Abstract
Microbiological and histological analysis of a sample from a swollen testicle of a 2-year-old Border Collie dog revealed a mixed infection of the fungus Blastomyces dermatitidis and the Gram-negative bacterium Aureimonas altamirensis. When subjected to an automated microbial identification system, the latter isolate was provisionally identified as Psychrobacter phenylpyruvicus, but the organism shared several biochemical features with Brucella canis and exhibited agglutination, albeit weakly, with anti–B. canis antiserum. Unequivocal identification of the organism was only achieved by 16S ribosomal RNA gene sequencing, ultimately establishing the identity as A. altamirensis. Since its first description in 2006, this organism has been isolated infrequently from human clinical samples, but, to the authors’ knowledge, has not been reported from a veterinary clinical sample. While of unknown clinical significance with respect to the pathology observed for the polymicrobial infection described herein, it highlights the critical importance to unambiguously identify the microbe for diagnostic, epidemiological, infection control, and public health purposes.
Keywords
Aureimonas (formerly Aurantimonas) altamirensis is a Gram-negative coccobacillus, a member of a bacterial genus that is most often recovered from aquatic environments and a species infrequently reported as a clinical isolate in human medicine.4,6–8,11 This organism was originally recovered from the complex microbial community whose white-colored colonization is a threat to the Paleolithic drawings on the walls of the Altamira Cave (Cantabria, Spain). 6 The organism is the type species of the genus, which belongs to the bacterial family Aurantimonadaceae of the order Rhizobiales.6,9 Since the first description of Aurantimonas altamirensis in 2006, subsequently reclassified as Aureimonas in 2011, 9 a limited number of publications have reported isolation of the organism from human clinical samples, including blood,4,8 pleural effusions, 11 contact lens, lens-cleansing solution, and foreign body–containing corneal culture. 7 Germane to the current communication, Aureimonas altamirensis has also been isolated from a human diabetic patient with scrotal swelling. 8 However, the clinical relevance of the isolate in all cases remains uncertain.
Aureimonas altamirensis is slow growing (2–5 days of culture at 35°C) and results in formation of yellow smooth colonies on trypticase soy agar. 6 The organism is a Gram-negative coccobacillus that is oxidase, catalase, and urease positive. 6 Except for the yellowish appearance (unobservable at 18 hr incubation), these attributes are not unlike those of the important canine reproductive pathogen, Brucella canis, which is not uncommonly isolated. 5 Brucella canis is a facultative intracellular bacterium that is a catalase, oxidase, KOH, and urease positive, aerobic, Gram-negative coccobacillary organism that forms small raised mucoid colonies on Brucella agar medium after 2–3 days of incubation at 35°C. 13 In the current work, the isolation of A. altamirensis from the testicle of a dog presenting with fever, sore toes, and scrotal swelling for which canine brucellosis was included as part of the differential diagnosis is described.
A 2-year-old, 19-kg male Border Collie dog repeatedly presented with fever, generalized lymphadenopathy, sore toes, and testicular swelling. The dog was found to be serologically negative for B. canis, treated empirically with doxycycline and carprofen a to alleviate symptoms, and was eventually neutered. The testicular tissue was forwarded to the University of Missouri, College of Veterinary Medicine Veterinary Medical Diagnostic Laboratory (Columbia, Missouri) for examination. The fixed and fresh tissues were examined by histology and bacterial and fungal culture, respectively.
Microscopically, 4–5-µm sections of the reproductive tissue stained with hematoxylin and eosin showed heavy infiltration of neutrophils into the spermatic tubules with a complete loss of spermatocytes and the presence of multifocal round to oval yeast forms 15–25 μm in diameter consistent with Blastomyces dermatitidis or a genetically distinct clade within B. dermatitidis 2 (Fig. 1). While uncommon, one of the extrapulmonary foci of B. dermatitidis infection in males (canine and human) is the genitourinary tract.10,12
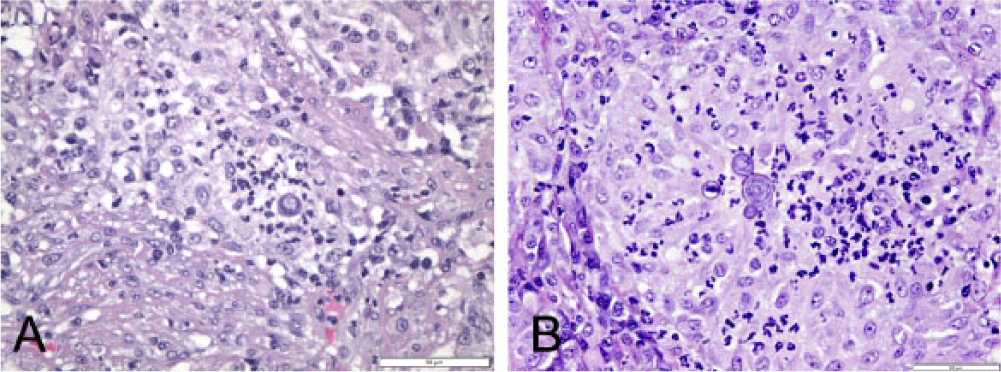
Canine testicle.
In the bacteriology laboratory section, as per standard operating procedure, the testicular tissue was aseptically cultured on blood, MacConkey, and Brucella (containing 5% defibrinated sheep blood) agars b for bacterial isolation and mycosel and Sabouraud agars b for fungal isolation. Bacterial cultures were incubated at 35°C (in the presence of 5% [v/v] CO2 and 95% air for the Brucella agar) for approximately 72 hr with daily examination; fungal cultures were incubated at room temperature for 30 days and examined weekly for fungal growth.
No growth was detected on the fungal culture plates, a finding that is not uncommon for B. dermatitidis (W.H. Fales, personal communication, 2014). After initial incubation of approximately 16 hr, a light growth of minute bacterial colonies was noted on the Brucella agar, which was further incubated for an additional 24 hr. After the second day of incubation, the small raised mucoid colonies were examined further and were found to be comprised of KOH, catalase, and oxidase positive Gram-negative coccobacilli. Furthermore, while delayed, the bacteria were urease positive (in approximately 40 min) in the rapid urea test commonly employed to initially identify B. canis and demonstrated weak but consistent agglutination with anti–B. canis antiserum. c While the scrotal swelling could be explained by the presence of B. dermatitidis (as determined by histology) and the inflammatory response to the same, given the origin of the specimen, the presentation of the patient, the differential diagnosis provided by the attending clinician, and the preliminary phenotypic results obtained during bacteriological examination, it was imperative to subject the bacterial isolate to further analysis to confirm or refute the tentative identification of B. canis and to identify the bacterial component of the dual infection.
In addition to the conventional tests above, the bacterial isolate was examined using automated systems for identification d and antimicrobial susceptibility profiling e of Gram-negative isolates, as per the manufacturer’s recommendations. The system identified the isolate as Psychrobacter phenylpyruvicus, finding it to be positive in 2 of the 32 tests utilized for Gram-negative identifications, urease, and the proprietary reaction FR7. However, B. canis is not included in the database for the identification system, and known Brucella isolates (as determined by 16S ribosomal RNA [rRNA] gene sequencing) are also identified as P. phenylpyruvicus by this identification system (TJ Reilly, unpublished results). The bacterial isolate, when retaining the automated identification, showed susceptibility to penicillin at ≤0.06 µg/ml; ampicillin, cefovecin, ceftiofur, enrofloxacin, marbofloxacin, and oxacillin at ≤0.25 µg/ml; trimethoprim–sulfamethoxazole at ≤0.50 µg/ml; gentamicin, imipenem, and rifampin at ≤1.00 µg/ml; cefoxitin, cefpodoxime, and doxycycline at ≤2.00 µg/ml; amikacin, amoxicillin–clavulanic acid, cefazolin, and chloramphenicol at ≤4.00 µg/ml; and ticarcillin and ticarcillin–clavulanic acid at ≤8.00 µg/ml. The isolate showed intermediate sensitivity and resistance to erythromycin and clindamycin at 4.00 µg/ml and >4.00 µg/ml, respectively.
To more definitively identify the organism, bacterial genomic DNA was prepared using a commercial kit, f according to the manufacturer’s protocol for Gram-negative bacteria and an aliquot used for polymerase chain reaction (PCR) amplification of a major portion of the gene(s) encoding 16S rRNA, using oligonucleotide primers g 16Sf (5′-AGAGTTTGATCMTGGCTCAG) and 16Sr (5′-GACGGGCGGTGTGTRCA) 3 and a PCR amplification kit. h The resulting amplicon was purified by spin chromatography i and the nucleotide sequence determined using fluorescent dye terminator chemistry j and an automated DNA sequencer k at the University of Missouri DNA Core facility (Columbia, MO). The 1,152-nucleotide sequence (deposited in GenBank under accession no. KF561899) was compared to entries in the nonredundant nucleotide (nr/nt) database (NCBI) using BLASTn 1 and was found to be identical to two 16S rRNA gene sequences from 2 A. altamirensis isolates and differed at only 4 nucleotides (99.65% identity) from the 16S rRNA gene sequence of the A. altamirensis type strain S21BT (database accession DQ372921). After A. altamirensis, the next best species matches were Aureimonas frigidaquae (14 differences to CW5T type strain, 98.8% identity) and Aurantimonas coralicida WP1T (type strain; 40 differences, 96.5% identity). The 16S rRNA gene sequence was 92% identical to that of the B. canis ATCC23365T type strain (92 differences/1,159 nucleotides). Based on the 16S rRNA gene phylogenetic analysis (BLASTn suite 1 ), the canine testicle isolate was identified as A. altamirensis.
As no additional A. altamirensis nucleotide sequences were available for further comparison, and no protein encoding gene sequences have been determined for this species, a random clone from a plasmid l insert library (StuI m restriction fragments) was selected, and nucleotide sequence determined by primer walking. The sequences of the metS gene (encoding methionyl tRNA synthetase), together with that encoding a TatD family DNAse, were unique, with greatest similarity to orthologs in Aureimonas ureilytica (80% and 69% amino acid identity, respectively). Downstream open reading frames for a putative hydrolase (64% identity to Aurantimonas manganoxydans), a glyoxal–bleomycin resistance family protein (63% identity to Pseudogulbenkiania sp.), and a fucose transport protein (51% identity to Herbaspirillum seropedicae) exhibited best matches to open reading frames from the environmental bacterial taxa indicated. The nucleotide sequence of this 4.2-kb insert has been deposited in GenBank under accession no. KF561900 and provides additional potential target sequences for molecular diagnostic and epidemiological studies wherein strain discrimination of A. altamirensis isolates may be important to ascertain.
The data reported in the current study describes the isolation of A. altamirensis from a canine diagnostic sample and the issues associated with the confident identification of a species that has been rarely previously encountered. These observations also raise the possibility that this species may be overlooked as part of a polymicrobial infection. Furthermore, because it is an unusually encountered organism, it may not be definitively diagnosed in a situation where the patient responds to antimicrobial therapy, especially since commercially available phenotypic tests do not provide adequate resolution of uncommon taxa. Indeed, commercial strain identification systems do not include such a rarely isolated organism in their databases. This absence from the databases is highlighted by the current case as well as the experiences of 2 other laboratories in which standard identification schemes failed to accurately identify the organism. In one report, the A. altamirensis isolate was initially identified as Brucella spp. or CDC group EO-3, depending on the typing approach employed, 8 and in a second study, the isolate was conflictingly identified as Ochrobactrum anthropi, Sphingomonas paucimobilis, or Shewanella putrefaciens 7 by different identification kits. In all cases described to date, accurate species identification was only achieved by 16S rRNA gene sequencing and comparison to reference 16S rRNA genes in the National Center for Biotechnology Information database. It is envisaged that greater awareness of the organism and the difficulties associated with identification by automated biochemical systems should lead to increased recognition of A. altamirensis and thus a more complete understanding of the pathogenic potential, or the possible status as an emerging infectious agent, of this bacterium.
Footnotes
a.
Novox Vedco Inc., St. Joseph, MO.
b.
Remel Inc., Lenexa, KS.
c.
D-Tec CB, Synbiotics Corp., San Diego, CA.
d.
GNID AP80, Trek Diagnostic Systems, Thermo Scientific, Cleveland, OH.
e.
Sensititre Aris 2X system, Trek Diagnostic Systems, Thermo Scientific, Cleveland, OH.
f.
DNeasy blood and tissue kit, Qiagen Inc., Valencia, CA.
g.
Integrated DNA Technology Inc., Coralville, IA.
h.
Expand long template PCR system, Roche Diagnostics Corp., Indianapolis, IN.
i.
QIAquick PCR purification kit, Qiagen Inc., Valencia, CA.
j.
BigDye terminator cycle sequencing kit, Life Technologies, Grand Island, NY.
k.
3730xl DNA analyzer, Life Technologies, Grand Island, NY.
l.
CloneJET PCR cloning kit, Thermo Scientific, Pittsburgh, PA.
m.
New England Biolabs Inc., Ipswich, MA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for this research, authorship, and/or publication of this article.
