Abstract
The proteasome beta-4 subunit is required for the assembly of 20S proteasome complex, forming a pivotal component for the ubiquitin–proteasome system. Emerging evidence indicates that proteasome beta-4 subunit may be involved in underlying progression and mechanisms of malignancies. However, the role of proteasome beta-4 subunit in melanoma is currently unknown. Here, we reported that proteasome beta-4 subunit was markedly upregulated in human melanoma tissues and cells, compared with normal skin samples. High proteasome beta-4 subunit levels were significantly associated with poor overall survival in patients with melanoma. Proteasome beta-4 subunit knockdown strongly decreased melanoma cell growth in vitro and in vivo. We further identified miR-148b as a negative regulator of proteasome beta-4 subunit. Enforced expression of miR-148b resulted in vitro growth inhibition of melanoma cells, whereas this inhibition was further abolished by enforced expression of proteasome beta-4 subunit. Our findings, for the first time, indicated that the miR-148b/proteasome beta-4 subunit axis contributed to the development of melanoma, revealing novel therapeutic targets for the treatment of melanoma.
Introduction
Melanoma is one of the most devastating skin cancers with highly increasing incidence these years. In a recent article, 76,380 new cases and 10,130 deaths were estimated in the United States during 2016. 1 Although various gene-targeted therapy and immune-based approach were introduced lately,2,3 not all the patients can benefit from these treatments due to the elusive malignant properties of melanoma. However, the understanding of melanoma is limited, and the potential mechanisms driving this malignant progress still need to be further studied to explore potential therapeutic strategies against melanoma.
It is well established that proteasome participates in regulating a variety of cellular functions by accelerating the proteolysis to degrade damaged, misfolded, or unwanted proteins. Proteasome beta-4 subunit (PSMB4), known as 20S proteasome subunit beta-7, is located in nucleus and cytoplasm 4 and contributes to the complete assembly of 20S proteasome complex. 5 Since emerging evidence indicates that the ubiquitin–proteasome system (UPS) plays an essential role in the development of various tumors, 6 PSMB4 may be involved with the oncocytes survival and tumor growth as it is a central component of UPS. Recent studies elucidated that PSMB4 is upregulated in multiple tumors and is involved in tumor progression and poor prognosis,7–9 making it as a potential oncogene. 10 However, the gene and protein expression of PSMB4 and its roles in the development of melanoma remain poorly understood.
MicroRNAs (miRNAs) are a family of short single-stranded non-coding RNA molecules containing about 22 nucleotides. MiRNAs can function in RNA silencing and regulate human genes expression by directly paring to the 3′-untranslated regions (3′-UTR) of messenger RNAs (mRNAs). 11 Increasing evidences suggest that many miRNAs are involved in several tumor occurrence, progression, and development by regulating their target genes, 12 indicating a new direction for tumor research and treatment. Cumulative evidences have demonstrated that miR-148b was aberrantly expressed and functions as a tumor suppressor by regulating multiple genes and signaling pathways in multiple tumors, including pancreatic cancer, 13 non–small cell lung cancer (NSCLC), 14 hepatocellular carcinoma, 15 and glioma. 16 However, the potential biological roles of miR-148b in melanoma are still not clear.
In this study, we investigated the expression of PSMB4 in melanoma tissues and cell lines, and the in vitro and in vivo effects of PSMB4 on melanoma growth were further evaluated. PSMB4 was then predicted and confirmed as a functional target gene of miR-148b, indicating that the miR-148b/PSMB4 axis may be a novel therapeutic target in melanoma.
Materials and methods
Tissue specimens and cell lines
Melanoma tissues (12 primary melanoma and 27 secondary metastasis) were obtained from untreated patients (male: n = 18, female: n = 21, average age: 57.45) during surgery, following informed consent and the Institutional Review Board (IRB) approval. A total of 12 patients had stage I/II melanoma and 27 patients had stage III/IV melanoma. Normal skin was adjacent tissues in surgical specimens. Wild-type (WT) BRAF (SK-MEL-1, SK-MEL-13. and SK-MEL-3), BRAF mutated (A375, G-361, SK-MEL-24, SK-MEL-28, HT144, A2058, and WM-115) melanoma cell lines, and 293T cells were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA) and were cultured in RPMI-1640, supplied with 10% fetal bovine serum (FBS; Hyclone, Logan, UT, USA) and maintained in an incubator at 37°C with 5% CO2.
Quantitative reverse transcription polymerase chain reaction for miRNAs and mRNAs
Total RNA from melanoma tissues and cells was extracted with the TRIzol Reagent (Invitrogen, Life Technologies, Carlsbad, CA, USA). Complementary DNA (cDNA) was obtained using the SuperScript First-Strand Synthesis System (Invitrogen). SYBR Green PCR Master Mix and ABI Prism 7500 Fast System (Applied Biosystems, Foster City, CA, USA) were used to validate the gene expression. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) is an internal control. The primers used were as follows—PSMB4: 5′-GAAGCGTTTTTGGGGTCGC-3′ (forward primer) and 5′-GAGTGGACGGAATGCGGTA-3′ (reverse primer); GAPDH: 5′-ACAACTTTGGTATCGTGGAAGG-3′ (forward primer) and 5′-GCCATCACGCCACAGTTTC-3′ (reverse primer). For the analysis of miR-148b, TaqMan microRNA assay (Applied Biosystems) was performed as previously described. 13 MiR-148b levels were normalized by U6 small nuclear RNA.
Plasmid constructs and transfection assays
The human PSMB4 (Gene ID: 5692; NM_002796.2) short hairpin RNA (shRNA) lentiviral particles (sc-76269-V; Santa Cruz Biotechnology, Santa Cruz, CA, USA) and the non-target control lentivirus (sc-108080; Santa Cruz Biotechnology, CA, USA) were used to infect A375 and SK-MEL-1 cells. After selection with 5 µg/mL puromycin for 3 weeks, the stable transfectants were generated. Enforced miR-148b expression was carried out by transfecting plasmid pEZX-MR04 expressing pre-miR-148 (HmiR0185-MR04; GeneCopoeia, Guangzhou, China) and scrambled control (CmiR0001-MR04; GeneCopoeia). Lipofectamine 3000 Reagent (Invitrogen) was used to perform cell transfection according to the manufacturer’s protocol.
Western blotting and immunohistochemistry
Tissues from patients and cells after treatment were lysed in radioimmunoprecipitation assay (RIPA) buffer containing protease inhibitors (Sigma, St. Louis, MI, USA). Then, the whole-cell lysates were subjected to Western blotting analysis using PSMB4 primary antibody (1:1000; Santa Cruz Biotechnology), with anti-GAPDH (1:1000; Abcam, San Francisco, CA, USA) as an internal control. For immunohistochemistry (IHC), tissues from patients and tumors from mice were fixed in 4% formaldehyde. After epitope retrieval (ethylenediaminetetraacetic acid (EDTA) buffer, pH 8.8) and blocking steps, the 3 µm thick sections were treated with PSMB4 antibody (1:100; sc-390878; Santa Cruz Biotechnology) or Ki67 antibody (1:100; ab16667; Abcam). The immunohistochemical staining was observed and photographed under an inverted light microscope after counterstaining with hematoxylin.
Cell proliferation assay
Cell proliferation assay was performed using methyl tetrazolium salt (MTS) assay (Sigma, St. Louis, MO, USA) according to the manufacturer’s protocol. Briefly, 3000 A375 and SK-MEL-1 cells were seeded onto 96-well culture plates after transfection, and the optical density (OD; at 592 nm) values were detected at 24, 48, and 72 h.
In vivo animal models
For xenograft models, A375 cells were transfected with PSMB4 shRNA lentivirus (sh-PSMB4) or negative control (sh-NC), and the stable transfectants were selected with puromycin. A volume of 2 × 106 A375 cells expressing sh-PSMB4 or sh-NC were injected subcutaneously into the back of 6-week-old male BALB/c mice (n = 10). Electronic calipers were used to examine the length and width of tumors; then, the tumor volume V (mm3) was calculated by the equation: (Length × Width2)/2.
Dual luciferase assay
For reporter assays, WT or mutant (MT) 3′-UTR of PSMB4 (length: 115 bp) was amplified and cloned into pTZ57R/T vector (K1213; Fermentas, Amherst, NY, USA). After digestion with XhoI and NotI, the fragment was inserted into psiCHECK™-2 Vector (C8021; Promega, Madison, WI, USA). Cells were co-transfected with 0.5 µg psiCHECK™-2 vector harboring PSMB4 3′-UTR and pre-miR-148 expression vector. Dual-Glo® Luciferase Assay System (E2940; Promega) was used to perform luciferase assays 24 h after transfection, according to the manufacturer’s protocol.
Statistical analysis
Each experiment was performed three independent times, and the data were showed as mean ± standard error of the mean (SEM) Statistical significance between groups was evaluated by t test, one-way analysis of variance (ANOVA), or log-rank test using GraphPad Prism software (GraphPad Software, San Diego, CA, USA).
Results
PSMB4 was upregulated in malignant melanoma and associated with poor overall survival
To investigate the association of PAMB4 with melanoma tumorigenesis, we first explored the expression of PAMB4 in 22 normal skins and 39 melanoma tissues. After comparison of these samples by quantitative reverse transcription polymerase chain reaction (qRT-PCR), we found a significant increase in the levels of PAMB4 in melanoma tissues (p < 0.001; Figure 1(a)). Similar results were obtained by reanalyzing a previously published dataset (GEO# GSE 3189; Figure 1(b)), indicating that PSMB4 may be involved in melanoma development and progression. Abnormal expression of PSMB4 in melanoma tissues was further confirmed by immunohistochemical assays (Figure 1(c)). Furthermore, overall survival (OS) analysis for 30 patients was performed, and the results indicated that patients with high PSMB4 expression suffered a poorer prognosis than those with low levels (p = 0.0323; Mantel–Cox log-rank test; Figure 1(d)). In addition, PSMB4 was also overexpressed in a variety of melanoma cell lines (Figure 1(e)).

PSMB4 is upregulated in malignant melanoma and is associated with poor overall survival. (a) The values of PSMB4 expression in 22 normal skin and 39 melanoma tissues were measured by quantitative RT-PCR analysis (qRT-PCR). (b) The previously published dataset, GSE3189, was obtained from National Center for Biotechnology Information (NCBI; https://www.ncbi.nlm.nih.gov), and reanalysis of PSMB4 expression was performed. (c) Immunohistochemical staining of PSMB4 expression in normal skin and melanoma tissues was performed, and the representative graphs were shown (scale bar: 50µm). (d) The overall survival for patients with melanoma according to PSMB4 expression was analyzed using Mantel–Cox log-rank test (p = 0.0323). (e) The expression of PSMB4 in 4 normal skin tissues and 10 typical melanoma cell lines were examined by qRT-PCR.
Silencing PSMB4 inhibited melanoma cell growth in vitro and in vivo
To further explore the roles of PSMB4 in melanoma development, RNA interference (RNAi) was used to specifically silence the expression of PSMB4 with lentiviral shRNA in the two typical melanoma cell lines, A375 and SK-MEL-1. As shown in Figure 2(a), PSMB4 knockdown significantly inhibited the proliferation of A375 melanoma cells, and the efficiency of silence was confirmed by Western blotting. Similar results were obtained by using SK-MEL-1 cells (Figure 2(b)). The effects of PSMB4 on colony-forming ability were further measured. Slicing PSMB4 resulted in a significant inhibition of colony formation of A375 and SK-MEL-1 cells (Figure 2(c)), indicating that PSMB4 may play an important role in the growth of melanoma.
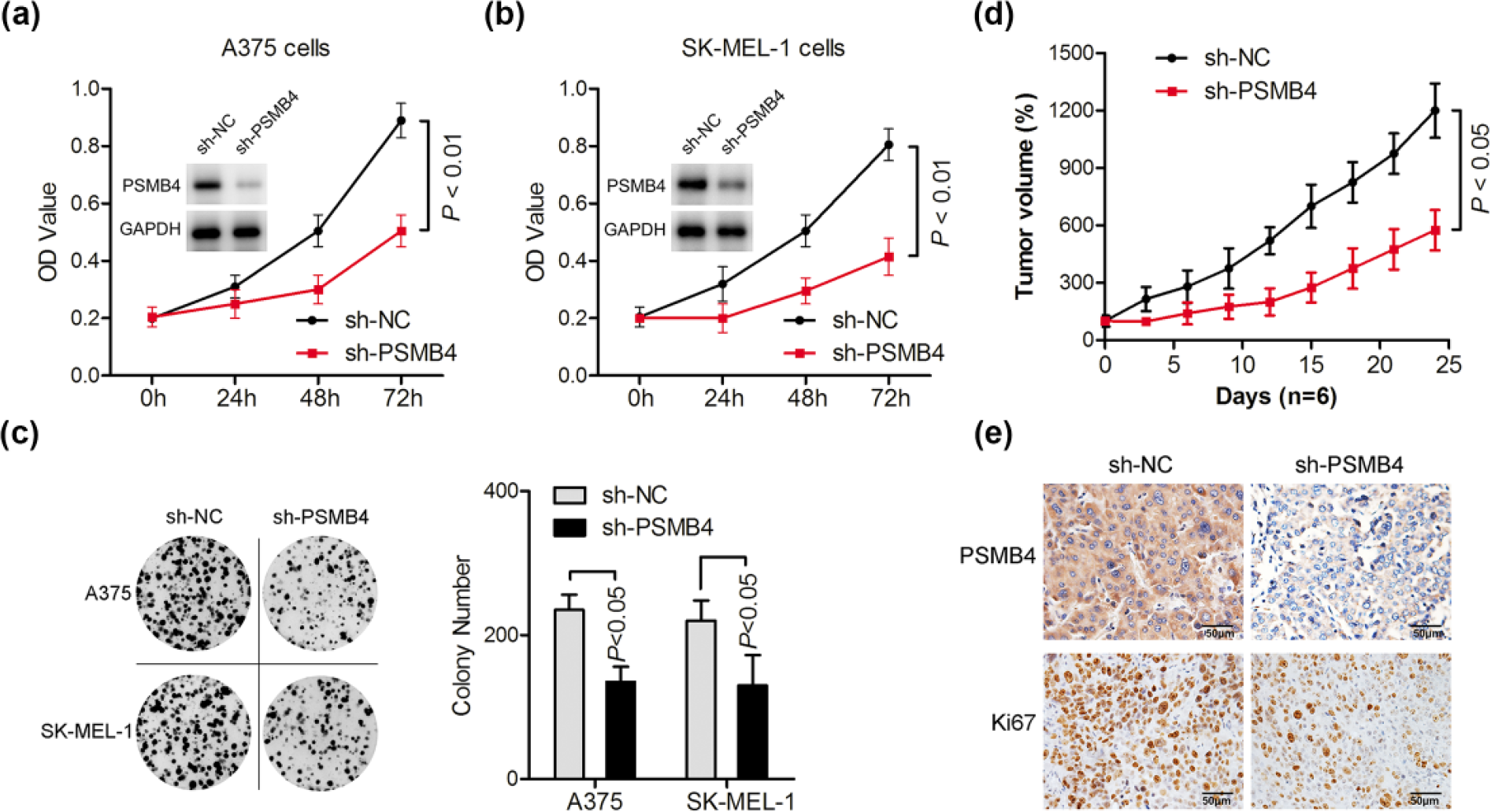
Silencing PSMB4 inhibited melanoma cell growth in vitro and in vivo. (a) A375 cells were transduced with negative control (sh-NC) or PSMB4-specific shRNA (sh-PSMB4); melanoma growth was evaluated by detecting the OD (592nm) values at 24, 48, and 72 h. The protein levels of PSMB4 were assessed by Western blotting. (b) After transduction, the effects of PSMB4 on the proliferation of SK-MEL-1 melanoma cells were determined. The expression of PSMB4 was detected by Western blotting. (c) Cells were transduced with PSMB4-specific shRNA or negative control (sh-NC), and the effects of PSMB4 on melanoma colony formation were evaluated. Colony numbers were then counted. (d) After transduction with PSMB4-specific shRNA or negative control (sh-NC), A375 cells were inoculated subcutaneously into mice (n = 10). Tumor growth was evaluated by monitoring the tumor volume for every 3 days using the equation (Length × Width2)/2. (e) PSMB4 and Ki67 immunohistochemical staining of A375 tumor xenografts (scale bar: 50 µm).
Effect of PSMB4 on tumorigenic potential of melanoma cells was evaluated in vivo. After transduction with lentivirus containing sh-PSMB4 or the empty vector (sh-NC), A375 cells were inoculated subcutaneously in mice (n = 10), and tumor growth was monitored. Deletion of PSMB4 reduced the growth of A375 melanoma xenografts compared with the control group (p < 0.05; Figure 2(d)). Moreover, immunohistochemical staining of excised tumors demonstrated decreased PSMB4 and Ki67 levels in sh-PSMB4-transduced tumors compared with controls (Figure 2(e)) that were associated to reduced proliferation in vivo.
MiR-148b regulated PSMB4 expression by binding to its 3′-UTR
Since the levels of human oncogenes are normally influenced by naturally occurring miRNAs, we further explore which miRNAs regulated PSMB4 expression in melanoma. The TargetScan program was used to search the potential miRNAs targeting PSMB4 gene. There were conserved binding sites for miR-148b in the 3′-UTR region of the PSMB4, as shown in Figure 3(a). We examined the regulating effect of miR-148b on PSMB4 mRNA using dual luciferase report system. A reduced luciferase activity was found in 293T cells co-transfected with 3′-UTR of PSMB4 and miR-148b, while there was no change with MT PSMB4 3′-UTR or negative control miRNA (miR-NC; Figure 3(b)). Similar results were observed in A375 or SK-MEL-1 cells transfected with 3′-UTR sequence of PSMB4 together with miR-148b but not with the negative control (Figure 3(c)). Moreover, Western blotting analysis demonstrated that overexpression of miR-148b significantly repressed the expression of PSMB4 in A375 or in SK-MEL-1 cells (Figure 3(d)), indicating that PSMB4 was regulated by miR-148b in melanoma cells. In addition, a significant negative correlation was found between PSMB4 and miR-148b (p < 0.01; r = −0.4975) in melanoma tissues (Figure 3(e)).

MiR-148b regulated PSMB4 expression by binding to its 3′-UTR. (a) PSMB4 3′-UTR containing the wild-type (WT) and mutated (MT) binding sequences for miR-137 are shown. (b) Dual luciferase assay of 293T cells transfected with luciferase reporter vector along with the miR-148b expression vectors or empty vector (miR-NC) as control. (c) After transfection with miR-148b for 48 h, the relative luciferase activities were examined in A375 and SK-MEL-1 cells. (d) The protein levels of PSMB4 in A375 and SK-MEL-1 cells were detected by Western blotting after transfection with miR-148b or negative control (miR-NC) for 48 h. (e) The correlation between PSMB4 and miR-148b levels in melanoma tissues was measured.
MiR-148b suppressed tumor growth through inhibiting PSMB4 expression
Some studies reported that the miR-148b frequently affects tumor development as a suppressor gene; we investigated whether miR-148b was able to influence melanoma growth. A375 and SK-MEL-1 cells were transfected with miR-148b, and the proliferation was examined by calculating the average absorbance (OD value) at 592 nm. Transfection of miR-148b led to an obvious inhibition in melanoma cell growth (Figure 4(a)). Notably, the ability of cell proliferation was further restored by re-expression of PSMB4 in A375 and in SK-MEL-1 cells (Figure 4(b)). Although overexpression of miR-148b markedly decreased the number of colony-forming cells, re-expression of PSMB4 in A375 and in SK-MEL-1 cells, as confirmed by Western blot analysis (Figure 4(c)), rescued the ability of tumor cell colony formation (Figure 4(d) and (e)).

MiR-148b suppressed tumor growth through inhibiting PSMB4 expression. (a) A375 and SK-MEL-1 cells were transfected with vectors expressing miR-148b or control (miR-NC), and melanoma cell proliferation was determined by detecting the OD (592 nm) values at 24, 48, and 72 h. (b) After transfection with expression vectors of miR-148b and PSMB4 for 72 h, cell proliferation was examined in A375 and SK-MEL-1 cells, (c) and PSMB4 expression was evaluated by Western blotting. After treatment as described above, (d) colony formation assay was performed and (e) the representative graphs were shown.
Discussion
Melanoma is responsible for about 74% of skin cancer–related deaths. 1 Although inhibiting mutated BRAF (V600E) has shown an OS benefit in patients with melanoma, 17 targeted therapy against melanoma with WT BRAF remains an outstanding issue. Recently, the U.S. Food and Drug Administration (FDA) approved ipilimumab (Yervoy) and nivolumab (Opdivo) in clinical treatment of BRAF V600 WT melanoma based on the results from CheckMate-066.18,19 Also, immune checkpoint therapy and chimeric antigen receptor T-cell immunotherapy (CAR-T) have been used for the treatment of melanoma recently.20,21 Since durable responses are not common during such regimens, more study is still underway to explore other valuable targets that may provide more effective treatments against melanoma.
In this study, we first compared the expression of PSMB4 in melanoma and in normal skin. The results showed that PSMB4 in melanoma tissues was higher than that in normal skin (Figure 1(a)). We further found that high PSMB4 expression was significantly related to poor OS (p < 0.05; Figure 1(d)), indicating that PSMB4 may act as a potential cancer driver gene in melanoma. Recent studies have also reported the upregulation of 26S Proteasome Subunit PSMB4 in several tumors and association with cancer growth and poor prognosis.8–10 To assess the contribution of PSMB4 in melanoma development, we studied the effects of silencing PSMB4 on the growth of melanoma cells. Indeed, here we found that targeting PSMB4 triggered marked growth-inhibitory effects in melanoma cells in vitro and in vivo (Figure 2), representing a target amenable of therapeutic intervention for melanoma.
It is well known that miRNAs are able to downregulate oncogenes involved in tumor development by directly binding to the 3′-UTR sequences. 22 In this regard, miR-148b was further algorithmically predicted to be a potential upstream regulator of PSMB4 (Figure 3(a)). We confirmed the regulatory function of miR-148b by luciferase reporter assay together with Western blotting analysis, demonstrating that miR-148b was able to downregulate PSMB4 expression (Figure 3). A negative correlation between miR-148b and PSMB4 was also found in melanoma tissues (Figure 3(e)). Functionally, overexpression of miR-148b resulted in marked suppression of PSMB4 expression and cell proliferation (Figure 4(a)), consistent with previous findings, shown that miR-148b functions as a tumor suppressor in several cancers.13–15 It has been reported that enhanced miR-148b expression inhibited the dissemination of melanoma or breast cancer cells, 23 indicating the contribution of miR-148b/PSMB4 axis in melanoma metastasis, which need be further studied. Notably, these suppressive effects on cell growth by miR-148b were further abolished by enforced expression of PSMB4 (Figure 4). These results revealed a miR-148b/PSMB4 axis that may contribute to melanoma growth, making it a potential target in targeted therapy against melanoma.
In conclusion, our findings highlighted the effects of the miR-148b/PSMB4 axis in regulating melanoma cell growth and in providing novel scenarios and new insights into the development of melanoma. Since oncogenic functions of PSMB4 and tumor suppressor roles of miR-148b were elucidated in this study, it would be worth investigating further to explore more effective targeted intervention strategies against melanoma.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
