Abstract
Background
Pyrogallol, a bioactive compound derived from natural sources, is the focus of this study, aiming to investigate its anti-cancer properties against A549 lung cancer cells and uncover the underlying mechanisms.
Purpose
This study seeks to elucidate the therapeutic potential of pyrogallol as a novel anti-cancer agent for NSCLC by targeting aerobic glycolysis and signal transducer and activator of transcription 2 (STAT2) signaling pathway.
Background
Pyrogallol, a bioactive compound derived from natural sources, is the focus of this study, aiming to investigate its anti-cancer properties against A549 lung cancer cells and uncover the underlying mechanisms.
Materials and Methods
A range of pyrogallol doses was administered to A549 cells, followed by a comprehensive analysis utilizing bioinformatic tools like Gene Expression Omnibus (GEO), GEO2R, WebGestalt, and the Search Tool for Interactions of Chemicals database. Assessments of cytotoxicity were conducted using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) and trypan blue assays post-treatment of A549 cells with pyrogallol. Cell cycle progression and cell death were evaluated via flow cytometry using Annexin V-fluorescein isothiocyanates/propidium iodide double staining. Network analysis helped identify pertinent signaling pathways, while RT-PCR validated mRNA expression changes in A549 cells. Auto dock docking studies were employed to gauge the binding affinity of pyrogallol with targets associated with the STAT2-mediated pathway.
Results
Pyrogallol notably impeded the progression of A549 cells by prompting cell cycle arrest in the G0/G1 phase and fostering apoptosis. Network analysis highlighted its role in regulating metabolism, apoptosis, and STAT2 targets linked to lung tumorigenesis. RT-PCR validation affirmed the downregulation of genes associated with cell cycle and apoptosis targets. Furthermore, docking studies indicated a robust binding affinity between pyrogallol and the signaling targets within the STAT2 pathway. Pyrogallol displays the potential to halt the growth of lung cancer cells, indicating its promise as a viable treatment for this condition.
Conclusion
These discoveries emphasize the need for additional research and clinical studies to fully harness its therapeutic benefits for lung cancer treatment.
Keywords
Introduction
Lung cancer stands as the most frequently diagnosed cancer worldwide, with an alarming estimate of 2.3 million cases and a concerning rise in mortality rates, as reported by Sung et al. (2021). The disease is underscored by statistics that surpass those of other prevalent cancers such as colorectal, pancreatic, and breast cancer. Despite ongoing efforts, the survival outcome for lung cancer remains dismally poor, reflecting the complex interplay of various mechanistic factors contributing to cancer progression (Nooreldeen & Bach, 2021; Ramos et al., 2021). A multitude of challenges drugs the battle against lung cancer, including the presence of cancer stem cells, intricate tumor microenvironmental, resistance to the chemotherapeutics, metastasis tendencies, and invasion characteristics. These factors collectively create a formidable barrier to successful treatment intervention. In-depth investigations have unveiled numerous signaling pathways intricately linked with the relentless progression of lung cancer, emphasizing the need for targeted therapeutic strategies to disrupt these pathways and impede tumor development (Quail & Joyce, 2013). Crosslinking with bioinformatic analysis constitutes an advanced strategy in disease research, providing valuable insights into the complex molecular interactions that drive pathological processes. Fundamentally, crosslinking entails chemically stabilizing biomolecular interactions within cellular contexts, thereby conserving transient or fragile interactions for further scrutiny (An et al., 2019).
The protein signal transducer and activator of transcription 2 (STAT2) play a crucial role in governing gene activity and participating in various cellular functions. Operating actively in tissues across the body, STAT2 is instrumental in regulating processes such as cell growth and division, cell mobility, and the programmed self-destruction of cells (Loh et al., 2019). Moreover, STAT2 has the potential to involve chemoresistance in cancer cells via influencing cellular metabolisms, particularly glycolysis (Tošić, & Frank 2021; Wang et al., 2022). The inappropriate activation of STAT2 in cancer cells leads to persistent transcription of cell growth factors and anti-apoptotic molecules, pivotal for sustaining cell growth and survival. Additionally, STAT2 amplifies the malignancy of tumors by fostering invasion, migration, metastasis, and angiogenesis (Tolomeo & Cascio, 2021). The study of Zhou et al. (2018) revealed that downstream signaling pathways of STAT3 are associated with Apoptosis and cell cycle arrest. Another study conducted by Wei et al. (2013) demonstrates that STAT3 interacts with the Skp2/p27/p21 pathway to regulate cancer motility and invasion. Research evidence suggests that targeting STAT2 is a potential therapeutic option in cancer treatment.
Plant-derived drugs exhibit the promising potential to treat various diseases without inducing toxicity in normal cells. The attraction of scientists to this field stems from the unique properties of phytochemicals which encompass the ability to alter cell cycle control, evade apoptosis, and influence processes such as angiogenesis and metastases. Over the last two decades, there has been a substantial rise in the utilization of complementary alternative medicines in several countries. Notably, around 60% of anticancer agents trace their origins to medical plants and other natural resources (Aruchamy & Duraisamy, 2021). Pyrogallol is resourced by heating gallic acid with water. Gallic acid from galls and barks of several trees. Pyrogallol is a product of the decarboxylation of gallic acid. It’s a phenol family (Britannica, 2018). Pyrogallol has the ability to be an antioxidant (Sutanto et al., 2019), anti-inflammatory (Nicolis et al., 2008), and anti-proliferative activity (Khan et al., 2002). In this study, we aim to demonstrate that pyrogallol inhibits the STAT3, and their downstream signaling pathways to inhibit lung cancer progression.
Materials and Methods
Data Mining
A precise approach to locate relevant lung cancer datasets within the Gene Expression Omnibus (GEO) database. Using the search criteria (“OSCC”[MeSH Terms] OR lung cancer [All Fields]) AND “Homo sapiens”[porgn], we successfully pinpointed the GSE2514 and GSE7670 datasets for detailed examination. Afterward, we acquired and downloaded this dataset from the GEO database. GSE2514 comprises 20 normal tissue samples and 19 tumor tissue samples and GSE7670 contains 27 pairs of adjacent normal and tumor tissue samples. Notably, it was generated using the GPL570 Affymetrix platform, specifically the Human Genome U133 Plus 2.0 Array. We conducted analyses on the dataset utilizing Gene Set Enrichment Analysis (GSEA), Comparative Toxicogenomics Database (CTD), and WebGestalt. For visualizing biological functions, pathway predictions, and Venn diagrams, we utilized the Funrich tool.
Functional Annotation by Search Tool for Interactions of Chemicals (STITCH)
Advancements in understanding the complex interactions between pharmaceutical compounds and proteins rely heavily on computational methods to predict precise protein targets of drug molecules. The STITCH database, curated meticulously by the European Molecular Biology Laboratory, plays a crucial role in facilitating these explorations. It serves as a comprehensive repository housing verified and potential interactions between drugs and proteins.
Cell Lines
A549 human lung cancer cells were sourced and underwent careful cultivation and maintenance in a controlled setting, specifically within a CO2 incubator maintained at a stable temperature of 37°C. To facilitate cell proliferation, a growth medium comprising DMEM, supplemented with 10% FBS, and 1% penicillin-streptomycin antibiotics was utilized.
Cell Viability Assay
To assess pyrogallol’s impact on the cytotoxicity of the A549 cell line, we utilized the MTT assay (Jayaraman et al., 2023b). The procedure began with seeding an initial population of 1 × 104 cells, allowing them to incubate overnight. Following this incubation, we introduced the MTT reagent into the cell culture and solubilized it using dimethyl sulfoxide, measuring absorbance at 590 nm. Additionally, images of A549 morphology at 10× magnification were observed and captured using an Olympus inverted microscope.
Glucose Uptake Assay
A total of 1 × 104 A549 cells were individually placed in 96-well plates and allowed to grow for 24 hours. Following this, they were treated with pyrogallol for 48 hours. After the treatment, the cells underwent two washes with Krebs-Ringer-phosphate-HEPES buffer. Glucose uptake was quantified using a specialized colorimetric kit as per the manufacturer’s instructions. Cell lysates were extracted, incubated with 2DG enzymatic mix, and then measured at an optical density of 450 nm after an hour.
Flow Cytometry Analysis
Flow cytometric analysis was employed to conduct cell cycle analysis and evaluate cell death (Jayaraman et al., 2023a). At the outset, A549 cells were seeded in six-well plates at a density of 1 × 106 cells per well. Following the treatment period, the cells underwent fixation, and flow cytometric analysis was conducted utilizing a BD FACS machine. The data obtained from these analyses were subsequently processed using Cell Quest Pro V 3.2.1 software, BD, USA.
mRNA Expression Analysis
Spectrometric analysis was utilized to quantify the RNA content in the samples (Prasad et al., 2023). The study employed specific primers (5′–3′) listed such as STAT2: F-GAGCCTTTGTGGTAGAAA, R-CCTGTCAATGGAGACTTC (Guo et al., 2012), cyclin-dependent kinase inhibitor 1A (CDKN1A): F-GGAAGGGACACACAAGAAGAAG, R-AGCCTCTACTGCCACCATCTTA (Wang et al., 2022), B-cell lymphoma 2 (Bcl-2)-associated X-protein (BAX): F-TTCTGACGGCAACTTCAACTG, R-TGAGGAGTCTCACCCAACCA (Chen et al., 2015), BCL2: F-GACGCTTTGCCACGGTGGTG, R-GGGGCAGGCATGTTGACTTCAC (Chen et al., 2015). The analysis included a melting step, and an invariant control served as a reference. The findings from this assessment were presented as fold changes in comparison to the control.
Molecular Docking
Ligand and Protein Docking Analysis
The study investigates the binding interactions of pyrogallol (CID: 1057) with various proteins associated with cancer cell regulation. These proteins included STAT3 (PDB ID: 6UX2), p21 (PDB ID: 1AXC), Bcl-2 (PDB ID: 6O0K), and BAX (PDB ID: 4S0O), and their crystal structures were obtained from the Protein Data Bank (PDB) (
Statistical Analysis
The statistical findings of the study were presented as mean values along with their corresponding standard deviations. GraphPad Prism 8 software was employed for this data analysis. Regarding t-tests, significance levels were defined at p values of 0.05, 0.01, 0.001.
Results
Pyrogallol Efficiency in Lung Cancer by Bioinformatic Analysis
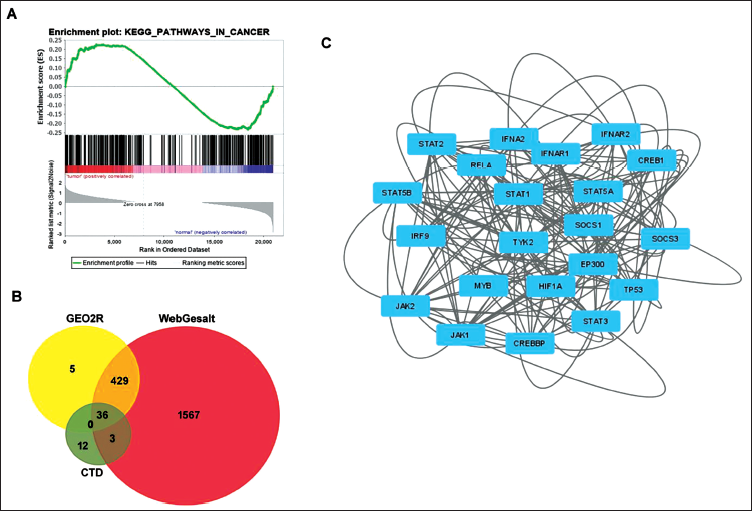
In this study aiming to identify crucial genes influencing lung cancer regulation, an array of bioinformatic tools played a pivotal role. Initially, the GSE2514 and GSE7670 datasets, containing patient samples, underwent GSEA, highlighting a significant portion of genes associated with cancer-related pathways, emphasizing their relevance in lung carcinogenesis (Figure 1A). To pinpoint differentially expressed genes, drug-based databases were employed. Additionally, co-regulating genes interacting with pyrogallol were extracted from the GEO2R, CTD Database, and WebGestalt databases. The Venn diagram (Figure 1B) distinctly illustrated 36 genes of substantial importance, displaying strong interactions with pyrogallol in lung cancer cells. Moreover, an evaluation of the nodal strength and interactive networks of these 36 genes was carried out through the STITCH server, depicted in Figure 1C.
Identification of Oncogenic Genes Associated with Pyrogallol Using Bioinformatic Tools. (A) GSE2514 and GSE7670 Dataset Has Been Retrieved from GEO Omnibus and Indicated Enrichment Plot from KEGG Shows Pathways in Cancer Using GSEA Software, (B) Venn Diagram from Funrich Tool Shows the 36 Overlapping from the Various Databases Analysis of GEO2R, WebGestalt, and CTD Database Analysis, (C) STITCH Server and Cytoscape Software Helps for Drug-protein Interacting for Pyrogallol and Overlapping Targets.
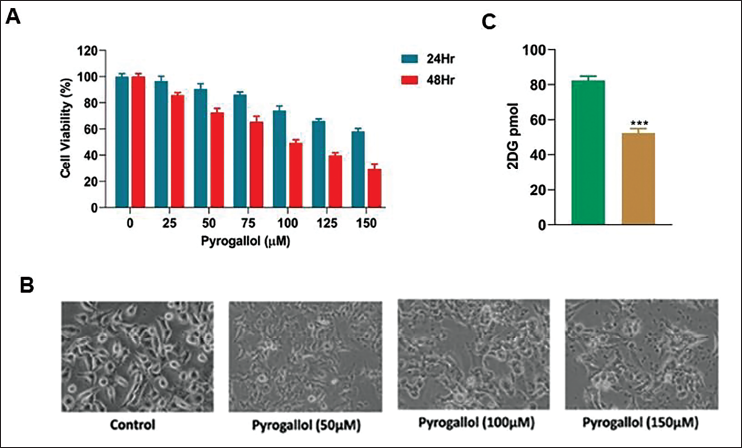
Pyrogallol Inhibits Cell Growth in A549 Cells
The evaluation of pyrogallol’s impact on A549 cells within the context of lung cancer involved conducting MTT and trypan blue exclusion assays. The results, depicted in Figure 2A, unmistakably demonstrated a significant, dose-dependent decrease in cell viability in response to pyrogallol exposure, exhibiting a time-dependent pattern. The half-maximal inhibitory concentration (IC50) for pyrogallol was determined as 95.23 µM after 48 hours of exposure. Furthermore, the impact of different concentrations of pyrogallol (50, 100, 150 µM) for 48 hours in A549 cells was examined, revealing noticeable cellular damage and cell blebbing in A549 cells (10×), shown in Figure 2B. We next investigated how pyrogallol impacts aerobic glycolysis and observed significant outcomes. In A549 cells, pyrogallol notably reduced glucose uptake activity for 48 hours of treatment, as illustrated in Figure 2C. These impactful results collectively indicate that pyrogallol strongly inhibits both cell proliferation and aerobic glycolysis in A549 cells. These compelling findings underscore pyrogallol’s potent cytotoxic properties against A549 cells, highlighting its potential therapeutic role in managing lung cancer.
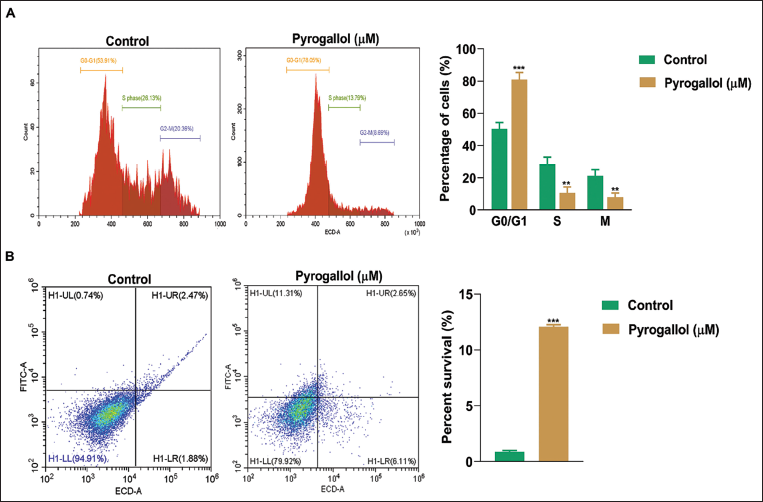
Pyrogallol Induces G1 Phase Arrest and Promotes Apoptosis in A549 Cells
For a deeper understanding of pyrogallol’s biological effects, we analyzed cell cycle regulation and cell death in A549 cells using flow cytometry. Consistent with the bioinformatic data, our flow cytometry analysis showed that after a 48-hour pyrogallol treatment, the cell population was arrested in the G0/G1 phase, notably increasing to 78.05% compared to the untreated group’s 53.91% (Figure 3A). Additionally, we conducted an Annexin V-FITC dual staining assay to measure apoptosis, and induction by pyrogallol in A549 cells. The assay demonstrated a significant increase in cell death after 48 hours of pyrogallol treatment, with an apoptotic rate of 12.26%, as displayed in Figure 3B. These findings strongly support pyrogallol’s potent anticancer activity by stimulating apoptosis in A549 cancer cells.
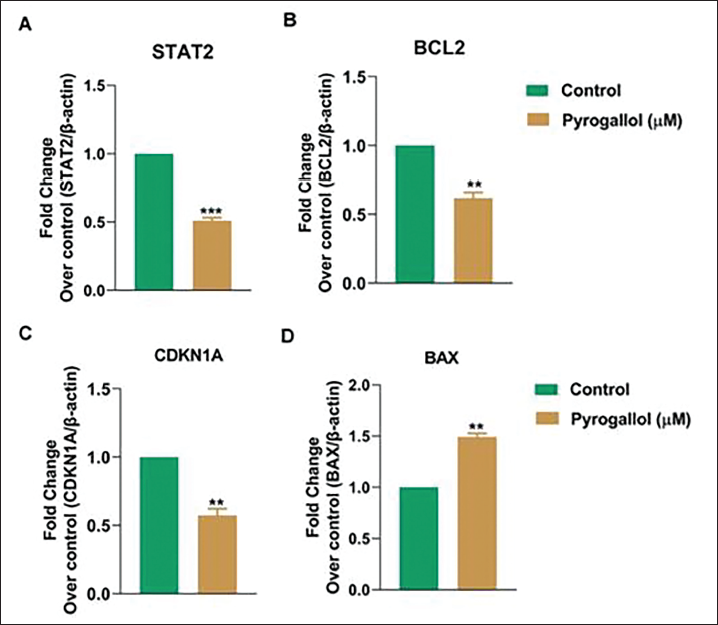
Pyrogallol Inhibits Cell Proliferation-mediated STAT2
Moreover, the study employed Real-time PCR to evaluate the mRNA expression levels of several genes crucial to apoptosis in Lung cancer A549 cells following exposure to pyrogallol at IC50 doses for 48 hours. The outcomes, illustrated in Figure 4A–D, unveiled several noteworthy findings. Primarily, the mRNA expression of key genes influenced apoptosis and cell cycle (STAT2, CDKN1A, BAX, and BCL2) demonstrated a gradual decrease in response to pyrogallol treatment within the 48-hour timeframe. Notably, these genes, including STAT2, CDKN1A, BAX, and BCL2 are linked to the inhibition of protein synthesis, cell proliferation, cell cycle progression, and induce apoptosis. These collective results suggest that pyrogallol treatment in A549 cells exerts regulatory effects and is involved in the STAT2 signaling pathway.
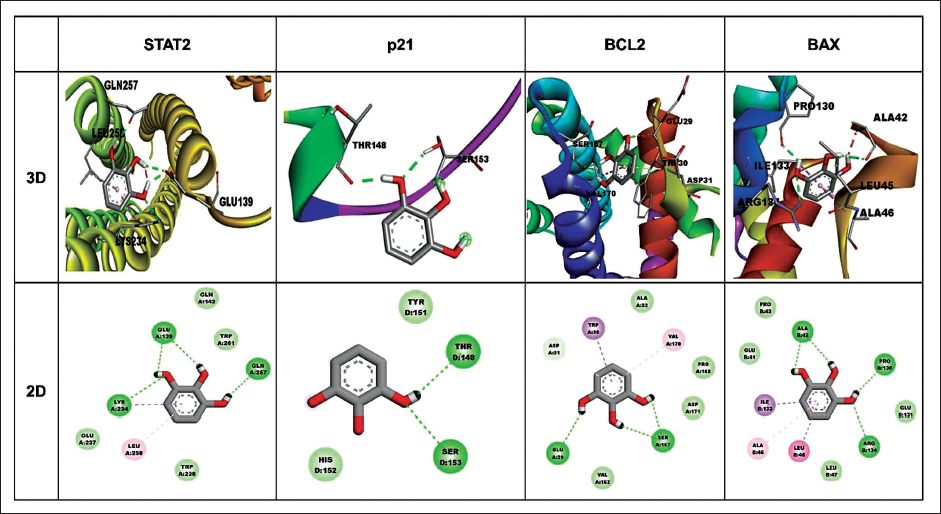
Molecular Docking Pyrogallol and Signaling Targets
Molecular docking techniques are crucial components in drug discovery, due to the easy to identify the binding energy and affinity of the drug and target proteins. We employed AutoDockTools version 1.5.6 to analyze the binding energy of pyrogallol and target proteins. 2D and 3D view was visualization utilized in the Discovery Studio 2021 as depicted in Figure 5. The binding energy of the target STAT2, p21, Bcl-2, and BAX has the values of −4.06, −3.79, −4.29, and −4.76 kcal/mol presented in Table 1. The target BAX has a lower binding energy has −4.76. The pyrogallol has introduced the 3 hydrogen bonds with ALA42, PRO130, and ARG 134 residues of BAX. These interactions regulate the cell death in cancer cells to inhibit lung cancer proliferation.
Molecular Docking Analysis Between Pyrogallol and Target Proteins. 3D, and 2D Visualization of Molecular Interaction of Pyrogallol with Targets Proteins (STAT2, p21, BCL2, and BAX).
Molecular Docking Results.
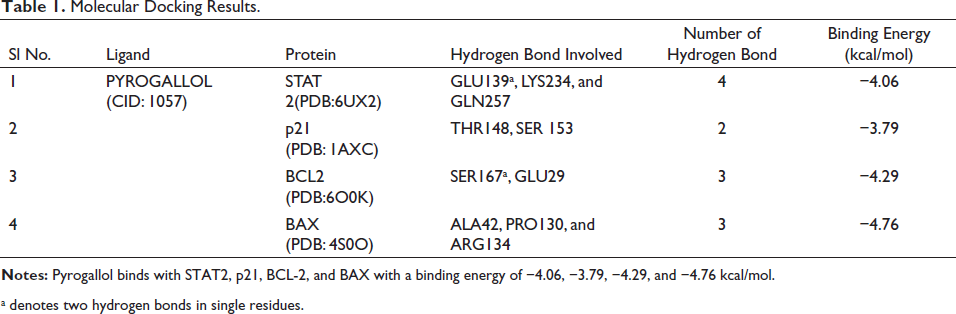
a denotes two hydrogen bonds in single residues.
Discussion
In the current research work, we investigated the anticancer mechanism of pyrogallol in the A549 human lung cancer cell line. Our findings demonstrate that the anticancer efficacy of pyrogallol is attributed to its ability to induce apoptosis, as evidenced by the presence of the apoptosis marker Annexin V. Given the significance of apoptosis induction in thwarting human malignancies, particularly those stemming from defects in apoptosis pathways, the identification of agents capable of promoting apoptosis in cancer cells holds substantial promise for chemotherapy (Chaudhary et al., 2016; Sznarkowska et al., 2017). Pyrogallol was selected for this investigation due to its derivation from the heating of gallic acid, and previous research has already established its cytotoxic effects on various human cell lines, including human erythromyeloid K562, B-lymphoid Raji, Erythroleukemic HEL cell line (Khan et al., 2002). Our study contributes novel insights into the anticancer mechanisms of pyrogallol specifically in human lung cancer cells, shedding light on its potential as a valuable agent in the ongoing pursuit of effective cancer therapies.
Initially, bioinformatics analysis assisted in creating a GSEA plot that highlighted pathways highly enriched in cancer. Additionally, the analysis of the GEOR2, CTD, and WebGestalt database revealed 36 overlapping targets strongly associated with pyrogallol against lung cancer. Furthermore, the protein-protein interaction analysis illustrated the interrelationship between these targets. We conduct a cell viability assay of pyrogallol against the human lung cancer A549 cell line using the MTT assay method. The increased concentration of pyrogallol resulted in decreased viability in the lung cancer A549 cell line. Pyrogallol-induced morphology changes in lung cancer A549 cells, as shown in Figure 2B. From the result, pyrogallol demonstrated the ability to induce cytotoxicity in the lung cancer A549 cell. Moreover, an elevated concentration of pyrogallol at 150 µM has prooxidant activity. This prooxidant effect can disrupt mitochondrial function, which could account for the heightened cytotoxicity observed at higher doses (Fernandes et al., 2021). Additionally, pyrogallol induces morphological changes at higher concentrations in treated groups, changes remarkably similar to the hallmark of apoptosis observed in flow cytometry analysis. Previous studies demonstrate that a higher concentration of pyrogallol induces cytotoxicity in liver cancer cell lines like Hep G2 and HUH7 (Ahn et al., 2019). Another study report suggests that pyrogallol could enhance apoptosis by Modulating Akt/GSK-3β/β-catenin signaling Pathway in prostate cancer (Arjsri et al., 2023). In our subsequent study, we examined the effects of pyrogallol on aerobic glycolysis and found substantial results. Specifically, in A549 cells, pyrogallol significantly decreased the activity of glucose uptake, as illustrated in Figure 2C.
The author further observed that inhibition of the STAT2 protein might suppress cancer cell proliferation. Molecular docking results demonstrate the binding energy of pyrogallol and STAT2 proteins has a value of −4.06 kcal/mol. Then, pyrogallol suppresses STAT2 and its downregulated signaling pathways. These signaling pathways are intricate with cell cycle and apoptosis events. From our data, pyrogallol inhibits STAT2, downregulates their downstream signaling cascade to arrest the cell cycle via p21, and inhibits the anti-apoptosis molecules Bcl-2, as shown in Figure 4. Pyrogallol can induce DNA damage in breast cancer cells. DNA damage can trigger cell cycle arrest or apoptosis as cells attempt to repair the damage (Dutta & Khanna, 2016). Previous findings suggest the potential of pyrogallol as an anti-cancer agent, further research is needed to fully understand its mechanisms of action. The CDKN1A gene encodes the protein CDKN1A, also known as p21. CDKN1A is s cell cycle regulator that controls cell growth, proliferation, differentiation, DNA repair, and apoptosis. It is a key regulator of the cell cycle that ensures genomic stability. However, CDKN1A is often deregulated in human cancer. It involves in G2/M checkpoint that contributes to resistance to Taxol-induced apoptosis in ERBB2-overexpressed breast cancer cells (Kreis et al., 2019). Moreover, CDKN1A has demonstrated important prognostic significance and may be used as an independent prognostic factor for patients with RGA (Lin et al., 2017). In our study, we analyzed the molecular interaction of pyrogallol with p21 protein, with a binding energy value of −3.79 kcal/mol, and in vitro study shows that pyrogallol suppresses the expression of p21. Meanwhile, our investigation strongly recommends the inhibition of anti-apoptosis molecules like Bcl-2 to promote the cell death mechanisms. From the result of our study, the molecular interaction of pyrogallol has a higher affinity and lower binding energy with Bcl-2, with a value at −4.29 kcal/mol, introducing two hydrogen bonds with SER167, and one hydrogen bond with GLU29 to increase stability. Yang et al. (2009) previously conducted a study demonstrating that pyrogallol induces G2-M arrest in human lung cancer cells and inhibits tumor growth in an animal model. Moreover, previous research investigations, licochalcone H (Park et al., 2022), celecoxib (Liu et al., 2012), ursolic acid (Li et al., 2017), and AG490 have the potential to induce the cell cycle arrest in the G1 phase and induce the apoptosis in Human skin cancer cells (Xiong et al., 2008), nasopharyngeal carcinoma cell line, renal and breast cancer, and colorectal cancer are respectively. These findings suggest that pyrogallol has great potential to inhibit cancer progression via targeting STAT2 signaling to induce cell cycle arrest and cell death in lung cancer.
Conclusion
Taken together, our study unveils the potent anticancer properties of pyrogallol in human lung cancer cells, highlighting its ability to induce apoptosis and disrupt key signaling pathways, like STAT2, p21, and Bcl-2. This finding underscores pyrogallol’s potential as a valuable agent in the development of target therapies against lung cancer. Further research and clinical exploration are warranted to harness the full therapeutic benefits of pyrogallol in the battle against this formidable disease.
Abbreviations
BAX: Bcl-2-associated X-protein; Bcl-2: B-cell lymphoma 2; CDKN1A: Cyclin-dependent kinase inhibitor 1A; CTD: Comparative Toxicogenomics Database; GEO: Gene Expression Omnibus; GSEA: Gene Set Enrichment Analysis; KEGG: Kyoto Encyclopedia of Genes and Genomes; MTT: 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide; STAT2: Signal transducer and activator of transcription 2; STITCH: Search tool for interactions of chemicals.
Authors’ Contributions
HL and YC: Data acquisition, analysis, and interpretation of data.
LJ: Conception and design, drafting the manuscript, review key revision, and approval publishing.
