibility of NTM from aquatic organisms has been investigated. For example, different strains of M. marinum isolated from zebrafish required high concentrations of trimethoprim and sulfamethoxazole for inhibition, indicating resistance to these drugs. 11 The same M. marinum strains were susceptible to amikacin and clarithromycin treatments. 11 M. fortuitum isolated from aquaria and zebrafish facilities had resistance to macrolides, streptomycin, isoniazid, rifampicin, and ethambutol.6,11 Tigecycline and clarithromycin have been used to treat M. chelonae in zebrafish, reducing the severity of the disease but not eliminating the pathogen. Nine antimicrobials were tested against 99 NTM isolates from diseased ornamental fish, and M. abscessus, M. chelonae, M. fortuitum, M. mucogenicum, M. saopaulense, and M. szulgai isolates had multidrug resistance profile with ≥ 3 different classes of antimicrobials. 28 In that same study, the growth of most NTM isolates was inhibited by amikacin and kanamycin, with only a few isolates with a multidrug resistance profile that rendered these antimicrobials ineffective. Therapeutic strategies for treating NTM infections are yet to be standardized. Therefore, the best option for the control of mycobacteriosis remains prophylaxis.
Conclusion
Mycobacteriosis is an emerging disease in aquaculture and aquarium settings. Accurate diagnosis is critical for rapid implementation of loss-minimization measures and disease surveillance. Because mycobacteriosis can cause a wide variety of clinical presentations, it is important to enhance the awareness of field personnel as well as utilize the most efficient and accurate laboratory testing procedures. Despite the limitations described in our review, targeted gene sequencing is considered the gold standard approach, and it can be applied directly from clinical specimens or from pure NTM isolates. Traditional bacterial culture continues to be relevant for NTM identification, especially with the advent of MALDI-TOF MS technologies used for genus and species confirmation. Therapeutic strategies for treating NTM infections have not been standardized; hence, the best option for the control of mycobacteriosis in aquatic organisms remains prophylaxis. However, in vitro susceptibility testing is still relevant for NTMs isolated from ornamental, zoo, or research aquatic organisms. The most widely accepted protocol for in vitro susceptibility testing of NTM recommends the broth microdilution method as the gold standard technique; nevertheless, there are no current breakpoints established for NTM of aquatic organisms. It is evident from the literature that the rapid detection and characterization of NTM remains challenging, not only given the variety of disease presentations, but also because of the range of NTM species, and potentially subspecies, involved.
Footnotes
Acknowledgements
We thank the Athens Veterinary Diagnostic Laboratory (University of Georgia, Athens, GA, USA) for providing agar plates and acid-fast slides for illustration purposes. We also thank Dr. Alvin C. Camus (Department of Pathology, University of Georgia) for providing images of histologic lesions.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
