Abstract
Background
Ziziphi spinosae semen (ZSS), dry mature seeds of Ziziphus jujuba Mill. var. spinosa (Bunge) Hu ex H.F. Chow., are widely used in the treatment of insomnia, fright palpitations, and profuse dreaming in medical treatments. The counterfeit species of ZSS, Ziziphi mauritianae semen (ZMS), and Hovenia dulcis Thunb (HDT) are seeds from Ziziphus incurva Roxb. and Hovenia acerba Lindl., respectively. However, the pharmacopeia is not recorded for ZMS and HDT. In this article, we reviewed the identification methods of ZSS, ZMS, and HDT.
Materials and Methods
The random amplified polymorphic DNA (RAPD) technique was used to identify the ZSS and its copies. Plant genome DNA was extracted from Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. Random primers were designed to amplify the genome. The amplified products were detected by agarose gel electrophoresis.
Results
By analyzing the polymorphism of the amplified product DNA fragment, the DNA fingerprint maps of Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. were successfully constructed.
Introduction
Characteristics of Ziziphus jujuba Mill., Ziziphus incurva Roxb., and Hovenia acerba Lindl.
Ziziphus jujuba Mill., a member of the genus Ziziphus in the family Rhamnaceae, is widely distributed in China, mainly growing in the hills and mountains. It has been used in Traditional Chinese medicine for more than 2000 years and has a high medicinal value. According to “Shen Nong’s Herbal Classic” records, its fruit can treat sleeplessness, pain above and below the umbilicus, sweating, and polydipsia. It can also tonify the liver, make the ribs strong, nourish the yin, and make us fat. In recent years, pharmacological research has found that Z. jujuba Mill. contains various bioactive compounds, such as polysaccharides, flavonoids, saponins, triterpenoids, alkaloids, and tannins, which have several health benefits. Modern medical research has shown that Ziziphi spinosae semen (ZSS) contains triterpenoids, organic acids, flavonoids, polysaccharides, amino acids, and trace elements (Xie et al., 2021). ZSS can prevent cerebral diseases (Liu et al., 2022). It could protect against lipopolysaccharide-induced blood–brain barrier dysfunction (Liu et al., 2022). In addition, ZSS exerted sedative and hypnotic effects mainly through the GABAergic and serotonergic systems (Bian et al., 2021; Zhou et al., 2018). Sleep was improved by regulating the expression levels of the GABA receptor subunit alpha-1 (GABAARα1) and GABA acid receptor subunit gamma-2 (GABAARγ2) receptors in tissue sections of the hypothalamus and hippocampus. ZSS can exert a neuroprotective effect on rats with p-chlorophenylalanine-induced insomnia via activation of the GABAA receptor (Xiao et al., 2022). ZSS can also inhibit biofilm formation, improve the bacterial pH environment, and eliminate the hydrophobic effect of reactive oxygen species and flavonoids (Miao et al., 2020). In recent years, the increasing economic value of Z. jujuba Mill. has garnered interest. Z. jujuba Mill. juice, Z. jujuba Mill. leaf tea, and other related products have been developed. Additionally, the domestic market demand for ZSS is increasing. The contradiction between supply and demand is becoming increasingly prominent, and its market price has gradually soared from 30 yuan per kilogram to approximately 500 yuan per kilogram.
Many drug-related units sell and use imported jujube kernels instead of ZSS. The imported jujube kernel, also known as Ziziphus incurva Roxb., is derived from the dry, mature seeds of Hovenia acerba Lindl. It is cheaper than ZSS, but its medicinal traits are similar to those of ZSS, with a sweet and neutral taste. Jujube kernels can treat insomnia, fright palpitations, and agitation. Wu et al. (2021) established a stable and reliable quality evaluation method for ZSS and Ziziphi mauritianae semen (ZMS), which provides a reference for quality marker components and a scientific basis for the feasibility of ZMS as a substitute for ZSS (Wu et al., 2021). While the source of ZMS is not consistent with the ZSS contained in the Chinese Pharmacopeia, it cannot replace the ZSS.
Hovenia dulcis Thunb (HDT), the fruit of Hovenia acerba Lindl., is very similar to ZSS. HDT can also be used as a medicine with significant effects on liver protection (Je et al., 2021; Xu et al., 2023), anti-hepatic fibrosis (Geng et al., 2008), and anti-fatigue (Jia et al., 2009). HDT contains flavonoids (Yang et al., 2019), and ten known flavonoid C-glycosides were purified (Xu et al., 2020). Because the shapes of HDT and ZSS are very similar, confusion can occur during drug transportation and storage.
Research Progress of Identification of ZSS, ZMS, and HDT
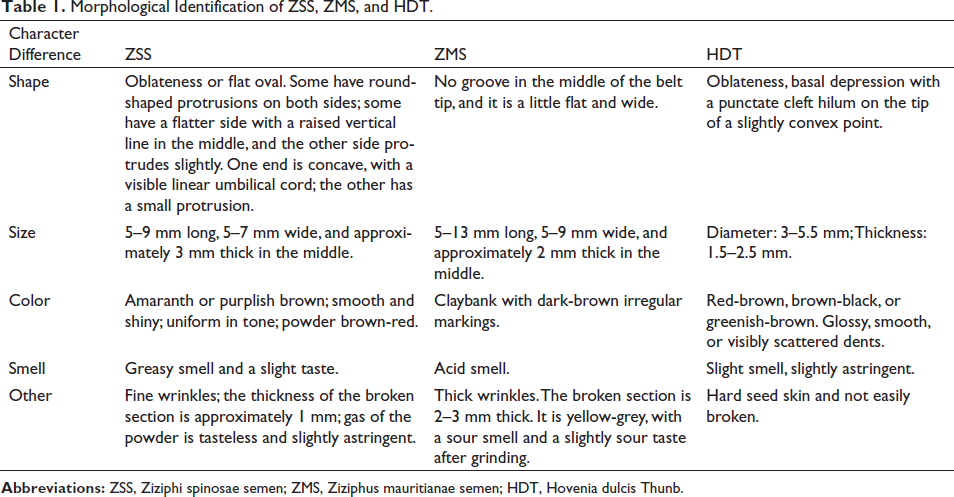
Traditional Chinese Medicine identification methods mainly rely on appearance and shape characteristics. Morphological identification of shape, size, color, smell, taste, texture, and other traits (Table 1) is simple and direct. However, this method is highly subjective. Its accuracy entirely depends on the experience of the detector. It is challenging to identify processed fragments or powdered materials. With the development of science and technology, a more objective Traditional Chinese medicine identification standard has been established from the perspectives of biological taxonomy (fundamental identification), histology (microscopic identification), and chemistry (physical and chemical identification; Table 2).
Morphological Identification of ZSS, ZMS, and HDT.

Microscopic and Physico-chemical Identification of ZSS, ZMS, and HDT.
High-performance liquid chromatography (HPLC), ultra-performance liquid chromatography (UPLC), and thin-layer chromatography (TLC) are mainly used to identify ZSS and ZMS. Hongshuai et al. (2020) performed HPLC fingerprint analysis and identification studies of ZSS and ZMS, indicating ten common fingerprint peaks and two different peaks. HPLC can be used for the quality control of ZSS (Hongshuai et al., 2020; Sun et al., 2014).
There was no significant difference in the infrared spectra of ZSS and ZMS because they contained similar chemical components. However, near-infrared spectroscopy technology can be used to study the qualitative and quantitative detection of ZSS and its three counterfeits, namely, Ziziphus mauritiana Lam., HDT, and Lens culinaris (Zhao et al., 2022).
The multiplex allele-specific polymerase chain reaction (PCR) can accurately identify ZSS, Hovenia acerba semen, and ZMS simultaneously (Li et al., 2023). The multiplex ligation-dependent probe amplification technique can also identify Z. jujuba var. spinosa seeds and adulterants (Wang et al., 2022). The source, efficacy, in vivo process, and traditional properties were probed to predict the Q-markers of ZSS based on the concept of Q-markers (Yan et al., 2019).
In short, the traits and microstructures of ZSS, ZMS, and HDT are different. However, ZSS and HDT can only be identified if they have an intact appearance. Several physical and chemical identification methods have been developed. These methods are easy to manipulate and should be applied in conjunction with other auxiliary methods to prevent discrimination. However, for Chinese patent medicines with more complex components, different components may affect the results of the physical and chemical identification methods.
Compared to these previous standards, DNA molecular labeling techniques have many advantages (Zhang et al., 2021). The DNA mapping method is based on the genetic polymorphism of Chinese medicinal materials using random amplified polymorphic DNA (RAPD) analysis technology to identify Z. jujuba Mill. The RAPD-PCR method had high sensitivity and yielded clear results when used to identify Z. incurva Roxb. and H. acerba Lindl. This method is not affected by external factors (e.g., environmental conditions), biological ontogeny stages, or organ and tissue differences. The genome contains almost an infinite amount of information. All organisms, including animals, plants, and micro-organisms, have several common molecular properties. Some basal metabolic genes and enzymes have similar functions. Therefore, the corresponding level of genetic variation between organisms and classification levels can be directly compared using DNA molecular marker data. DNA is more stable than proteins and can be detected in vivo. In addition, trace amounts of extracted DNA from old specimens can be used to study DNA molecular markers. The RAPD-PCR fingerprint-specific bands can be used for cultivar identification. RAPD-PCR can be used to identify the authentic properties of Chinese medicinal materials and easily mixed varieties (Li et al., 2022; Moon et al., 2015).
In this study, we used DNA fingerprint technology and a random primer PCR amplification method to establish the DNA fingerprint of Taishan Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. We expect to develop a new method with reasonable cost, accuracy, high throughput, and suitability for large-scale applications.
Materials and Methods
In this study, genomic DNA was extracted from the leaves of Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. Random primers were designed, and the genome was amplified with double primers using RAPD-PCR. Grayscale analysis was performed with ImageJ software.
Sample Collection
To identify Z. jujuba Mill. and Z. incurva Roxb., fresh leaves of Z. jujuba Mill. and Z. incurva Roxb. were collected from Shandong Medicine Technician College and Yunnan, respectively. Experts identified the samples.
In addition, leaves of Z. jujuba Mill. and H. acerba Lindl. were collected near Taishan Red Gate in Tai’an City and Shandong Agricultural University, respectively. Professors Zhang and Yongqing identified these samples.
DNA Extraction
The leaves were placed in a pre-cooled mortar, followed by the addition of liquid nitrogen, and ground as soon as possible. Leaf powder (1 g) was added to 10 mL of cetyltrimeth-ylammonium bromide (CTAB) buffer (2% CTAB, 1.4 M NaCl, 20 mM EDTA·2Na, 100 mM pH 8.0 Tris–HCl, and 0.2% β-mercaptoethanol) and preheated to 65°C. The leaf powder and CTAB buffer mix were kept at 65°C in a water bath for 30 min. Next, an equal volume of chloroform-isoamyl alcohol (24:1) was added. The mixture was then gently mixed and placed on ice for 10 min. After centrifugation at Hema Medical Instrument Co., Ltd., Zhuhai, at 4,000 rpm for 10 min, the supernatant was discarded. Then, a two-thirds volume of isopropyl alcohol (Shanghai Chemical Reagent Factory, China) was added, and the mixture was gently mixed at 4°C overnight. The next day, the mixture was centrifuged at 4,000 rpm for 2 min, and the supernatant was removed. The precipitate was washed three times with 75% ethanol. Finally, it was dried at room temperature and dissolved in TE buffer (TGL-18R). Genomic DNA was subjected to electrophoresis on a 1% agarose gel.
RAPD-PCR of Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl.
To identify Z. jujuba Mill. and Z. incurva Roxb., the primers indicated in Table 3 were used, and all the primers were synthesized (Dalian Dabao Biotechnology Company, China). S1-1, S1-2 … S10-1, and S10-2 were used to identify Z. jujuba Mill. and H. acerba Lindl. (Table 3). S11-1, S11-2, S12-2-1, S12-2 … S20-1, and S20-2 were used to identify Z. jujuba Mill. and Z. incurva Roxb. (Table 3). The primers were screened to prevent amplification between the two primer pairs.
Random-Primer Numbers and Sequences Used to Identify Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl.

A PCR was designed and run on a My-Cycler (Bio-Rad, USA) with the following components: 1U Taq enzyme (Ex Taq, Dalian Biology, RR001Q), dNTP mix (10 mM of each dNTP), 2.5 µL 10× Taq buffer, 1 µM primer, and 200 ng DNA template, supplemented with sterilized water to 25 µL. The amplification cycles were as follows: denaturation at 95°C for 30 s, 42°C for 30 s, and extension at 72°C for 1 min. The amplified products were electrophoresed (DYCP-31D, Beijing Liuyi Instrument Factory) on 1% agarose gel (Nanjing Shengxing Biotechnology Co., Ltd., Spain).
Results
DNA Extraction
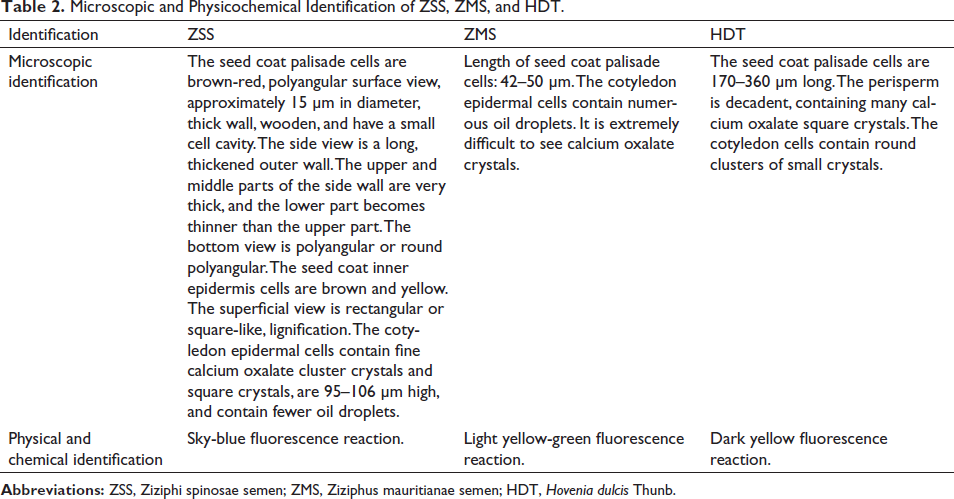
Genomes were extracted from the young leaves of Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. The extracted DNA was visualized using UV spectrophotometry after running on a 1% agarose gel. The extracted genomic results are shown in Figures 1 and 2. The band intensities are measured using ImageJ software and the data is available in the Supplementary Material.
Z. jujuba Mill. and Z. incurva Roxb. Genomic DNA. M: DL2000 DNA marker; 1–3: Z. jujuba Mill. Genomic DNA; 4–6: Z. incurva Roxb. Genomic DNA.
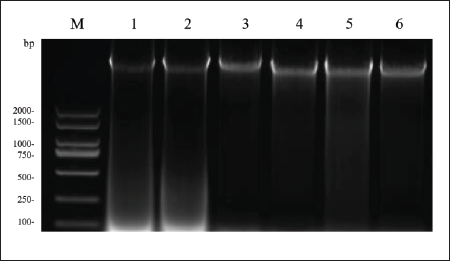
Z. jujuba Mill. and H. acerba Lindl. Genomic DNA. M: 15000 bp DNA Marker; 1: Z. jujuba Mill. Genomic DNA; 2: H. acerba Lindl. Genomic DNA.
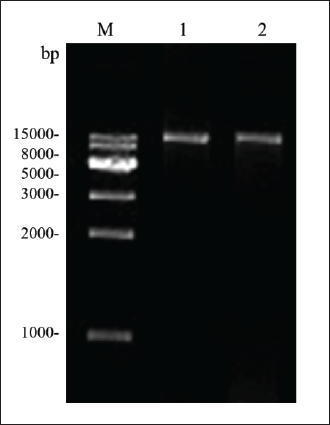
RAPD-PCR Results of Z. jujuba Mill. and Z. incurva Roxb.
DNA samples from Z. jujuba Mill. and Z. incurva Roxb. were amplified using random primers. In lanes 1, 3, 5, 7, 9, and 11, we used Z. jujuba Mill. as the template, while in lanes 2, 4, 6, 8, 10, and 12, we used Z. incurva Roxb. as the template. Some primers did not amplify the band, and the amplified RAPD polymorphism was absent. Both samples had the same amplified bands in the S11, S12, S13, and S15 primer groups. In the S14 group, a dark band was amplified for Z. jujuba Mill., whereas Z. incurva Roxb. did not amplify this band. The two DNA samples were different, but the differences were not significant. Therefore, the S14 primer was unsuitable for distinguishing Z. jujuba Mill. from Z. incurva Roxb. Grayscale scans showed no statistically significant difference in the bands amplified by S11, S12, S13, and S15 primers, which are indicated by the letter a (Figure 3B). In the S17 group, Z. jujuba Mill. showed a bright band of approximately 240 bp. However, Z. incurva Roxb. had two distinct bands of approximately 250 bp (Figure 3A). The grayscale scans of each lane showed a significant difference between lanes 11 and 12 (Figure 3B). The letters a and b denote statistical differences. Identification of Z. jujuba Mill. and Z. incurva Roxb. can be conducted. Finally, a pair of discriminating primers were selected: S17-1 and S17-2.
From the DNA fingerprint, some primers amplified a similar band, indicating that the two genes were similar. Only the S17 primers amplified characteristic bands, showing specific differences. Therefore, these primers were selected as characteristic primers to identify Z. jujuba Mill. and Z. incurva Roxb.
RAPD-PCR Results of Z. jujuba Mill. and H. acerba Lindl.
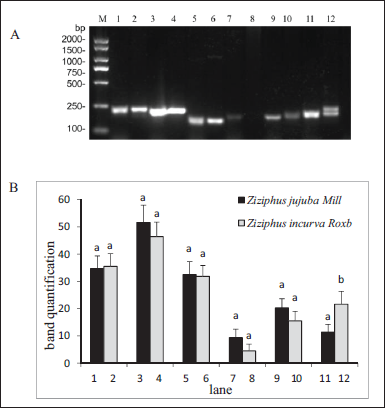
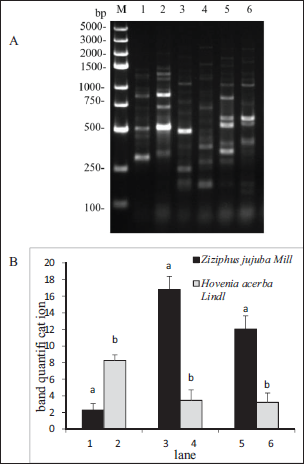
Amplification occurred between the S2-1 and S2-2 and the S7-1 and S7-2 primer pairs. Therefore, the two primer sets were discarded. The other eight primer pairs resulted in bands being amplified. Of these groups, S1, S4, S6, S9, and S10 of the two-sample amplified bands were identical. The other three amplified bands (S3, S5, and S8) were different. On lanes 1, 3, and 5, we used Z. jujuba Mill. as the template, while on lanes 2, 4, and 6, we used H. acerba Lindl. as the template. In group S3, there were bright bands at about 500 bp and 750 bp in H. acerba Lindl., while the band of Z. jujuba Mill. was unclear. In the S5 group, Z. jujuba Mill. had a distinct band at about 500 bp, but H. acerba Lindl. had no band. In group S8, Z. jujuba Mill. had three bands between 250 bp and 500 bp, and one band at approximately 800 bp. However, only one band was found between 250 bp and 500 bp, and no band at approximately 800 bp in H. acerba Lindl. (Figure 4A). The grayscale scans of each lane showed a significant difference (Figure 4B). The letters a and b denote statistical differences. Using the above specific bands, we identified Z. jujuba Mill. and H. acerba Lindl. Three primer pairs were selected: S3-1 and S3-2; S5-1 and S5-2; and S8-1 and S8-2.
Discussion
Currently, the majority of ZSS identification is derived from microscopic identification and thin-layer chromatography. The microscopic identification was subjective. Chromatography is influenced by spot sample size and other factors. Further experimental verification is necessary for ZSS identification.
In this study, RAPD was used to establish the DNA fingerprint map of Taishan Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. It is a new method that is sensitive, accurate, has high throughput, and is suitable for large-scale popularization and application. TCM uses genetic polymorphism to map DNA. Using DNA markers and other advanced technology to establish a complete and accurate Chinese medicinal materials identification standardization system can protect the existing resources, develop new resources, and improve the quality of medicinal materials. In particular, DNA molecular markers are rarely limited by biological individuals’ growth and development stages, the state of the material examined, or the length of preservation time. It directly identifies the genetic differences within samples; therefore, it is widely used in researching germplasm resources and various identification methods of Chinese medicinal materials. In this study, genomic DNA was extracted from fresh leaves of Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. using the CTAB method. The RAPD-PCR amplification method was performed. The fingerprint of Z. jujuba Mill. was completely different from that of Z incurva Roxb. and H. acerba Lindl. Identifying a DNA fingerprinting method to distinguish Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. has filled this research gap in China.
Babu et al. (2021) presented the applications of RAPD in genetic diversity analysis (Babu et al., 2021). RAPD technology provides a new alternative for cultivar identification and classification in celery (Yang & Quiros, 1993), date palm cultivars (Al-Khalifah & Shanavaskhan, 2017), and Lilium (Yamagishi, 1995). The authentication of the medicinal plant Senna angustifolia also utilized the RAPD technique (Khan et al., 2011). RAPD-derived DNA markers can also be used to authenticate Bulbus fritillariae cirrhosae (Xin et al., 2014).
However, there are some limitations to using the RAPD analysis technique in TCM. We tested pure products, but synthetic products are encountered in real-world situations. The next step requires the identification of the synthetic mixture. In recent years, the study of Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl has attracted the attention of scholars at home and abroad. Researchers have been mainly interested in the chemical composition, pharmacological effects, and product development of these three plant species. Molecular identification technology is now widely used in germplasm resources, genetic diversity research, classification, kinship identification, and authenticity identification, but only rarely in jujube plant biological activity and resource development and utilization. In the next step, a comprehensive investigation of jujube plant resources can be conducted. Molecular identification technology can be used to conduct scientific seed selection and systematic cultivation of jujube varieties growing naturally in the wild to provide resource guarantee for their rational utilization and continuous development, combined with different molecular identification techniques to establish genetic maps unique to other species of Jujube. This innovation will gradually become one of the most important methods for studying jujube plants as new molecular identification technology is developed and applied.
Conclusion
This study used a simple, reproducible, and reliable RAPD analysis method for Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. that can readily be utilized for species identification and quality control of Z. jujuba Mill. and its products. Several pairs of primers were selected, and fingerprints were obtained. Plant genomes were extracted from fresh leaves of Z. jujuba Mill., Z. incurva Roxb., and H. acerba Lindl. The DNA quality was determined by UV spectrophotometry. RAPD-PCR amplification was then performed by agarose gel electrophoresis. One primer pair was selected based on the electrophoresis results to identify Taishan ZSS and ZMS. The amplification of three pairs of primers made the fingerprint of ZSS completely different from that of HDT.
This could be an opportunity to conduct an investigation and seed selection for ZSS, strengthen resource protection, establish a cultivation base of fine varieties, and finally strengthen its research and development. This will increase the overall use of Taishan ZSS and provide economic benefits to the local pharmaceutical industry. Furthermore, this research protects drug quality, patient safety, and public health and has some social promotion and application value.
Footnotes
Summary
DNA fingerprint technology and the RAPD method were used in this study to distinguish ZSS from its fake alternatives. It is a new method with a reasonable cost and fast, accurate, and high-throughput identification. This approach offers an opportunity to conduct an investigation and seed selection for ZSS resources. It is beneficial for strengthening the protection of ZSS resources, establishing the cultivation base of excellent ZSS varieties, and strengthening the development of ZSS processing. It will play a role in promoting the comprehensive utilization of real Taishan ZSS and increasing the economic benefits of the local pharmaceutical industry. In addition, this research plays a protective role in ensuring the quality of drugs, the safety of patients, and people’s health, and it has a particular social promotion and application value.
Abbreviations
CTAB: cetyltrimethylammonium bromide; HDT: Hovenia dulcis Thunb; HPLC: high-performance liquid chromatography; PCR: polymerase chain reaction; RAPD: random amplified polymorphic DNA; TLC: thin-layer chromatography; UPLC: ultra-performance liquid chromatography; ZMS: Ziziphus mauritiana semen; ZSS: Ziziphi spinosae semen.
Acknowledgments
We thank Professor Zhang Yongqing for his kind assistance.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research was supported by the Tai’an Science and Technology Development Plan Project (grant number: 201340629) and Ningxia key research and invention program of science and technology cooperation of the East and the West (No. 2017BY084).
