Abstract
Picornaviruses (PVs) of different terrestrial tortoise species, previously designated as Virus “X,” have been frequently detected from various tissues by virus isolation in Terrapene heart cell culture as the preferred laboratory method for diagnosis. Here, we describe the development of 2 diagnostic reverse transcription (RT)-PCR–based assays for the identification and characterization of tortoise PVs belonging to the tentative genus Topivirus. To test the novel diagnostic systems, PVs were isolated from swab and tissue samples collected in Germany, Italy, and Hungary between 2000 and 2013. All 25 tested isolates gave positive results with both novel consensus primer sets. Sequencing of the amplified products confirmed that all studied viruses were members of the new proposed genus Topivirus. Phylogenetic analyses clearly distinguished 2 lineages within the genus. Based on sequence analysis, no association was observed between the geographic distribution and genetic relatedness. Furthermore, no strict host specificity was indicated. The PCR-based diagnosis may provide a time-saving and sensitive method to detect tortoise PVs, and evaluation of PV presence in these animals may help control virus spread.
The family Picornaviridae includes small nonenveloped viruses with a positive-sense, single-stranded RNA genome. 9 This genetically diverse family is currently divided into 26 genera (http://www.ictvonline.org/) and contains a number of additional unassigned picornaviruses (PVs). PVs infect humans, several orders of mammals and birds, and have been reported in lower vertebrates including reptiles and fish.2,5,6,10,13,15 With some exceptions, PVs cause mild or subclinical infections and are strictly host specific, inducing disease only in a single or a few closely related host species. 9
Picornaviruses of tortoises have been detected in several instances by virus isolation in Terrapene carolina heart cells (TH-1; ATCC CCL-50) based on their typical lytic cytopathogenic effect. Such viruses were referred to as Virus “X.” 11 PV infection is usually diagnosed in spur-thighed tortoises (Testudo graeca)—one of the most popular species among exotic animal keepers—but has also been confirmed in other terrestrial species, including marginated tortoises (T. marginata), Hermann’s tortoises (T. hermanni), leopard tortoises (Stigmochelys pardalis; syn. Geochelone pardalis), and Egyptian tortoises (T. kleinmanni). 11 Clinical signs reported in tortoises with picornaviral infection include softening of the carapace of young animals, 8 diphtheroid-necrotizing stomatitis and pharyngitis, rhinitis, pneumonia, conjunctivitis, enteritis, and ascites, but such infections have also been detected from clinically healthy animals (RE Marschang, unpublished observations, 1996–2015). In several cases, PVs have been diagnosed as coinfections, together with other pathogenic agents of tortoises, mainly herpesviruses and Mycoplasma spp. (RE Marschang, unpublished observations, 1996–2015). The near complete genome sequences of a dozen tortoise PV strains have been determined and characterized. 5 Sequence analyses of these genomes showed that tortoise PVs do not belong to any recognized genera, but most probably are the first members of a new genus in the Picornaviridae family, provisionally named Topivirus. 5
Another group of tortoise PVs, rafivirus A was described in 2015 in Sulawesi tortoises (syn. Travancore tortoises, Indotestudo forstenii) in connection with an adenovirus outbreak and mass mortality event. 13 This virus proved to be phylogenetically distinct from topiviruses and was more closely related to members of the Gallivirus/Kobuvirus/Meg-rivirus/Oscivirus/Passerivirus/Sakobuvirus/Salivirus/ Sicinivirus supergroup.
Virus isolation in cell culture is an effective tool for diagnosing topivirus infection in tortoises and offers several advantages, including its ability to detect infectious viral particles as well as to detect other infectious viruses (e.g., herpesviruses). However, this method is time-consuming and can result in false-negative results caused by handling of samples, possible differences in susceptibility of the applied cell lines to various virus strains, and coinfections with different viruses, which may also affect isolation in cell culture. Therefore, we felt that development of a diagnostic reverse transcription (RT)-PCR assay for in vitro detection and preliminary characterization of these viruses would be a useful laboratory diagnostic tool.
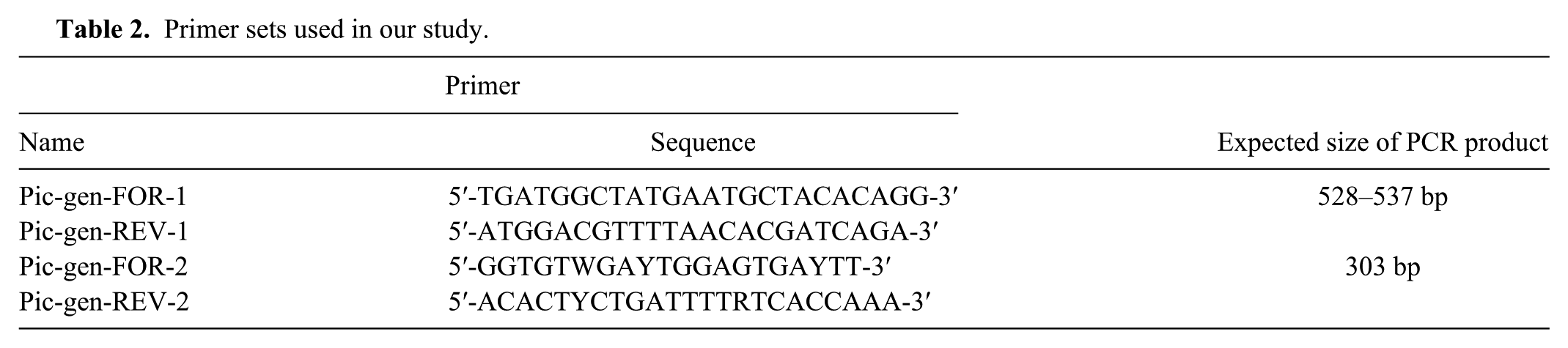
Samples processed in our study were collected from different terrestrial tortoise species in Germany, Italy, and Hungary between 2000 and 2013. Oral, pharyngeal, and/or cloacal swabs were tested from live animals, whereas internal organ specimens were collected for virus isolation from dead animals (Table 1). In all cases virus isolation was performed on the TH-1 cell line as described previously. 1 RNA from virus isolates with a typical lytic cytopathic effect (CPE) was prepared by freezing at −20°C and thawing the inoculated cell culture. Purification of RNA from virus isolates and pathological samples (Supplementary Table 1; Supplementary Fig. 1; available online at http://vdi.sagepub.com/content/by/supplemental-data) homogenized in phosphate buffered saline was performed through high-speed shaking in plastic tubes with stainless beads a using a commercial kit b with TRI reagent c according to the manufacturer’s instructions. Prior to the RT step, RNA was denatured (95°C, 5 min) in the presence of 1 μL of random hexamer (50 μM), and then cooled on ice slurry for 5 min. After cooling the RNA, a reaction mixture containing 1 μL of Avian myeloblastosis virus reverse transcriptase d (10 U/μL) and its buffer, 0.5 μL of RNase inhibitor e (40 U/μL), 1 μL of deoxyribonucleotide triphosphate (dNTP) mixture f (10 mM each), and nuclease-free water was added to reach the final volume of 25 μL. The complementary DNA synthesis was performed applying the following temperature profile: 25°C for 10 min, 42°C for 60 min, and heat inactivation at 75°C for 15 min. Three microliters of complementary DNA was used in the PCR assay. For specific detection of tortoise PVs, 2 primer sets were designed on the basis of the nucleotide alignment of tortoise PV complete genome sequences determined in our previous study (Table 2). The 25-μL reaction mixture contained 1 μL of forward and reverse primer (20 μM each), 0.15 μL of DNA polymerase e (5 U/μL), 2.5 µL of 10× buffer, 0.5 μL of dNTP mixture (10 mM each), and nuclease-free water. The PCR conditions for amplification included an initial denaturation at 95ºC for 5 min followed by 40 cycles of denaturation at 94°C for 30 s, annealing at 45°C for 30 s, extension at 72°C for 40 s, followed by final extension at 72°C for 8 min. In all RT-PCR assays, nuclease-free water was used as negative control.
Animals and samples processed in our study.*
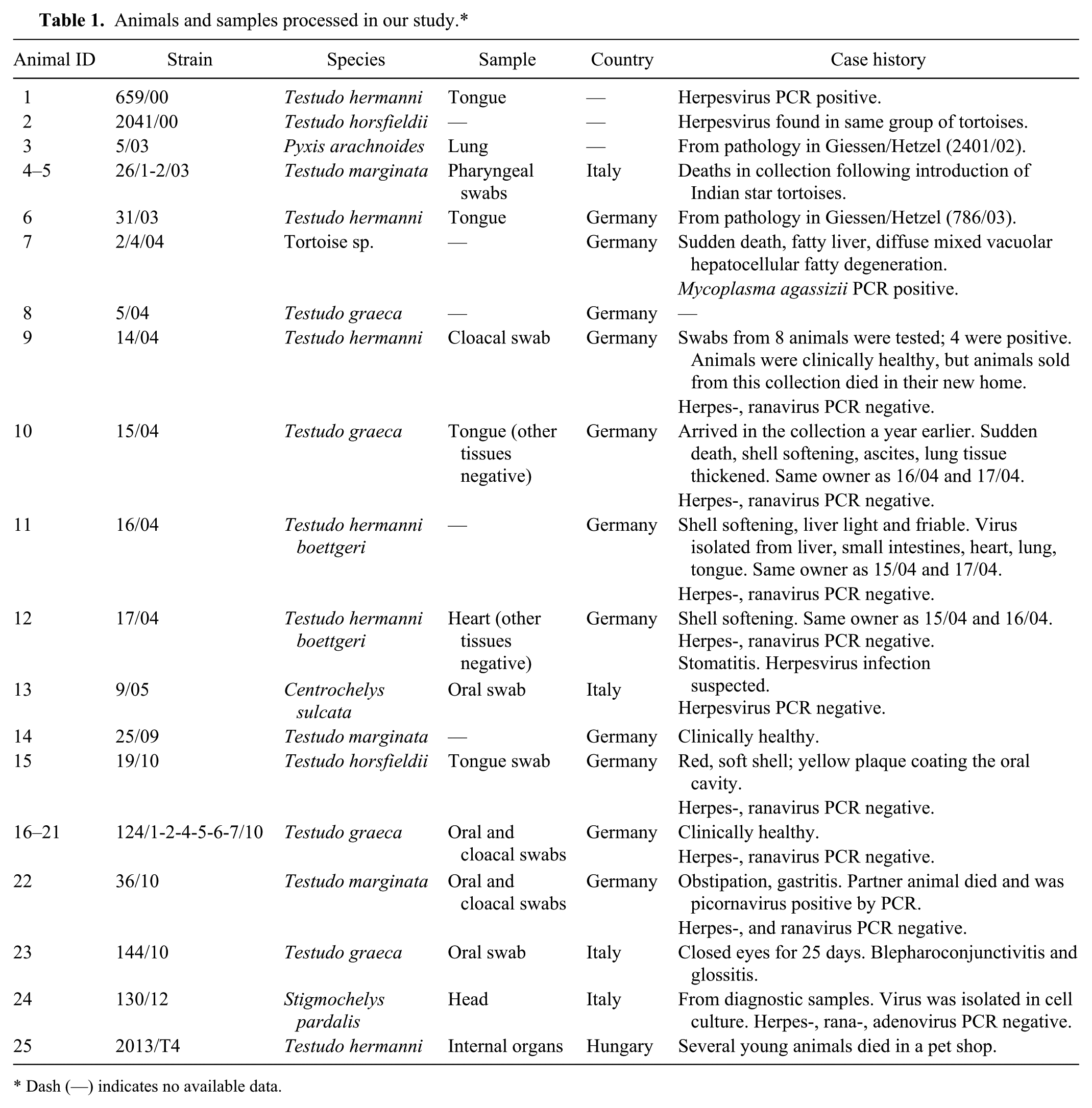
Dash (—) indicates no available data.
Primer sets used in our study.
Analytical performance of the assay was tested by using representative reptile virus strains, including tissue culture supernatant of strains 5/03 (titer 2.81E+04 TCID50/mL) and 14/04 (titer 5.00E+05 TCID50/mL), as well as a variety of clinical and pathological samples (previously tested for adenoviruses, chelonian herpesviruses, ranaviruses, and Mycoplasma agassizii by PCR3,12,17,18). To confirm the PCR results, amplicons of appropriate size were excised and purified with a commercial kit f then directly sequenced g according to the manufacturer’s instructions, but in a reduced final volume (10 μL).
Sequence data was processed using MultAlin online software 4 and were edited in GeneDoc. 14 Nucleotide and amino acid sequence identity values were calculated using BioEdit version 7.2.0. 7 Phylogenetic analysis was performed applying the MEGA version 6 package. 16 Maximum-likelihood and neighbor-joining trees were generated; the best-fit models were selected based on the Bayesian information criterion, and tree topologies were validated by bootstrap analysis as implemented in MEGA 6.
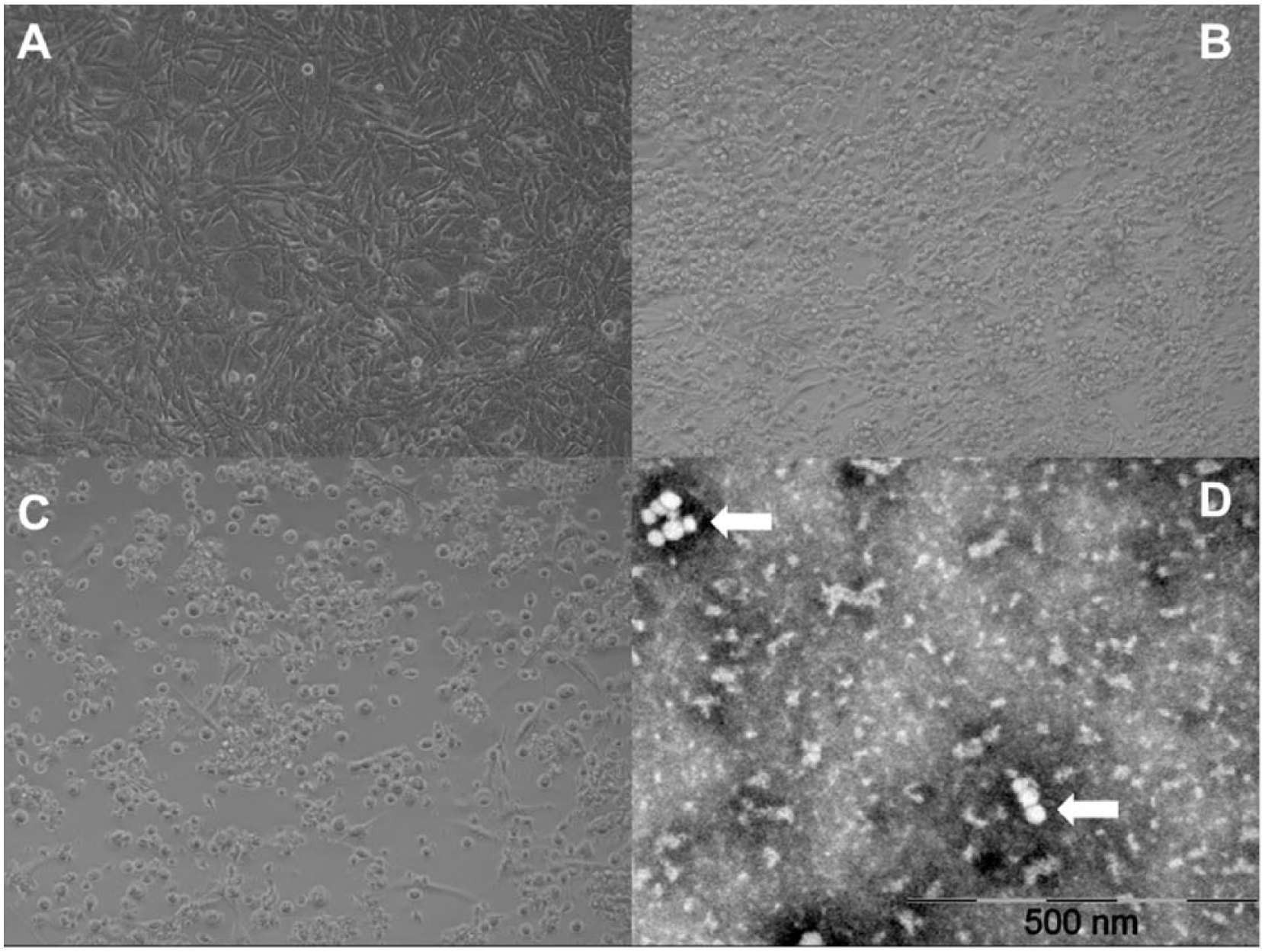
Tortoise PVs were isolated from swab samples collected from live animals as well as from internal tissues from dead animals (Fig. 1; Table 1). Virus isolation was successful from apparently healthy individuals and animals displaying different clinical signs and lesions, including stomatitis, glossitis, gastritis, obstipation, pneumonia, blepharoconjunctivitis, shell-softening, fatty liver or other lesions in the liver, ascites, and death (Table 1). Positive oral and cloacal swab samples of healthy tortoises indicate that the virus may also be shed by asymptomatic animals, which may contribute to unnoticed spread of the virus in populations and collections. In a few cases along with the tortoise PVs, herpesvirus and M. agassizii coinfection was also detected by PCR. Electron microscopic examination of the prototype strain 14/04 revealed small viral particles ~30 nm in diameter (Fig. 1).

To test the specificity of the newly developed RT-PCR assay, a total of 25 virus isolates inducing CPE, resembling that of Virus “X,” in TH-1 cell lines were tested for the presence of PV RNA using 2 different RT-PCR assays. The tortoise PV–specific primer sets amplified 2 regions of the genome, containing a portion of the 5′-untranslated region, the Leader peptide, and the VP4 capsid protein, and a portion of the picornaviral RNA-dependent RNA polymerase (3D), respectively (Table 2). All PV isolates proved to be positive with both assays; in contrast, no specific bands were obtained when DNA or RNA from reptilian-origin reo- (strain CH 1197/96), adeno- (strain IBD 27/00), irido- (strain 100/01), rana- (strain CH8), herpes- (strain 4295), and paramyxovirus (strain 9/5/03) isolates were used as templates in the tests.
Analytical sensitivity was tested by serial dilutions of topivirus isolates representing various genetic lineages. The highest dilution of 5/03 strain RNA yielding positive result was 10−4 and 10−3 by RT-PCR with the first or second primer set, respectively (Fig. 3). Thus, the sensitivity of primer set 1 was comparable to that of the routine cell culture–based detection, whereas the sensitivity of primer set 2 was ~10 times lower. Both primer sets were able to detect PV RNA from pooled organ samples of tortoises examined in Hungary from which strains 2013/T4, 2013/T11, and 2011/247 could be isolated (Supplementary Table 1; Supplementary Fig. 1). The assay was found to be highly reproducible as evidenced by duplicate and/or triplicate runs in independent assays.
Partial sequences of the detected PV strains were deposited in GenBank (accessions KR363962–KR363997). Respective nucleotide and amino acid sequence identity values (Leader peptide: 67.9–100% and 67.3–100%; VP4: 79.2–100% and 94.2–100%; 3D: 77.5–100% and 86.9–100%) revealed a close genetic relationship among the studied strains, and beside the typical CPE characteristic to these viruses, analysis of the obtained partial sequence data also confirmed that all strains belong to the prospective genus Topivirus and that none of the detected viruses were related to the other tortoise PV, the tortoise rafivirus (data not shown). A 3-aa long deletion in the Leader peptide was observed in strains 5/03 and 124/1-7/10. In the phylogenies, 2 main lineages could be clearly distinguished; most of the strains belonged to lineage 1, whereas lineage 2 was formed by strains 5/03 and 124/1-7/10 (Fig. 2; Supplementary Table 1; Supplementary Fig. 1). Viruses isolated and processed in our study were collected from tortoises in 3 different European countries (Germany, Italy, and Hungary), but no association could be observed between the geographic distribution and the genetic relatedness of the strains, presumably a result of the high levels of international trade of exotic infected animals. Similar to our previous observations, no strict host specificity was identified based on the examination of the topivirus isolates. 5
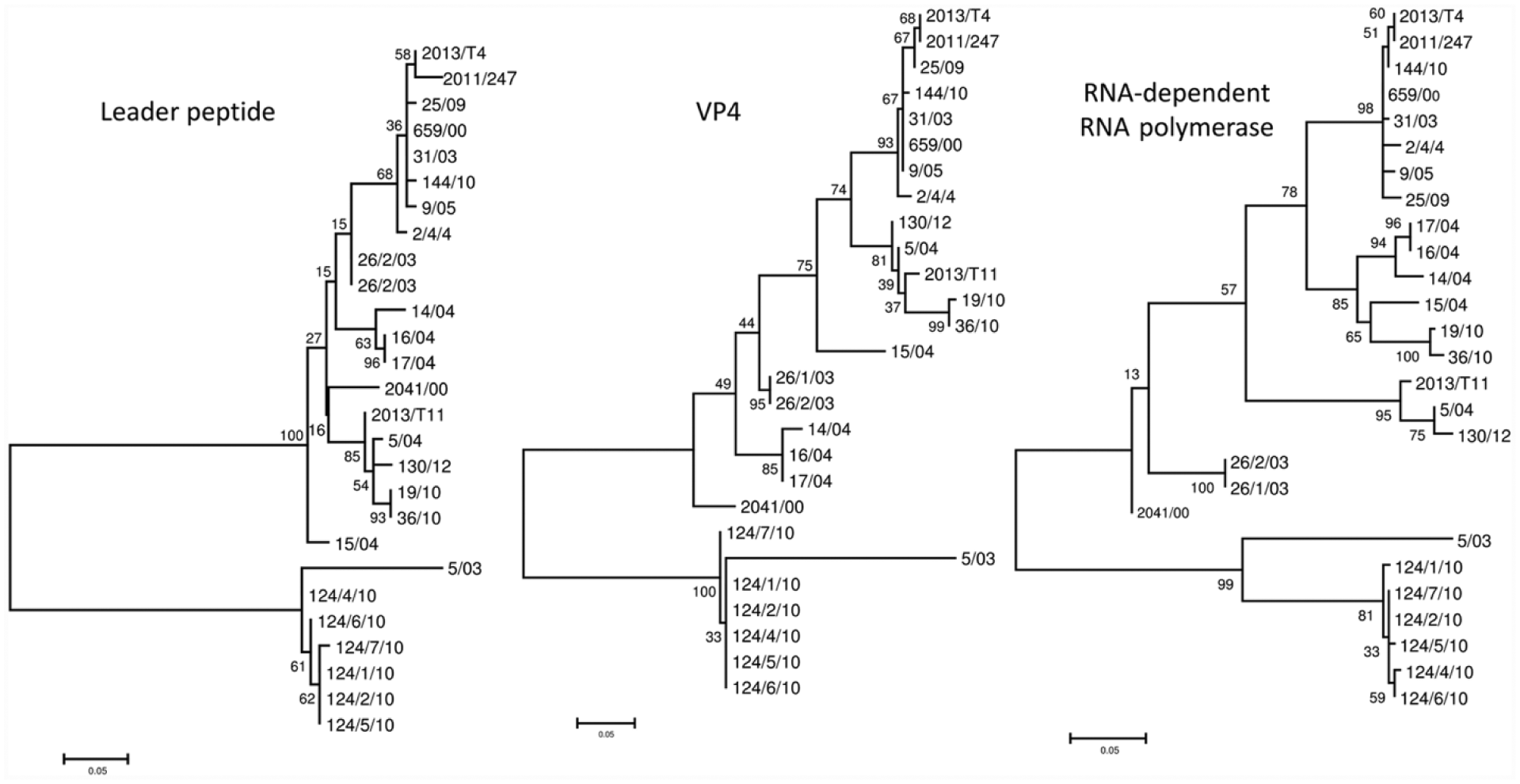
Unrooted phylogenetic trees of the Leader peptide, VP4, and partial RNA-dependent RNA polymerase nucleotide sequences obtained from the studied tortoise picornaviruses. Phylogenetic calculations were carried out applying maximum likelihood method. The length of the branches indicates the phylogenetic distance between different viruses; the scale bar is proportional to the genetic distance.
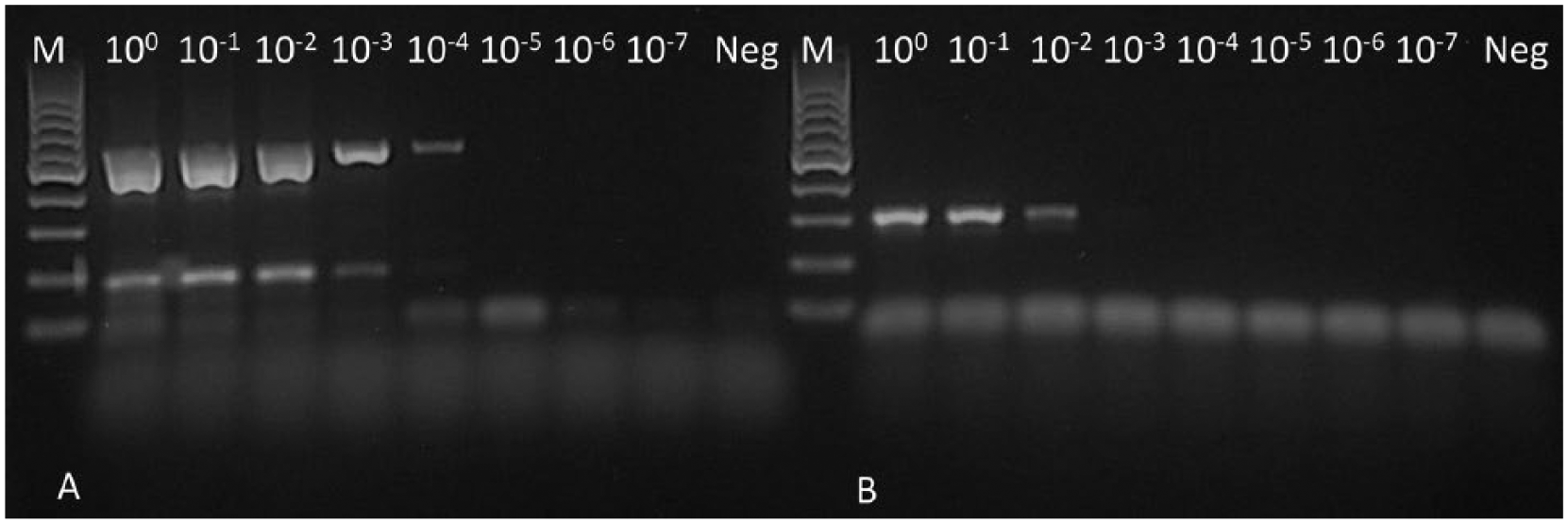
Reverse transcription PCR products of strain 5/03 visualized by gel electrophoresis performed with (
Despite the frequent diagnosis of topivirus infection in tortoises, no studies, to our knowledge, have been performed to analyze the connection between the lesions and the presence of these viruses. Detection of tortoise PVs in clinically healthy animals does not rule out their pathogenic role in this animal group, and asymptomatic infections may lead to extensive spread of these viruses, especially in collections where usually several animals are housed together. Until the etiological role of these viruses is better understood, it is prudent to observe and test newly acquired animals during a quarantine period to avoid introducing potential pathogens into tortoise collections or populations. We believe that the PCR assay developed in this study provides an equally sensitive, but more specific and time-saving, alternative to cell culture–based detection of topivirus infections.
Footnotes
Acknowledgements
We thank Renáta Pop (Szent István University, Faculty of Veterinary Science, Department of Pathology) for performing the electron microscopic examination.
Authors’ contributions
All authors contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved. RE Marschang, K Ihász, R Kugler, G Lengyel, S Marton, K Bányai, T Aqrawi, and SL Farkas drafted the manuscript.
a.
TissueLyser II, Qiagen GmbH, Hilden, Germany.
b.
Direct-zol RNA MiniPrep, Zymo Research Corp., Irvine, CA.
c.
TRI Reagent, Sigma-Aldrich, St. Louis, MO.
d.
AMV reverse transcriptase, Promega Corp., Madison, WI.
e.
RiboLock, DreamTaq; Thermo Scientific, Waltham, MA.
f.
Gel/PCR DNA fragments extraction kit, Geneaid Biotech Ltd., New Taipei City, Taiwan.
g.
Big Dye Terminator V1.1 cycle sequencing kit, Life Technologies, Carlsbad, CA.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: One author (Marschang) is currently employed by a commercial laboratory offering diagnostic testing for veterinarians.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Momentum Program of the Hungarian Academy of Sciences.
