Abstract
Brucellosis is a widespread zoonotic disease caused by Brucella spp. Immunization with attenuated vaccines has proved to be an effective method of prevention; however, it may also interfere with diagnosis. Brucella abortus strain A19, which is homologous to B. abortus strain S19, is widely used for the prevention of bovine brucellosis in China. For effective monitoring of the control of brucellosis, it is essential to distinguish A19 from field strains. Single-nucleotide polymorphism–based assays offer a new approach to such discrimination studies. In the current study, we developed a cycleave PCR assay that successfully distinguished attenuated vaccine strains A19 and S19 from 22 strains of B. abortus and 57 strains of 5 other Brucella species. The assay gave a negative reaction with 4 non-Brucella species. The minimum sensitivity of the assay, evaluated using 10-fold dilutions of chromosomal DNA, was 7.6 fg for the A19 strain and 220 fg for the single non-A19/non-S19 Brucella strain tested (B. abortus 104M). The assay was also reproducible (intra- and interassay coefficients of variation: 0.003–0.01 and 0.004–0.025, respectively). The cycleave assay gave an A19/S19-specific reaction in 3 out of 125 field serum samples, with the same 3 samples being positive in an alternative A19/S19-specific molecular assay. The cycleave assay gave a total of 102 Brucella-specific reactions (3 being the A19/S19-specific reactions), whereas an alternative Brucella-specific assay gave 92 positive reactions (all also positive in the cycleave assay). Therefore, this assay represents a simple, rapid, sensitive, and specific tool for use in brucellosis control.
Keywords
Introduction
Brucellosis is a disease that has been a global concern for many years, as it causes substantial economic losses and threatens human health. Although human brucellosis can be treated, disease control in livestock involves culling of infected individuals (“test and slaughter”) and/or vaccination of healthy animals. 4 Although attenuated vaccines do not provide complete protection against brucellosis, those that are currently available are considered to be the most cost-effective tools for disease control. 19
Brucella abortus strain 19 was originally isolated as a virulent strain from a Jersey cow in 1923, and was found to have become attenuated after being kept in the laboratory at room temperature for over a year. 20 It was named B. abortus A19. Although B. abortus A19 has low virulence in cattle, vaccination of pregnant cows can result in abortion rates of 1–2.5%.19,20 B. abortus A19 can also infect humans, causing typical features of brucellosis, including acute febrile illness.16,19 To reduce this problem, an erythritol-sensitive strain, unable to grow in the presence of erythritol, was developed from B. abortus A19. It was named B. abortus S19 and has been used for vaccination since 1956.3,15 Currently, S19 is the vaccine strain used in most countries of the world. However, the S19 strain was not introduced into China, where B. abortus A19, which is homologous to B. abortus strain S19, is still widely used. 5
For the effective monitoring of brucellosis, it is essential to distinguish B. abortus A19 vaccine strains from field strains. Because of the smooth nature of B. abortus A19 and the strong antibody response against the O-side chain, it is difficult to distinguish infected animals from those that are vaccinated using serologic assays. 1 Different types of PCR assays for the detection of single-nucleotide polymorphisms (SNPs) have been developed and are widely used in the discrimination of species and biotypes of bacteria as well as virulent and nonvirulent forms of bacteria.7,8,10,11 The cycleave PCR method amplifies the template with a pair of primers and a chimeric DNA–RNA–DNA probe. 12 When the probe hybridizes to the complementary target DNA, RNaseH cleaves the probe at the RNA linkage resulting in marked fluorescence. 12 By measuring the intensity of the emitted fluorescence, the amount of amplified product can be measured. 12 Hence, cycleave PCR represents a highly sensitive and specific method for the detection of SNPs.12,21 In our study, a new cycleave PCR assay based on an A19/S19-specific SNP was developed for the rapid detection of B. abortus A19 in China.
Materials and methods
Strains and samples
The reference and vaccine strains used in our study are listed in Table 1. For A19, 12 isolates collected at different times and from different sites were used. Field isolates that consisted of B. abortus (3 isolates of biovar 4; 10 isolates of unknown biovar), B. melitensis (1 isolate of biovar 1; 10 isolates of unknown biovar), B. suis (10 isolates of biovar 3; 23 isolates of unknown biovar), and B. canis (1 isolate) were also used. As well, 4 nontarget organisms—Escherichia coli K99, Pasteurella multocida C48-1, Pseudomonas aeruginosa DI-1, and Streptococcus suis ST171—were used. The Brucella strains were cultured on tryptose agar at 37°C, with 5–10% CO2 when required, for 48–72 h in a biosafety level 3 laboratory. Bacteria were then washed with normal saline containing 0.5% formaldehyde, and inactivated at 37°C for 24 h. The 4 non-Brucella species were cultivated as described previously, 13 and then harvested and inactivated as described above.
Reference and vaccine strains of Brucella used in the current study and the results of the cycleave PCR assay specific for Brucella abortus strain A19.*
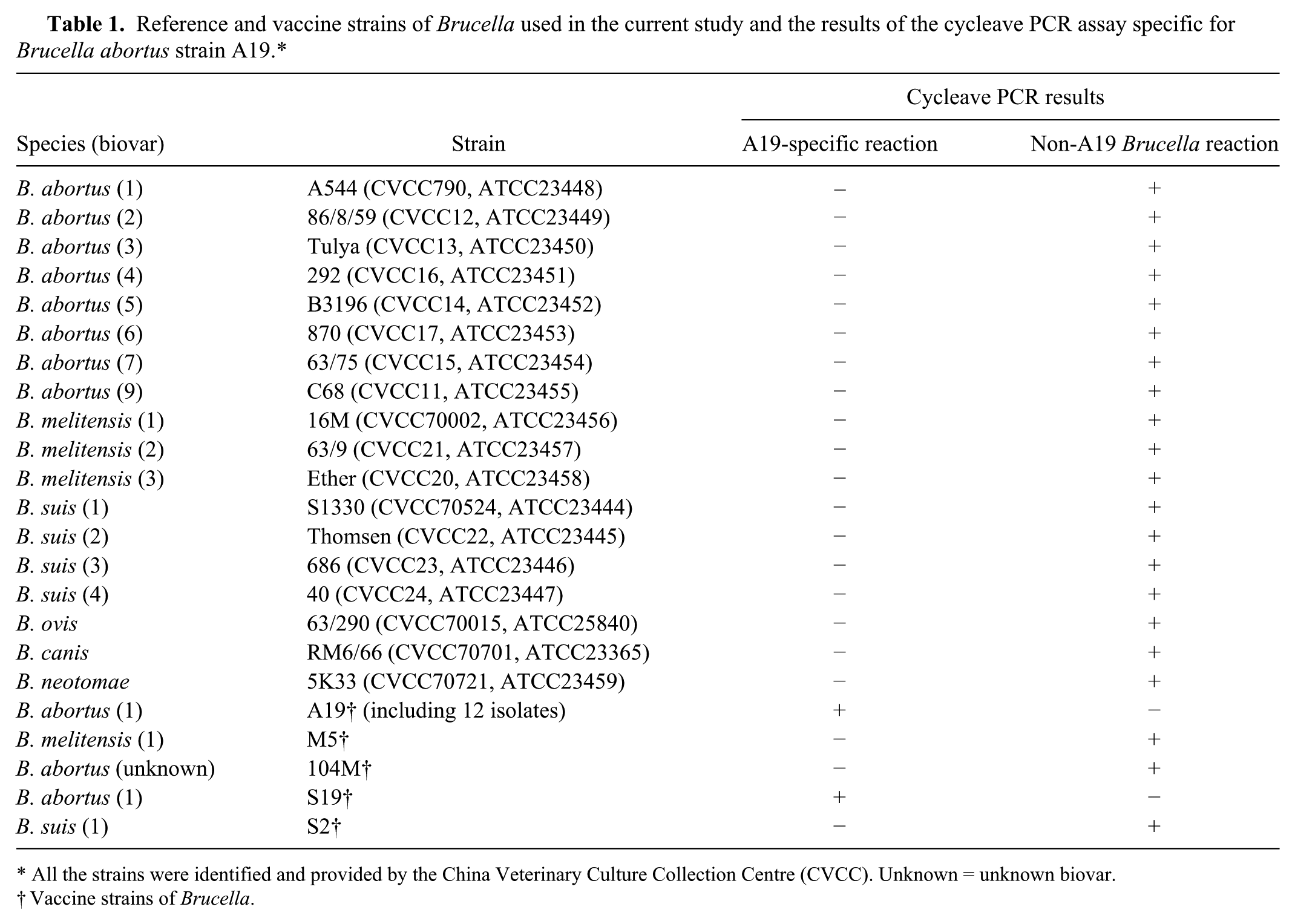
All the strains were identified and provided by the China Veterinary Culture Collection Centre (CVCC). Unknown = unknown biovar.
Vaccine strains of Brucella.
A total of 125 serum samples taken from 58 pigs, 43 cows, and 24 goats that had clinical brucellosis or were from herds infected with brucellosis, were obtained. All samples tested positive for brucellosis infection both with Rose Bengal plate test (RBT) and with commercial ELISA kits.a,b Unless specified, genomic (g)DNA was extracted c according to the manufacturer’s instructions. The concentration of the extracted DNA was measured by a commercial system. d
Cycleave PCR
For distinguishing B. abortus A19, a SNP C587–T587 in BAbS19_I07270 (arginyl–transfer RNA–protein transferase) was selected. This SNP was identified by comparison of the B. abortus S19 draft genomic sequence with the sequences of B. abortus 9-941, B. melitensis M28, B. suis S1330, B. pinnipedialis B2/94, B. microti CCM 4915, B. canis ATCC 23365, and B. ovis ATCC 25840. A set of primers and probes were designed based on this SNP (Table 2).
Sequences of primers and probes used in the current study.*
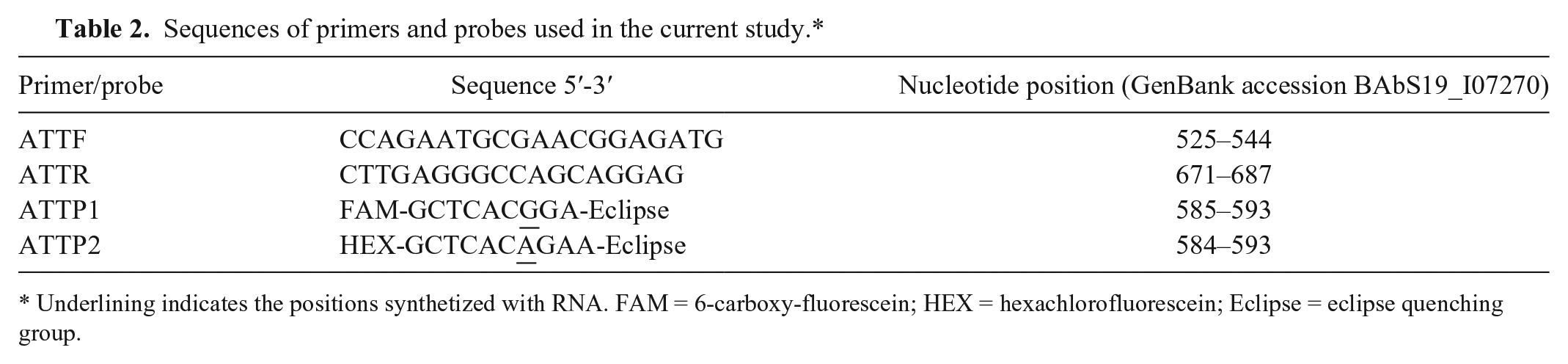

Underlining indicates the positions synthetized with RNA. FAM = 6-carboxy-fluorescein; HEX = hexachlorofluorescein; Eclipse = eclipse quenching group.
The assay was performed using a commercially available kit, e in a reaction volume of 25 µL containing deoxyribonucleotide triphosphates (300 µM), MgCl2 (5 mM), 6-carboxyfluorescein (FAM)- and hexachlorofluorescein (HEX)-labeled probes (200 nM each), primers (400 nM each), Tli RNaseH II (100 units), Ex Taq HS (1.25 units), and the DNA template (2 µL). The reaction conditions were as follows: 95°C for 10 s followed by 40 cycles of 95°C for 5 s, 55°C for 10 s, and 72°C for 20 s. Amplifications were performed using a commercial real-time PCR system. f The real-time PCR products were electrophoresed on a 1.5% agarose gel, and purified and directly sequenced g in both the forward and reverse directions.
Sensitivity, specificity, and reproducibility
The sensitivity of the cycleave PCR was evaluated using 10-fold serial dilutions of gDNA from B. abortus A19 for FAM fluorescence, and B. abortus 104M for HEX fluorescence. Each dilution was included in the assay to determine the minimum discriminatory amount of gDNA detected in the assay.
The specificity of the cycleave PCR was evaluated using the reference and vaccine strains and field isolates of Brucella. Together, these included almost all common species and biovars of Brucella and the vaccine strains used in China. In addition, 4 non-Brucella spp. (E. coli K99, P. multocida C48-1, P. aeruginosa DI-1, and S. suis ST171) were tested.
Assay reproducibility was determined by calculating the intra- and interassay coefficients of variation (CVs), using at least 3 replicates of each of the 10-fold serial dilutions of gDNA, and a standard curve was produced. The efficiency of the assay was determined using the following calculation: efficiency = 10(–1/slope) – 1. The threshold line was determined by an overall consideration of the amplification curve, coefficient of determination (R2), slope, and efficiency.
Analysis of serum samples
Initially, the 125 RBT-positive serum samples were evaluated by an established B4/B5 PCR 2 for the detection of Brucella DNA. The assay was performed using a commercially available kit h in a reaction volume of 25 µL containing 2× Premix Ex Taq (12.5 µL), primers (800 nM each), and the DNA template (2 µL). The reaction conditions were as follows: 95°C for 3 min followed by 30 cycles of 95°C for 30 s, 60°C for 30 s, and 72°C for 30 s, then 72°C for 10 min. Amplifications were performed using a commercial PCR system. i
All the samples were then analyzed in parallel using both the cycleave PCR method and the previously established BruAB1_0060 minor grove binding (MGB) assay, 10 which is capable of distinguishing B. abortus A19 and S19. The BruAB1_0060 MGB assay was performed using a commercially available kit h in a reaction volume of 25 µL containing 2× Premix Ex Taq (12.5 µL), FAM-labeled probe (400 nM), primers (800 nM each), and the DNA template (2 µL). The reaction conditions were as follows: 95°C for 3 min followed by 40 cycles of 95°C for 10 s and 60°C for 30 s. Amplifications were performed using a commercial real-time PCR system. f The results of the cycleave PCR were compared using a chi-square test by commercial software j with those obtained in the B4/B5 PCR and BruAB1_0060 MGB assays.
Results
Assay sensitivity
Using the 10-fold serial dilutions of B. abortus A19 gDNA ranging from 3.8 ng/µL to 0.38 fg/µL, the minimum sensitivity for the assay was in the range of 7.6 fg per reaction (Table 3). Using the 10-fold serial dilutions of B. abortus 104M gDNA ranging from 1.1 ng/µL to 0.11 fg/µL, the minimum sensitivity for the assay for non-A19/non-S19 Brucella strains was in the range of 220 fg per reaction (Table 3). These results indicated that the assay was highly sensitive for the detection of A19 and non-A19/non-S19 Brucella gDNA in a single reaction.
Results of mean quantification cycle (Cq) values using the cycleave PCR.*
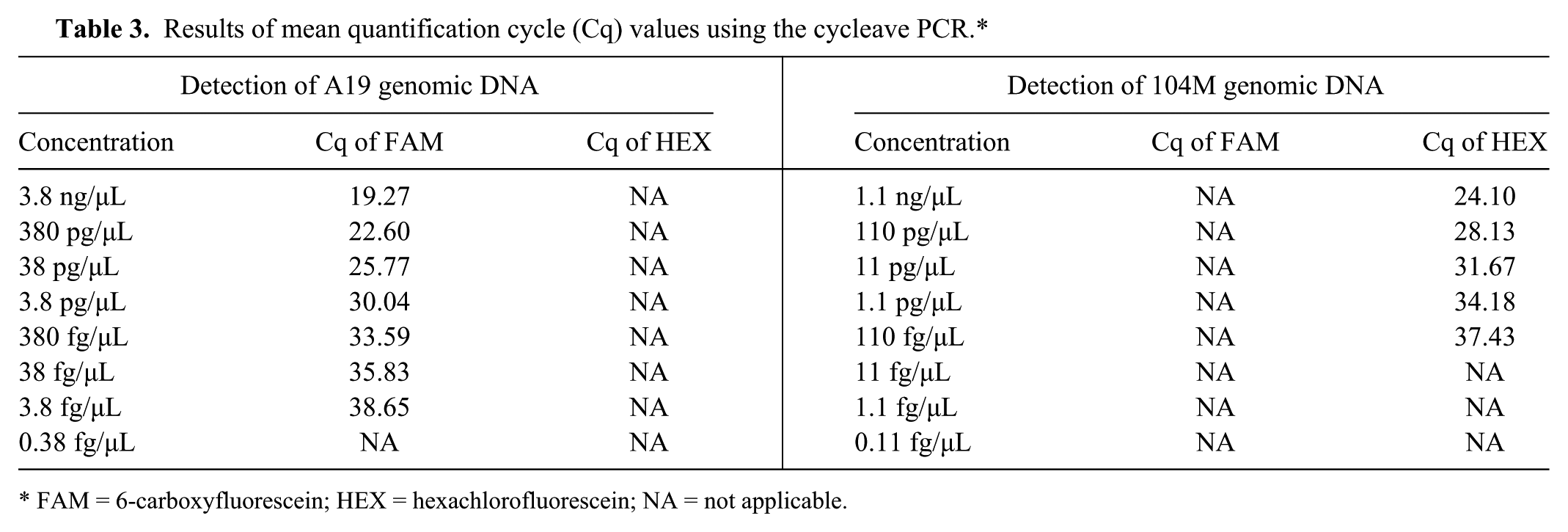
FAM = 6-carboxyfluorescein; HEX = hexachlorofluorescein; NA = not applicable.
Assay specificity
Using our cycleave PCR assay, 18 representative strains of Brucella (including B. abortus biovars 1–7 and 9, B. melitensis biovars 1–3, B. suis biovars 1–4, B. canis, B. neotomae, and B. ovis), 3 Brucella vaccine strains (S2, M5, 104M), and 58 Brucella field strains were all positive for HEX fluorescence and negative for FAM fluorescence, indicating that these were non-A19/non-S19 Brucella strains. Only the vaccine strains B. abortus A19 and S19 tested positive for FAM fluorescence and negative for HEX fluorescence, indicating that these were A19/S19 strains. Four non-Brucella species (E. coli K99, P. multocida C48-1, P. aeruginosa DI-1, and S. suis ST171) all tested negative for both FAM and HEX fluorescence. These results indicated that our cycleave PCR assay was specific for detection of B. abortus A19/S19 and non-A19/non-S19 Brucella gDNA.
Assay reproducibility
The standard curve generated using gDNA was linear over a wide range of dilutions (R2 = 99.4 and slope = −3.3 for FAM fluorescence; R2 = 99.4 and slope = −3.279 for HEX fluorescence). The assay was reproducible, with an intra-assay CV range of 0.003–0.01 and interassay CV of 0.004–0.025. The efficiency of the assay was 100.9% for FAM fluorescence and 101.8% for HEX fluorescence. In our study, this was referenced to determine the threshold line.
Analysis of clinical serum samples
A total of 125 serum samples obtained from animals with brucellosis were tested with the cycleave PCR assay and the BruAB1_0060 MGB assay. 10 The same 3 samples tested positive for A19/S19, showing concordance between the results of the 2 assays.
In the B4/B5 PCR, 2 92 samples tested positive for Brucella DNA, whereas 102 positive samples were detected using our cycleave PCR assay (including 3 FAM fluorescence positive and 99 HEX fluorescence positive). All of these 92 B4/B5 PCR–positive samples also tested positive in the cycleave assay. The 10 divergent samples were confirmed to be Brucella DNA–positive by sequencing. By chi-square test, the results of these 2 assays showed a significant difference (p < 0.01).
Discussion
Compared to B. abortus A19, the B. abortus S19 genome has a 702–base pair deletion within the eryCD locus. 5 By targeting this locus, B. abortus S19 can be easily distinguished from field strains of B. abortus by conventional PCR assays,3,9 although these methods are not suitable for distinguishing B. abortus A19. To date, studies on direct discrimination of B. abortus A19 are scarce, although the recent development of SNP-based assays offers a new approach for this purpose. SNP-based MGB assays have been used for rapid identification of the 3 Brucella vaccine strains (B. abortus strains S19 and RB51, and B. melitensis Rev1) as recommended by the World Organization for Animal Health (OIE). 10
In our study, we developed a new cycleave PCR assay that can successfully distinguish A19/S19 strains from other bacterial strains, with a sensitivity of 7.6 fg, equating to ~3 cells; this is ~10-fold more sensitive than the sensitivity reported for existing MGB assays. 10 Furthermore, our cycleave PCR assay detected non-A19/non-S19 Brucella strains in a single reaction with a sensitivity of 220 fg, equating to ~60 cells. The assay allowed accurate and reliable discrimination of A19/S19 and non-A19/non-S19 Brucella strains from common species and biovars of Brucella, Brucella vaccines, and other bacteria strains. Therefore, our assay represents a simple, rapid, sensitive, and specific tool for use in the control of brucellosis.
The cycleave PCR assay developed in our study was shown to detect A19/S19-specific clinical serum samples rapidly and accurately. For Brucella DNA, the positive detection rates were significantly higher (81.6%, 102/125) with the cycleave PCR assay than with the B4/B5 PCR assay (73.6%, 92/125); this may due to the lower detection limit of the new method for the detection of Brucella DNA. Although all serum samples tested positive by RBT and commercial ELISA kits, not all tested positive for Brucella DNA. In other studies, the sensitivities of PCR and real-time PCR assays for the detection of Brucella DNA in human blood or serum samples varied from 58% to 100%.14,17,18,22 In a previous report of the analysis of human serum samples of brucellosis, most patients tended to test positive in real-time PCR assays in the early course of brucellosis. 6 Therefore, the presence of Brucella DNA in serum samples during different infection phases should be evaluated systematically in further studies. However, the use of the cycleave PCR assay developed in this study combined with immunological tools such as RBT, standard agglutination tests, and complement fixation tests could be beneficial in areas where brucellosis prevention and control programs are applied.
Footnotes
Authors’ contributions
WL Nan contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and critically revised the manuscript. YY Zhang contributed to acquisition and analysis of data, and drafted the manuscript. PF Tan contributed to acquisition and analysis of data. ZL Xu contributed to design of the study, and contributed to analysis of data. YQ Chen and KR Mao contributed to acquisition of data. YP Chen contributed to conception and design of the study; contributed to interpretation of data; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Brucellosis antibody test kit, IDEXX Montpellier SAS, Montpellier, France.
b.
Ingezim Brucella Compac 2.0, Inmunologia Y Genetica Aplicada SA, Madrid, Spain.
c.
QIAamp DNA mini kit, Qiagen GmbH, Hilden, Germany.
d.
Qubit 2.0, Invitrogen, Life Technologies, Grand Island, NY.
e.
Cycleave PCR core kit, TaKaRa Biotechnology Co. Ltd., Dalian, Liaoning Province, China.
f.
MiniOpticon, Bio-Rad Laboratories Inc., Hercules, CA.
g.
TaKaRa Biotechnology Co. Ltd., Dalian, Liaoning Province, China.
h.
Premix Ex Taq, TaKaRa Biotechnology Co. Ltd., Dalian, Liaoning Province, China.
i.
Mycycler, Bio-Rad Laboratories Inc., Hercules, CA.
j.
IBM SPSS Statistics 19, IBM (China) Co. Ltd., Beijing, China.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Science and Technology Program of Qingdao (grant 14-2-4-79-jch).
