Abstract
Erysipelothrix rhusiopathiae causes erysipelas in swine and is considered a reemerging disease contributing substantially to economic losses in the swine industry. Since an attenuated live vaccine was commercialized in 1974 in Japan, outbreaks of acute septicemia or subacute urticaria of erysipelas have decreased dramatically. In contrast, a chronic form of erysipelas found during meat inspections in slaughterhouses has been increasing. In this study, a new strain-typing method was developed based on nucleotide sequencing of a hypervariable region in the surface protective antigen (spaA) gene for discrimination of the live vaccine strain from field isolates. Sixteen strains isolated from arthritic lesions found in slaughtered pigs were segregated into 4 major patterns: 1) identical nucleotide sequence with the vaccine strain: 3 isolates; 2) 1 nucleotide substitution (C to A) at position 555: 5 isolates; 3) 1 nucleotide substitution at various positions: 5 isolates; and 4) 2 nucleotide substitutions: 3 isolates. Isolates with the same nucleotide sequence as the vaccine strain were further characterized by other properties, including the mouse pathogenicity test. One strain isolated from pigs on a farm where the live vaccine had been used was found to be closely related to the vaccine strain. The phylogenetic tree constructed based on the spaA sequence suggests that the evolutionary distance of the isolates is related to the pathogenicity in mice. The new strain-typing system based on nucleotide sequencing of the spaA region is useful to discriminate the vaccine strain from field isolates.
Erysipelothrix rhusiopathiae is a small Gram-positive rod bacterium that causes erysipelas in swine and a variety of diseases in other animals, as well as erysipeloid, a skin disease of humans. 19 Since swine erysipelas reappeared as a clinical problem in pigs in the Midwestern United States, it has been considered as a reemerging disease that contributes substantially to economic losses in the swine industry. 12 In Japan, an attenuated strain, Koganei 65–0.15, which was obtained by 65 passages on agar media containing 0.15% of an acriflavine dye, has been used as a seed strain for production of a live vaccine. 14 Since commercializing and distributing this live vaccine nationwide, outbreaks of acute septicemia or subacute urticaria of erysipelas have dramatically decreased, 16 thus demonstrating the efficacy of this live vaccine. In contrast, a chronic form of erysipelas, including polyarthritis and endocarditis, found during meat inspections in slaughterhouses, has been increasing. According to the abattoirs law of Japan, a carcass with a lesion of the chronic form of erysipelas from which E. rhusiopathiae could be isolated, must be entirely condemned. Therefore, the economic losses caused by the chronic form of erysipelas are not negligible. A previous report 2 describes that E. rhusiopathiae strains isolated from chronic arthritis are closely related to the vaccine strain, arousing suspicion that the live vaccine strain itself may partly be involved in the occurrence of the chronic form of erysipelas.
To address this issue, the development of a new technique that can differentiate the vaccine strain from other field strains is needed. Because the vaccine strain Koganei 65–0.15 is resistant to acriflavine during its growth on agar media, this property could be used as a marker of the vaccine strain, but because of the presence of spontaneous acriflavine-resistant Erysipelothrix strains in the field, 5 this property may not be a reliable marker for the vaccine strain. Differentiation of Erysipelothrix strains can be achieved by DNA fingerprinting 5,10 or pulse-field gel electrophoresis 11,12 ; however, discrimination of the vaccine strain from field isolates cannot be accomplished by these methods. Using several genetic typing techniques, including randomly amplified polymorphic DNA (RAPD) typing, ribotyping, and restriction fragment length polymorphism (RFLP) in combination with acriflavine resistance and mouse pathogenicity tests, field isolates of E. rhusiopathiae have been classified into several groups. 2 A RAPD 1–2 type of E. rhusiopathiae serovar 1a may prove to be the vaccine strain, 2 and it has been suggested that some of the cases of chronic swine erysipelas in Japan can be ascribed to a side effect of the use of a live vaccine.
In the present study, a new strain-typing method was developed based on the nucleotide sequencing of a hypervariable region in the surface protective antigen (spaA) gene, which has recently been revealed to have variations among Erysipelothrix serovar strains. 18 A total of 16 E. rhusiopathiae isolates originating from slaughtered pigs with arthritis were used in the present study. Two strains were isolated from pigs raised in farms A and B, and 6 strains each were isolated from farms C and D. The farms were located in the Chubu area in central Japan. These strains were isolated between May 2001 and August 2002. Pigs from farms A, B, and D had not received any live or killed vaccine against E. rhusiopathiae. Pigs from farm C were given an attenuated live vaccine.
A live vaccine strain, Koganei 65–0.15, and a virulent wild-type strain, Fujisawa, were used as reference strains in an acriflavine resistance test and a pathogenicity test in a mouse model. Vector plasmid pGEM-T Easy a was used to clone the spaA gene. Escherichia coli XL1-Blue was used as a host strain for replication of the vector plasmid. Erysipelothrix strains were grown in tryptose phosphate broth b supplemented with 1% proteose peptone 3 c and 0.1% Tween 80 d (pH 7.8). The E. coli strain was grown in Luria Bertani medium. When appropriate, the medium was supplemented with ampicillin (100 μg/ml).
The procedure used to determine the serovars of the isolates was similar to that described previously 3 with some minor modifications. Briefly, colonies of each strain grown for 48 hr on agar plates were harvested and suspended in saline. Bacterial cells were washed 3 times with saline and suspended in distilled water. The cell suspension was autoclaved for 1 hr at 121°C and clarified by centrifugation. Rabbit antisera against all known serovars, 1–23 and type N, was made in the authors' institute. The heat-extracted solution of the bacterial cells was probed with the rabbit antisera against all serovars of E. rhusiopathiae in 1% Noble agar with 150 mM of sodium chloride−10 mM phosphate buffer (pH 7.2). After reacting for 24 hr at room temperature, formation of the precipitation line was observed.
The acriflavine resistance test of E. rhusiopathiae isolates was performed by inoculation of the isolates on agar plate supplemented with 0.01% of acriflavine. e The growth of the colonies of each isolate was recorded after incubation at 37°C for 48 hr.
In previous work by the authors, it was found that the N-terminal half of the spaA gene, which has been shown to be involved in immunoprotection, is highly variable in both nucleotide and deduced amino acid sequences. 18 The following polymerase chain reaction (PCR) primers, which are located at the conserved sequence among the published nucleotide sequences of the spaA gene from serovars 1a, 1b, 2, 5, 8, 9, 12, 15, 16, 17, and N but are located across a hypervariable sequence, were designed. Erko-1F (5′-GTGAAACACCGTATTTTAGTA-3′) and Erko−2R (5′-TTCAAGAAGTTCCTGTAGTTT-3′) are located at nucleotide positions 502 to 522 and 933 to 913, respectively, of the spaA gene.
Chromosomal DNA of E. rhusiopathiae was prepared as previously described. 18 Polymerase chain reaction was performed as described elsewhere 9 under the following conditions: a denaturation step at 94°C for 3 min and then 25 cycles of denaturation at 94°C for 45 sec; annealing at 55°C for 1 min and extension at 72°C for 2 min; and extension of 5 min for the final cycle. Thermostable DNA polymerase f with high proofreading activity (3′-5′ exonuclease activity) was used. The PCR product was ligated into the plasmid pGEM-T Easy, and then transformed into E. coli XL-1 Blue by electroporation g at a setting of 2.5 kV, 25 mF, and 400Ω. Both strands of DNA of the cloned PCR product were sequenced on a genetic analyzer h using a kit. i The sequencing reactions were performed with M13 vector primers. At least 3 plasmid clones were sequenced, and finally, the PCR products were directly sequenced to confirm the accuracy of the data. The nucleotide sequence was analyzed with programs GENETYX-MAX j and SeqMan. k Alignments and phylogenetic tree analysis of the highly variable region of the spaA gene were performed with the CLUSTAL W program 17 and the UPGMA method using the MEGA program, 4 respectively.
Information on the results of RAPD typing and ribotyping of the 16 E. rhusiopathiae isolates was provided by Dr. Yumiko Imada (National Institute of Animal Health, Tsukuba, Japan). These typing procedures have been described elsewhere. 2 In the previous study, 492 E. rhusiopathiae isolates of serovar 1 and 2 showed 7 major banding patterns (RAPD types of 1, 1–1, 1–2, 1–4, 1–6, 2–4, and 12). The RAPD type of the isolates was determined by comparing the given banding pattern with those of the 7 RAPD types. A RAPD 1–2 type was suggested to be the marker of the vaccine strain. As for the ribotyping, 524 E. rhusiopathiae isolated showed 7 major ribotypes (1, 2, 3, 6, 7, 8, and 11), and the vaccine strain exhibited ribotype 1.
Nucleotide substitutions in a hypervariable region on the spaA gene of 16 Erysipelothrix rhusiopathiae field strains isolated from slaughtered pigs with chronic arthritis compared with the corresponding sequence of the live vaccine strain Koganei 65–0.15.
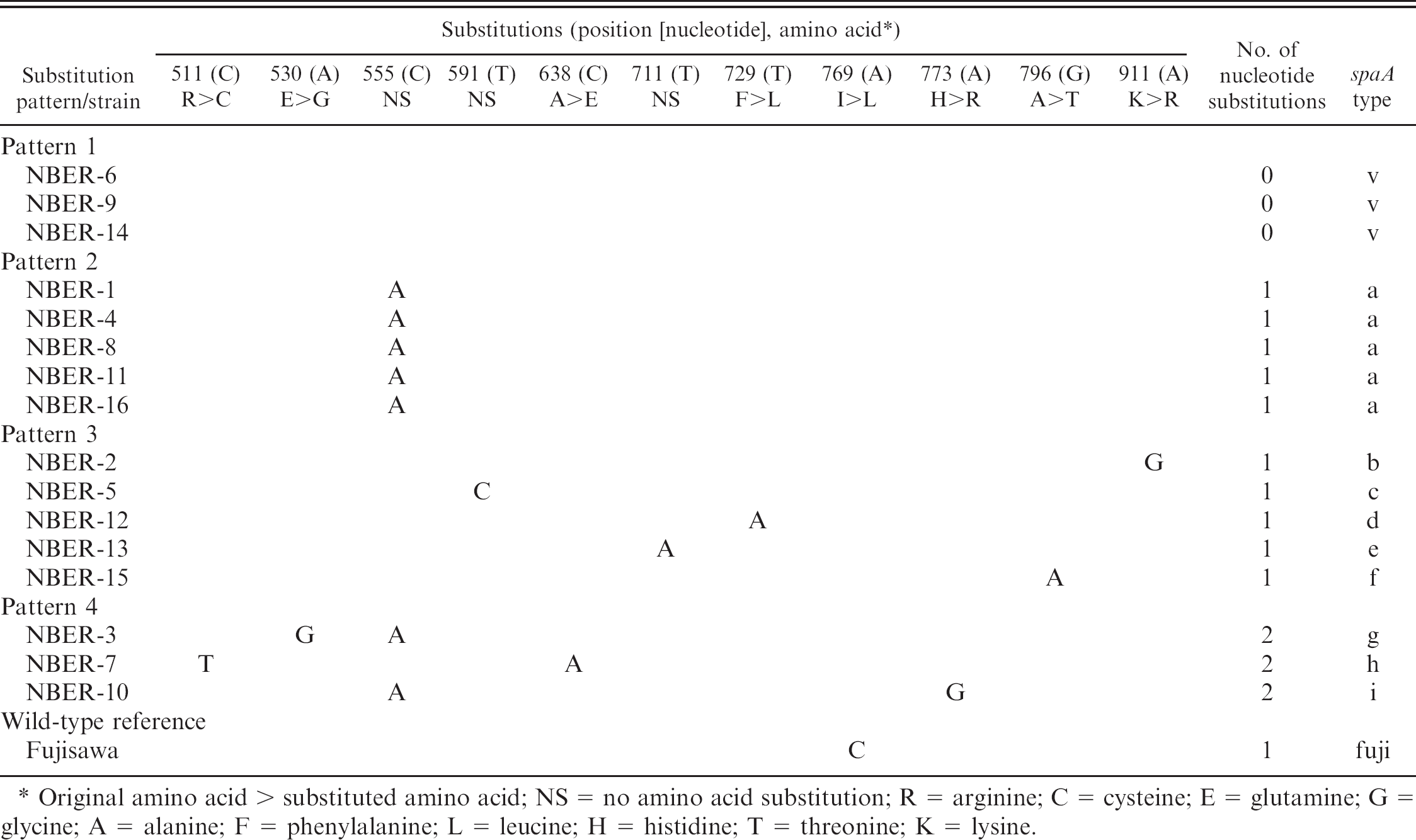
Original amino acid > substituted amino acid; NS = no amino acid substitution; R = arginine; C = cysteine; E = glutamine; G = glycine; A = alanine; F = phenylalanine; L = leucine; H = histidine; T = threonine; K = lysine.
The mouse pathogenicity test was carried out according to the procedure described in the Japanese Standard of Veterinary Biological Products. Briefly, 10, 4-week-old female ddY mice l were injected subcutaneously in the right inguinal region with 0.1 ml of approximately 10 7 colony-forming units (CFU) of the isolate or the reference strain. A group of 10 mice served as a nontreated control. Mice were observed daily to detect clinical signs of the disease for the subsequent 10 days. The standard provides that the vaccine strain will not kill mice but will induce arthritis in more than 80% of animals. For classification of virulence levels of E. rhusiopathiae strains, the isolate that caused mortality in more than 1 of 10 mice was considered to have high virulence, and the isolate that caused no mortality and induced arthritis in less than 8 of 10 mice was considered to have low virulence. The animals used in the present study were cared for in accordance with the guidelines for animal treatment of Nippon Institute for Biological Science, which conform to the standard principles of laboratory animal care.
A comparison was made of the nucleotide sequences of the spaA gene from various E. rhusiopathiae strains that were available in the EMBL/GenBank/DDBJ database, and most of the nucleotide substitutions were found to be located in the N-terminal half of the gene. Thus, an attempt was made to analyze the nucleotide sequences of this hypervariable region at positions 502 to 933 nucleotides (432 bp) of the spaA gene from the 16 E. rhusiopathiae strains isolated from slaughtered pigs with arthritis. The different nucleotide sequences in the spaA region of the 16 field isolates and the wild-type reference strain Fujisawa compared with the corresponding sequence of the attenuated live vaccine strain Koganei 65–0.15 are shown in Table 1. Three of the 16 isolates showed an identical nucleotide sequence with that of the vaccine strain in the analyzed genetic region (pattern 1, spaA-type v). Ten of the 16 isolates showed 1 nucleotide difference from that of the vaccine strain. In strains with a 1-nucleotide difference, 5 isolates had the same nucleotide substitution at position 555 where C was changed to A (pattern 2, spaA-type “a”). The remaining 5 isolates with the 1-nucleotide difference had a substitution at various positions in the spaA region (pattern 3, spaA-type “b,” “c,” “d,” “e,” and “f”). Three of the 16 isolates had 2 nucleotide substitutions at various positions (pattern 4, spaA-type “g,” “h,” and “i”); however, 1 of the 2 substitutions was located at nucleotide position 555, where C was changed to A, in 2 of the 3 isolates. The wild-type reference strain Fujisawa had 1-nucleotide substitutions at position 769 (spaA-type “fuji”).
Of the 11 positions of nucleotide substitutions shown in Table 1, 8 (positions 511, 530, 638, 729, 769, 773, 796, and 911) induced amino acid exchange in the SpaA protein, but the remaining 3 (positions 555, 591, and 711) did not induce the change of amino acids.
spaA types and other biological and genetic properties of 16 Erysipelothrix rhusiopathiae isolates compared with the live vaccine strain.
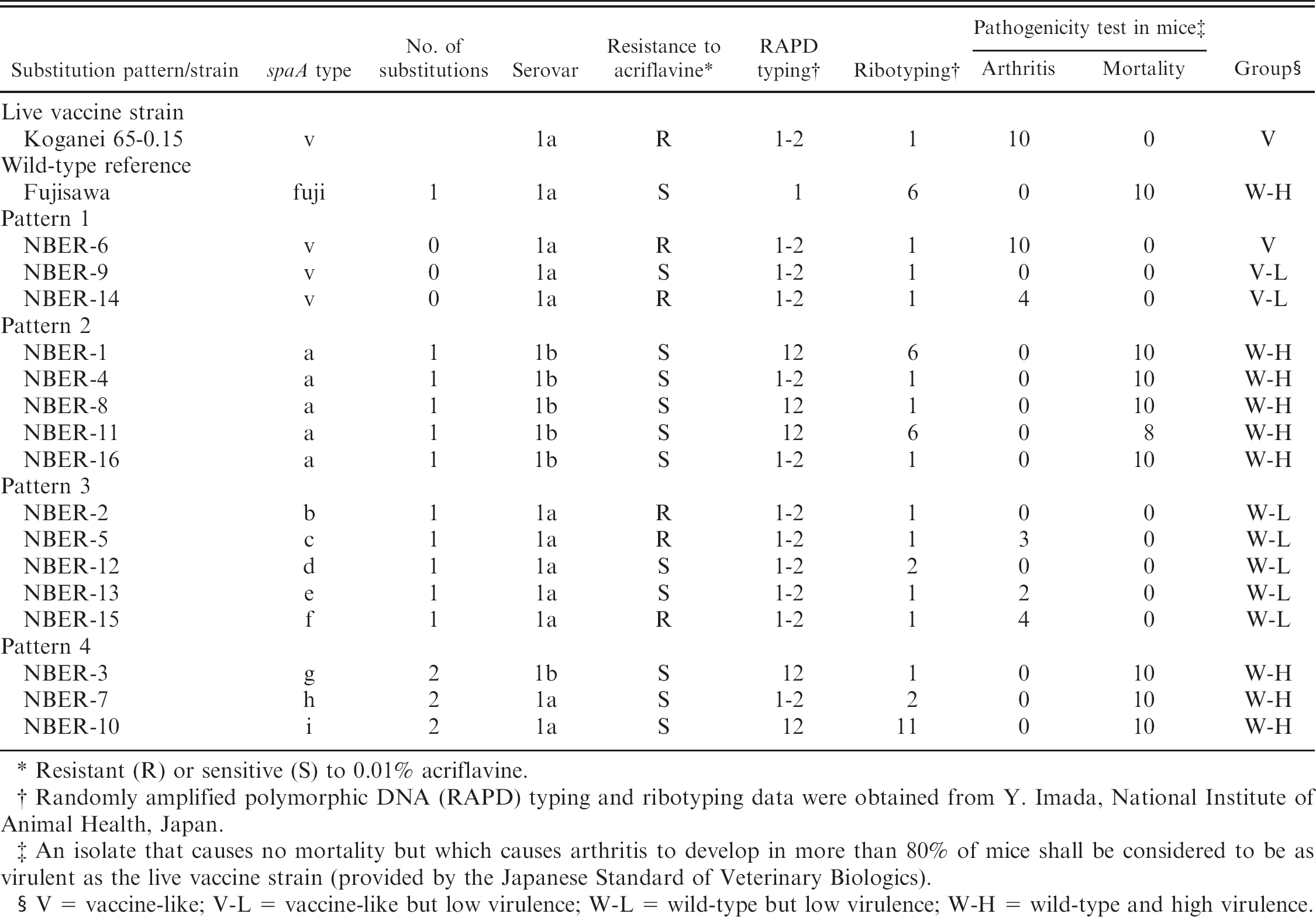
Resistant (R) or sensitive (S) to 0.01% acriflavine.
Randomly amplified polymorphic DNA (RAPD) typing and ribotyping data were obtained from Y. Imada, National Institute of Animal Health, Japan.
An isolate that causes no mortality but which causes arthritis to develop in more than 80% of mice shall be considered to be as virulent as the live vaccine strain (provided by the Japanese Standard of Veterinary Biologics).
V = vaccine-like; V-L = vaccine-like but low virulence; W-L = wild-type but low virulence; W-H = wild-type and high virulence.
The spaA types as well as other biological and genetic properties, including serotyping, resistance to acriflavine, RAPD, ribotyping, and pathogenicity in mice, of the 16 isolates are listed in order of spaA types (Table 2). The 3 isolates showing spaA-type v (substitution pattern 1) belonged to serovar 1a and had a RAPD pattern 1–2 and a ribotyping pattern 1. Two (NBER-6 and 14) of the 3 isolates were resistant to acriflavine at a concentration of 0.01%; however, the remaining isolate (NBER-9) was sensitive to acriflavine. In the pathogenicity test in mice, none of the isolates caused mortality; however, 2 of the 3 isolates induced arthritis in 10 (NBER-6) and 4 (NBER-14) mice. Accordingly, only 1 isolate, NBER-6, was serologically, genetically, and biologically identical to the vaccine strain. NBER-6 was tentatively assigned to the V (vaccinelike) group. Isolates NBER-9 and NBER-14 were serologically and genetically similar to the vaccine strain; however, their virulence in mice was lower than that of the vaccine strain. NBER-9 and NBER-14 were tentatively assigned to the V-L (vaccine-like but low virulence) group.
The 5 isolates showing spaA-type a (substitution pattern 2) belonged to serovar 1b and were sensitive to acriflavine. Their RAPD pattern was 12 or 1–2, and the ribotyping pattern was 6 or 1. In the pathogenicity test in mice, the isolate NBER-11 caused mortality in 8 of 10 mice, and the other 4 isolates caused mortality in all mice. From this data, the isolates were tentatively assigned to the W-H (wild-type and high virulence) group.
Isolates having 1 nucleotide substitution at various nucleotide positions in the spaA gene (substitution pattern 3, spaA-type b, c, d, e, and f) belonged to serovar 1a and showed a RAPD pattern 1–2 and ribotyping pattern of 1 or 2. Three of the 5 isolates were resistant to acriflavine, and the remaining 2 were sensitive to acriflavine. These isolates did not cause mortality in mice, but 3 of 5 isolates induced arthritis in 3 (NBER-5), 2 (NBER-13), or 4 (NBER-15) of the 10 mice. Thus, isolates showing these spaA types were assigned to the W-L (wild-type but low virulence) group.
Isolates with 2 nucleotide substitutions in the spaA gene, showing spaA-type g, h, and i (substitution pattern 4), belonged to serovar 1b or 1a, and showed a RAPD pattern 12 or 1–2 and ribotyping pattern 1, 2, or 11. These isolates were sensitive to acriflavine. They caused mortality in all of the inoculated mice, indicating that they were more virulent in mice than the vaccine strain. These isolates with 2 nucleotide substitutions were tentatively assigned to the W-H (wild-type and high virulence) group.
The wild-type reference strain Fujisawa had 1 nucleotide substitution in the spaA region, belonged to serovar 1a, was sensitive to acriflavine, showed RAPD pattern 1 and ribotyping pattern 6, and caused mortality in all of the inoculated mice. Thus, the Fujisawa was assigned to the W-H group. The 16 isolates and their biological and genetic properties are listed in order of the farms where they were originally isolated (Table 3). The E. rhusiopathiae isolates assigned to group V and V-L were isolated only from farm C, where the live vaccine against erysipelas had been regularly used. In farms A, B, and D, where the live vaccine against erysipelas was not used, most of the isolates were assigned to the W-H group.
Biological and genetic properties of the 16 Erysipelothrix rhusiopathiae isolates sorted according to the farm where the strain was originally isolated.
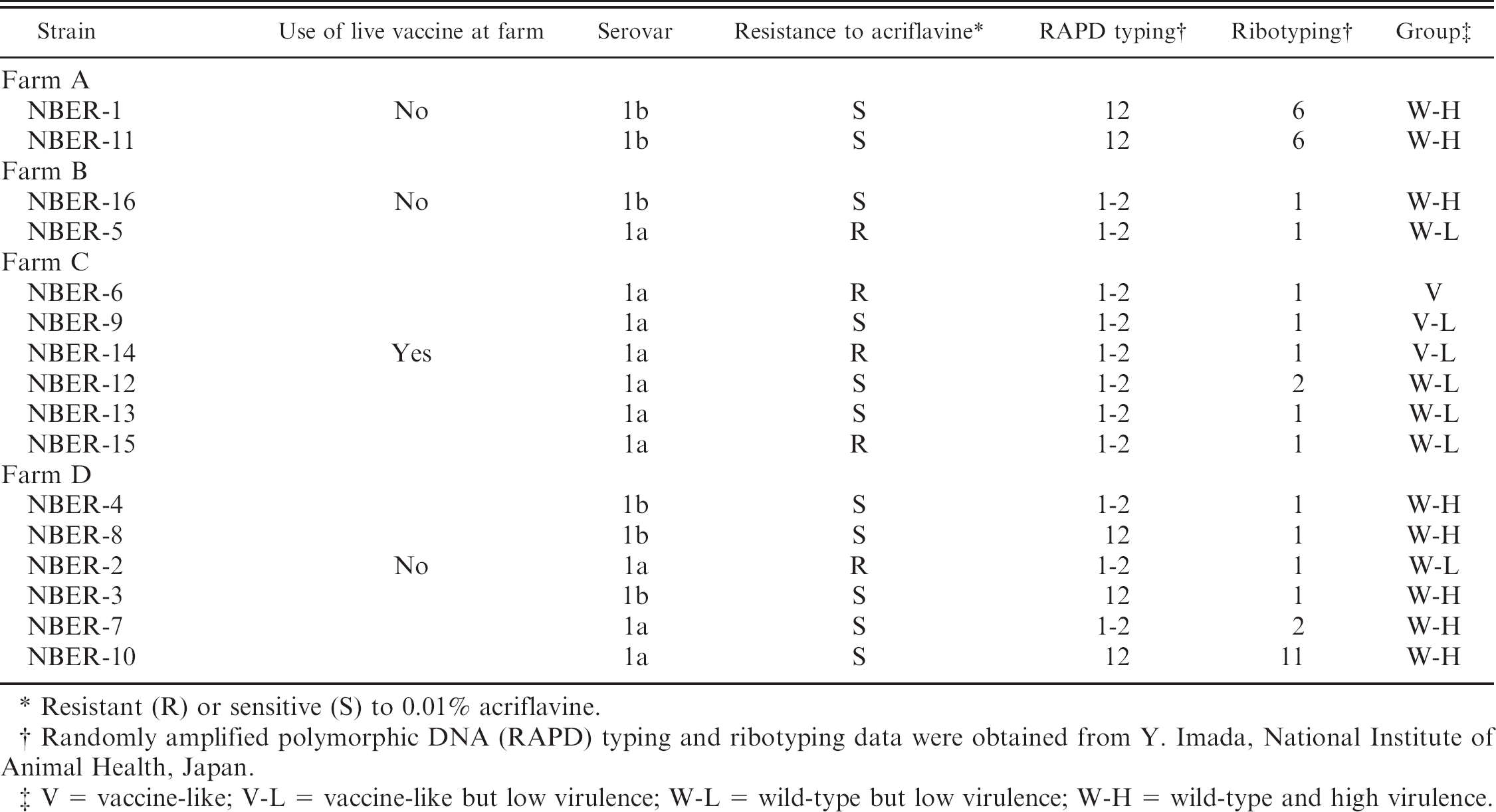
Resistant (R) or sensitive (S) to 0.01% acriflavine.
Randomly amplified polymorphic DNA (RAPD) typing and ribotyping data were obtained from Y. Imada, National Institute of Animal Health, Japan.
V = vaccine-like; V-L = vaccine-like but low virulence; W-L = wild-type but low virulence; W-H = wild-type and high virulence.
A phylogenetic tree was constructed based on the nucleotide sequence of the hypervariable region in the spaA gene (Fig. 1). The farm where the strain was originally isolated, the spaA type, and a group based on the pathogenicity in mice, are also noted in the tree. The isolates from farm C, where the live vaccine was routinely used, were located on the same branch as the vaccine strain (NBER-6, NBER-9, and NBER-14) or on nearby branches (NBER-13 and NBER-12). Interestingly, the tree branched mainly into 2 stems near the base: 1 included isolates assigned to V, V-L, and W-L groups, and the other included isolates assigned to the W-H group. The nucleotide substitution at position 555 was the key nucleotide in making the phylogenetic tree branch into 2 stems, each of which included high or low virulence isolates. The 7 of 8 field isolates that were assigned to the W-H group had the substitution of C to A at position 555 (spaA-types a, i, and g). This indicates that the pathogenicity in mice and phylogenetic systems based on analysis of the hypervariable region in the spaA gene are closely related among E. rhusiopathiae strains, which were originally isolated from herds located in this area of Japan.
Because concern arose that a recent increase in chronic arthritis caused by E. rhusiopathiae infection might be ascribed to the use of a live vaccine, 2 discrimination of strains in serovar 1a of E. rhusiopathiae has become an important issue. The live vaccine strain Koganei 65–0.15 used in Japan was obtained by 65 passages on an agar plate containing 0.15% acriflavine dyes. Thus, resistance to acriflavine appears to be a suitable marker of the vaccine strain; however, acriflavine resistance was found to be lost by passage in medium without an acriflavine, 8 and spontaneous acriflavine-resistant strains are sometimes isolated from unvaccinated pigs. 5 In the present study, there existed an acriflavine-sensitive strain showing spaA-type v, and an acriflavine-resistant strain was isolated from a pig in a herd that was not given the live vaccine. As a result, acriflavine resistance seems to be an unsuitable marker for discriminating the live vaccine strain from other E. rhusiopathiae field isolates.
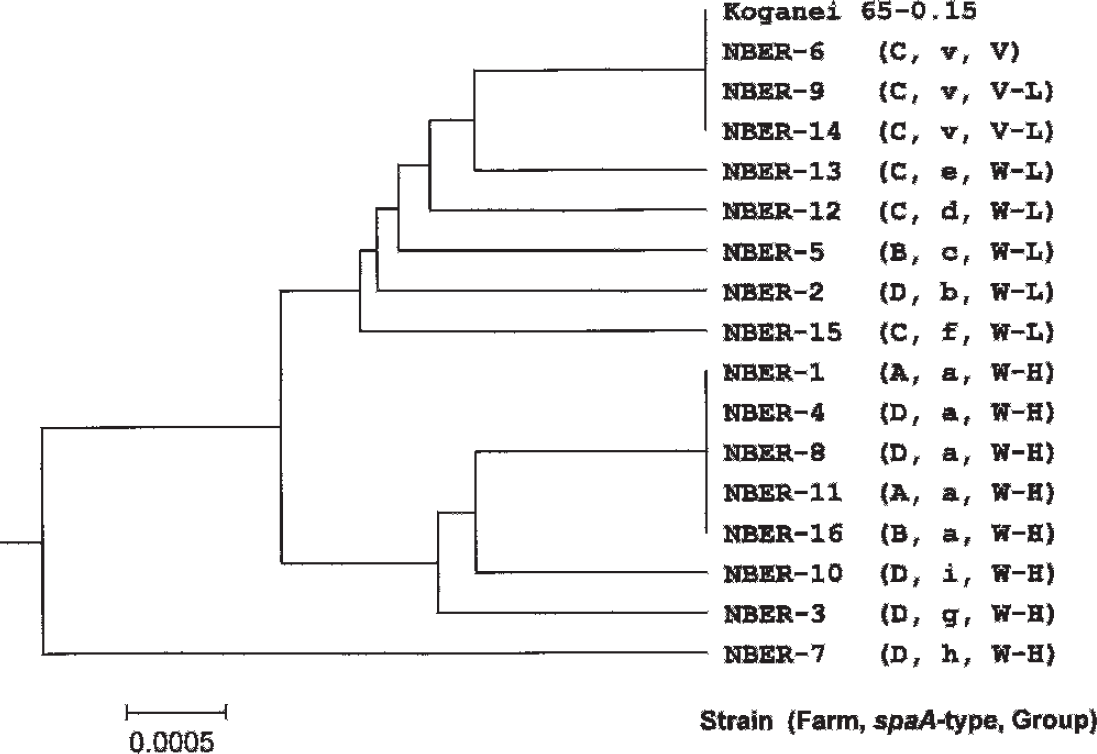
A neighbor-joining tree produced using the nucleotide sequence of a hypervariable region in the spaA gene of Erysipelothrix rhusiopathiae isolates. The name of the isolate (NBER-1 to 16), the farm where the strain was originally isolated (A, B, C, or D), the spaA type (v, a, b, c, d, e, f, g, h, or i), and the group (V, V-L, W-L, or W-H) to which the relevant isolate was assigned based on the genetic properties and virulence in mice, are indicated. V = vaccine-like; V-L = vaccine-like but low virulence; W-L = wild-type but low virulence; W-H = wild-type and high virulence.
Spa protein is synthesized in the bacterial cellular cytoplasm, secreted outside of bacterial cells, and then bound to the bacterial cell surface by a special anchoring mechanism common to Gram-positive bacteria. 6 Spa protein is considered to be involved in the virulence of Erysipelothrix strains. 1,7,15,18 The genetic region analyzed in the present study is located at the domain that is involved in immunoprotection in the SpaA molecule. The present analysis suggested that the nucleotide substitution of C to A at position 555 may link to the high-virulence phenotype of the isolates. However, because this nucleotide substitution did not induce amino acid exchange in the SpaA protein and because the virulent wild-type reference strain Fujisawa did not have this mutation at the nucleotide position 555, it is probable that the high-virulence phenotype could not be ascribed to this mutation in the spaA gene. Further studies are needed to clarify whether the difference in virulence level of each strain would relate to variation in other domain of the SpaA protein or variation in other unknown virulence factors.
The 3 isolates showing an identical nucleotide sequence to the vaccine strain were isolated from herds where the live vaccine had been used; however, 2 of the 3 isolates showed low virulence in mice, which does not meet the criteria for the live vaccine strain described in the Japanese Standard of Veterinary Biological Products. The 2 isolates cannot, therefore, be identified as a vaccine strain, but the possibility that the vaccine strain may lose virulence in mice during passage in the swine population cannot be dismissed. One isolate (NBER-6), which was genetically and biologically identical to the vaccine strain, seemed to be closely related to the live vaccine strain. In a recent study, 18 strains isolated from arthritic lesions in pigs at slaughter were analyzed, and 1 isolate was found to be very similar to the vaccine strain. 13 The rate of detection of the vaccine-related strain in the previous study was very similar to the present study.
The most important advantage of the present procedure for discrimination of the E. rhusiopathiae strain is that there is no ambiguity in the results obtained. Nucleotide sequence analysis affords the same results irrespective of the laboratory involved. As most clinical laboratories in Japan are equipped with an auto-DNA sequencer, the survey and characterization of E. rhusiopathiae strains by nucleotide sequence analysis can be easily performed. In addition, as the number of nucleotides to be determined in this procedure is confined to 432 bp, both DNA strands of this genetic region can be determined in only 2 reactions. In this study, for the purpose of accuracy of the sequencing results, the authors sequenced the genetic region using the cloned DNA fragment and, afterwards, directly sequenced the amplified fragment to confirm the substitutions. When this technique is applied to analyze large number of isolates, it should be easier to perform by direct sequencing only.
In summary, the following procedure to detect vaccine-related strains is recommended. The first screening should be by serotyping (serovar 1a) and RAPD analysis (type 1–2). The second screening should be by sequence analysis of the spaA gene (spaA-type v). Finally, the candidates on the short list should be inoculated into mice to see whether they meet the category of the vaccine strain as described in the Japanese Standard of Veterinary Biological Products. A large and nationwide survey using the above procedure may allow diagnosticians to clarify whether the vaccine strain can induce chronic arthritis in swine.
Acknowledgement. The authors thank Dr. Yumiko Imada, National Institute for Animal Health, Tsukuba, Japan, for conducting RAPD and ribotyping of the E. rhusiopathiae isolates.
Footnotes
a.
pGEM-T Easy, Promega, Madison, WI.
b.
Tryptase phosphate broth, Difco Laboratories, Detroit, MI.
c.
Proteose peptone No. 3, Difco Laboratories, Detroit, MI.
d.
Tween 80, Kanto Chemical Co., Inc., Tokyo, Japan.
e.
Acriflavine, Wako Pure Chemical Industries, Osaka, Japan.
f.
Expand High Fidelity, Boehringer Mannheim Biochemicals, Indianapolis, IN.
g.
Gene Pulser and Pulse Controller, Bio-Rad, Richmond, CA.
h.
ABI PRISM 310 genetic analyzer, Applied Biosystems, Foster City, CA.
i.
Big Dye Terminator cycle sequencing kit, Applied Biosystems, Foster City, CA.
j.
GENETYX-MAX programs ver. 12, SDC, Tokyo, Japan.
k.
SeqMan program of the DNA star software package, DNASTAR Inc., Madison, WI.
l.
Female ddY mice, Nippon SLC, Shizuoka, Japan.
