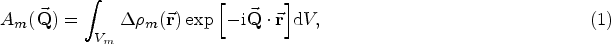Abstract
In this work, we present the development of small-angle scattering components in McStas that describe the neutron interaction with 70 different form and structure factors. We describe the considerations taken into account for the generation of these components, such as the incorporation of polydispersity and orientational distribution effects in the Monte Carlo simulation. These models can be parallelized by means of multi-core simulations and graphical processing units. The acceleration schemes for the aforementioned models are benchmarked, and the resulting performance is presented. This allows the estimation of computation times in high-throughput virtual experiments. The presented work enables the generation of large datasets of virtual experiments that can be explored and used by machine learning algorithms.
Keywords
Introduction
McStas1,2 is an open-source package for Monte Carlo (MC) neutron ray-tracing simulations widely used by the neutron scattering community for instrument design in neutron facilities and also for educational purposes. A recent trend in the use of MC simulation software, inspired by increasing computing power, is the generation of large virtual experimental datasets. These datasets can then be explored using machine learning algorithms to learn underlying structures that may provide insight into novel data analysis techniques and instrument design optimization.
3
If the system is anisotropic, then the expression for the small-angle scattering (SAS) model now depends on the orientation of the particle relative to the incident beam, and therefore a shape-specific modulation of the scattering pattern as a function of the scattering vector
Large-scale datasets can be constructed by varying simulation parameters in virtual experiments of neutron instruments. The information that can be extracted from such datasets depends greatly on the possibility that the MC software grants in exploring the simulation’s hyper-parameter space. This space consists mainly of parameters related to the instrument configuration in a given MC simulation, as well as parameters that describe the neutron interaction with the sample. In McStas, the complexity of the simulation’s parameter space is determined by the available components that describe the instrument and sample in a virtual experiment. This work aims to expand the latter by introducing 70 new SAS model components that can be used in McStas to describe a sample in a virtual neutron experiment.
In a first approximation and for simple problems where only first-order interactions and no secondary productions are taken into account, 5 MC neutron ray-tracing simulations are embarrassingly parallel. 6 That is, each neutron generated from the source distribution by means of MC choices is independent of the previously generated neutrons, and therefore the task of generating neutrons can be parallelized. The propagation of the neutrons in a beamline and their interaction with instrument components can also be parallelized because they are also independent. If only single scattering events are considered, the inclusion of a form factor or structure factor model for the sample description does not change this characteristic. Therefore, large dataset generation can have a significant speed-up if parallelization strategies are used in the MC neutron virtual experiments that generate the data.
This work describes the inclusion of 70 SAS models in McStas, which are optimized for parallel execution. A benchmark of the computation time of these models is also presented to emphasize and give estimates of the computing times necessary to build up big datasets.
Methods
Form factor and structure factor models
In scattering theory, the form factor
In the most general case of scattering from a scattering object of the model
A total of 70 SAS models have been developed as independent components in McStas
1
and are available since version 3.4 onward. Tables 1 and 2 present a summary of the models, classified into seven similarity groups labeled A to G. The components have also been imported to the X-ray counterpart McXtrace
9
as they describe the SAS characteristics that depend on the sample and not on the incident particle. Each model is described by two functions written in C language: one defining the scattering amplitude
Anisotropic small-angle scattering models included in McStas, inherited from SasView.
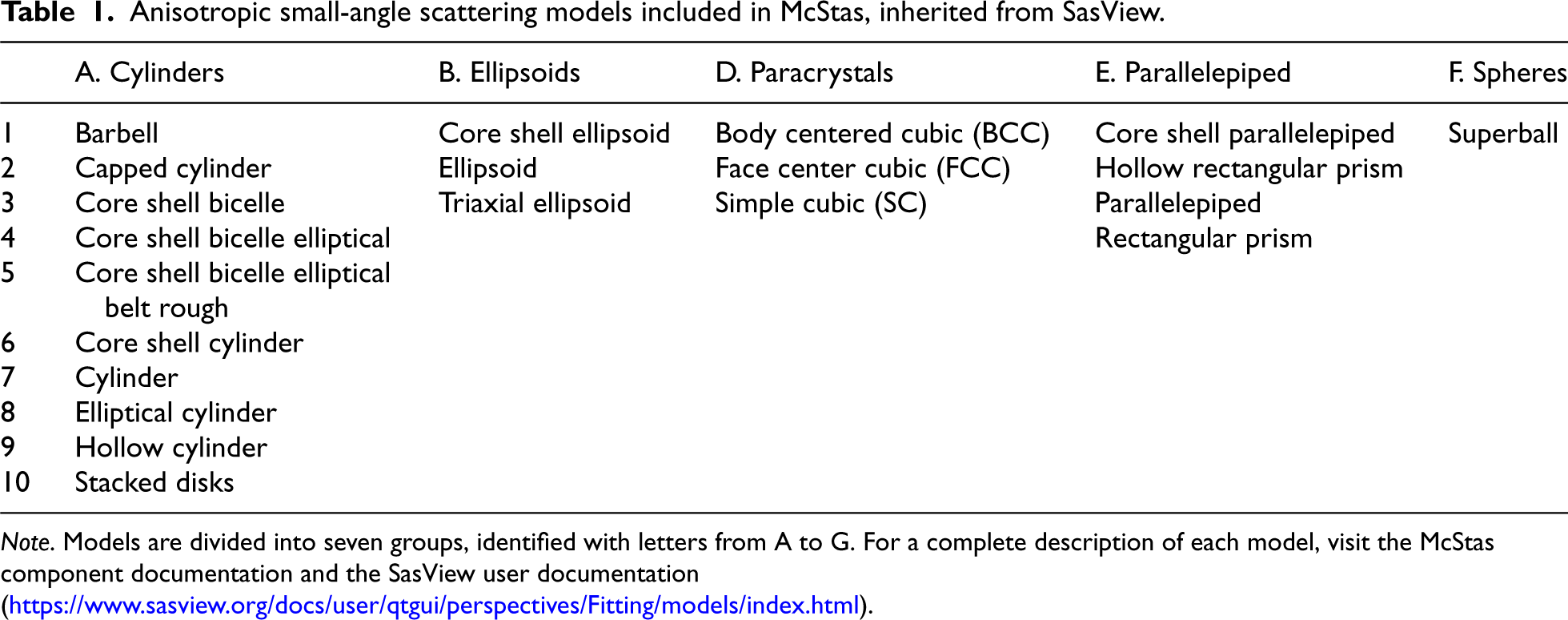

Note. Models are divided into seven groups, identified with letters from A to G. For a complete description of each model, visit the McStas component documentation and the SasView user documentation (https://www.sasview.org/docs/user/qtgui/perspectives/Fitting/models/index.html).
Isotropic small-angle scattering models included in McStas, inherited from SasView.
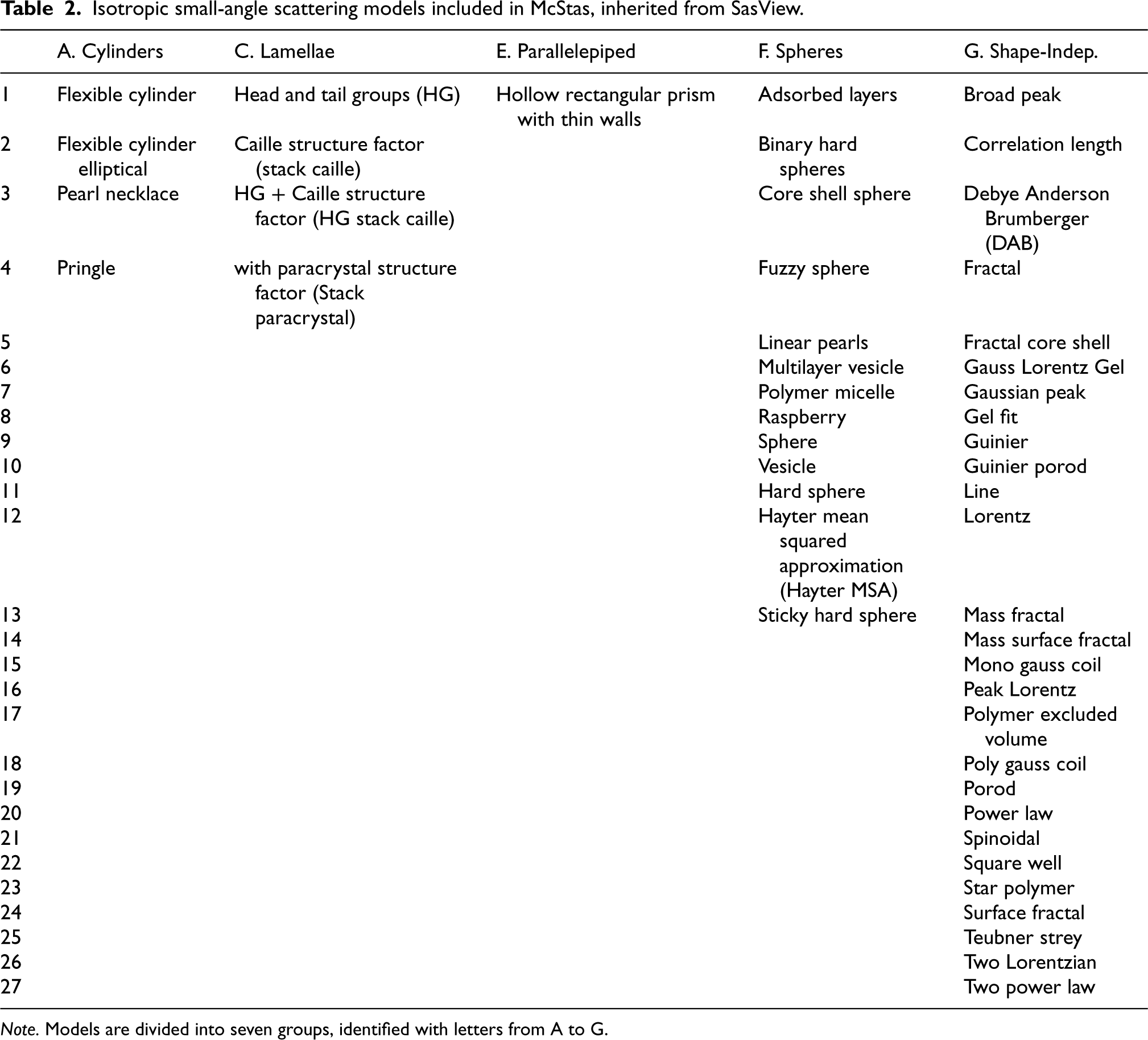
Note. Models are divided into seven groups, identified with letters from A to G.
A total of 21 models exhibit anisotropic analytical models, which are indicated in Table 1. In an isotropic scattering model, the measured intensity 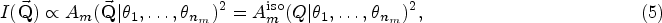
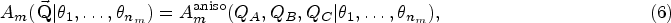
If the form of the scattering object has one symmetry axis, suppose 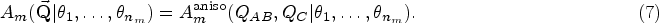
To test the developed models, we simulated the KWS-1 small-angle neutron scattering (SANS) beamline at FRM-II in McStas. This simulation consisted of a monochromatic source, guides and slits to propagate the neutron beam until the sample with defined divergence, one of the SANS modules listed in Tables 1 and 2, and a 2D position sensitive detector at a defined distance from the sample. For an accurate description of the McStas instrument, the simulation description can be found in the supplementary material of Robledo et al. 3
The components presented in this work offer the possibility of defining distributions for some model parameters in which this can be relevant. This includes polydispersity in all distances and radii parameters and orientational distributions in
For the moment, the polydispersity of a parameter 
The orientational distributions are also defined on each neutron interaction by random sampling from uniform distributions of the angles 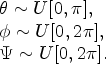
The developed components allow multi-thread parallelization by means of a message passing interface (MPI)
11
and can also be deployed to the graphical processing unit (GPU) for higher-order acceleration. This is managed internally in each component using OpenACC.
12
Schematic of a simulation flow in McStas, given a simulation statistic of 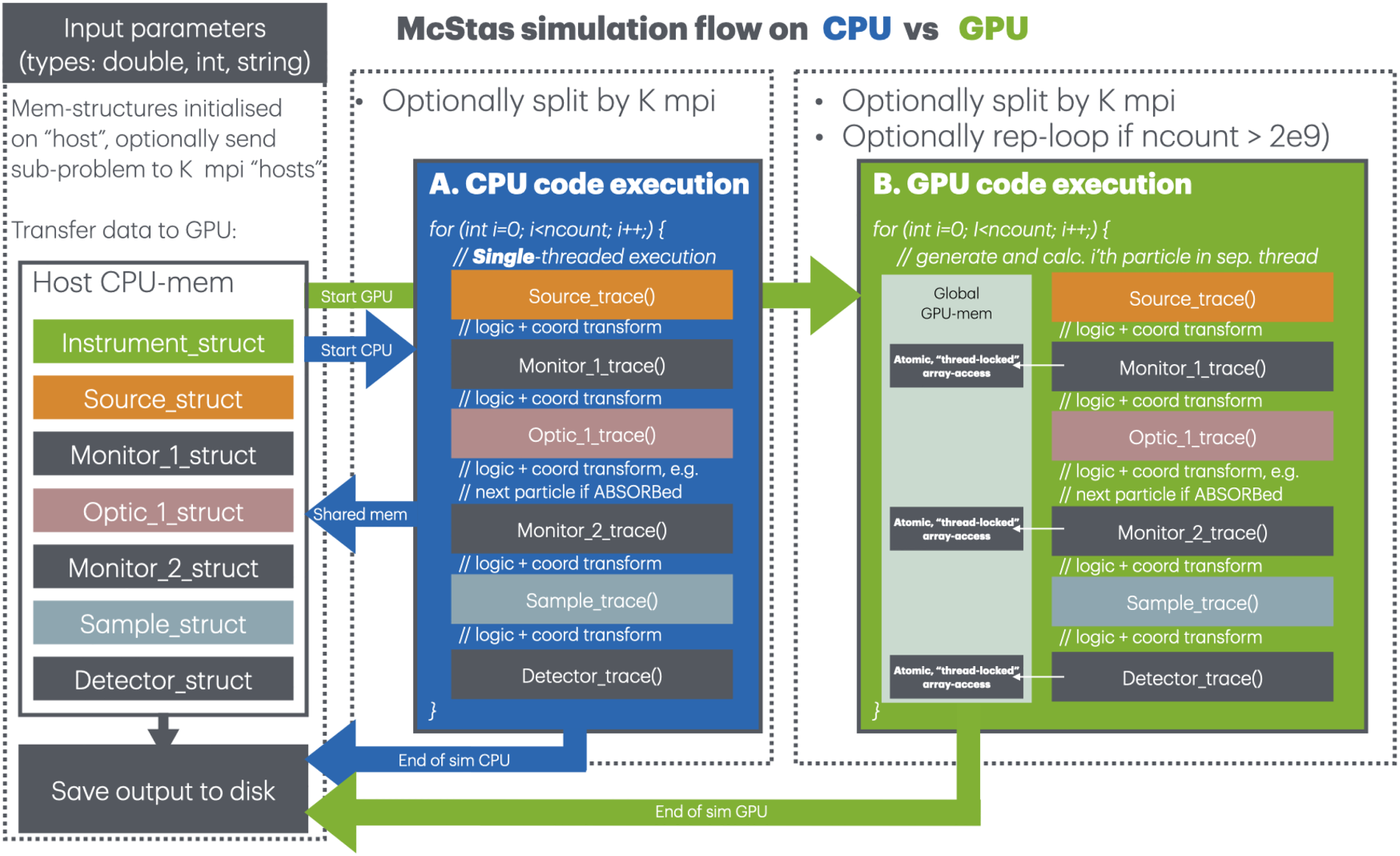
Figure 1 illustrates the McStas simulation flow during execution in three settings: single-threaded CPU, multi-threaded MPI, and parallel GPU execution. Given that McStas neutron rays are treated in a fully independent manner, a simulation problem of
To compare the acceleration capabilities, three different simulation schemes were tested:
a single-core simulation, to be used as reference; a Multi-threading approach with a GPU approach with an NVIDIA A100.
For each one of these schemes, all 70 models (listed in Tables 1 and 2) were used for the comparison of the acceleration schemes. Each one was simulated in an independent virtual experiment with a different number of incident neutrons (
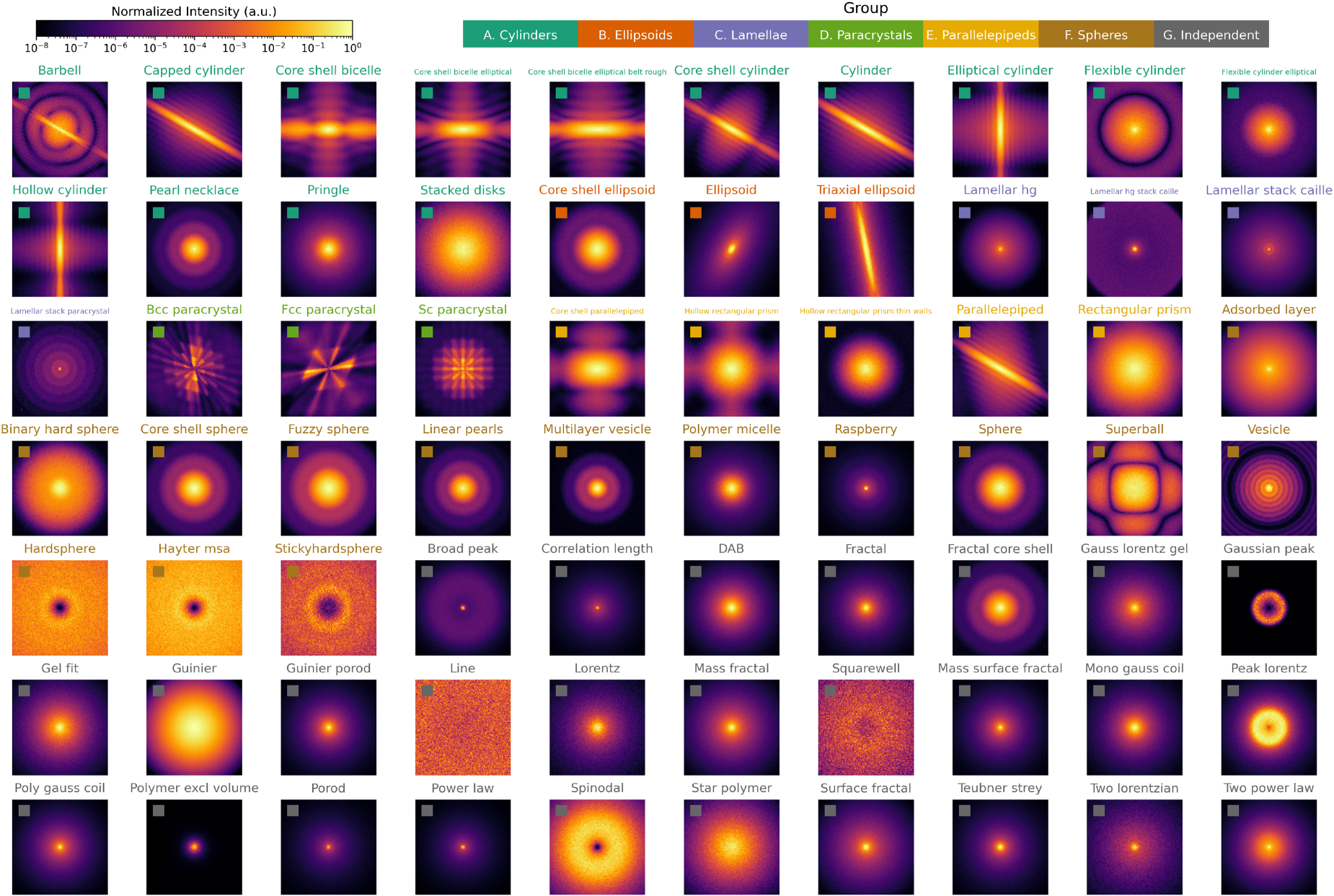
Normalized scattering pattern results from small-angle neutron scattering (SANS) simulations in McStas for all models developed in this work. Each image corresponds to one simulation of the same instrument configuration but changes in the sample description. All samples were defined by setting the default parameters for each SAS model and no polydispersity. For all anisotropic models, no orientation distribution was included, resulting in perfectly oriented scattering objects to emphasize anisotropic scattering features. Rectangles indicate the groups in Tables 1 and 2.
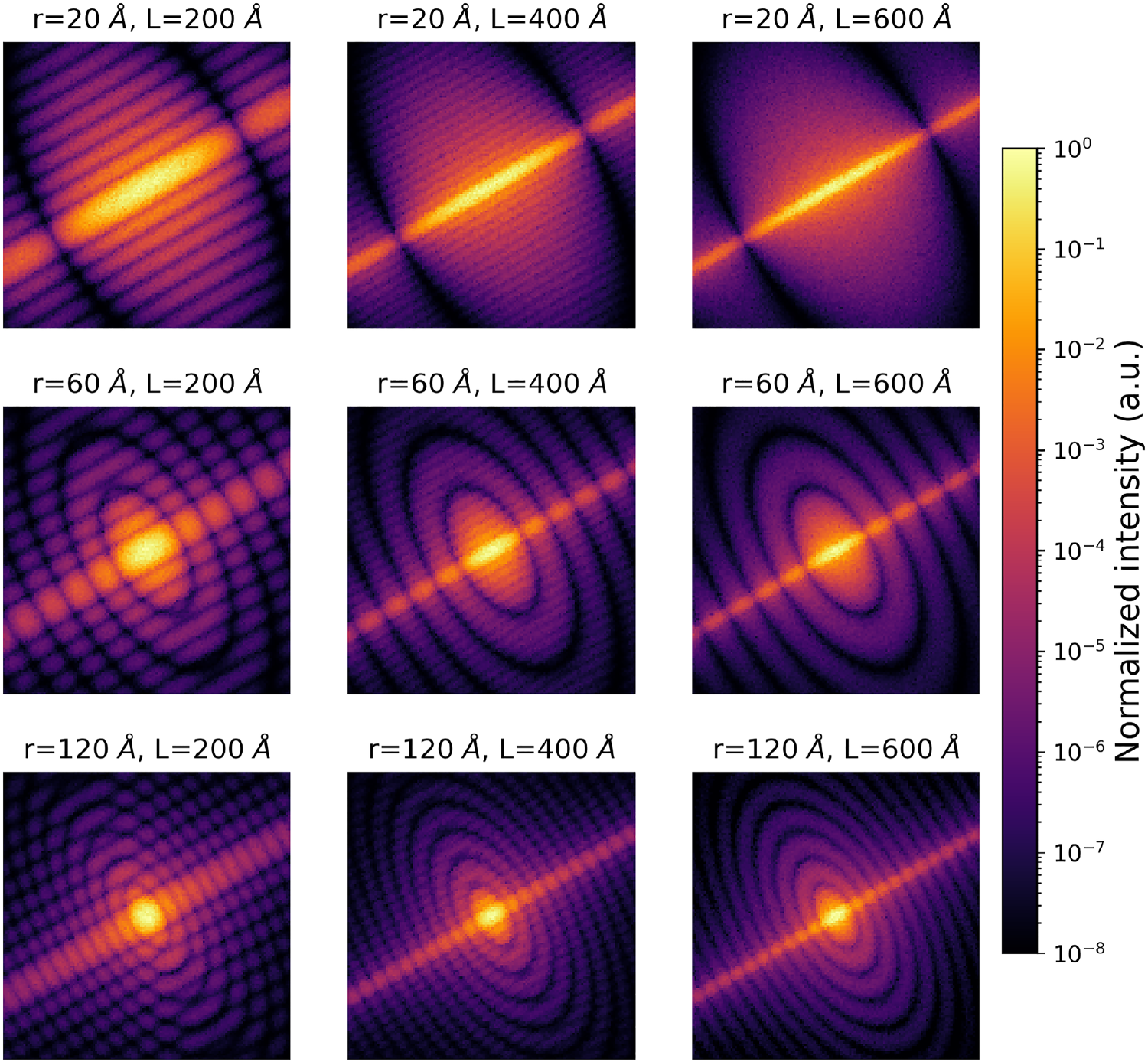
Example of intra-class variability for a cylinder form factor model with anisotropy. Each image corresponds to one simulation where the shape of the scattering objects is defined by the radius
SAS models
The results of 70 simulations in which the instrument configuration was fixed and the sample description was changed in all of the models presented in this work are shown in Figure 2. Each scattering pattern corresponds to testing one of the SAS models 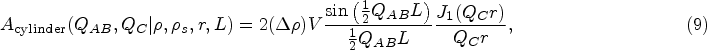
As can be seen in both, Figures 3 and 4, the scattering patterns corresponding to a given model may be very different given that all parameters express themselves in different features on the 2D scattering pattern. The variation of model parameters
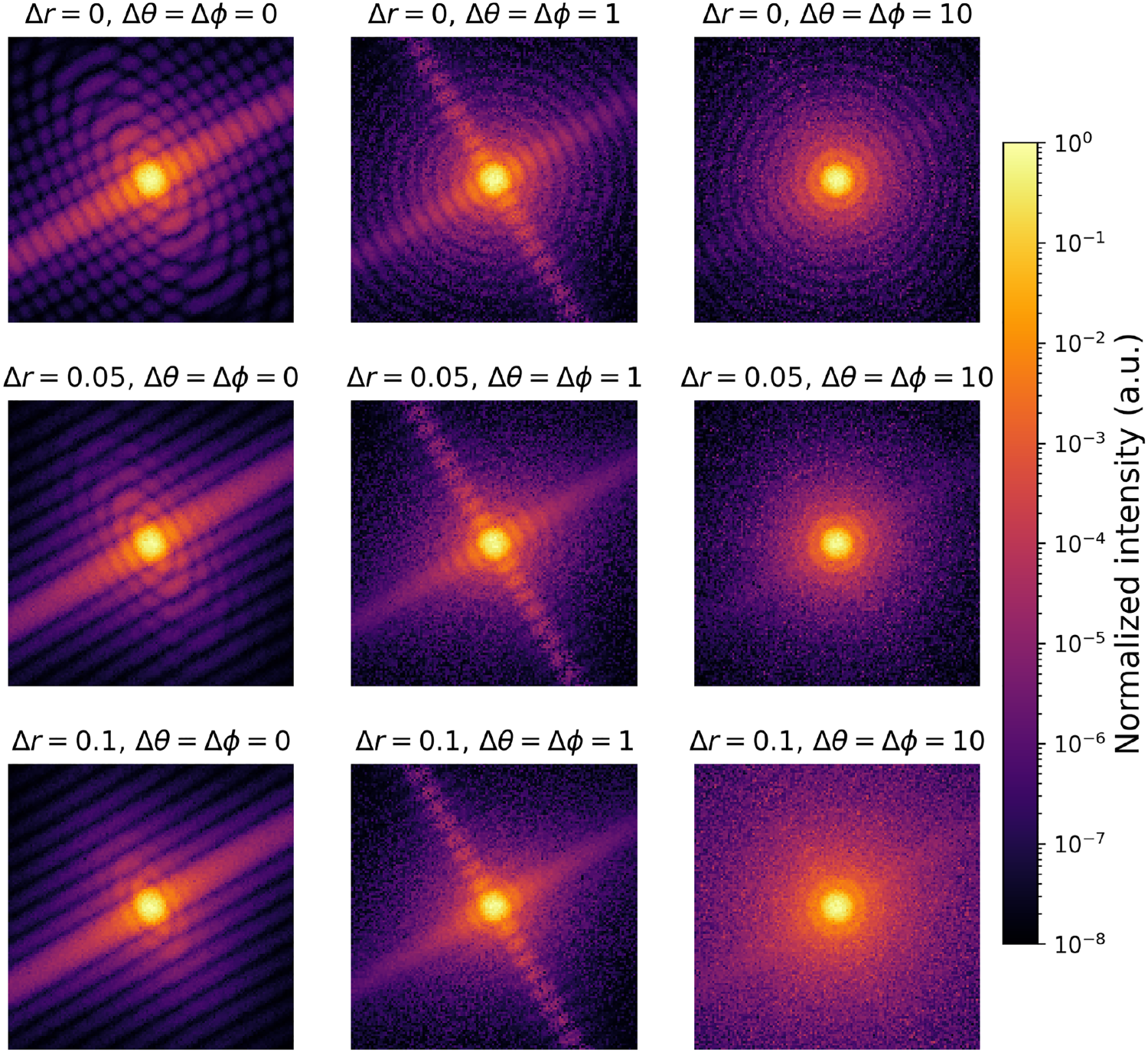
Example of intra-class variability due to polydispersity in radius and orientational distributions for a cylinder form factor model with anisotropy. Every image corresponds to a different simulation with varying parameters as described above the image, and common parameters mean radius
Finally, we benchmark the cylindrical model with an experimental data curve of cylindrically shaped micelles measured at the SANS instrument KWS-1 located at FRM-II.
15
This dataset has been widely studied since it is part of the Lab-course material of the Jülich Centre for Neutron Science.
16
The sample consists of amphiphilic polymers POO
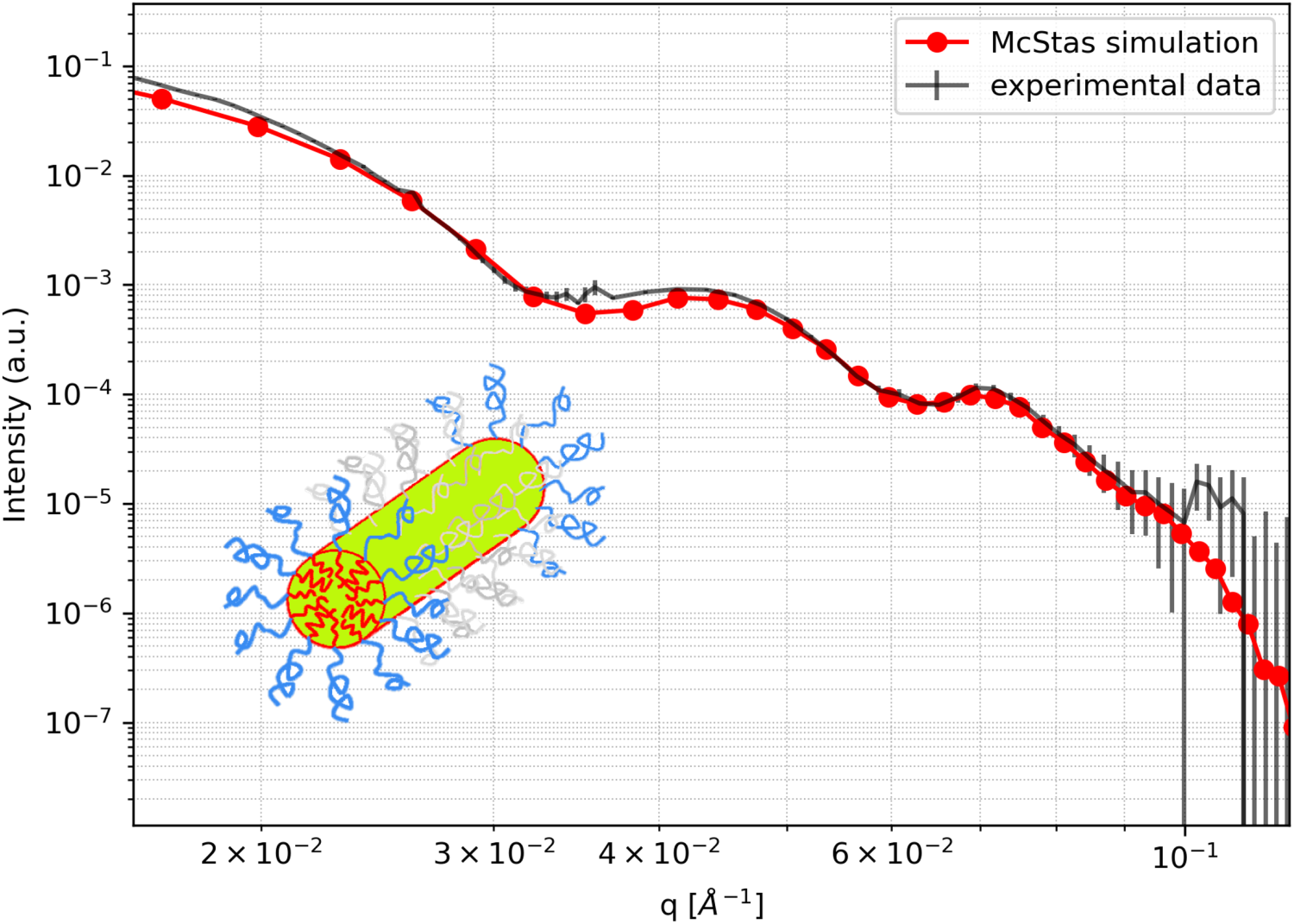
Comparison of small-angle scattering curves of a cylindrical model McStas simulation with experimental data measured at KWS-1, FRM-II. The inlay image depicts the amphiphilic polymers POO
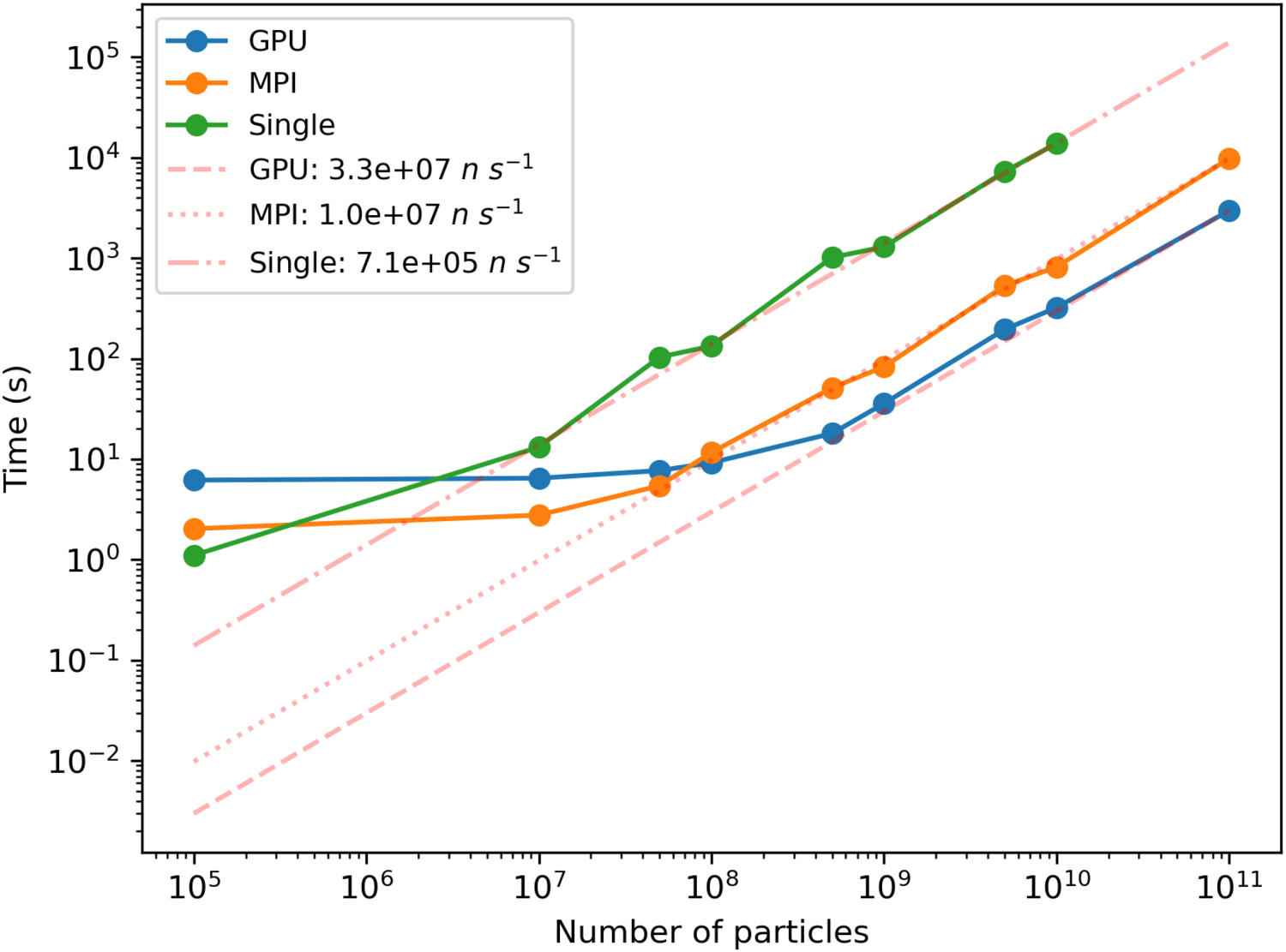
Median time of the models presented in this work as a function of the number of simulated neutrons, using the GPU, MPI, or single-core (single). The inverse of the slope, in neutrons per second, of the linear regression for high number of particles is given in the labels for each type of simulation. GPU: graphical processing unit; MPI: message passing interface.
All models described in Tables 1 and 2 were tested in simulations with different numbers of incident neutrons
When either the GPU or all of the cores available get filled with trajectories, then the simulation time increase becomes linear with increasing
It is interesting to observe that the median times of the distributions cross somewhere between
Figure 6 also shows that simulations accelerated by GPU (blue curve) take roughly the same time as those accelerated by MPI (orange curve) for
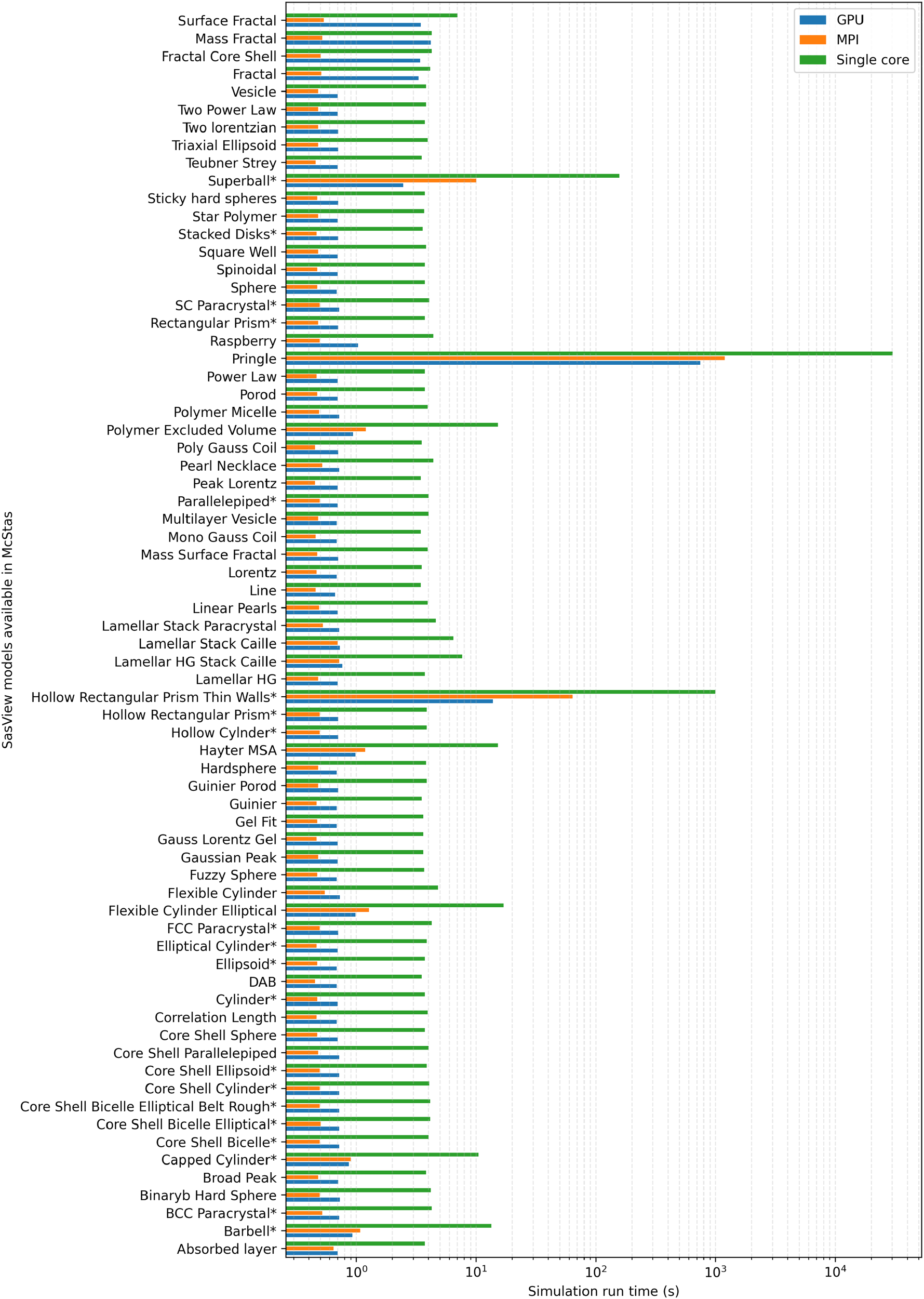
Simulation time of
This work describes the optimized SAS sample components that are readily available in McStas, which opens up the landscape for the possible small-angle neutron scattering virtual experiments that can be performed with the software. All the SAS components have the effects of polydispersity as well as orientational distribution included. Components can be used under acceleration schemes, according to the necessity. For single virtual experiments, where statistics is not very important, rather qualitative features, single-core simulations are adequate. When increasing the number of particles in a given simulation, multi-core and GPU parallelization of the particle traces becomes relevant. The GPU parallelization outperforms the multi-core simulation presented in this work when the number of neutrons in the simulation is higher than
The parallelization of traces in MC particle ray-tracing algorithms can be deeply exploited in simulations with large hyper-parameter spaces to explore a vast region of this space and generate large datasets of MC simulations. Simulation datasets can help in data augmentation, 18 as well as to learn from them through machine learning and then to infer on small datasets of real experiments (which is the usual case in neutron scattering experiments). 19 With the recent advances in generative adversarial networks that allow mapping from a simulated data distribution to an experimental data distribution in a bijective manner, 20 this type of mechanism for dataset generation can become very useful.
Footnotes
Acknowledgements
We would like to acknowledge the Helmholtz Artificial Intelligence group by providing computational resources through HAICORE project 27114.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement no. 101034266. This work benefited from the use of the SasView application, originally developed under NSF award DMR-0520547. SasView contains code developed with funding from the European Union’s Horizon 2020 research and innovation programme under the SINE2020 project, grant agreement no. 654000.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.