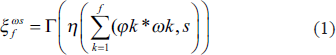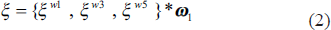Abstract
Background
Automatic lung segmentation is a crucial initial step in computer-aided lung computed tomography (CT) diagnosis. However, existing methods struggle to achieve accurate segmentation in the presence of dense abnormalities, limiting their clinical reliability.
Objectives
This research aims to improve lung segmentation accuracy by proposing a novel generative adversarial network (GAN)-founded method, named LungCTSeg-Net, designed to effectively segment lungs even in the presence of dense abnormalities.
Materials and Methods
The projected LungCTSeg-Net processes input lung CT pieces through a series of encoders, converting them into feature plots. A specialized multi-scale dense feature extraction (MSDFE) module abstracts multi-scale features from these encoded maps. The segmentation map is generated using decoders, with repeated down-sampling and upsampling to ensure invariance to the size of dense anomalies. The method is tested on the publicly available interstitial lung disease (ILD) dataset.
Results
Experimental results demonstrate that LungCTSeg-Net achieves robust performance regardless of the presence of dark anomalies in lung CT scans, outperforming existing approaches in segmentation accuracy.
Conclusion
The proposed LungCTSeg-Net approach improves lung segmentation accuracy in challenging cases with dense anomalies. The combination of multi-scale feature extraction and GAN architecture enhances the model’s capability to capture composite lung constructions, supporting more reliable computer-aided lung CT diagnosis.
Keywords
Introduction
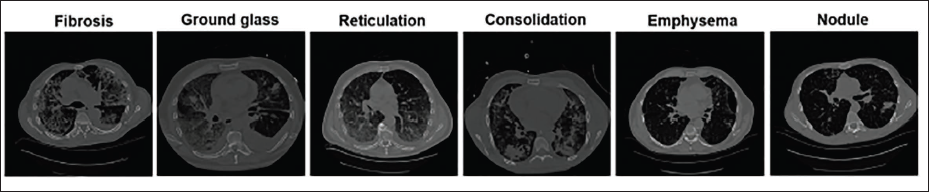
One of the main causes of death is lung cancer. 1 According to the American Cancer Society’s most recent data, 135,000 novel cases of lung cancer are malignant, out of an estimated 228,000 novel cases of lung cancer overall. 1 Research indicates that in a small number of cases that are detected early on, lung cancer is curable. 2 Mostly, a late diagnosis and treatment lead to death. 3 Thus, increasing the patient’s chances of survival requires early lung cancer detection. A computed tomography (CT) scan is the recommended technique for diagnosing lung diseases. Because current CT scanners create many CT scans, physical lung disease analysis and marking are tedious and labor-intensive, even for skilled radiologists. An automatic computer-aided diagnosis (CAD) of the lung CT scan is a useful remedy for this, with the intention of assisting radiologists. The process of accurately automatically diagnosing a lung CT image involves multiple steps. The robustness of the lung segmentation process controls the overall strength of the CAD scheme for the analysis of lung illness. Interstitial lung disease (ILD) states a collection of over 150 dark lung parenchymal anomalies. 4 The majority of these result in lung tissue scarring, which eventually affects breathing. On high-resolution lung CT (HRCT) imageries, different ILDs typically display different combinations of tissue patterns. 5 The various forms of dark irregular lung tissue patterns categorized such as ILD are shown in Figure 1. The presence of dense irregular lung tissue patterns leads to erroneous lung segmentation since their intensity is comparable to that of the chest region. 6 As a result, the majority of the algorithms in use do not segment the lung to account for these dense irregularities. According to the study, the majority of the automatic lung segmentation algorithms currently in use ignore dense anomalies and mislead medical professionals regarding lung diagnosis. 7 Patient death may result from erroneous lung segmentation and dense irregularities. The facts mentioned above inspired us to suggest a unique method for precise lung segmentation. This study proposes conditional generative adversarial networks (c-GANs)-founded technique for precise lung segmentation.
The Various Forms of Dense Irregular Lung Tissue Forms Categorized as Interstitial Lung Disease (ILD).
Related Work
It is easy to discern among the lung and non-lung areas on a normal lung CT scan due to the notable variation in attenuation. 8 As a result, simple gray-level thresholding-based algorithms were used in early techniques for lung segmentation from a non-lung area.9–14 Using a region-growing method based on gray value, gradient magnitude, and region homogeneity, Sun et al. segmented the lung region. 11 The inner voids were subsequently sealed off using a morphological closing technique. The standard thresholding-based and/or manually constructed filtering-based methods do not include ILD patterns like lung nodules, ground glass, fibrosis, and so on, in the lung segmentation. The aberrant lung tissue patterns are incorporated into methods published in the literature to minimize lung segmentation error. 7 An adaptive border marching procedure has been suggested by Pu et al. that hikes along the segmented lung border and corrects it in order to find the inner lung border. 6 The purpose of this procedure is to segment the lung by including the juxtapleural nodules.15–18 Larger abnormal lung tissue patterns, however, should not be taken into account. Sluimer et al. proposed segmenting the lung region comprising most of the clinical anomalies using an image registration technique. 19 Furthermore, Prasad et al. suggested leveraging the ribcage’s curvature in an adaptive thresholding method to integrate lung regions that demonstrate higher diminution. 20 Ye et al. proposed a shape-based technique for segmenting the irregular patterns of the lungs. 21 They proposed a contour correction technique based on chain codes, wherein significant key sites were identified along the lung border and subsequently connected to produce an accurate lung boundary. However, when more aberrant lung structure is included, this technique may result in excessive segmentation. Choi et al. proposed genetic programming-based feature transform and classification for the automatic detection of pulmonary nodules on computed tomography image. 22 Shen et al. have recommended a support vector machine (SVM) classifier-based strategy for nodule inclusion as well as an enhanced bi-directional chain coding method. 23 System performance is impacted by the classifier, adaptive threshold, and scan quality due to the cascading structure of the process. A multi-step method for juxtapleural node addition has been presented by Zhou et al. 17 A level-set founded method for concurrently segmenting and recording the 3D lung CT volume has been presented by Swierczynski et al. 24 Furthermore, level sets for lung nodule segmentation have been proposed by Farag et al. 25 They handled the statistical intensity representation of the background regions and nodules using a non-parametric density estimation technique. Furthermore, a mixed geometric active contour model for lung segmentation has been established by Zhang et al. 26 Active contour modeling was used by Filho et al. to segment lung pictures. 27 However, these methods require previous knowledge of the shape. The corpus of existing literature has proposed many intensity-based approaches for automatic lung segmentation.28–41 Robust feature extraction is a key factor in the efficacy of intensity-based lung segmentation techniques. Acquiring the required characteristics Hosseini-Asl et al. employed the conventional k-means clustering method after applying a non-negative matrix factorization (NMF) algorithm for lung segmentation. 28 Additionally, authors improved the NMF-founded method by gradually adding restrictions to it. 29 When there are severe abnormalities in lung tissue patterns, methods that employ juxtapleural nodules and pulmonary arteries to great effect become ineffective.28, 29 Researchers examined the efficacy of convolution neural networks for lung CT scan examination.30–40 The NMF algorithm fails because of its unsupervised nature, which makes it incapable of identifying the aberrant lung tissue pattern. 42
Along with natural image processing, several convolutional neural network (CNN)-based techniques for medical image analysis have been presented.43–53 Simonyan et al. have demonstrated the benefits of using small convolutional filters for image recognition applications. 47 In general, choosing the appropriate filter size is essential to building a deep network. There is no universal criterion that can identify the optimal combination of filter sizes. The inception module, a parallel convolution filter bank developed by Szegedy et al., has been proposed as a solution to this problem. 48 Huang et al. have proposed DenseNet, arguing that deep networks can be taught more successfully. 49 He et al. presented the ResNet for optimizing deep networks with respect to depth and other network parameters. 51 The deep deconvolutional residual network (DDRN)-based technique for lung nodule segmentation from CT images was first presented by Singadkar et al. 52 For lung pathology segmentation, the multi-path deep network proposed by Harrison et al. was demonstrated. 36 They took advantage of the deep network’s capacity to recognize various ILD tissue patterns. Moreover, it proposes the division of the recovered patch into lung and non-lung regions using a tiny CNN model. 38 The performance of this approach is restricted to the native region. A V-Net for 3D lung segmentation has been created by Negahdar et al. 39 Additional approaches for comprising irregular patterns of lung tissue, in particular lung nodules, consist of the central pooling technique, multi-view CNN, enhanced U-Net, and residual U-Net.30–33 While current lung segmentation algorithms have been becoming better over time, there is still room to propose a more generic lung segmentation method that can take into account different kinds of ILD patterns. As mentioned earlier, the size, shape, and textural characteristics of the different ILD patterns vary, which influences the segmentation technique. A c-GAN-based approach to the image-to-image style transfer problem has been put out by Isola et al. 54 They predicted the absent textural elements in an image by utilizing the generative models’ potent learning capabilities. Ronneberger et al and Howard et al proposed convolutional networks for biomedical image segmentation.55, 56 Pawar et al. proposed a generative model using a generative adversarial network. 57
This study proposes LungCTSeg-Net, a precise lung segmentation approach based on c-GAN. The salient aspects of the projected technique are outlined below,
The projected LungCTSeg-Net analyzes the input in the form of a lung CT piece and generates a lung segmentation map deprived of additional post-processing. The multi-scale dense feature extraction (MSDFE) module of the proposed LungCTSeg-Net abstracts the reliable multi-scale features. The suggested LungCTSeg-Net is efficient in segmenting the lungs, and it performs well even when there are dense abnormalities in the lung CT image.
Materials and Methods
It is commonly recognized that the attenuation of the lung parenchyma and nearby tissue differs significantly in a normal lung parenchyma. Therefore, the easiest method for obtaining the lung field mask is threshold-based lung segmentation. But as Figure 1 illustrates, there are aberrant lung tissue patterns associated with ILDs, such as ground glass, fibrosis, reticulation, consolidation, lung nodule, emphysema, and others. The CAD system performs worse as a result of under-segmentation frequently brought on by these aberrant lung tissue patterns. From the standpoint of image processing, lung segmentation deteriorates more when ILD features such as fibrosis, consolidation, and nodules are present because there is a reduced connection between the lung tissues. Furthermore, the size, shape, and texture of these dense anomalies can vary. 58 Therefore, lung segmentation needs a robust learning process that can both adjust to changes in the appearance of various ILD patterns and be invariant to the severity and size of the ILD.
Encoder/Decoder Block
The non-linear activation function, or rectified linear unit (ReLU), comes after the convolution/deconvolution layer in the encoder/decoder block. The network feature maps have been normalized through the application of instance normalization. 59 3 × 3 convolution/deconvolution filters make up every encoder/decoder block, which has one per convolution/deconvolution layer. Here, the input lung CT slice is encoded into feature maps using an encoder block, and the encoded feature maps are then down-sampled. The encoder block’s downsampling action aids in expanding the receptive field. By upsampling the feature maps, the decoder block’s upsampling factor of two maintains the symmetry in the projected generator network.
MSDFE Module
As was previously mentioned, dark anomalies on lung CT scans can vary in shape, size, and feel. Variations in the look of dense anomalies affect the enactment of the segmentation procedure. As a result, the majority of current processes do not take into consideration these dark anomalies, which are mainly abundant at the lung border (samples are given in Figure 1). The juxtapleural and pulmonary arteries are included in lung segmentation by researchers using border marching algorithm and various size morphological filters in the literature currently under publication. However, while modifying the hyper-parameters, the nodule’s dimensions and form must be considered. These drawbacks drive our desire to extract multi-scale features, which will allow dense abnormalities to be included in lung segmentation while maintaining appearance invariance.
For reliable feature extraction, an MSDFE module is suggested. As seen in Figure 2, the projected MSDFE unit is made up of four inception blocks joined successively by dense networks in between. The purpose of the thick connections between inception blocks is to increase the variety in the input of successive inception blocks by reusing the learned features of the first inception blocks. Compact connections between initial blocks facilitate the extraction of multi-scale information and the optimal path toward precise lung segmentation for the projected network.
Input feature maps are processed by the inception block using three convolution layers with different filter sizes, namely, 1 × 1, 3 × 3, and 5 × 5. Subsequently, ReLU and instance normalization are applied. As illustrated in Figure 2, these convolution layers are coupled in a parallel paradigm. The inception block can be expressed mathematically as follows:
where, 𝝋 is the input to the inception block having f feature maps, 𝞈s denotes the convolution filter of s × s spatial size and f channels, 𝜂 is the instance normalization, Г represents the ReLU. Further, these filtered feature maps are concatenated and processed through a simple 1 × 1 convolution filter given in Equation (2).
where {·} represents the feature map concatenation process and 𝞷 is the feature response of the inception block.
Six MSDFE modules are coupled to form the proposed LungCTSeg-Net, as seen in Figure 2. The multi-scale features produced from the first three MSDFE modules are processed using a simple filtration layer with a 3 × 3 filter size and a stride factor of two. For the next three MSDFE modules, the acquired multi-scale feature maps are handled using a basic deconvolution layer with upsampling factor of two, in order to preserve network symmetry. Making the suggested LungCTSeg-Net invariant to the appearance of the thick anomalies on lung CT slice is the driving force behind this.
The Network Architecture of the Projected Generator.
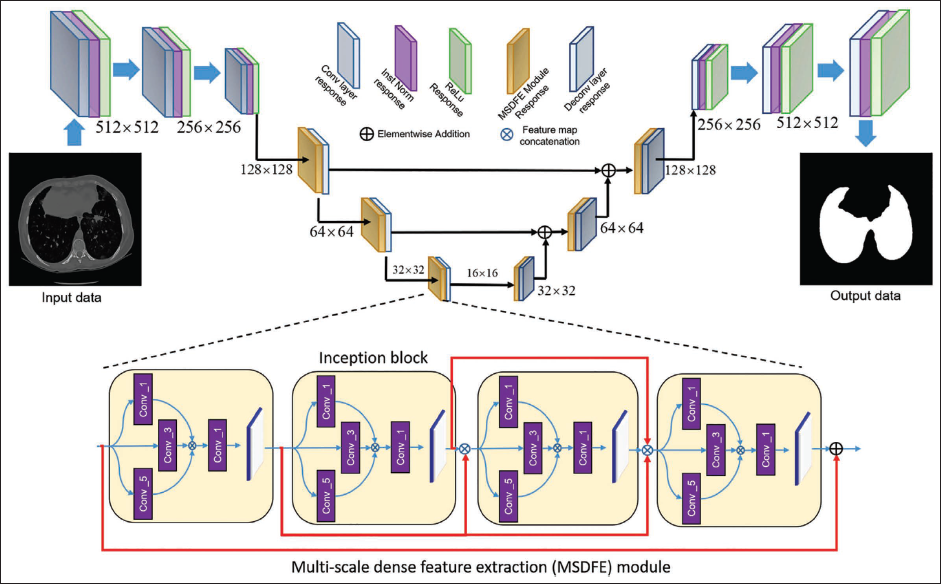
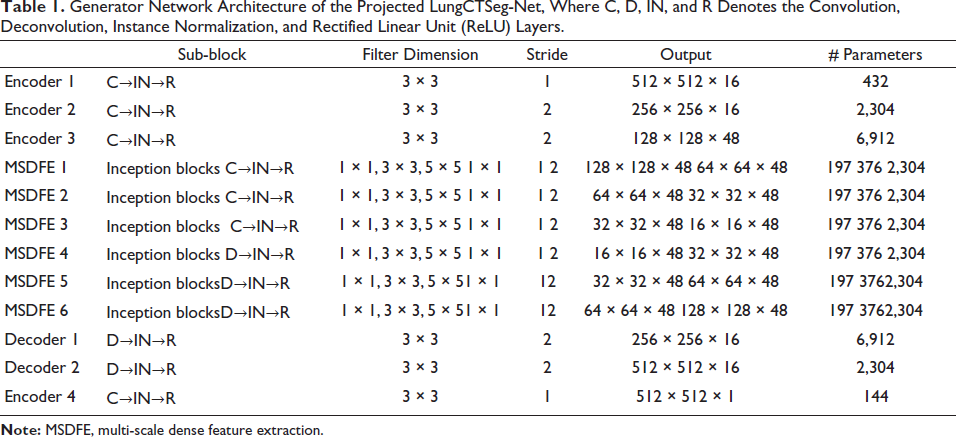
The proposed LungCTSeg-Net makes a matching facility for iterative down- and upsampling. Through skip connections, the feature maps acquired in the first MSDFE modules are also pooled throughout the network while creating the symmetric network design. The suggested LungSeg-Net is able to produce the dominant edge info in the lung segmentation map output. Table 1 provides the projected LungCTSeg-Net’s parameter specifications.
Generator Network Architecture of the Projected LungCTSeg-Net, Where C, D, IN, and R Denotes the Convolution, Deconvolution, Instance Normalization, and Rectified Linear Unit (ReLU) Layers.
Network Loss Function
The generator and discriminator networks make up the two networks in the suggested method for precise lung segmentation. The generator network’s job in this case is to figure out the mapping G: {x, z} ↓ y. On the other hand, x and y stand for the lung segmentation map and input lung CT picture, respectively. Here, we update the network weight values using adversarial loss. 31 Equation (3) provides the conditional GAN’s objective.
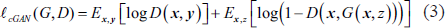
where discriminator D seeks to maximize adversarial loss, while generator G seeks to decrease it. Along with the adversarial loss, we considered old ℓL1 loss as given in Equation (4) to apprise the network weight factors.
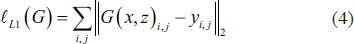
where, G (
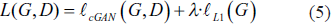
Therefore, the cGAN’s goal for precise lung segmentation is,
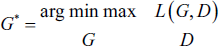
Training Details of the Proposed Network
The benchmark ILD dataset is taken into consideration in this work in order to produce the labeled data needed to accurately train the suggested LungCTSeg-Net for lung segmentation. Depeursinge et al. developed the ILD database to offer a standard framework for evaluating automated ILD analysis methods. One hundred and eight high-resolution CT images of the lungs (HRCT) with annotated lung field maps are included in the database. Twenty lung CT slices are typically included in each lung CT scan. These scans are recognized as one of the ILD based on lung tissue forms such as consolidation, fibrosis, ground glass, reticulation, lung nodules, emphysema, and so on.
Fifty of the 108 HRCT scans from the ILD dataset are used to create the training dataset. Still, the remaining lung CT scans are used to confirm the projected lung segmentation model, LungCTSeg-Net. In order to enhance the size of the training dataset, the flip left, flip right, flip up-down, and picture transpose operators are taken into consideration when applying the data augmentation approach. A total of 4,000 lung CT slices with any kind of ILD are built together with the matching lung field segmentation maps in order to train the projected LungCTSeg-Net for precise lung segmentation. Training criteria include adversarial loss (ℓ𝑐𝐺𝐴𝑁) and L1 (ℓL1) loss parameters.
To train the suggested network for precise lung field segmentation, a personal computer (PC) equipped with an NVIDIA GTX 1080 8 GB GPU and an Intel Core i7 processor running at 4.20 GHz is utilized.
Result and Discussion
To assess the accuracy of lung segmentation, a thorough experimental examination of the suggested LungCTSeg-Net is conducted in this section. There are two sections to this experimental investigation:
Quantitative investigation and Qualitative investigation.
Quantitative Investigation
The quantitative investigation is performed by equating the two assessment metrics of the projected and current advanced networks, the Jaccard index (J) and dice similarity coefficient (DSC). These are expressed mathematically as follows:
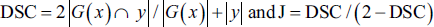
where,
G (
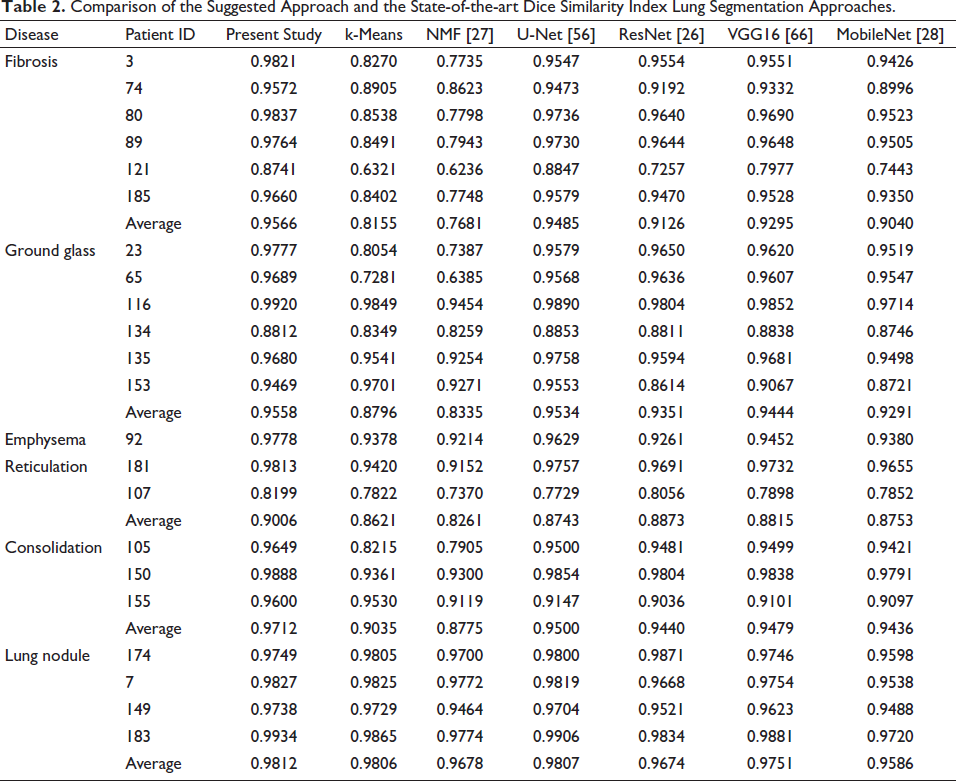
Twenty-two lung CT scans from the ILD databank have had their lung segmentation quantitatively analyzed. 5 One thing to note is that the lung CT slices that were part of the training set and the testing set did not overlap. The suggested LungCTSeg-Net’s performance is compared to that of the current cutting-edge deep networks, including VGG16, ResNet, U-Net, and MobileNet. To provide a fair comparison, the proposed LungCTSeg-Net is trained on the same training set-up and training dataset as the existing U-Net, ResNet, VGG16, and MobileNet, through adversarial training. 51 Additionally, the suggested LungCTSeg-Net for lung segmentation is compared with the currently available unsupervised segmentation methods, such as k-means clustering and non-matrix factorization-based method.25, 26 Table 2 provides a quantitative assessment of the suggested and current DSC methods. It is evident that the type of ILD affects how well the current lung segmentation techniques operate. More precisely, when compared to current lung segmentation techniques, the suggested method performs noticeably better in the presence of ground glass, fibrosis, consolidation, and reticulation. Conversely, Tables 2 and 3 show that the accuracy of the current lung segmentation techniques is unaffected by the existence of a lung nodule. This is because it is challenging to accurately reflect, in a quantitative analysis, the exclusion of the lung nodule using a specific technique.
Comparison of the Suggested Approach and the State-of-the-art Dice Similarity Index Lung Segmentation Approaches.
Comparison of the Suggested Approach to the State-of-the-art Lung Segmentation Techniques Using the Jaccard Index (J).
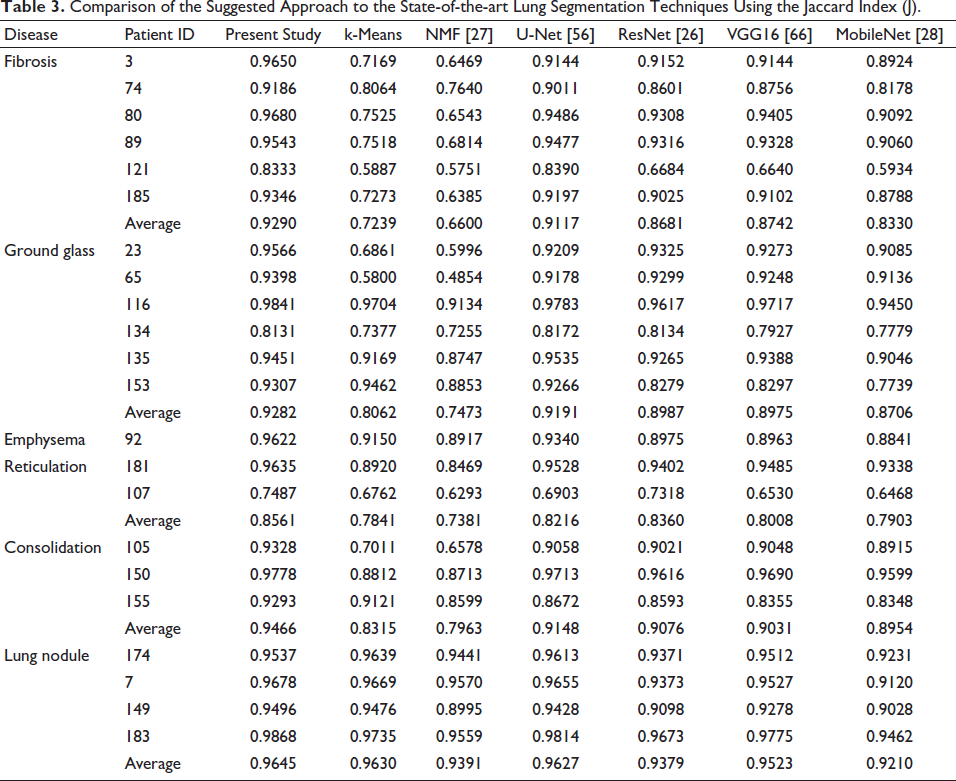
When equated to the current techniques for lung segmentation, Table 2 shows a significant increase in the DSC of the projected LungCTSeg-Net when the input lung CT scan comprises dense abnormalities. This demonstrates how the dense anomalies can be included in the lung segmentation process using the suggested LungCTSeg-Net. Additionally, Table 3’s higher Jaccard index for the suggested LungCTSeg-Net in comparison to the techniques currently in use for lung segmentation shows the least amount of over segmentation. Table 3 shows that the suggested network performs better for precise lung field segmentation than the other approaches that are currently in use.
Qualitative Investigation
Lung CT scans from the ILD databank are used for the proposed LungCTSeg-Net’s qualitative investigation. Dense anomalies of various sizes and textures, positioned at different points inside the lungs, are intentionally chosen for visual inspection in lung CT images. Figures 3 through 4 depict the lung segmentation map produced by the suggested LungCTSeg-Net and current cutting-edge techniques when the input lung CT slice has various ILD tissue patterns (dense abnormalities). It is evident that these extensive anomalies in lung segmentation are entirely absent from the current unsupervised approaches. The rationale behind this is that in order to distinguish between the lung- and non-lung regions, they only take into account the pixel intensity. Likewise, current supervised methods such as U-Net and ResNet only slightly incorporate dense abnormalities in lung segmentation; nonetheless, they remain significantly disconnected from the corresponding ground truth lung segmentation map. However, regardless of the kind, size, texture, or connectivity of the anomalies, the suggested LungSeg-Net efficiently incorporates them into the segmentation of the lungs. The newly proposed transform-based technique and the suggested LungCTSeg-Net’s performance are also contrasted. Conversely, the aberrant pattern in lung segmentation is accurately included in the proposed LungCTSeg-Net.
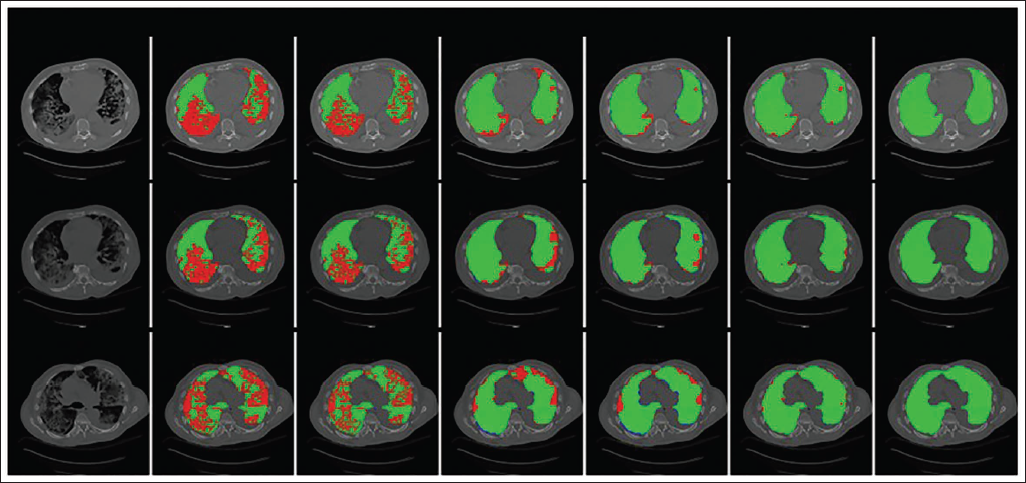
Graphic Assessment Between the Projected and Current Methods for Segmentation of Lung Computed Tomography (CT) Slice that Show the Fibrosis Form. (a) Input Lung CT Slice (b) k-means Clustering (c) Non-negative Matrix Factorization (NMF) (d) U-Net (e) ResNet (f) Projected LungCTSeg-Net (g) Ground Truth.
Lung Segmentation Outcomes of the Projected and Current Methods When Input Lung Computed Tomography (CT) Slice Comprise Lung Nodules. (a) Input Lung CT Slice (b) k-means Clustering (c) Non-negative Matrix Factorization (NMF) (d) U-Net (e) ResNet (f) Projected LungCTSeg-Net (g) Ground Truth.
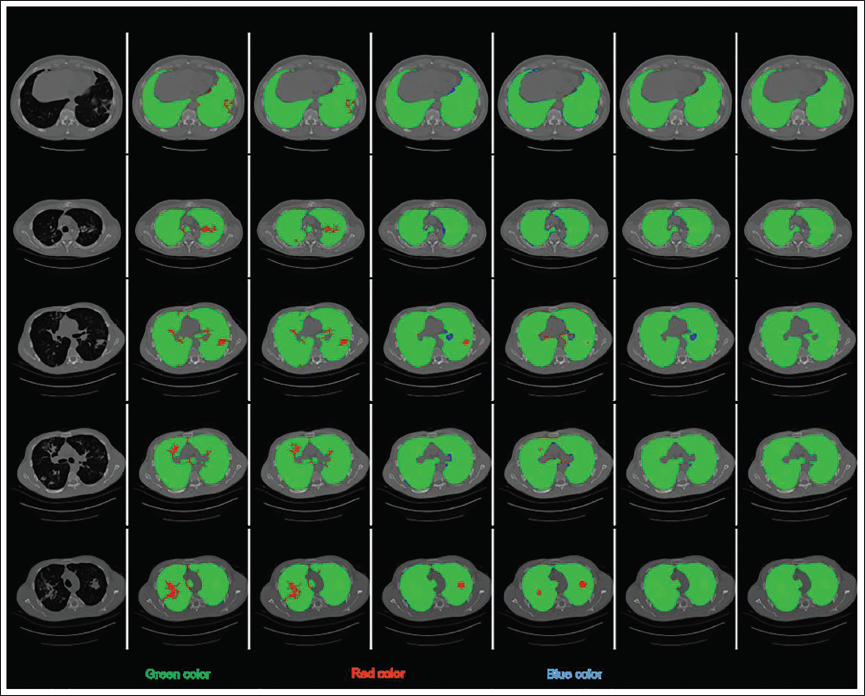
According to a qualitative investigation, the suggested LungCTSeg-Net performs better for lung segmentation than the current methods.
Conclusion
In this work, we suggest a lung segmentation technique based on c-GANs. The suggested generator network processes the input lung CT slice via the three series of encoders to encode it into a set of feature maps. Furthermore, six MSDFE modules are used to extract the multi-scale features that accurately identify the shape, size, and texture of lung nodules as well as other patterns of lung tissue. Lastly, the lung segmentation map is obtained by using a trail of two decoders. The experimental study was performed on the ILD database, which is accessible to the public, in order to segment aberrant patterns of lung tissue, such as fibrosis, ground glass, reticulation, emphysema, consolidation, and micro-nodules. In order to compare the proposed network’s quantitative and qualitative performance to the currently used lung segmentation techniques, an experimental analysis is being conducted. DSC and Jaccard index are used to measure the quantitative performance. While the segmentation concert of all the cases is improved by the suggested method, the segmentation of the cases with fibrosis, ground glass, reticulation, and consolidation abnormalities is reported to have improved significantly. The benefits of the suggested algorithm, which takes into account both minimal over segmentation and dense anomalies in lung segmentation, are amply demonstrated by the performance analysis. Because of their bigger size, previous approaches for lung segmentation fail to include the lung nodule; however, the suggested method successfully includes the lung nodule in the lung segmentation map.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent
Not applicable.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.