Abstract
Background
Osteoporotic fractures (OFs) are a significant public health issue, which can lead to pain and impaired mobility. The underlying mechanisms of OFs remain unclear, but recent studies have suggested that the circRNA-miRNA-mRNA pathway may play a crucial role.
Purpose
This study aimed to investigate the potential involvement of the circHIPK3/miR-378a-3p/HDAC4 pathway in the pathogenesis of OFs.
Methods
We collected tissue and serum samples from 10 patients with OFs and 10 healthy controls. The expression levels of circHIPK3, miR-378a-3p, and HDAC4 were measured by qPCR and WB. Additionally, we quantified the serum levels of bone metabolism-related indicators (ALP, OC, TRAP, OCIF, ODF) using ELISA.
Results
Our results revealed significant upregulation of circHIPK3 and HDAC4 in both tissue and serum samples from OF patients compared with controls. Simultaneously, we detected a lower expression level of miR-378a-3p in OF tissues and serum than that in the control group. Furthermore, the serum levels of bone metabolism-related indicators ALP, TRAP, OCIF, and ODF were significantly higher in OF patients than in the control group. Interestingly, the serum level of OCIF was lower in OF patients than in the control group.
Conclusion
Our study provides important evidence for the involvement of the circHIPK3/miR-378a-3p/HDAC4 pathway in the pathogenesis of OFs. The upregulation of circHIPK3 and HDAC4 and downregulation of miR-378a-3p observed in OF patients suggests their potential regulatory effects on bone metabolism. Meanwhile, abnormal expression of serum bone metabolism-related indicators may contribute to the development of OFs by disrupting the balance of bone remodeling.
Materials and methods
Introduction
Osteoporotic fractures (OFs) are a common health problem, especially among the elderly.1,2 They can lead to chronic pain, decreased mobility, and increased morbidity and mortality rates. 3 To date, the underlying mechanisms of OFs remain unclear, but emerging evidence suggests that the circRNA-miRNA-mRNA pathway may play a crucial role in bone metabolism regulation.4–6
CircRNAs are a class of endogenous transcripts that function as regulators of gene expression through various mechanisms, including acting as miRNA sponges,7–9 interacting with RNA-binding proteins10–12 and translating into proteins in a cap-dependent manner.13–15 The circular RNA hsa_circ_0000284 (termed circHIPK3) is a recently identified circRNA that has been reported to play an important role in several human diseases.16–19 Additionally, emerging studies suggest that circHIPK3 may also be involved in the pathogenesis of osteoporosis and related fractures.
miRNAs are small non-coding RNAs that play crucial roles in post-transcriptional gene regulation.20,21 miR-378a-3p is a member of the miR-378 family that has been reported to have multiple functions in various biological processes.22–24 Recent studies have indicated that miR-378a-3p is involved in the regulation of osteoblast differentiation and bone formation.25,26
HDAC4 is a histone deacetylase that plays important roles in the regulation of gene expression and protein function. 27 It has been shown to regulate osteoblast differentiation and bone formation by inhibiting Runx2 transcriptional activity. 28 Recent studies have suggested that HDAC4 may also be involved in the pathogenesis of osteoporosis and related fractures.29,30
Bone metabolism-related indicators, including alkaline phosphatase (ALP), tartrate-resistant acid phosphatase (TRAP), osteoclastogenesis inhibitory factor (OCIF), osteoclast differentiation factor (ODF), and osteocalcin (OC), are commonly used as diagnostic markers for bone metabolism disorders. ALP is a marker of osteoblast activity, 31 while TRAP is a marker of osteoclast activity. 32 OC is a marker of bone formation, 33 whereas OCIF and ODF play crucial roles in osteoclast differentiation and activation. 34
In this study, we aimed to investigate the potential involvement of the circHIPK3/miR-378a-3p/HDAC4 pathway in the pathogenesis of OFs. Our findings may provide new insights into the molecular mechanisms underlying OFs and suggest potential therapeutic targets for this common health problem.
Materials and methods
Clinical specimens
We collected bone tissue and serum samples from 10 patients with osteoporotic fractures (OF group) and 10 healthy controls (Control group). Tissue samples were collected and frozen in liquid nitrogen immediately after surgical resection. The inclusion criteria for both groups were patients undergoing interlocking intramedullary nail fixation for bone fractures. Patients with pathological fractures caused by open fractures, old fractures, space-occupying lesions, inflammation, bone tumors, severe cardiovascular, liver or kidney diseases, abnormal coagulation function, secondary osteoporosis caused by drugs or diseases, rheumatoid arthritis, and those who had taken drugs that affect bone metabolism within 6 months were excluded from the study. The participants in the study provided informed consent, and the study was approved by the Clinical Research Ethics Committee of The First people's Hospital of Hefei, China.
RNA extraction and reverse transcription
Total RNA was extracted from bone tissue using TRIzol reagent according to the manufacturer's instructions and quantified by Nanodrop ND-1000 spectrophotometer (Thermo Scientific, Wilmington, USA). Reverse transcription of RNA into cDNA was performed using a PrimeScript RT reagent Kit (Takara, Dalian, China).
qPCR analysis: qPCR was conducted using SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA) on an ABI 7500 Real-Time PCR system (Applied Biosystems, Foster City, CA, USA). The primer sequences used in this study were as follows: circHIPK3 Forward: 5'-GCAGCCTTACAGGGTTAAAGT-3', Reverse: 5'-TTGTGAGGCCATACCTGTAGTA-3'; HDAC4 Forward: 5'-GAGAGCGATGAGGAAGAGGC-3', Reverse: 5'-GGTAGTTCCTCAGCTGGTGG-3'; miR-378a-3p Forward: 5'-CGCGACTGGACTTGGAGTCA-3', Reverse: 5'- AGTGCAGGGTCCGAGGTATT-3',RT:5'-GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGCCTTC-3'; GAPDH Forward: 5'-GCCGTCTAGAAAAACCTGCC-3', Reverse: 5'-CCACCTGGTGCTCAGTGTAG-3'.
U6 Forward: 5'-CTCGCTTCGGCAGCACA-3', Reverse: 5'-AACGCTTCACGAATTTGCGT-3'.
GAPDH was used as an internal control for circHIPK3 and HDAC4.U6 was used as an internal control for hsa-miR-378a-3p.
Western blot analysis
Total protein was extracted from bone tissue using RIPA lysis buffer (Beyotime, Shanghai, China) and quantified using a BCA protein assay kit (Thermo Fisher Scientific, Waltham, MA, USA). Equal amounts of protein were separated by SDS-PAGE and transferred to nitrocellulose membranes. The membranes were probed with primary antibodies against HDAC4 (Abcam, Cambridge, UK) and GAPDH (Abcam, Cambridge, UK), followed by HRP-conjugated secondary antibodies. Protein bands were visualized using an enhanced chemiluminescence detection kit (Millipore, Billerica, MA, USA).
ELISA analysis
The serum levels of bone metabolism-related indicators, including ALP, TRAP, OCIF, ODF, and OC, were measured using ELISA kits (CUSABIO, Wuhan, China) according to the manufacturer's instructions.
Statistical analysis
Data were presented as mean ± standard deviation (SD) and analyzed using GraphPad Prism software version 8.0 (GraphPad Software, San Diego, CA, USA). Differences between the groups were evaluated by Student's t-test or one-way ANOVA. P values less than 0.05 were considered statistically significant
Results
In this study, we aimed to investigate the role of the circHIPK3/miR-378a-3p/HDAC4 pathway in osteoporotic fractures. We first assessed the expression level of circHIPK3 in both serum and tissue samples from patients with osteoporotic fractures and normal fracture cases using qPCR analysis. The qPCR results showed that circHIPK3 was upregulated in both serum and tissue samples from osteoporotic fractures patients compared to normal controls (Figure 1(a) and (b)). Given that circRNAs have been shown to regulate gene expression through the ceRNA mechanism, we predicted potential miRNAs that could bind to circHIPK3 using the starbase database (https://starbase.sysu.edu.cn/). Our prediction revealed that miR-378a-3p has a binding site with circHIPK3 (Figure 1(c)). We then investigated the expression level of miR-378a-3p in serum and tissue samples from both groups of patients using qPCR analysis. The results showed that miR-378a-3p was downregulated in both serum and tissue samples from patients with osteoporotic fractures compared to those from normal fracture cases (Figure 1(d) and (e)), which is consistent with the hypothesis that circHIPK3 may function as a competing endogenous RNA (ceRNA) by sequestering miR-378a-3p. The expression of circHIPK3 and hsa-miR-378a-3p showed a negative correlation in samples from patients with osteoporotic fractures. (a) The expression level of circHIPK3 in serum samples was quantified using qPCR. (b) The expression level of circHIPK3 in tissue samples was quantified using qPCR. (c) The starbase database predicted the presence of binding sites between circHIPK3 and hsa-miR-378a-3p. (d) The expression level of hsa-miR-378a-3p in serum samples was quantified using qPCR. (e) The expression level of hsa-miR-378a-3p in tissue samples was quantified using qPCR.
We further used starbase database to predict the downstream target of miR-378a-3p and found that HDAC4 might be a potential target (Figure 2(a)).We carried out qPCR analysis to assess the expression levels of HDAC mRNA, and our results showed that HDAC4 was upregulated in serum and bone tissue samples from patients with osteoporotic fractures compared to those from normal fracture cases (Figure 2(b) and (c)). At the protein level, we also performed Western blotting analysis on bone tissue samples from both groups of patients and observed a significant increase in HDAC4 protein levels in bone tissue samples from patients with osteoporotic fractures (Figure 2(d)). As a downstream target gene of hsa-miR-378a-3p, HDAC4 was found to be highly expressed in samples from patients with osteoporotic fractures. (a) The starbase database predicted the presence of binding sites between HDAC4 and hsa-miR-378a-3p. (b) The expression level of HDAC4 in serum samples was quantified using qPCR. (c) The expression level of HDAC4 in tissue samples was quantified using qPCR. (d) Western blot analysis was performed to investigate the expression levels of HDAC4 protein in tissue samples obtained from patients with osteoporotic fractures and normal fracture patients.
Additionally, we measured several bone metabolism-related markers including ALP, TRAP, OCIF, ODF, and OC in serum samples from both groups of patients using ELISA assays. The results showed that ALP, TRAP, OCIF, ODF were significantly elevated in serum samples from patients with osteoporotic fractures compared to those from normal fracture cases (Figure 3(a)–(d)). On the contrary, the level of OC was found to be decreased in serum samples from patients with osteoporotic fractures compared to those from normal fracture cases (Figure 3(e)). The concentrations of bone metabolism-related markers ALP (alkaline phosphatase), TRAP (tartrate-resistant acid phosphatase), OCIF (osteoclastogenesis inhibitory factor), ODF (osteoclast differentiation factor), and OC (osteocalcin) were measured using ELISA in serum samples. (a) The concentration of ALP in serum samples was measured using ELISA. (b) The concentration of TRAP in serum samples was measured using ELISA. (c) The concentration of OCIF in serum samples was measured using ELISA. (d) The concentration of ODF in serum samples was measured using ELISA. (e) The concentration of OC in serum samples was measured using ELISA. (f) A schematic model displaying the role of circHIPK 3/miR-378a-3p/HDAC4 axis in osteoporotic fractures.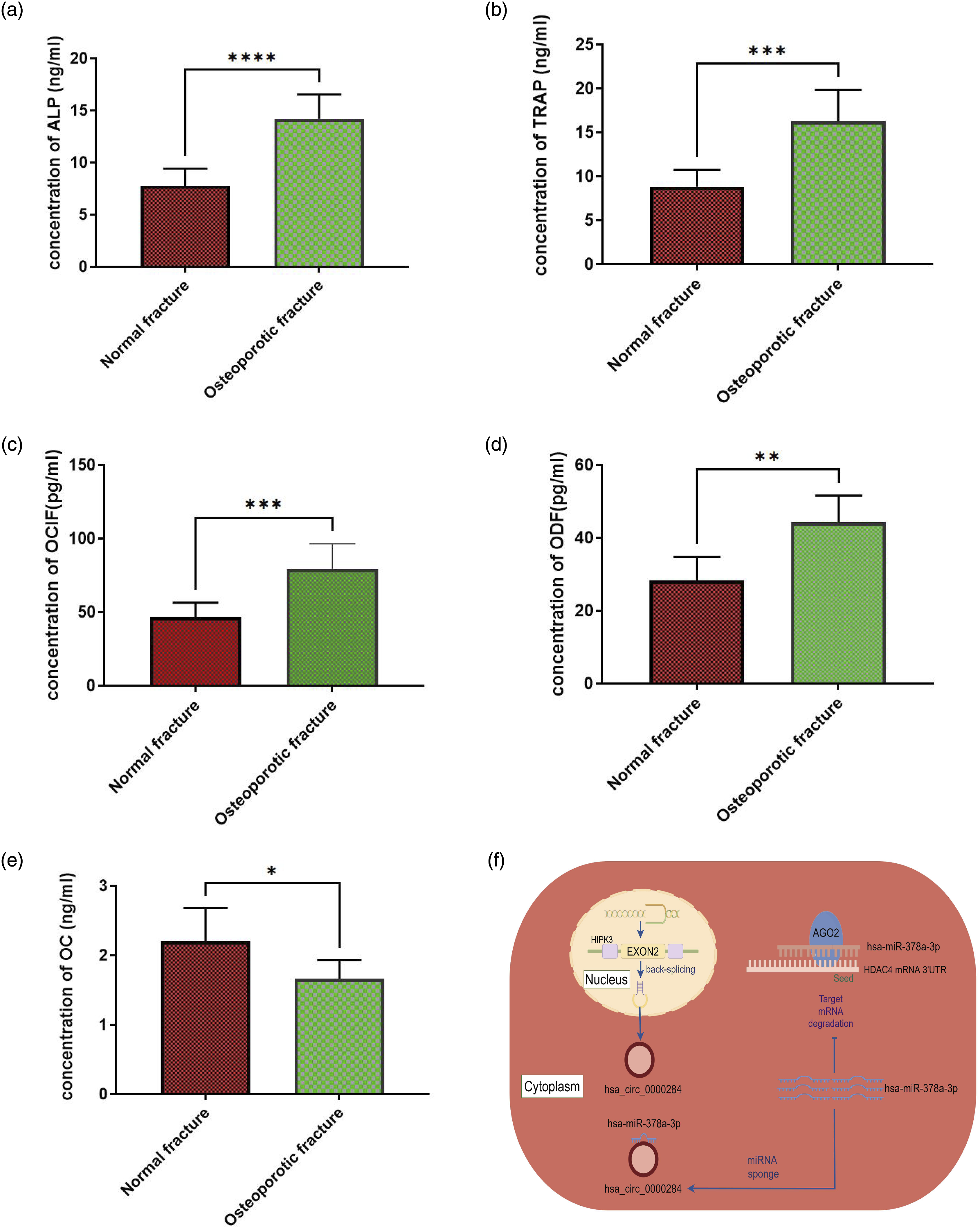
Taken together, our results suggest the involvement of the circHIPK3/miR-378a-3p/HDAC4 pathway in the pathogenesis of osteoporotic fractures. The upregulation of circHIPK3 and HDAC4 and downregulation of miR-378a-3p observed in our study may have clinical implications for developing novel therapeutic strategies for managing osteoporotic fractures. Moreover, our findings regarding the changes in bone metabolism-related markers provide new insights into the mechanisms underlying osteoporotic fractures.
Specifically, the upregulation of ALP, TRAP, OCIF, and ODF in serum samples from patients with osteoporotic fractures suggests an increase in bone turnover rate and osteoclast activity. On the other hand, the decrease in OC levels in serum samples from patients with osteoporotic fractures indicates a decrease in osteoblast activity. These observations are consistent with the underlying pathophysiology of osteoporosis, which is characterized by an imbalance between bone formation and resorption. Overall, our study provides new insights into osteoporotic fractures.
Discussion
Osteoporotic fractures (OFs) are a significant public health issue, particularly among older adults. OFs can lead to chronic pain, decreased mobility, and increased morbidity and mortality rates. Although many studies have investigated the pathogenesis of OFs, the underlying mechanisms remain unclear.
In this study, we aimed to investigate the potential involvement of the circHIPK3/miR-378a-3p/HDAC4 pathway in the pathogenesis of OFs. Our results showed that the expression levels of circHIPK3 and HDAC4 were significantly higher in both tissue and serum samples from OF patients compared with healthy controls. Simultaneously, miR-378a-3p expression was significantly lower in OF tissues and serum than that in the control group. Additionally, several serum bone metabolism-related indicators showed abnormal expression in OF patients, indicating their potential involvement in the development of OFs.
Our findings suggest that circHIPK3 and HDAC4 may play critical roles in the regulation of bone metabolism. CircRNAs have been shown to regulate gene expression through various mechanisms, including sponging miRNAs, regulating transcription, and interacting with RNA-binding proteins. In addition, HDAC4 has been shown to regulate osteoblast differentiation and bone formation by inhibiting transcriptional activity.35,36
Meanwhile, miRNAs have been shown to play crucial roles in bone metabolism. Specifically, miR-378a-3p has been reported to be involved in the regulation of osteoblast differentiation and bone formation. 37 The downregulation of miR-378a-3p observed in OF tissues and serum may disrupt the balance of bone remodeling, contributing to the development of OFs.
Additionally, our study revealed that several serum bone metabolism-related indicators showed abnormal expression in OF patients. These indicators, including ALP, TRAP, OC, OCIF, ODF, have been previously reported to play crucial roles in bone remodeling. 38 For example, ALP and TRAP are markers of osteoblast and osteoclast activity, respectively, while OC is a marker of bone formation. Therefore, their abnormal expression may contribute to the development of OFs by disrupting the balance of bone remodeling.
Our study has some limitations that should be considered when interpreting our results. First, the sample size was relatively small, and larger studies are needed to validate our findings. Second, we only measured the expression levels of circHIPK3, miR-378a-3p, and HDAC4 in OF tissues and serum, but we did not investigate their functional roles directly.Additionally, we did not perform direct functional validation or molecular-level verification of their direct interactions, such as utilizing a dual-luciferase reporter gene system. Further studies are needed to elucidate the exact molecular mechanisms underlying the circHIPK3/miR-378a-3p/HDAC4 pathway in OFs.
In conclusion, our study provides important evidence for the involvement of the circRNA-miRNA-mRNA interaction in the pathogenesis of OFs. Our findings suggest that circHIPK3/miR-378a-3p/HDAC4 pathway may play a critical role in the regulation of bone metabolism and provide new insights into the molecular mechanisms underlying OFs. Further research is needed to explore potential therapeutic targets for OFs
Footnotes
Authors’ contributions
Dr. ZS was responsible for collecting serum and tissue samples. Dr. LW was responsible for drafting the manuscript and data analysis. Dr. TY was responsible for designing and performing the experiments. Furthermore, Dr. TY has contributed to data interpretation, editing and critical revision of the manuscript. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Applied Medical Research Project of Hefei Municipal Health Commission (HWK2022yb021).
