Abstract
Epidermal growth factor receptor (EGFR)-tyrosine kinase inhibitors (TKIs) are first-choice treatments for advanced non-small-cell lung cancer patients harboring EGFR mutations. Although EGFR mutations are strongly predictive of patients’ outcomes and their response to treatment with EGFR-TKIs, early failure of first-line therapy with EGFR-TKIs in patients with EGFR mutations is not rare. Besides several clinical factors influencing EGFR-TKI efficacies studied earlier such as the Eastern Cooperative Oncology Group performance status or uncommon mutation, we would like to see whether semi-quantify EGFR mutation gene expression calculated by 2−ΔΔct was a prognostic factor in EGFR-mutant non-small cell lung cancer patients receiving first-line EGFR-TKIs. This retrospective study reviews 926 lung cancer patients diagnosed from January 2011 to October 2013 at the Kaohsiung Chang Gung Memorial Hospital in Taiwan. Of 224 EGFR-mutant adenocarcinoma patients, 148 patients who had 2−ΔΔct data were included. The best cutoff values of 2−ΔΔct for in-frame deletions in exon 19 (19 deletion) and a position 858 substituted from leucine (L) to an arginine (R) in exon 21 (L858R) were determined using receiver operating characteristic curves. Patients were divided into high and low 2−ΔΔct expression based on the above cutoff level. The best cutoff point of 2−ΔΔct value of 19 deletion and L858R was 31.1 and 104.7, respectively. In all, 92 (62.1%) patients showed high 2−ΔΔct expression and 56 patients (37.9%) low 2−ΔΔct expression. The mean age was 65.6 years. Progression-free survival of 19 deletion mutant patients with low versus high expression level was 17.07 versus 12.04 months (P = 0.004), respectively. Progression-free survival of L858 mutant patients was 13.75 and 7.96 months (P = 0.008), respectively. EGFR-mutant lung adenocarcinoma patients with lower EGFR gene expression had longer progression-free survival duration without interfering overall survival.
Keywords
Introduction
Lung cancer is the leading cause of cancer-related death worldwide and remains a major health problem, with an incidence that is increasing in Taiwan.1,2 Non-small-cell lung cancer (NSCLC) is the most common subtype of lung cancer and accounts for 80% of all lung cancer patients, the majority of whom have disease at an advanced stage at the time of diagnosis. 3 Among NSCLC patients, 60% have subtype histology corresponding to adenocarcinoma. Recently, targeted therapy with tyrosine kinase inhibitors (TKIs)—based on the presence of tumor molecular markers—has become a treatment option for patients with advanced stage NSCLC. This strategy targets the epidermal growth factor receptor (EGFR). 4 Among the 20%–60% of NSCLC patients harboring EGFR gene mutations, the response rate to TKIs is approximately 75%. Therefore, detection of EGFR mutations is important for predicting the response to TKIs in first-line therapy.5,6 Previous studies have shown that for patients with TKI-sensitive EGFR mutations, progression-free survival (PFS), overall survival (OS), and quality of life are better in patients treated with TKIs compared with conventional chemotherapy.7,8 As large tumor specimens are often difficult to obtain in patients with advanced stage NSCLC, a highly sensitive method for analysis of small samples is very important. 9 The Scorpion amplification refractory mutation system (ARMS) is a highly sensitive technique, with a sensitivity of about 1% for the detection of EGFR mutations. This method involves analysis of polymerase chain reaction (PCR) products generated from small tumor specimens, such as pleural effusion fluid or tissue samples obtained by transbronchial biopsy. 10 The Scorpion-ARMS method is an effective, rapid, and unique technique that has been approved by the US Food and Drug Administration (FDA) for the detection of EGFR mutations. 11 The two most common EGFR mutations in NSCLCs are deletions in exon 19 and a point mutation in codon 858 of exon 21 (L858R). These two mutations have been shown to account for 90% of all EGFR mutations in those who show good response to TKI treatment.12,13 Real-time quantitative PCR has been shown to be a better method for quantification of gene expression than immunohistochemistry and fluorescence in situ hybridization immunostaining. Using real-time semi-quantitative PCR, relative expression of target genes can be calculated as 2−ΔΔct.14,15 The detection of EGFR mutations may influence treatment and prognosis, especially in NSCLC patients who received TKIs as first-line tumor therapy.16,17 The relationship between quantitative expression levels of the different mutant EGFR genes and prognosis in patients who receive TKI treatment is still unknown, and represents an area for further investigation. We therefore conducted a retrospective analysis to investigate the impact of EGFR gene expression levels in advanced stage lung adenocarcinoma patients. We also examined if baseline clinical parameters affect EGFR gene expression levels.
Materials and methods
Patient and clinical characteristics
The present retrospective study reviewed consecutive patients in the period from January 2011 to October 2013 at the Kaohsiung Chang Gung Memorial Hospital, a tertiary medical center in southern Taiwan. Patients were followed-up until March 2015. Patients eligible for this study were adults aged ≥18 years with NSCLC, confirmed histologically or cytologically as stage III or IV adenocarcinoma, with a deletion in exon 19 or L858R mutation of the EGFR gene, and who received EGFR-TKIs as first-line cancer therapy. Patients were excluded if they had received other chemotherapies, immunological therapies, or targeted therapy. All patients had to have completed assessment of the clinical parameters, blood biochemistry analysis, serum carcinoembryonic antigen (CEA), chest radiography, chest computed tomography, brain magnetic resonance imaging, and bone scintigraphy 4 weeks before initial treatment. Clinical parameters included age, sex, smoking status, body mass index (BMI), and Eastern Cooperative Oncology Group (ECOG) performance status (PS). This study was approved by the Institutional Review Board of Kaohsiung Chang Gung Memorial Hospital. The requirement to obtain informed consent from patients was waived.
EGFR mutation testing
Tumor specimens were obtained by bronchoscopy, computed tomography (CT)-guided biopsy, pleural effusion cytology, or surgical procedures. The specimens were fixed with formalin and embedded in paraffin. EGFR mutational analyses were performed using Scorpion-ARMS PCR. Using fragments amplified from genomic DNA extractions (using an EGFR RGQ PCR Kit, Qiagen), real-time PCR reactions were performed in order to detect specific EGFR gene mutations.18,19 Deletions in exon 19 and L858R mutations were the two most common types of EGFR gene mutation. To determine the expression of target genes in different specimens, normalization based on internal reference gene expression was performed, and the average of the normalized cycle threshold value (Δct) was defined as the peak of the cycle at maximum curvature of the second derivative growth curve. For real-time semi-quantification of target gene expression, 2−ΔΔct was calculated, where Δct = ct (target gene) − ct (internal reference gene) that was used for interpretation of the results. This calculation assumes 100% efficiency for all PCR reactions.
EGFR-TKI treatment response evaluation
To assess disease status and tumor treatment response, patients were routinely followed up by chest radiography for several weeks and chest CT was performed every 2–3 months. Disease progression was evaluated by a clinician, based on Response Evaluation Criteria in Solid Tumors (RECIST) criteria 1.1. 20 PFS was defined as the time interval between the first day of treatment with EGFR-TKIs and either the date on which disease progression was established, the date on which the patient died (from any cause), or the date of the final study visit at the end of follow-up. OS was defined as the time interval between the first day of treatment with EGFR-TKIs and either the date of death, the date of the final study visit at the end of follow-up, or loss to follow-up.
Statistical analyses
SPSS statistical software, version 21.0 (SPSS Inc., Chicago, IL, USA), was used for statistical analysis. The relationships between EGFR status and clinical characteristics were analyzed with Fisher’s exact test, independent t-tests, and Mc Nemar and Kappa tests. MedCalc (version 14.10.2) was used for analyses. Receiver operating characteristic (ROC) curves and Youden’s index were used to evaluate the optimal cutoff values for 2−ΔΔct, in order to determine the prognostic value of EGFR gene expression levels. To understand the influences of baseline clinical parameters on EGFR gene expression levels, we used a chi-square test to compare categorical variables, and an independent samples t-test to compare continuous data.
The survival curves of PFS and OS were prepared using the Kaplan–Meier method, and survival for different patient subgroups was compared by the log-rank test. Statistical significance was defined as P < 0.05 (two-tailed).
Results
Patient characteristics
Of 926 patients who were diagnosed with lung cancer from January 2011 to October 2013, 442 patients with advanced lung adenocarcinoma were screened for EGFR mutations. Among these, 224 (50.7%) had EGFR gene mutations. A total of 148 patients were ultimately diagnosed with EGFR mutations that are common in advanced lung adenocarcinoma, using the Scorpion-ARMS and 2−ΔΔct method. The clinical and pathological characteristics of these 148 patients are summarized in Table 1. The mean age of the study population was 65.6 years, 55.5% were females, and 68.2% never smoked. ECOG PS was less than 2 in 88.5% of patients, and 85.9% had cancer at stage IV. The mean CEA value was 184. Exon 19 deletions accounted for 48.6% of EGFR gene mutations, while the L858R mutation accounted for 51.4%. PFS was greater than 12 months for both subtypes of patient.
Clinical characteristics and therapy responses of all 148 patients.
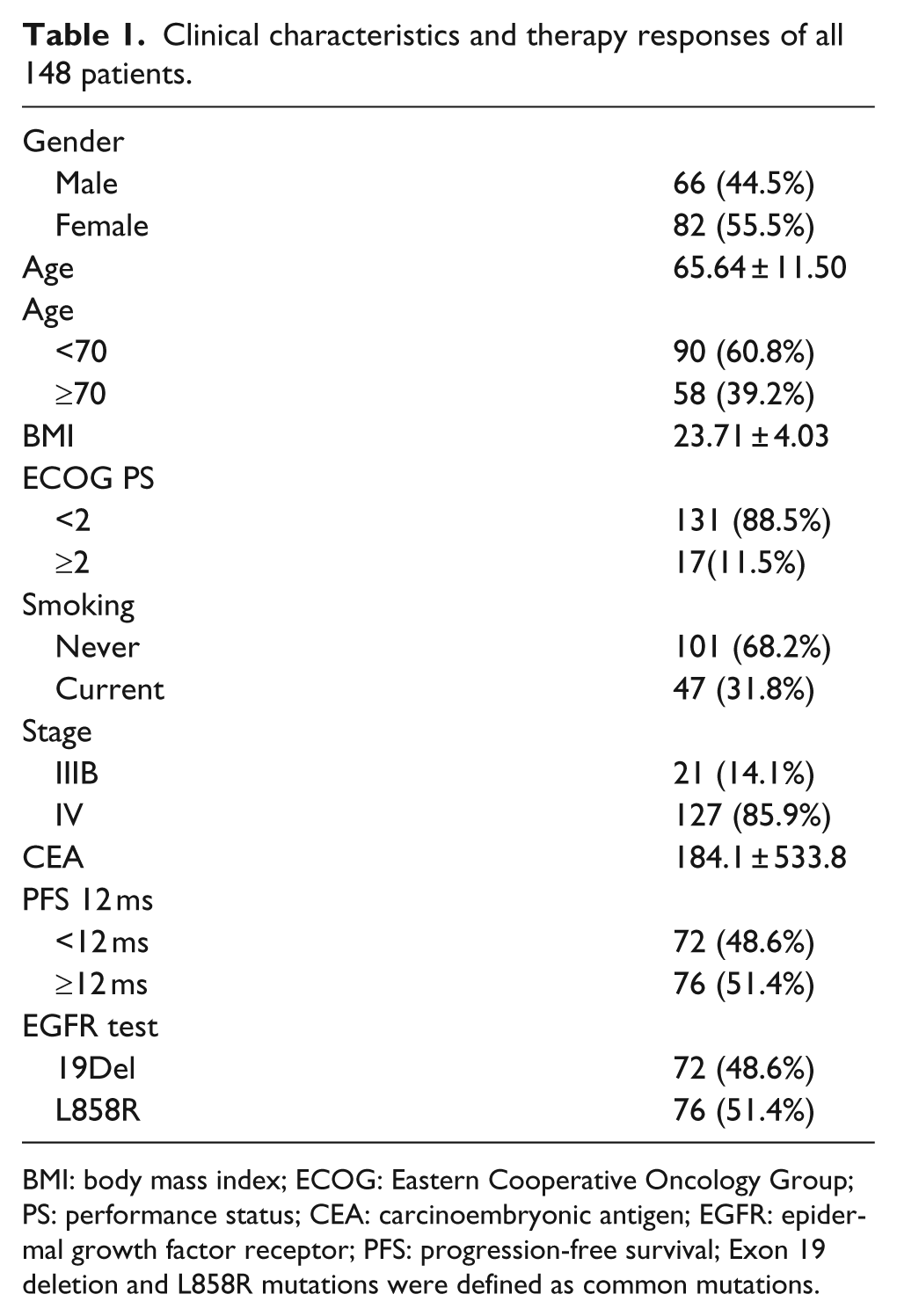
BMI: body mass index; ECOG: Eastern Cooperative Oncology Group; PS: performance status; CEA: carcinoembryonic antigen; EGFR: epidermal growth factor receptor; PFS: progression-free survival; Exon 19 deletion and L858R mutations were defined as common mutations.
Median PFS for patients with a deletion in exon 19 and patients with the L858R EGFR mutation was 13.2 and 12.3 months, respectively. For this reason, we used 12 months of PFS to further analyze the best cutoff points for the 2−ΔΔct value for exon 19 deletions and the L858R mutation; these were determined by ROC curves and Youden’s Index as 31.1 and 104.7, respectively. Patients were classified as having high or low 2−ΔΔct values based on the above cutoff values, linked to the intensity of EGFR gene expression. In all, 92 patients (62.1%) had high 2−ΔΔct expression and 56 (37.9%) had low 2−ΔΔct expression. Comparisons of groups with high and low 2−ΔΔct expression did not reveal an influence for baseline clinical parameters including sex, BMI, ECOG PS, smoking history, disease stage, and CEA values on 2−ΔΔct expression (Table 2).
Univariate and multivariate analyses of clinical factors of the different 2−ΔΔct expression.
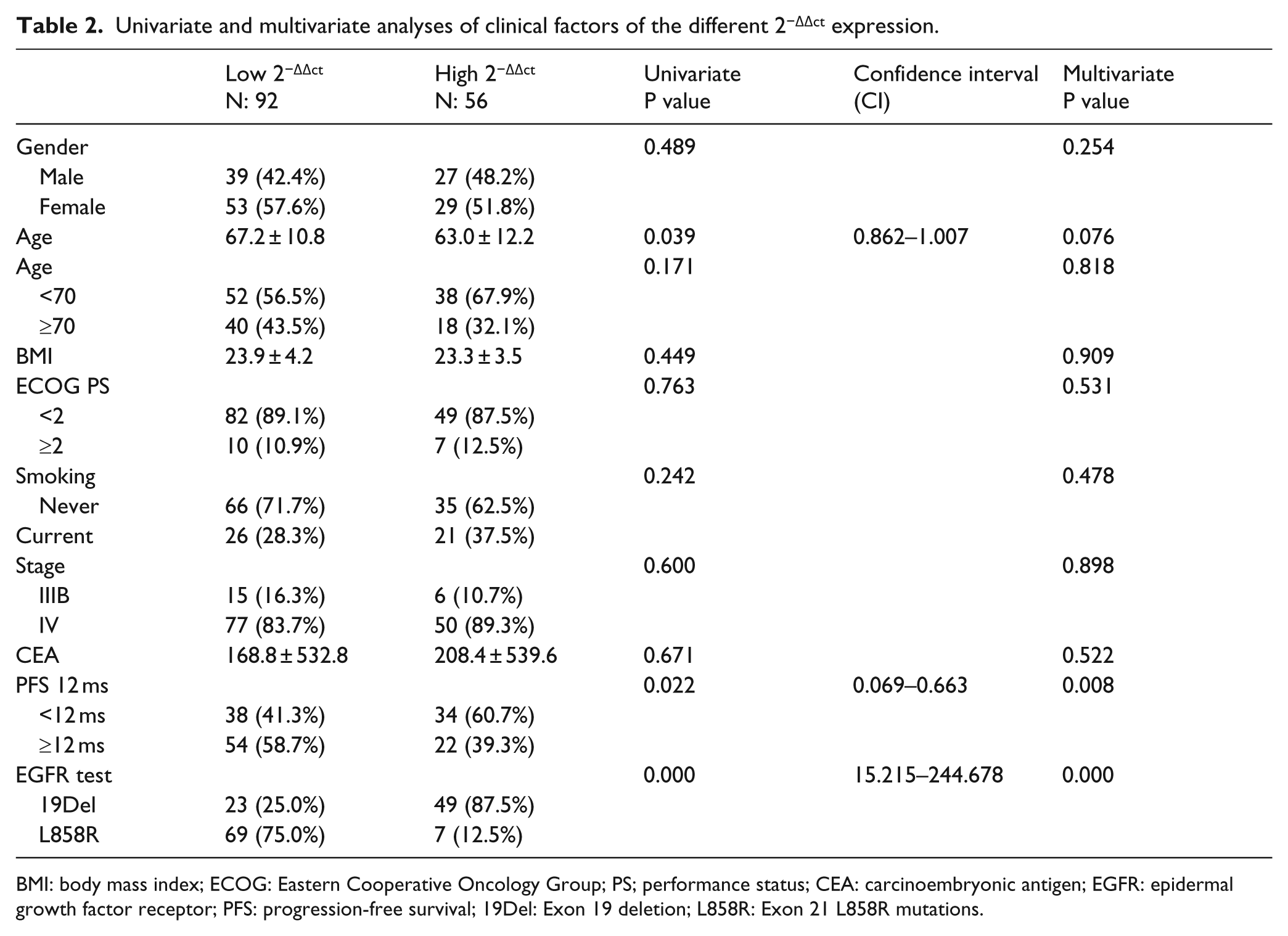
BMI: body mass index; ECOG: Eastern Cooperative Oncology Group; PS; performance status; CEA: carcinoembryonic antigen; EGFR: epidermal growth factor receptor; PFS: progression-free survival; 19Del: Exon 19 deletion; L858R: Exon 21 L858R mutations.
Relationship between PFS and clinical factors
Overall, PFS in patients with low 2−ΔΔct was longer than in the high expression group (14.93 and 10.89 months, respectively; P = 0.01; Figure 1). The effects of specific types of EGFR gene mutation on PFS were also analyzed. For analysis of PFS, patients were allocated to one of the four groups: (1) patients with deletions in exon 19 and low 2−ΔΔct expression, (2) patients with an L858R gene mutation and low 2−ΔΔct expression, (3) patients with deletions in exon 19 and high 2−ΔΔct expression, and (4) patients with an L858R gene mutation and high 2−ΔΔct gene expression. For patients with deletions in exon 19, those in the low expression group had better PFS than those in the high expression group (17.07 vs 12.04 months; P = 0.004; Figure 2(a)). Similar effects on PFS were found for patients with the L858R mutation, with low expression associated with greater PFS, compared with high expression (13.75 and 7.96 months, respectively; P = 0.008; Figure 2(b)). Patients with low gene expression of the exon 19 deletion had greater PFS than those with low expression of the L858R mutation (17.79 vs 13.75 months, respectively; P = 0.044; Figure 2(c)). Similarly, PFS was better in patients with high expression of the exon 19 deletion, compared with those with high expression of the L858R mutation (12.04 vs 7.96 months, respectively; P = 0.019; Figure 2(d)). PFS in the high expression exon 19 and L858R mutations groups and the low expression exon 19 and L858R mutations groups was 17.07, 13.75, 12.04, and 7.96 months, respectively (P = 0.001; Figure 3).
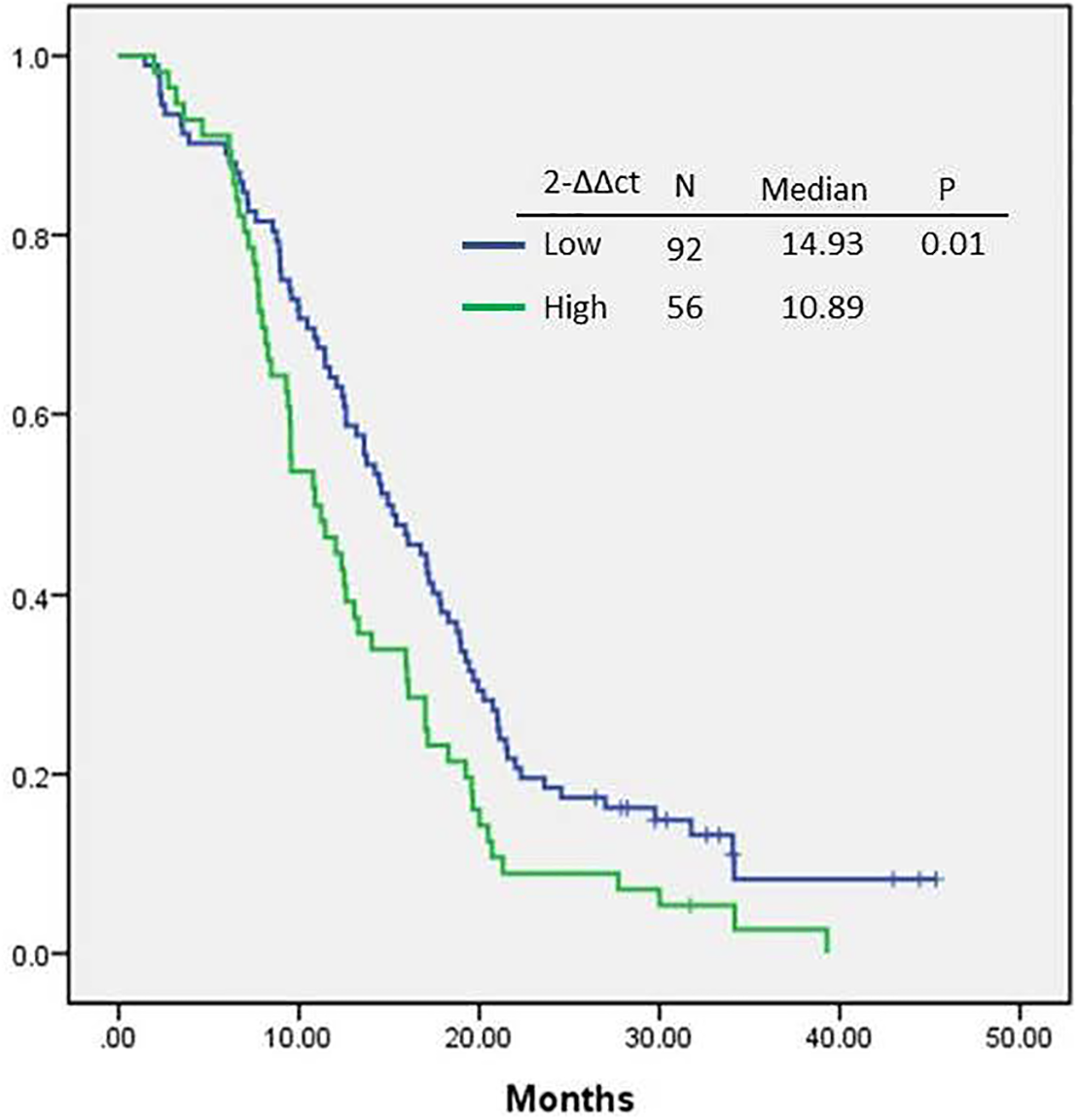
Progression-free survival according to low and high 2−ΔΔct gene expression among patients with EGFR-mutant adenocarcinoma of lung cancer who were treated using first-line tyrosine kinase inhibitors.

Progression-free survival of EGFR-mutant adenocarcinoma of lung cancer patients treated with first-line tyrosine kinase inhibitors: (a) impact of EGFR mutation gene expression on the outcome of 19 deletion patients; (b) impact of EGFR mutation gene expression on the outcome of L858R patients; (c) impact of EGFR mutation gene expression on the outcome of low gene expression patients; and (d) impact of EGFR mutation gene expression on the outcome of high gene expression patients.

Combination of different kinds of EGFR mutation gene expression impacted patient’s PFS.
Discussion
Preclinical studies have shown that the EGFR-associated signaling pathway has roles in cell functions important for cancer development and progression, including activation of tumor cell proliferation, angiogenesis, induction of apoptosis, and metastasis. 21 The anti-tumor activity of TKIs (via inhibition of EGFR phosphorylation) depends on the concentrations of the drug in tumor tissue.22,23 Consequently, its anti-tumor effects are determined by the efficiency of drug delivery to the tumor. In this study, we found that in patients with advanced adenocarcinoma of the lung, and who have EGFR mutations, the level of EGFR expression was correlated with the response to therapy. Moreover, a previous study found that high levels of EGFR gene expression in tumors influenced EGFR autophosphorylation and enhanced the activity of downstream signaling pathways (involving Akt, ERK 1/2, STAT, Shc, etc.)24–26 associated with cancer progression. Thus, greater drug concentrations in tumors with high EGFR expression may be needed to inhibit tumor development. Based on these findings, we hypothesized that variations in drug efficacy in tumors with the same EGFR mutations are due to differences in autophosphorylation and activation states of downstream signaling components. 27 This scenario is consistent with the results of this study, showing that the group with high EGFR gene expression had reduced PFS, compared with the group with low EGFR gene expression. Although previous studies found no association between EGFR gene expression, PFS, and overall survival, 28 but these studies were no further analyzed median level of EGFR expression and assess ROC value in both subtypes of patients. Tumor sensitivity was markedly influenced by the type of EGFR mutation, most likely as a result of the effects of different mutations on receptor structure and binding affinity for the inhibitor.29,30 Indeed, deletions in exon 19 map to one end of the C-helix (which is sensitive to conformational changes), while the L858R mutation lies in the N-terminal portion of the activation loop A of the kinase. 31 Differences in patterns of EGFR autophosphorylation were found between cell lines harboring deletions in exon 19 and those with an L858R point mutation in exon 21. 29 High intracellular adenosine triphosphate (ATP) concentration and other mechanisms may be an accurate reflection of a greater objective response rate to EGFR-TKIs.32,33 Several studies demonstrated a consistent PFS benefit across all clinical subgroups in patients with deletions in exon 19 and those with the L858R mutation in exon 21.34,35 Therefore, the essential diagnostic strategy was to identify EGFR-activating mutations that are closely related to the efficacy of EGFR-targeted drugs. To our knowledge, this is the first study to investigate if EGFR gene expression levels are a useful prognostic factor for advanced stage adenocarcinoma of the lung, in patients with different types of EGFR mutation receiving EGFR-TKIs. The 2−ΔΔct method was used to calculate EGFR gene expression in order to determine the feasibility of using EGFR as a marker gene for predicting clinical outcome. Further studies are needed to validate our findings. However, this study has several limitations. First, this study is retrospective and based on a small patient population; a prospective trial with a large sample size is needed to verify our findings. Second, we have not demonstrated the mechanisms that account for differences in PFS in patients with the same mutation. In addition to the above-mentioned mechanisms that may affect tumor response, heterogeneity and complexity within the tumor microenvironment may also result in distinct biochemical characteristics and acquired resistance may affect drug efficacy.36–38 Although it is possible that variation in outcomes is due to differences in EGFR gene expression levels, a previous cell line study did not find changes in therapeutic response with variations in phosphorylation levels of EGFR downstream signaling molecules. 39 Prospective studies are therefore needed to determine how EGFR alterations influence outcomes in patients with advanced NSCLC treated with EGFR-TKIs.
Conclusion
EGFR-mutant lung adenocarcinoma patients with lower EGFR gene expression had longer PFS duration without interfering OS.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
