Abstract
Clostridium botulinum is an important pathogen that causes botulism in humans and animals worldwide. C. botulinum group III strains, which produce a single toxin of type C or D or a chimeric toxin of type C/D or D/C, are responsible for botulism in a wide range of animal species including cattle and birds. We used unbiased high-throughput RNA sequencing (i.e., metatranscriptomics) to identify a strain of group III C. botulinum from a deceased Mongolian wild ass (Equus hemionus). The strain was closely related to some European strains. Genetic analysis of the recovered bacterial sequences showed that the C. botulinum strain identified might represent a type C/D strain of group III. Infection by C. botulinum producing the mosaic toxin of type C/D is the most likely cause of the death of the wild ass.
Clostridium botulinum is a gram-positive, spore-forming, anaerobic bacterium that has the ability to produce botulinum neurotoxin (BoNT), which causes severe neuroparalytic illness in humans and animals known as botulism. 7 In domesticated animals, botulism has caused significant economic losses worldwide. 7 Based on the toxin antigenic specificity of BoNT, C. botulinum strains are divided into 7 distinct serotypes (A–G). 15 A novel serotype (BoNT/H) has been reported and remains to be defined. 4 The bacteria can also be classified into 4 physiologic groups (I–IV). Group I strains produce 1 or sometimes 2 toxins of type A, B, or F; group II strains produce a single toxin of type B, E, or F. Both group I and group II bacteria cause botulism mainly in humans. 8 Group III strains, which produce a single toxin of type C or D or more commonly a chimeric toxin of type C/D or D/C, are able to cause botulism in a wide range of animal species. 13 Although group IV strains, which are rare and less well characterized, can produce type G toxin, no association with botulism have been reported. Notably, the classification based on biochemical and physiologic properties was supported by the phylogenetic analyses based on 16S ribosomal RNA (rRNA) sequences of C. botulinum strains. 2
The Mongolian wild ass (syn. onager; Equidae, Equus hemionus) is one of the most endangered large mammal species in the world. 19 It was listed as a class I endangered and protected species in China by the national government in 1988, and also listed as near threatened by the International Union for Conservation of Nature (IUCN) in 2015. 9 Although once widespread throughout the steppes and desert steppes of the Russian Federation, Mongolia, northern China, northwest India, Central Asia, and the Middle East, wild asses are now found mainly in the Mongolian Gobi and the adjacent border areas in northern China. 10
In March of 2017, a deceased adult Mongolian wild ass was found in the Urad Saxaul Forest–Mongolian Wild Ass National Nature Reserve in the Inner Mongolia Autonomous Region of China. An autopsy was performed after the carcass thawed, and samples of heart, liver, spleen, lung, kidney, and small intestine were collected. No obvious gross pathologic changes were observed in these major organs. The tissue samples were stored in dry ice and transported to the National Institute for Communicable Disease Control and Prevention, China Center for Disease Control and Prevention (CDC) in Beijing.
An unbiased, high-throughput RNA sequencing approach was used to identify potential pathogens associated with the death of the wild ass. Total RNA was extracted from 30-mg samples of spleen, liver, and kidney (RNeasy Plus minikit; Qiagen, Hilden, Germany) according to the manufacturer’s instructions; the RNA samples were pooled in equimolar amounts. The host rRNA was removed (Ribo-Zero-Gold, human-mouse-rat kit; Illumina, San Diego, CA), and a sequencing library was constructed (TruSeq total RNA library preparation kit; Illumina). Library sequencing generated 45,382,828 paired-end sequencing clean reads after passing default quality control filters on the Illumina platform and removing the adapter sequences.
After de novo assembly using Trinity, clean reads generated 711,849 contigs of 201–73,585 nucleotides (nt). The count of these contigs estimated by RSEM 11 (precise accounting of the number of reads mapping to the contig) was 0.23–2,209,088; the mean sequencing depths (as calculated by the number of reads per nucleotide position in relation to the total length of the contig) was 0.001× – 8,796×. These contigs were compared to the nonredundant nt and protein reference database deposited in National Center for Biotechnology Information (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to screen for viral, bacterial, and fungal pathogens by using blastn and blastx searches. Consequently, 365,079 contigs, 1,981 contigs, and 43 contigs were annotated as eukaryote, bacteria, and virus, respectively; 344,746 contigs were labeled as N/A that has no taxonomic information. Contigs annotated as eukaryote were identified to be of host RNA source, and no convincing hits to viral or fungal pathogens were found. Strikingly, C. botulinum was the most abundant, representing ~80% of identified bacterial contigs. The sequencing depths of contigs identified as C. botulinum were 0.01× – 8,796×. Other less abundant bacteria were species from the genera Bacillus, Bacteroides, and Streptococcus. Using the complete genome of C. botulinum strain BKT015925 (NC_015425.1) as the reference sequence, 12,767,214 reads in the metatranscriptomic library were remapped to this reference sequence and provided 25.2% genome coverage (697,434 of 2,772,964 nt) at a mean depth of 10×.
The complete coding region sequences of 3 reference genes (groEL, recA, and rpoB) were obtained from the assembled contigs; the nearly full-length (1,378 nt) of the 16S rRNA sequence was recovered from the DNA extractions of the liver, spleen, and kidney by nested PCR. All sequences obtained have been deposited in GenBank under accessions MK263740–MK263742, and MK256318. Blast analysis showed that the nucleotide similarity of the recA, groEL, rpoB, and 16S rRNA genes that we obtained and the corresponding sequences of C. botulinum available in GenBank were 79.5–98.9%, 80.0–98.5%, 79.9–98.3%, and 98.9–99.8%, respectively. The strain was most closely related to strain BKT015925 (NC_015425.1). In the phylogenetic trees based on 16S rRNA or the 6,392 nt concatenated sequences of 3 reference genes (groEL, recA, and rpoB), which were constructed using the maximum-likelihood (ML) method implemented in PhyML v.3.0, 6 the C. botulinum strain that we identified fell into group III (Fig. 1). Notably, in the multilocus phylogenetic comparison based on the concatenated sequence of the groEL, recA, and rpoB genes, the strain was closely related to those strains isolated from specimens of animal botulism outbreaks in Europe.12,16,17 Additionally, the strains BKT015925, Sp77, and 16868 produce the toxin of type C/D, type C, and type D, respectively.
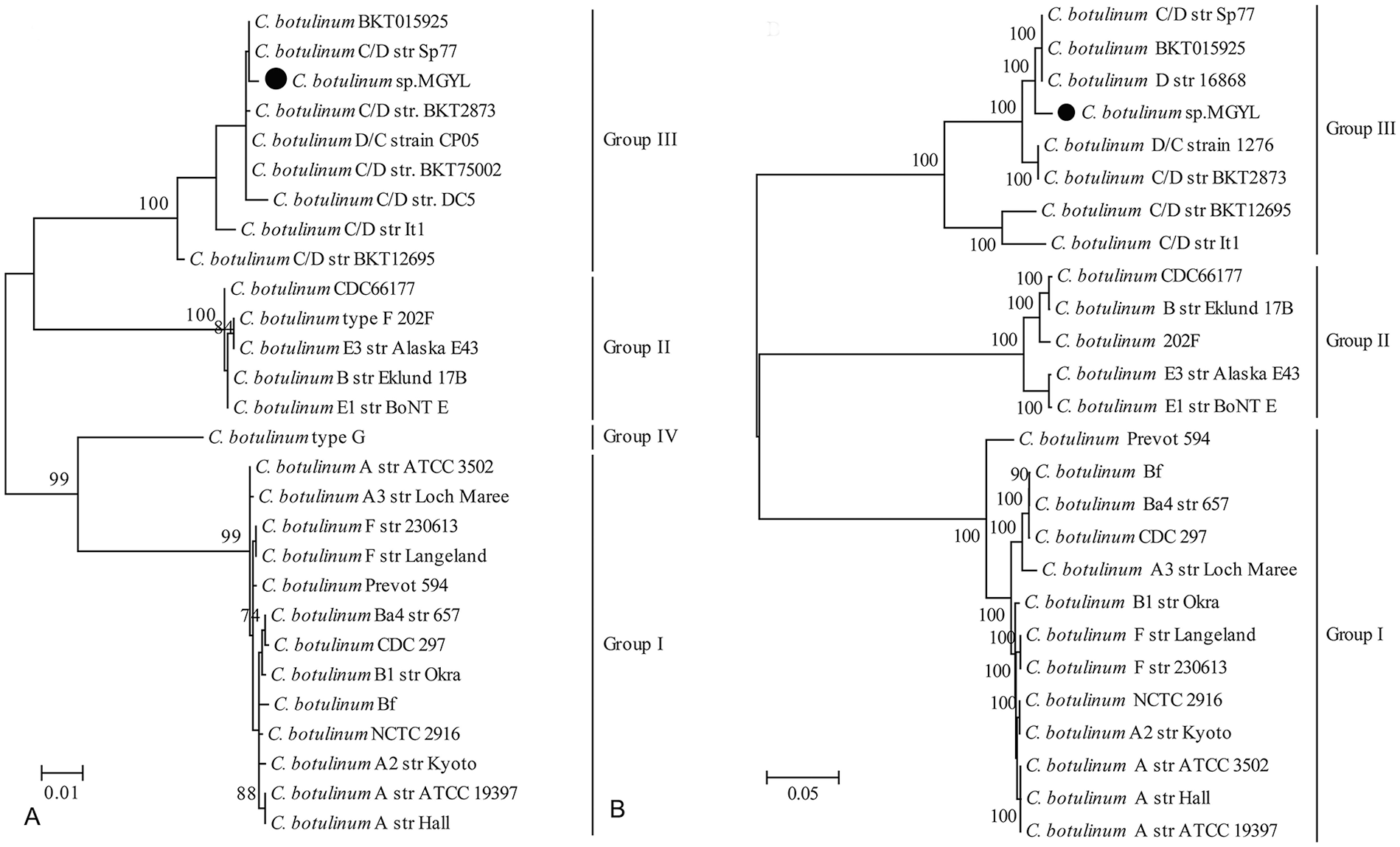
Phylogenetic trees reconstructed based on nucleotide sequences of the16S rRNA gene
To better characterize the C. botulinum strain that we identified, we attempted to find the BoNT gene from the metatranscriptomic data. However, only part of the BoNT gene sequence (≤ 1,143 nt) was obtained from the assembled contigs. Hence, the complete BoNT gene was amplified by PCR from spleen, liver, and kidney according to methods described previously.14,18 The recovered BoNT gene (accession MK468593) had an open reading frame of 3,843 nt that encoded 1,280 amino acids. Genetic analysis revealed that the C. botulinum strain that we identified produced a mosaic toxin of type C/D, with 83.6–99.8% nucleotide similarity to those of C. botulinum strains available in GenBank. Notably, the newly identified strain shared the highest nucleotide similarity (99.8% nucleotide similarity) with strains BKT2873 (JENT01000159) and S19 (FN436022) isolated from chickens in Sweden, and strain BKT75002 (JENS01000121) isolated from chickens in Denmark.
However, there were no poultry present in the National Nature Reserve, to our knowledge. C. botulinum is also commonly found in natural environments, including soil, water, and sediments. 5 Additionally, botulism outbreaks in wild animals are commonly reported when the weather is hot. 1 However, it was cold (local temperature ~5℃) when the wild ass was found dead in March 2017 in the Inner Mongolia Autonomous Region, China. Therefore, the source of C. botulinum producing the mosaic toxin of type C/D remains unclear. Although it is possible that the C. botulinum invaded after the death of the wild ass, the high percentage of C. botulinum matching contigs with high sequencing depths supports the cause of death of the wild ass as infection by C. botulinum.
Metatranscriptomics has the potential to revolutionize routine microbiologic detection. 3 A key advantage of metatranscriptomics over traditional detection techniques is that it has the potential to find any pathogen that produces an RNA molecule (RNA viruses, DNA viruses, bacteria, fungi, and protists) in an unbiased manner. 20 The combination of metagenomics and traditional methods will no doubt be helpful in the diagnosis of infectious diseases.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the National Natural Science Foundation of China (grants 81672057 and 81861138003).
