Abstract
While TGFβ plays a critical role in tumor formation and progression, the role and contribution of its three different isoforms remain unclear. In this study, we aimed at elucidating the prognostic value of the TGFβ isoforms and assessed their expression levels in breast cancer patients at different stages of the disease. We found higher levels of TGFβ1 and TGFβ3 in cancer patients compared to normal tissues, with no significant changes in TGFβ2 expression. Similarly, TGFβ1 and TGFβ3, but not TGFβ2, showed higher expression levels in advanced lymph node–positive and metastatic tumors, suggesting different roles for the different isoforms in tumor progression and the metastatic process, while in the least aggressive molecular subtype (luminal A), expression of the three TGFβ isoforms significantly correlated with expression of both TGFβ receptors, such correlation only occurred between TGFβ1 and TGFβ3 and the TGFβ type II receptor (TβRII) in the highly aggressive basal-like subtype. Interestingly, a distinct and somehow opposite pattern was observed in HER-2 tumors, only showing significant association pattern between TGFβ2 and the TGFβ type I receptor (TβRI). Finally, the three TGFβ isoforms showed distinct association patterns with patient outcome depending on the different molecular subtype, highlighting context-dependent, differential prognostic values.
Introduction
The transforming growth factor β (TGFβ) growth factors are expressed in most of the tissues and orchestrate various physiological processes including hormonal and immune responses, tissue repair, cell growth, cell death, cell fate determination, among others.1,2 The TGFβ family comprises three isoforms TGFβ1, TGFβ2, and TGFβ3, each encoded by a different gene. Despite the fact that the three isoforms show structural similarities 3 and usually transduce their signals through the same signaling pathways,4,5 they also exhibit tissue-specific expression and have different affinities for TGFβ receptors.6,7
Interestingly, the three TGFβ isoforms also have distinct expression patterns and functions during normal mammary gland development. TGFβ1 expression is high in the mammary tissue of virgin and early pregnant mice, reduced during pregnancy, and up again after weaning.8,9 In contrast, TGFβ2 was found to be low in the mammary glands of virgin mice, gradually increased through pregnancy, and decreased during lactation. TGFβ3 expression is also increased during pregnancy, reaching maximum levels before parturition, and start falling during lactation before peaking again after weaning.8,10 This difference in the expression levels observed during the mammary cycle was also associated with some difference in the expression patterns of the three isoforms during alveolar morphogenesis. 11 For example, in contrast to both TGFβ1 and TGFβ2, TGFβ3 was shown to be more expressed in the ducts than the alveoli. 11 In addition, the function of each isoform was found to be distinct. TGFβ1 plays a role in different biological processes including cell growth, differentiation and epithelial–mesenchymal transition (EMT), 12 ductal development and epithelial proliferation, matrix remodeling, apoptosis, and suppression of ovarian hormones’ proliferative effects.13,14 TGFβ2 is involved in epithelial–stromal cross-talk and matrix remodeling. 11 In contrast, TGFβ3 has a role in the maturation of myoepithelium, suppression of milk secretion, 15 and the EMT process.16,17 In cancer, the three isoforms also play different functions. For example, TGFβ1 is well known for its role in cell motility, invasion, and metastasis and in promoting EMT process.1,18–23 In contrast, TGFβ2 was suggested to play a role in enhancing tumor proliferation and decreasing immune surveillance during tumor development. 24 However, TGFβ3 was proposed to play a role in the EMT process.16,17 Furthermore, recent reports suggested that different TGFβ isoforms might exert distinct functions in skin cancer tumorigenesis, according to tumor advancement and stage. 25
In breast cancer, TGFβ has a dual role ranging from a tumor suppressor function to an oncogenic role, depending on cell type and stage of the disease. 1 The mechanisms involved in this transition are still not fully understood.1,20,21,26–32 Notably, most studies on the role of TGFβ in cancer formation and progression were performed using TGFβ1 and few information is available with respect to the other isoforms in these processes. Previous report showed variable levels of TGFβ isoforms in the different breast cancer tumors. However, due to the limited number of cases examined, no detailed analysis was performed to evaluate the association between the expression of the different isoforms and clinicopathological parameters. 33 Another recent study showed a preferential expression of the different isoforms in malignant and premalignant breast cancer lesions compared to normal tissues. 34 These studies suggest different roles for the three TGFβ isoforms in the pathogenesis of the disease.
Thus, and considering the prominent role played by the TGFβ family in breast cancer formation and progression, it is important to decipher the specific functions and pattern of expression of each TGFβ isoforms in breast cancer. Indeed, further clarifying the role of TGFβ in tumorigenesis and our understanding of the molecular mechanisms underlying tumor formation and progression may prove useful for designing more specific and efficient therapeutic strategies. In this study, we analyzed the expression patterns of the three TGFβ isoforms in the different breast cancer molecular subtypes and their association patterns with different clinical and pathological features, using tissue samples, tissue microarrays (TMAs) as well as publicly available datasets on large cohorts of breast cancer patients. Furthermore, we also investigated and correlated the TGFβ isoform expression patterns with those of the TGFβ type I and type II receptors (TβRI and TβRII) as well as with the estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor-2 (HER-2) status of the tumors. Interestingly, our results indicate that breast cancer tissues show higher TGFβ1 and TGFβ3 levels compared to normal tissues. Moreover, we also found increased TGFβ1 and TGFβ3 and decreased TGFβ2 expression in advanced lymph node (LN)-positive and metastatic tumors. Our data also indicate that the different TGFβ isoforms have distinct patterns of co-expression with the TGFβ receptors, depending on the breast cancer molecular subtypes. In addition, our results also indicate that the prognostic values of each isoform vary and sometimes were opposite to each other according to the molecular subtypes.
In summary, our results define differential and distinct expression patterns for the different TGFβ isoforms according to the stage of tumor progression and the molecular subtype. These results strongly suggest that the different isoforms play distinct roles and have different functions in different stages of the disease and in the different molecular subtypes of breast cancer. A better understanding of their specific roles and functions should prove useful in establishing more specific therapeutic options by targeting specific isoforms in distinct molecular subtypes, which in turn will contribute to better and improved patient response and outcome.
Results
All TGFβ isoforms are essential for breast cancer tumorigenesis
Most studies on the role of TGFβ in breast cancer were performed using the TGFβ1 isoform. To start evaluating the effect of different TGFβ isoforms in breast carcinogenesis, we first analyzed their tumor initiating capacities using tumorsphere assay in the basal subtype breast cancer cell line (SUM159) as previously described.22,23 As shown in Figure 1, all TGFβ isoforms could significantly increase the number of cancer stem cells, as measured by the increased sphere-forming ability of these cells when stimulated by TGFβ1, TGFβ2, and TGFβ3. These effects were observed and quantified on both tumorsphere numbers (Figure 1(b)) and tumorsphere sizes (Figure 1(c)). The effects appear specific to the TGFβ family, as Activin A, a closely related TGFβ family member, 35 does not induce any change in tumorsphere size or numbers (Figure 1). These data further confirm the important and specific role played by TGFβ in regulating breast tumorigenesis and indicate that all isoforms are active in this process.

TGFβ isoform effects on tumor initiation capacity and expression levels in breast cancer. (a) Representative images of the effect of TGFβ1, TGFβ2, and TGFβ3 and Activin A on tumorsphere formation in triple-negative breast cancer (TNBC) cell line SUM159. (b) Representative histogram of the effect of TGFβ1, TGFβ2, and TGFβ3 and Activin A on the number of tumorsphere formation in TNBC breast cancer cell line SUM159. (c) Representative histogram of the effect of TGFβ1, TGFβ2, and TGFβ3 and Activin A on the diameter of tumorsphere formation in TNBC breast cancer cell line SUM159.
TGFβ1 and TGFβ3, but not TGFβ2, expression levels are increased in breast cancer compared to normal tissues
We next compared the three TGFβ isoform expression levels in malignant cases versus normal tissues using The Cancer Genome Atlas (TCGA) datasets of the large publicly available ONCOMINE database. As shown in Figure 2(a), we found a significant increase in TGFβ1 (1.630 fold; p = 2.30E−14) and TGFβ3 (1.5 fold; p = 4.61E−6) gene expression levels in malignant tissues (389 breast cancer patients) compared to their normal counterpart (61 normal breast tissues). In contrast, no detectable change in expression was observed for TGFβ2. These data highlight a selective upregulation of specific TGFβ isoforms, namely, TGFβ1 and TGFβ3, in breast cancer, suggesting a differential role for the different isoforms in breast cancer tumorigenesis.
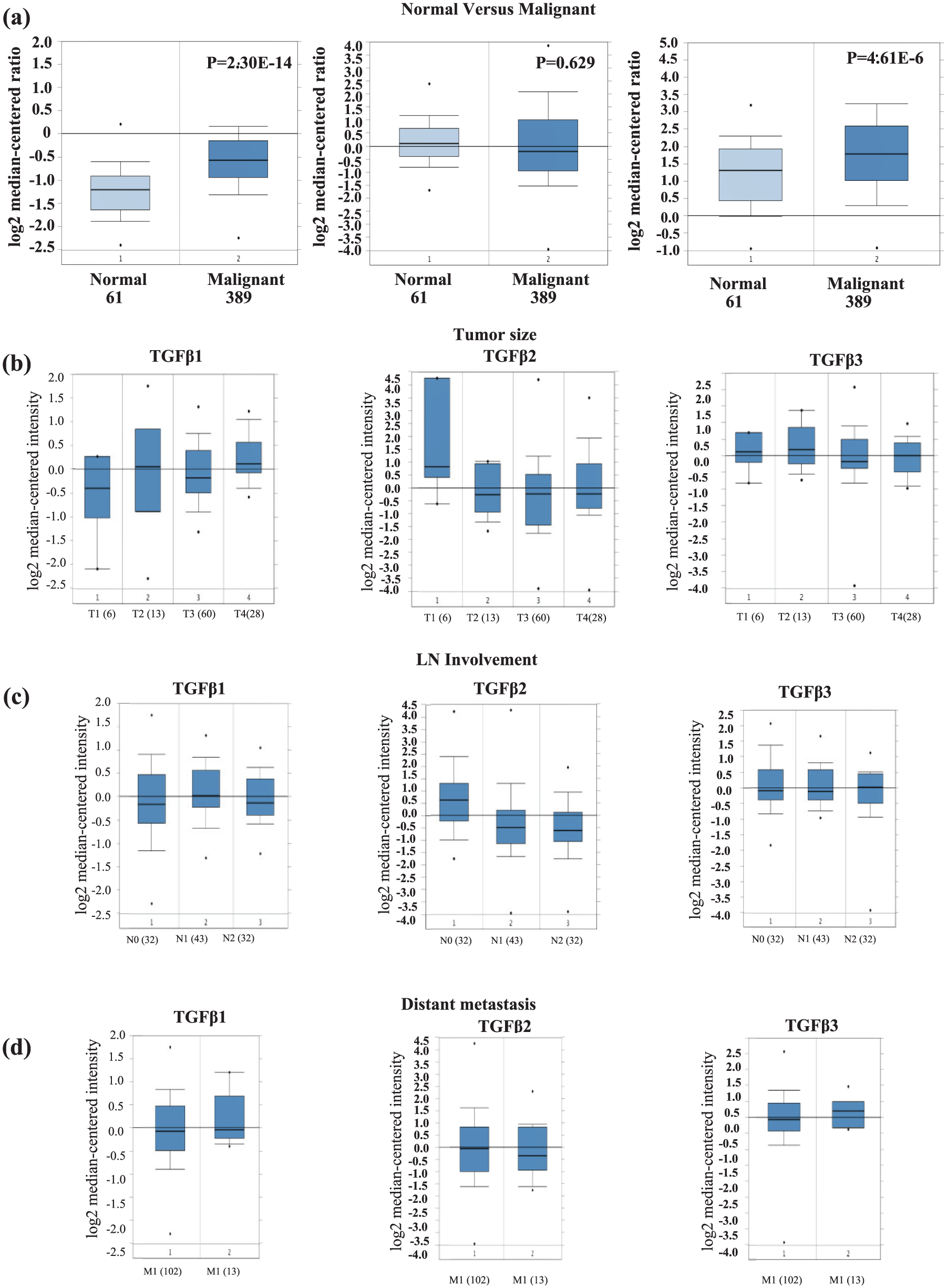
TGFβ isoform mRNA levels in association with different clinicopathological parameters. (a) TGFβ1, TGFβ2, and TGFβ3 mRNA expression levels in normal and invasive breast cancer cases using 593 cases of The Cancer Genome Atlas (TCGA) dataset in ONCOMINE database. (b) TGFβ1, TGFβ2, and TGFβ3 mRNA levels stratified according to tumor size using 167 cases of Sorlie dataset in ONCOMINE database. (c) TGFβ1, TGFβ2, and TGFβ3 mRNA levels stratified according to LN involvement using 167 cases of Sorlie dataset in ONCOMINE database. (d) TGFβ1, TGFβ2, and TGFβ3 mRNA levels stratified according to the absence or the presence of metastasis using 167 cases of Sorlie dataset in ONCOMINE database.
Advanced LN-positive and metastatic tumors are associated with higher TGFβ1 and TGFβ3 and lower TGFβ2 messenger RNA levels
To further evaluate the prognostic role of the different TGFβ isoforms in breast cancer and their role in tumor progression, we next assessed the association between their expression levels and tumor size, LN status, and the presence or absence of distance metastasis, using large datasets of the ONCOMINE database. While TGFβ1 messenger RNA (mRNA) levels were significantly higher in larger size tumor (T2, T3, and T4) compared to T1, TGFβ2 mRNA was significantly reduced in larger tumors compared to T1. In contrast, TGFβ3 showed no distinct pattern compared with size (Figure 2(b)). Moreover, and due to the fact that TGFβ also plays a critical role in promoting breast cancer invasion and metastasis, we next evaluated the association between expression of the different TGFβ isoforms and LN involvement as well as distant metastasis. Interestingly, higher mRNA expression levels of both TGFβ1 and TGFβ3 were observed in both LN-positive and metastatic tumors (Figure 2(c) and (d)). In contrast, TGFβ2 mRNA expression levels showed an opposite pattern, being significantly lower in LN-positive and metastatic tumors (Figure 2(c) and (d)). These results suggest that different TGFβ isoforms might play different roles during breast cancer metastasis and highlight TGFβ1 and TGFβ3 as markers of LN involvement and metastasis, while TGFβ2 levels remain low during LN involvement and metastasis.
TGFβ isoforms and their association with classical breast cancer markers
For breast cancer, expression of the classical biomarkers, including ER, PR, and HER-2 status, has an important clinical and prognostic value. Thus, we evaluated the association between TGFβ isoforms and the ER, PR, and HER-2 status of the tumors. Our analysis revealed that TGFβ1 and TGFβ3 mRNA expression levels were higher in ER-positive and/or PR-positive tumors compared to ER- and/or PR-negative tumors. TGFβ2 expression, however, was higher in the ER-negative tumors (p < 0.0001; Figure 3(a)–(c)). This distinct expression pattern of the TGFβ isoforms, in relation with the classical breast cancer markers, further highlight the differential behavior and potential roles played between the three TGFβ isoforms in different types of breast tumors.

TGFβ1, TGFβ2, and TGFβ3 and their association with classical biomarkers in breast cancer. Comparison of the three TGFβ isoform (TGFβ1, TGFβ2, and TGFβ3) mRNA levels and their association with (a) estrogen (ER), (b) progesterone (PR), and (c) HER-2 receptors status by immunohistochemistry (IHC) using bc-GenExMiner v4.0 (Breast Cancer Gene-ExpressionMiner v4.0) online dataset.
Breast cancer subtypes showed differential expression of TGFβ isoforms with more expression of TGFβ1 and TGFβ2 in the highly aggressive triple-negative breast cancer tumors
The different breast cancer molecular subtypes have distinct prognostic and predictive markers and values. 36 We assessed mRNA expression levels of the different TGFβ isoforms in 1881 breast cancer cases, using the GOBO database and Hu classification method. 37 We found higher TGFβ1 and TGFβ2 mRNA levels in basal tumors compared to other molecular subtypes (p ≤ 0.00001; Figure 4(a) and 4(b)). TGFβ3, however, showed a different expression pattern with the highest expression levels found in the luminal A tumors and the least expression in the basal-like subtype (Figure 4(c)). These results indicate that TGFβ isoforms are expressed at variable levels in the different molecular subtypes, suggestive of differential roles played by the different isoforms in the pathogenesis of the distinct molecular subtypes.
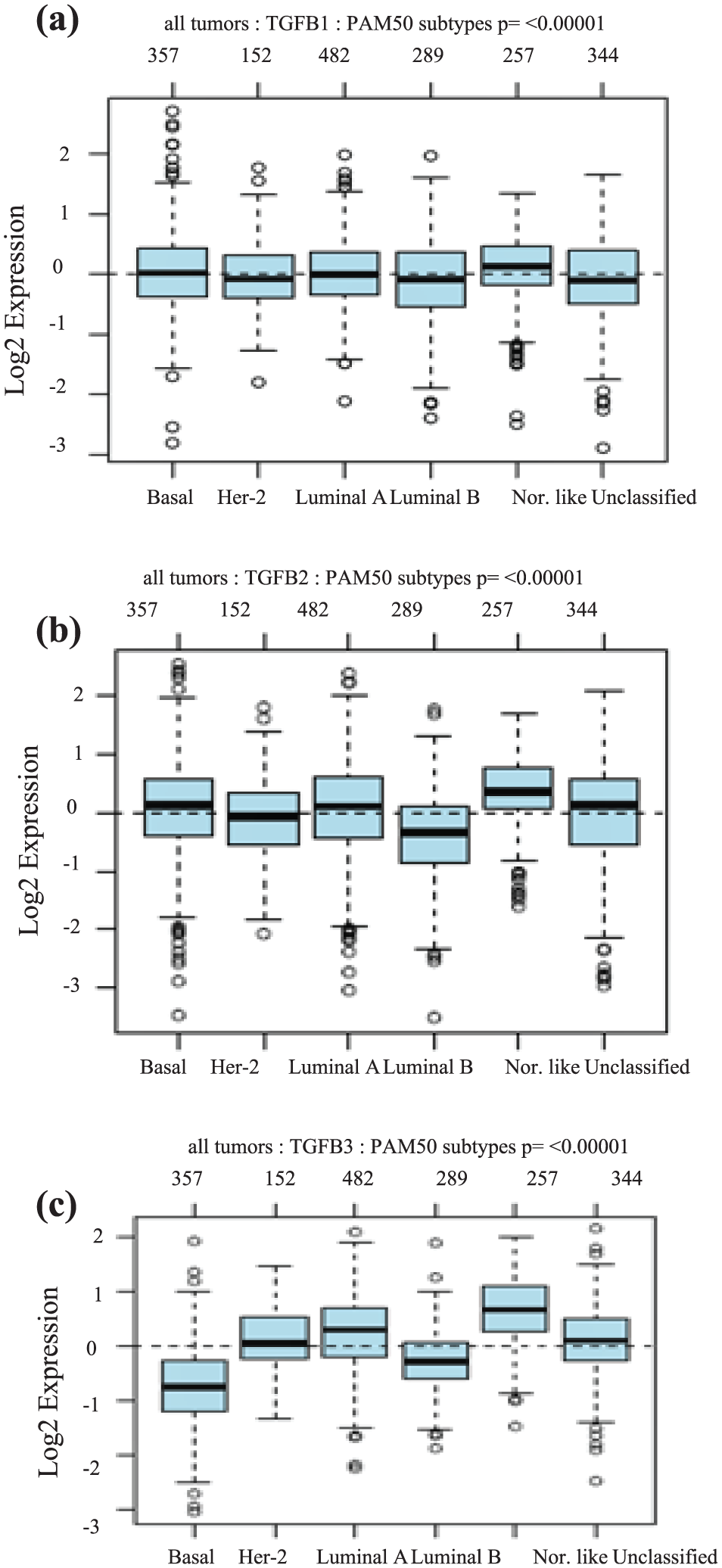
TGFβ1, TGFβ2, and TGFβ3 mRNA expression levels in different breast cancer molecular subtypes. (a) TGFβ1 mRNA levels in human tumor samples stratified according to PAM50 subtype using GOBO database including 1881 breast cancer patients. (b) TGFβ2 mRNA levels in human tumor samples stratified according to PAM50 subtype using GOBO database including 1881 breast cancer patients. (c) TGFβ3 mRNA levels in human tumor samples stratified according to PAM50 subtype using GOBO database including 1881 breast cancer patients.
TGFβ ligand and receptor expression patterns are molecular subtype-specific
We next analyzed whether expression of the TGFβ isoforms could correlate with the expression of the two TGFβ signaling receptors (TβRI and TβRII). Using the Breast Cancer Gene-Expression Miner v4.0 in a cohort of 5260 breast cancer patients, we found TGFβ1 mRNA levels to significantly correlate with TβRII mRNA expression (r = 0.17, p < 0.0001), while showing no association with TβRI mRNA expression (r = 0.01, p = 0.27). In contrast, we found TGFβ2 and TGFβ3 expression to be significantly associated with TβRI expression (r = 0.19, p < 0.0001; r = 0.13, p < 0.0001), while showing no association with TβRII (r = 0.24, p < 0.0001 for TGFβ3; r = 0.14, p < 0.0001 for TGFβ2; Figure 5(a) and (b)).

TGFβ1, TGFβ2, and TGFβ3 mRNA levels and its association with TβRII and TβRI and endoglin mRNA levels in different breast cancer subtypes. (a) TGFβ1, TGFβ2, and TGFβ3 mRNA levels and their association with TβRII mRNA levels in around 5000 human tumor samples using bc-GenExMiner 4.0 database. (b) TGFβ1, TGFβ2, and TGFβ3 mRNA levels and their association with TβRI mRNA levels in around 5000 human tumor samples using bc-GenExMiner 4.0 database. (c) Correlation between different TGFβ isoforms in addition to TβRII and TβRI mRNA levels in different breast cancer subtypes using bc-GenExMiner 4.0 database. (d) TGFβ1, TGFβ2, and TGFβ3 mRNA levels and their association with endoglin mRNA levels in around 5000 human tumor samples using bc-GenExMiner 4.0 database. (e) TGFβ1, TGFβ2, and TGFβ3 mRNA levels and their association with endoglin mRNA levels in 580 basal-like breast cancer subtypes using bc-GenExMiner 4.0 database.
We next more specifically investigated whether the observed association between the different isoforms and their receptors varied depending on the breast cancer subtypes, using the Breast Cancer Gene-Expression Miner v4.0 correlation tool. Interestingly, the association between expression levels of TGFβ1 and TβRII, observed in Figure 5(a), was found to be specific to the basal-like subtype (Figure 5(c)). A similar and significant association was also found between the TGFβ3 isoform and TβRII. However, the association between expression of TGFβ2 and TβRI could only be found in the HER-2 subtype. No clear or significant association could be established in the luminal subtypes, no matter the isoform (Figure 5(c)).
In summary, aggressive basal-like tumors showed significant correlation between TGFβ1 and TGFβ3 with TβRII, while HER-2 tumors only show significant correlation between TGFβ2 and TβRI. Altogether, these data highlight the complex network association between TGFβ ligands and their receptors, in a specific molecular subtype-dependent manner and further support the notion that TGFβ1 and TGFβ3 exhibit a different functional behavior compared to TGFβ2 in breast cancer progression and aggressiveness. These specific patterns of association between the TGFβ isoforms and their receptor may prove useful for the detection/prediction of specific subtypes of breast tumors.
TGFβ1 and TGFβ3, but not TGFβ2, are significantly associated with endoglin in highly invasive breast cancer tumors
Endoglin (CD105) is a TGFβ co-receptor and is found to be highly expressed in endothelial cells. 38 Moreover, endoglin expression was found to be increased in solid tumors and its expression in malignant cells increased their invasive capacity and migration potentials. 38 Interestingly, while endoglin has a constant affinity for both TGFβ1 and TGFβ3, it has a weak affinity for TGFβ2. 6 Thus, to further evaluate the role of each isoforms in breast cancer angiogenesis, invasion, and metastasis, we examined the association of each isoform with endoglin. In addition, analysis of a large cohort of breast cancer patients (over 5000 patients) revealed a strong association between endoglin and TGFβ1 and TGFβ3 (r = 0.2 and r = 0.14) while showing a weaker association with TGFβ2 (r = 0.08; Figure 5(d)). Moreover, patients diagnosed with the most aggressive basal-like subtypes showed a strong positive association between endoglin and TGFβ1 (r = 0.38) and endoglin and TGFβ3 (r = 0.30; Figure 5(e)). In contrast, no association was found between endoglin and TGFβ2 (r = 0.03) in this highly aggressive breast cancer subtype (Figure 5(e)).
These results further emphasize the strong association between expression of TGFβ1 and TGFβ3 with tumor progression and aggressiveness and also suggest that TGFβ2 may exert a different role during these processes. These also highlight the need for developing more specific therapeutic options that may specifically affect one or the other TGFβ isoforms.
TGFβ1 protein expression is upregulated in breast cancer cases compared to normal tissue and correlates with advanced-stage, larger size tumor, and LN-positive tumors
To further confirm our findings, we next evaluated the protein expression level of TGFβ1, the most abundant isoform in most tissues, using a TMA composed of 102 breast cancer cases. Our results showed a significantly higher TGFβ1 expression in invasive breast cancer cases (71%) compared to (20%) normal/benign and (17%) in situ carcinoma (Figure 6(a) and (b)). Next, and to further evaluate the role of TGFβ1 in breast cancer tumor progression, we investigated the association between TGFβ1 protein expression and tumor grade, size, LN status, and stage (Figure 6(a)). While TGFβ1 showed no significant difference with tumor grade, its expression was significantly increased (72%) in larger size tumors (>2 cm) compared to smaller size tumors (17%) and LN-positive tumors (73%) compared to LN-negative (62%) tumors. In addition, TGFβ1 was significantly higher (70.7%) in the more advanced stages II and III compared to earlier stages (17%; stages 0 and I).

TGFβ1 IHC expression and its association with different clinicopathological parameters. (a) Association of TGFβ1 protein expression levels with different clinicopathological parameters using immunohistochemistry. (b) Immunohistochemical expression of TGFβ1 in normal versus malignant breast cancer cases.
Due to the important role of the ER, PR, and HER-2 status in determining patient prognosis and therapeutic options, we also evaluated the association between TGFβ1 protein levels and ER, PR, and HER-2 status (Table 1). While TGFβ1 protein expression levels did not vary according to ER and HER-2 status, a clear association trend was observed with PR expression (Table 1).
Association of transforming growth factor β1 (TGFβ1) expression levels with TβRII and TβRI expression and other classical biomarkers.
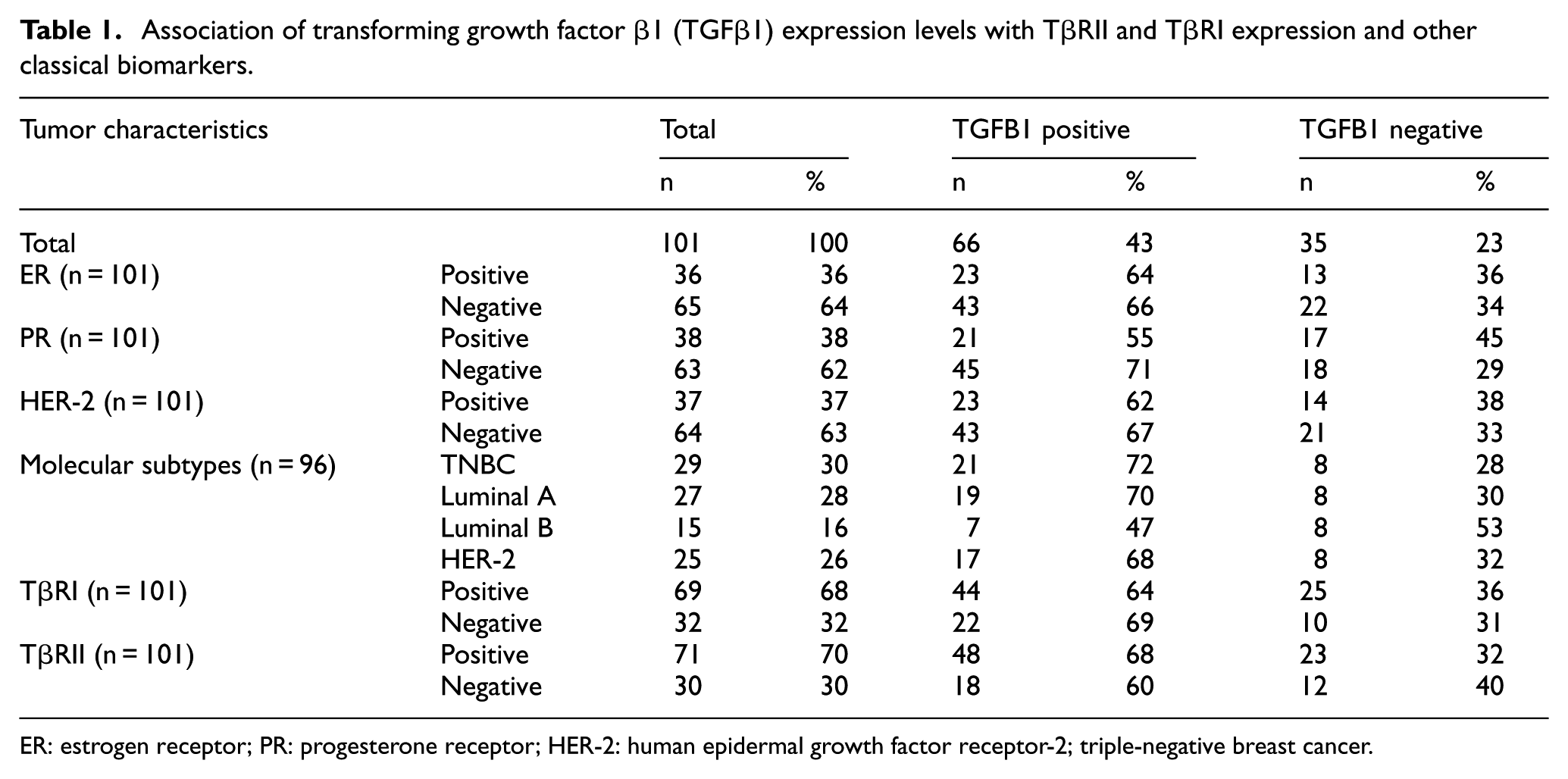
ER: estrogen receptor; PR: progesterone receptor; HER-2: human epidermal growth factor receptor-2; triple-negative breast cancer.
We also assessed the TGFβ1 expression levels in association with different molecular subtypes using the previously described classification method.39,40 As shown in Table 1, TGFβ1 highest expression was observed in the highly aggressive triple-negative breast cancer (TNBC) subtype (74%), while at the lowest in HER-2 (68%) and luminal B (47%) subtypes. Moreover, our immunohistochemistry (IHC) data in breast tumors revealed stronger association between TGFβ1 with TβRII, compared to TGFβ1 and TβRI expression (Table 1). Together, these data highlight the strong association between TGFβ1/TβRII axis and aggressive breast cancer tumors and further define TGFβ1 as a marker of tumor progression, LN involvement, metastasis, and aggressive phenotype in breast cancer.
Discussion
In breast cancer, TGFβ exerts a complex role acting as a tumor suppressor in early breast carcinogenesis while promoting tumor progression, EMT, and metastasis in more advanced breast cancers.1,20,21,26–32 This complexity was also observed with respect to the prognostic value of TGFβ receptors, which also display a dual prognostic role, depending on disease progression and molecular subtype. 39 This could be in part attributed not only to breast cancer heterogeneity but also to the fact the TGFβ family comprises more than one ligand. The three TGFβ isoforms have different tissue-specific expression and are encoded by distinct genes.6,7 Their specific and distinctive biological functions and prognostic values have not been addressed yet. TGFβ1 is the most abundant isoform and plays different physiological roles, including an important role in inducing and promoting EMT, cell motility, invasion, angiogenesis, metastasis, and stemness.18–23 In contrast, TGFβ2 was shown to play a role in reducing immune surveillance of tumor development and enhancing tumor growth. 24 TGFβ3 was shown to play a role in the morphogenesis of lung and normal palate and to regulate epithelial–mesenchymal interaction suggesting a role in the EMT process in cancer.16,17
In normal mammary development, it was shown that different TGFβ isoforms are expressed differentially during the different phases of development of the mammary gland. 8 Interestingly, while TGFβ1 was shown to be reduced during pregnancy, TGFβ2 and TGFβ3 were found to be upregulated during pregnancy. 8
These indicate that the TGFβ isoforms may play different roles during mammary gland development and during breast cancer formation and progression. In this study, we analyzed the expression levels of the TGFβ isoforms alone or in combination with their receptors in large cohorts of breast cancer patients, as well as in normal, benign, in situ, and invasive breast carcinoma of different grades and stages. Interestingly, we found TGFβ1 and TGFβ3, but not TGFβ2, to be expressed at higher levels in invasive breast cancer compared to normal tissues. These findings are in agreement with recent reports showing differential expression of the three isoforms in breast cancer 34 and in malignant grades of HaCaT keratinocytes. 25 Further analysis revealed that only TGFβ1 and TGFβ3 expression levels are higher in bigger size, LN-positive, and metastatic tumors. In contrast, TGFβ2 expression correlates with smaller tumor size, LN-negative tumors, and tumors with no distant metastasis. This is indicative of different functions played by the TGFβ isoforms in breast cancer tumorigenesis and of a differential behavior between the two groups (TGFβ1 and TGFβ3 vs TGFβ2) in association with disease progression. Our results also suggest that the two groups may have opposing role in the advanced-stage tumor compared to early tumors, thereby accounting in part for the dual role of TGFβ pathway in breast carcinogenesis.
Due to the heterogeneous nature of the breast cancer disease, we also evaluated the expression levels of the TGFβ isoforms and receptors in the different molecular breast cancer subtypes. Our results revealed expression correlations between TGFβ1 and TGFβ3 and the type II TGFβ receptor (TβRII) in the most aggressive basal-like breast tumors. In addition, both TGFβ1 and TGFβ3 were also strongly associated with endoglin, which is highly expressed in activated endothelium of tumor vessels and known to play a role in vascular invasion, further highlighting the prominent role for TGFβ1 and TGFβ3 in the promotion of breast cancer invasion. Interestingly, the HER-2-enriched subtype showed a complete different pattern of association between the different isoforms and the receptors, showing only a significant association between TGFβ2 and TβRI. This suggests a different TGFβ signaling interplay between the two most invasive breast cancer molecular subtypes (HER-2 and basal) and may help define more precise therapeutic strategies based on targeting specific TGFβ isoforms in specific breast cancer subtypes. Moreover, while TGFβ1 and TGFβ3 are expressed at higher levels in LN-positive and metastatic tumors, TGFβ2 levels are lower in these advanced, invasive tumors, raising the appealing hypothesis of having opposing roles played by the different isoforms in regulating breast cancer progression and metastasis.
Our results also highlight a distinct association between the different TGFβ isoform and the classical breast cancer biomarkers (ER, PR, and HER-2). This suggests potential differential roles for each of the isoforms in modulating the classical biomarkers’ expression. Thus, this work will provide an interesting experimental starting point for future studies to specially address the role of the different isoforms.
In summary, while we found all three TGFβ isoforms to be expressed and biologically active in breast cancer, we also showed that they exhibit differential expression as well as specific co-expression with their receptors. These distinct expression patterns may, in turn, dictate different roles, prognosis values, and disease outcomes between the isoforms, according to the stage, grade, metastatic status, and molecular type of the disease. Further investigation of the precise role and function of these isoforms in the different molecular subtypes will help refine more precise therapeutic options by targeting specific isoforms in a subtype-specific manner and lead to improved patient response and outcome.
Methods
TMAs
TMAs of 102 breast cancer patients were purchased from Pantomics, Inc. (Richmond, CA, USA; BRC1021). The cases include 5 normal/benign cases, 6 in situ carcinoma, and 91 invasive breast cancer tissues. The grade, stage, estrogen receptor (ER), progesterone receptor (PR), HER-2, and Ki67 protein expression levels were also provided. Based on the ER, PR, HER-2, and Ki67 expression and status, the cases were reclassified according to molecular subtypes as described. 40
IHC
The slides were first baked at 55°C for 30 min. This was followed by deparafinization in xylene and rehydration in 100%, 95%, 70%, and 50% ethanol and by heat-induced antigen retrieval in sodium citrate 10 mM, pH 6.0 buffer. This was further followed by hydrogen peroxide block incubation for 10 min and Ultra V Block. TGFβ antibody was added and incubated for 1 h at room temperature, and then, UltraVision LP Detection System HRP Polymer and DAB Plus Chromogen (Thermo Fisher Scientific, Fremont, CA, USA) were added. The slides were then scanned using Aperio XT slide scanner (Leica Biosystems, Concord, ON, Canada) and analyzed by an experienced pathologist.
IHC scoring
ER and PR were scored according to Allred scoring system, while HER-2 expression was classified according to ASCO/College of American Pathologists Guidelines for HER-2 testing in breast cancer. Ki67 cases were considered positive if they scored 14% or more and negative if less than 14%. TGFβ1 was scored in a semi-quantitative method with a score of 0 indicating undetectable expression, +1 if 1%–20% of tumor cells were positively stained, and +2 if >20% of tumor cells were positive. Both scores 0 and +1 was considered as negative, while +2 score was considered positive. The slides were scored independently by two investigators.
Data mining
Different large publicly available databases include ONCOMINE, GOBO, and Breast Cancer Gene-Expression Miner database. TCGA dataset of ONCOMINE database which includes 398 invasive breast cancer cases and 61 normal breast cancer tissues was used to investigate the difference in TGFβ isoform expression between normal and invasive breast cancer tumors. In addition, Sorlie dataset was also used to investigate the correlation between TGFβ mRNA levels and tumor size. Gene expression of the three TGFβ isoforms in different molecular subtypes of breast cancer was evaluated using GOBO database which include data of more than 1881 breast cancer patients. Finally, Breast Cancer Gene-Expression Miner v4.0 (bc-GenExMiner v4.0) online dataset (http:// bcgenex.centregauducheau.fr) comprising previously published gene expression datasets from 15 independent breast cancer studies totaling 2413 tumors was used to evaluate the association between different isoform expression and patients’ outcome.
All experimental protocols and procedures were performed in accordance to McGill University regulations. All experimental protocols and procedures were approved by McGill University.
Footnotes
Author contributions
Mahmood Y Hachim and Ibrahim Y Hachim performed experiments, bioinformatic analysis, and drafting of the article. Meiou Dai performed experiments. Suhad Ali contributed to the experimental design and corrections of the article. Jean-Jacques Lebrun is the principal investigator who initiated the research design and supervision of the project and writing of the article.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Canadian Institutes for Health Research (CIHR to J.-J.L.).
