Abstract
Moraxella oris was historically the only coccoid Moraxella identified in cultures of ocular fluid from cattle with infectious bovine keratoconjunctivitis (IBK) and could be morphologically and biochemically differentiated from Moraxella bovis. Moraxella bovoculi sp. nov. is a recently characterized Moraxella isolated from ulcerated eyes of calves with IBK in northern California in 2002. Like Moraxella ovis, M. bovoculi sp. nov. is a gram-negative coccus/diplococcus. All 18 original isolates of M. bovoculi sp. nov. possessed phenylalanine deaminase (PADase) activity and could therefore be differentiated from M. ovis and M. bovis. During the characterization of 44 additional isolates of hemolytic gram-negative cocci that were cultured from ulcerated eyes of IBK-affected calves, 2 PADase-negative isolates were identified that could not be differentiated biochemically from M. ovis; however, the DNA sequence of the 16S-23S intergenic spacer region (ISR) of the isolates matched the 16S-23S ISR DNA sequence of M. bovoculi sp. nov. To facilitate the identification of PADase-negative moraxellae, a polymerase chain reaction (PCR) coupled with restriction enzyme digestion analysis of amplified DNA was developed. Amplification of the 16S-23S ISR followed by AfaI digestion of amplified DNA could differentiate M. bovoculi sp. nov. from M. ovis and other moraxellae. The DNA sequence analysis of the amplified 16S-23S ISR from the 42 PADase-positive isolates of hemolytic gram-negative cocci indicated that all were M. bovoculi sp. nov. and all possessed an AfaI site. A PCR coupled with restriction analysis of amplified DNA can aid in identifying M. bovoculi sp. nov.
Infectious bovine keratoconjunctivitis (IBK; pinkeye) is the most common ocular disease of cattle; affected animals exhibit corneal ulceration leading to corneal scarring of varying severity. Koch's postulates have been satisfied for Moraxella bovis and IBK. 12 Pathogenic proteins of M. bovis include pilin that enable the organism to attach to corneal epithelial cells 4,16,18 and a pore-forming cytotoxin 9 (cytolysin; hemolysin) that lyses corneal epithelial cells 7 and host neutrophils resulting in a release of neutrophil-derived degradative enzymes into corneal stroma. 14,15,20 The M. bovis cytotoxin is an RTx (repeats in the structural toxin) toxin designated MbxA encoded by an RTxA gene (mbxA) 1 that is contained within an RTx operon (mbx operon) 2 that resides on a pathogenicity island in M. bovis. 13
Along with M. bovis, other gram-negative bacteria have been identified in ocular fluid from the eyes of normal and IBK-affected cattle. These bacteria have variously been designated hemolytic diplococci, 19 hemolytic Neisseria, 11 Neisseria spp., 5,6,21 Neisseria (Branhamella) catarrhalis, 22 or Moraxella ovis. 8,10 During an IBK antibiotic efficacy trial conducted in Northern California during the summer of 2002, hemolytic gram-negative cocci isolated from the ulcerated eyes of dairy and beef calves were further characterized. DNA sequence analyses and biochemical testing demonstrated that 18 characterized isolates were distinct from M. bovis and M. ovis and as such represented a novel species that was named Moraxella bovoculi sp. nov., within the genus Moraxella? M. bovoculi sp. nov. could be differentiated from M. bovis and M. ovis as well as other moraxellae isolated from animals on the basis of positive phenylalanine deaminase (PADase) activity.
During an IBK vaccine trial conducted during the summer of 2005, ulcerated eyes of calves were cultured, and a total of 44 additional isolates of hemolytic gram-negative cocci were characterized. Forty-two were PADase positive and were identified as M. bovoculi sp. nov.; however, 2 PADase-negative isolates (120L and 159R) could not be readily differentiated from M. ovis on the basis of PADase activity. To facilitate the molecular identification of PADase-positive or -negative hemolytic gram-negative coccoid moraxellae, a polymerase chain reaction (PCR) coupled with restriction enzyme analysis was developed.
Moraxella bovoculi sp. nov. (44 isolates), M. ovis, a M. bovis Tifton I, b Moraxella caprae, c and Moraxella boevrei d were propagated on 5% sheep blood agar e at 35°C; strains were stored frozen in 10% skim milk-50% glycerol at −80°C. To prepare genomic DNA, 1 loopful of bacteria was scraped from plates and resuspended in 500 μl of sterile deionized water, and genomic DNA was harvested according to the manufacturer's instructions. f Primers for amplifying the 16S-23S intergenic spacer region (ISR) were developed from the DNA sequence of the 16S rDNA gene, the 16S-23S ISR, and a portion of the 23S rDNA gene from M. bovis, M. ovis, and M. bovoculi sp. nov. 3 The PCR primers used were ISRdown (5‘-gtg aag tcg taa caa ggt agc cgt-3’) and ISRup (5‘-acc gac gct tat cgc agg cta tca-3’). The PCR reactions were made up to a final volume of 50 μl with ultra pure water 8 and contained 3 μl of genomic DNA template, 0.4 μl (2U) Taq DNA polymerase, 11 5 μl of 10− PCR buffer (the final 1− buffer was 1.5 mM MgCl2, 1.5 mM MgSO4, 10 mM KCl, 8 mM [NH4]2SO4, 10 mM Tris-HCl [pH 9.0], and 0.05% NP-40), 0.5 μl of 10 mM dNTP, i and 2.5 μl of each primer (10 pmole/μl). The reactions were run in a thermo-cycler j using an initial denaturation at 95°C for 1 minute, followed by 35 cycles of 95°C for 30 seconds, 50°C for 30 seconds, and 72°C for 30 seconds. Fifteen microliters of the resulting PCR reaction was mixed with 0.5 μl (4U) of AfaI k and incubated at 37°C for 1 hour. Following incubation, 5 μl of the original PCR reaction and 5 μl of the AfaI digested PCR reaction were electrophoresed on a 2% agarose gel l containing 0.25 μg/ml of ethidium bromide m in 1− TAE buffer (0.04 M Tris-acetate; 0.001 M EDTA). The gel was visualized on an ultraviolet lightbox 11 and photographed. o The ISRdown-ISRup amplicon generated from genomic DNA was sequenced directly with primers ISRdown and ISRup. p
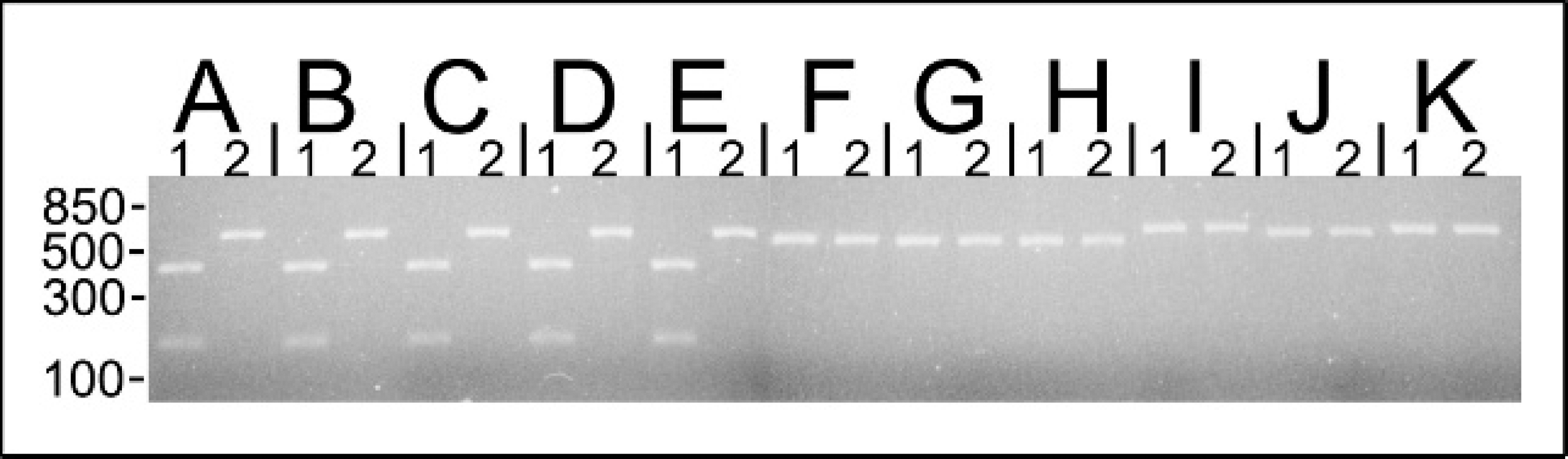
The genomic DNA from Moraxella bovoculi sp. nov. 38L (lane A), 117L (lane B), 120L (lane C), 159R (lane D), 237 (lane E), Moraxella ovis ATCC 33078 (lane F), Moraxella ovis ATCC 19575 (lane G), Moraxella ovis ATCC 19576 (lane H), Moraxella bovis Tifton I (lane I), Moraxella boevrei ATCC 700022 (lane J), and Moraxella caprae ATCC 700019 (lane K) was amplified with primers flanking the 16S rDNA-23S rDNA intergenic spacer region. Following amplification, an aliquot of the PCR reaction product was digested with AfaI at 37°C for 1 hour. Five microliters of restriction digested (lane 1) or undigested (lane 2) DNA was electorphoresed on a 2% agarose gel. The ∼600-bp amplified product from PADase-negative (isolates 120L and 159R) and PADase-positive (isolates 38L, 117L, and 237) M. bovoculi sp. nov. was cut once by AfaI into ∼450- and ∼150-bp fragments. The amplicon generated for the other Moraxella spp. were not cut by AfaI. Approximate molecular mass markers are indicated in bp.
ISRdown-ISRup amplified DNA exhibited a single band of ∼600 bp for M. bovoculi sp. nov. and M. ovis and ∼650 bp for M. bovis, M. boevrei, and M. caprae (Fig. 1). AfaI digestion of amplified DNA cut the ∼600 bp amplicon from M. bovoculi sp. nov. at 1 site, resulting in 2 bands of ∼450 and ∼150 bp. The original amplicons generated for M. bovis, M. ovis, M. boevrei, and M. caprae were not cut by AfaI and remained as a single band (Fig. 1). For the 44 isolates examined, the DNA sequence of the 16S–23S ISR amplified by the ISRdown-ISRup primers matched the previously reported 16S–23S intergenic spacer sequence for M. bovoculi sp. nov. (Genbank accession DQ153085). 3
The method described above for identifying hemolytic gram-negative cocci that are PADase negative as M. bovoculi sp. nov. is rapid and can be accomplished within a single day following colony purification and genomic DNA extraction. While the method described above was useful for differentiating PADase-negative M. bovoculi sp. nov. from M. ovis, it might also be useful in identifying hemolytic gram-negative cocci cultured from eyes of cattle in the absence of a PADase test result. The DNA sequencing results for the amplified ISR region in all 44 isolates of what were identified as M. bovoculi sp. nov. indicated that the region is well conserved and all 44 isolates possessed an AfaI site. While the selected primers amplified DNA from all 5 Moraxella spp., the restriction enzyme was chosen to recognize a site that was present only in the amplicon generated for M. bovoculi sp. nov. It is possible that mutations could lead to misidentification of M. bovoculi sp. nov. if PCR and restriction analysis were the only tests used for identification. Nevertheless, the 44 additional isolates of M. bovoculi sp. nov. from summer 2005 all had 16S–23S ISR DNA sequences that matched previously reported 16S–23S ISR DNA sequences of M. bovoculi sp. nov., and all had an AfaI site in the amplified ISR region.
During the original characterization of M. bovoculi sp. nov., the M. ovis chosen for comparison was a type strain of M. ovis (ATCC 33078) that was originally cultured from a sheep with keratoconjunctivitis. 17 Other species of gram-negative cocci have been isolated from cattle with IBK, and these have historically been referred to as Moraxella, Branhamella, or Neisseria spp./ovis. 5,6,8,10,11,21,22 It is possible that some agents historically identified as M. ovis might have fit the description of M. bovoculi sp. nov., and as such, future descriptions of bacteria identified as M. ovis or M. bovoculi sp. nov. should conclusively demonstrate speciation of these isolates. While biochemical testing has traditionally been used to differentiate moraxellae, a determination of the 16S–23S ISR DNA sequence may help to speciate the coccoid moraxellae, especially PADase-negative coccoid moraxellae isolated from cattle with IBK. Future studies on more geographically diverse isolates of M. bovoculi sp. nov. should help elucidate whether additional genetic and biochemical variability exists among isolates of this species. Until additional M. bovoculi sp. nov. are characterized at the molecular level, it may be most informative to use a combination of morphologic, biochemical, and molecular analyses to positively identify Moraxella bovoculi sp. nov.
Acknowledgement. Funding for completion of this work was provided by the University of California School of Veterinary Medicine Dean's Office Start Up Funds for New Faculty.
Footnotes
a.
ATCC 33078, ATCC 19575, and ATCC 19576, American Type Culture Collection, Manassas, VA.
b.
Provided by Lisle W. George, UC Davis, Davis, CA.
c.
ATCC 700019, American Type Culture Collection, Manassas, VA.
d.
ATCC 700022, American Type Culture Collection, Manassas, VA.
e.
Veterinary Medicine Biological Media Services, UC Davis, CA.
f.
DNEasy kit, Qiagen, Valencia, CA.
g.
Gibco UltraPure molecular biology grade water, Invitrogen, Carlsbad, CA.
h.
Choice Taq DNA Polymerase, Denville Scientific Inc., South Plainñeld, NJ.
i.
100 mM dNTP Set, Eppendorf, Westbury, NY.
j.
PTC-200 Peltier Thermal Cycler, MJ Research, Inc., Watertown, MA.
k.
AfaI (RsaI), Amersham Biosciences, Piscataway, NJ.
l.
GenePure LE Quick Dissolve Agarose, ISC BioExpress, Kaysville, UT.
m.
Sigma, St. Louis, MO.
n.
Dual Light Transilluminator FB-DLT-88A, Fisher Scientific, Pittsburgh, PA.
o.
Photodocumentation Handheld Camera, Fisher Scientific, Pittsburgh, PA.
p.
Davis Sequencing, Inc., Davis, CA.
